Abstract
Amelogenesis imperfecta (AI) comprises a group of rare, inherited disorders with abnormal enamel formation. Ameloblastin (AMBN), the second most abundant enamel matrix protein (EMP), plays a critical role in amelogenesis. Pathogenic biallelic loss-of-function
Introduction
Enamel is formed when ameloblasts secrete, then mineralize, an extracellular matrix (ECM) composed of enamel matrix proteins (EMPs), in a process known as amelogenesis. Throughout the secretory stage of amelogenesis, ameloblasts secrete the EMPs amelogenin (AMELX), ameloblastin (AMBN), enamelin (ENAM), and amelotin (AMTN) and the matrix modifier matrix metallopeptidase 20 (MMP20) (Lee et al. 1996). In the later maturation stage, a further matrix modifier, kallikrein-related peptidase 4 (KLK4), is also secreted (Smith et al. 2017; Pandya and Diekwisch 2021). EMPs play an essential role in the biomineralization and structural organization of enamel (Bartlett et al. 2006).
Amelogenesis imperfecta (AI) describes a heterogeneous group of Mendelian disorders causing abnormal amelogenesis, affecting all teeth of both dentitions (Smith et al. 2017). Reported prevalence ranges between 1 in 700 (Sweden) and 1 in 14,000 (United States) (Bäckman and Holm 1986; Witkop 1988). Poor aesthetics and early functional failure create considerable challenges for affected individuals and those providing care. AI can be isolated or part of syndromic conditions, with many genes implicated (Smith et al. 2017; Wright 2023).
AMBN, a phosphorylated glycoprotein, is the second most abundant EMP after AMELX. The
Here, we report 5 novel
Materials and Methods
Patients were recruited though UK dental clinics in accordance with the principles of the Declaration of Helsinki (ethical approval REC 13/YH/0028). Genomic DNA was isolated from saliva or from peripheral blood by standard approaches as detailed in the Appendix methods.
Proband genomic DNA was analyzed by short-read next generation sequencing of either whole-exome sequencing (WES) or single-molecule molecular inversion probes (smMIPs) data generated on HiSeq 3000, NextSeq 500, or NextSeq 2000 sequencers (Illumina). Further details of methods used in library preparation and sequence analysis are given in the Appendix methods. The pathogenicity status of detected variants was classified according to American College of Medical Genetics and Genomics (ACMG) guidelines using Franklin (https://franklin.genoox.com) (Richards et al. 2015).
Long-read sequencing was carried out on a Flongle flowcell (R.9.4.1), using a MinION (ONT) device running MinKNOW, to analyze long-range polymerase chain reaction (PCR) products amplified by SequelPrep polymerase (ThermoFisher Scientific), following the manufacturer’s guidelines. Methods used in sequence analysis are detailed in the Appendix methods. Haplotypes were defined after selection of reference and non-reference-matching nucleotides at the positions being examined, using the Jvarkit tool biostar214299 (http://lindenb.github.io/jvarkit/Biostar214299.html) (Lindenbaum 2015). Aligned sequence reads were visualized using the Integrated Genome Viewer (v.2.7.2) (Robinson et al. 2011).
Variants were confirmed and segregation tested by PCR amplification and Sanger sequencing on an ABI3130xl Genetic Analyser (Applied Biosystems). Electropherograms were analyzed using SeqScapeTM (v.2.5) (ThermoFisher Scientific).
Intact teeth were analyzed using a high-resolution micro–computed tomography (µCT) SkyScan 1172 (Bruker) scanner to quantify mineral density. Video showing the 3D internal and external features was created using CTVox (Bruker). Longitudinal mid-bucco slices of the teeth were imaged on an S-3400N scanning electron microscope (SEM) (Hitachi). Further details are in the Appendix methods.
Results
Probands in a large cohort of apparently unrelated patients/families with AI were subject to ongoing screening of AI-associated genes, either by targeted smMIPs or WES. Members of 11 families with likely pathogenic
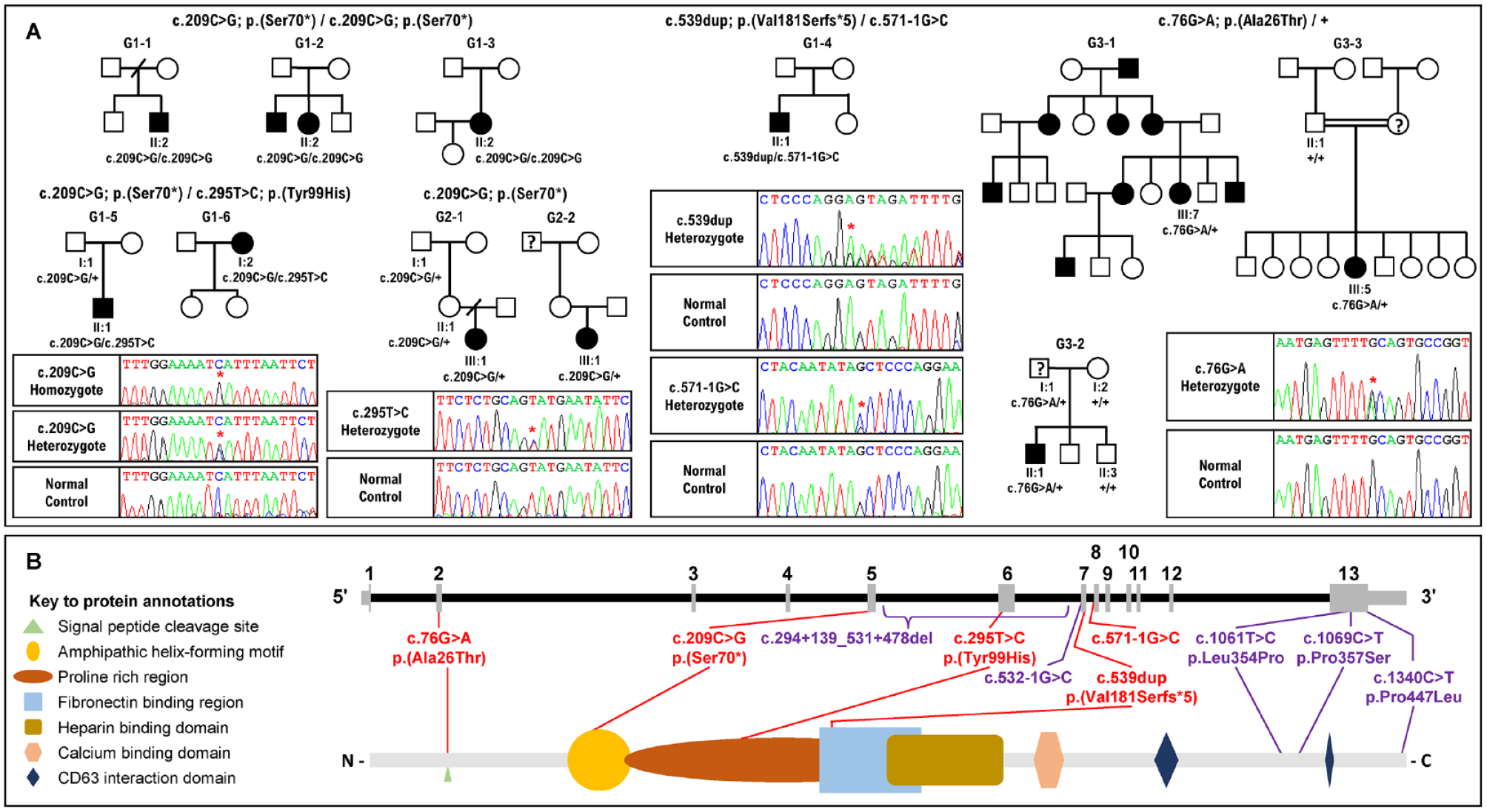
Pedigrees of the families recruited in this study, electropherograms of the variants identified and the location of the variants in the gene and protein. (
Genetic Findings
The 11 families could be sorted into 3 groups according to their
Details of AMBN Variants Detected in the Probands of 11 Recruited Families.
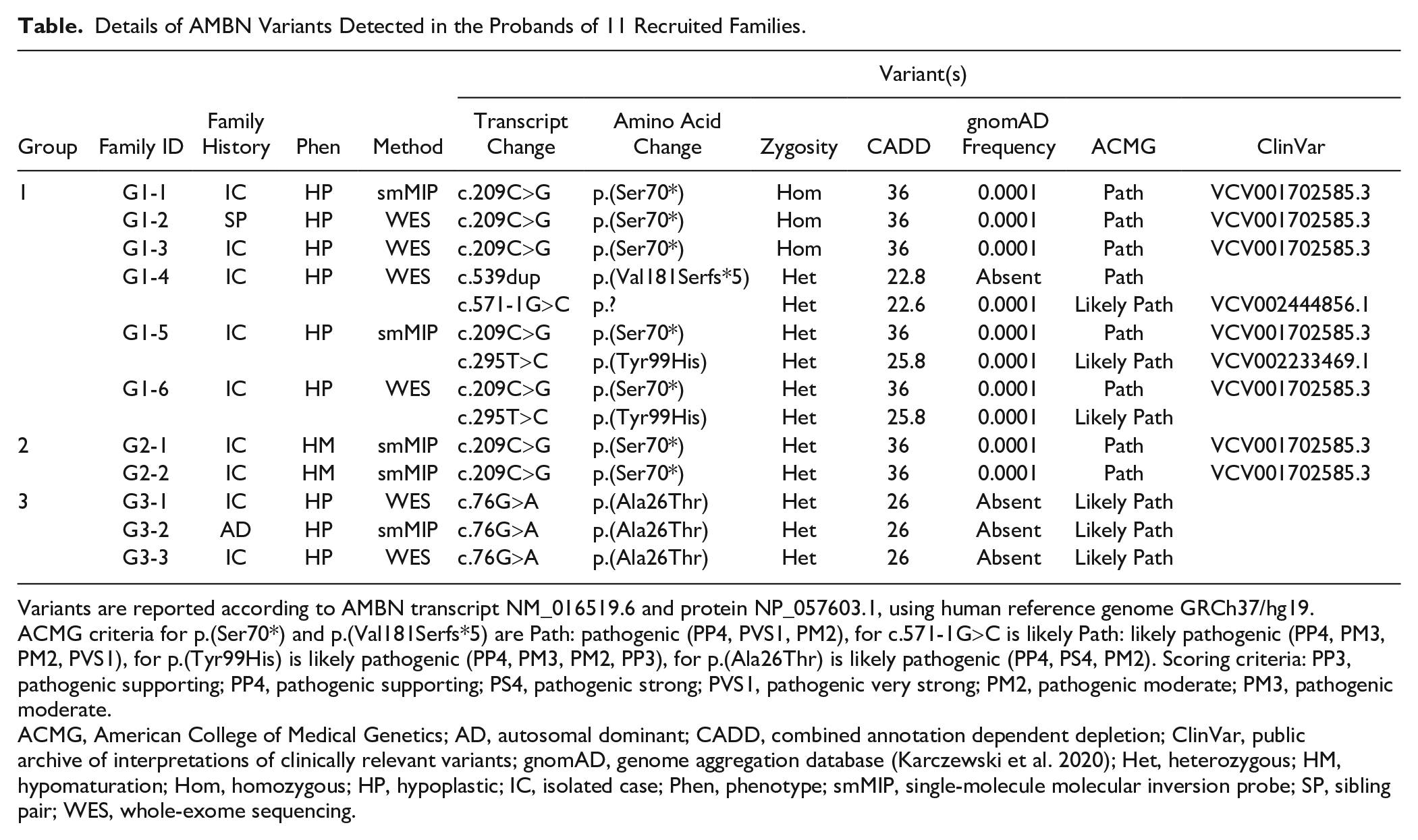
Variants are reported according to AMBN transcript NM_016519.6 and protein NP_057603.1, using human reference genome GRCh37/hg19. ACMG criteria for p.(Ser70*) and p.(Val181Serfs*5) are Path: pathogenic (PP4, PVS1, PM2), for c.571-1G>C is likely Path: likely pathogenic (PP4, PM3, PM2, PVS1), for p.(Tyr99His) is likely pathogenic (PP4, PM3, PM2, PP3), for p.(Ala26Thr) is likely pathogenic (PP4, PS4, PM2). Scoring criteria: PP3, pathogenic supporting; PP4, pathogenic supporting; PS4, pathogenic strong; PVS1, pathogenic very strong; PM2, pathogenic moderate; PM3, pathogenic moderate.
ACMG, American College of Medical Genetics; AD, autosomal dominant; CADD, combined annotation dependent depletion; ClinVar, public archive of interpretations of clinically relevant variants; gnomAD, genome aggregation database (Karczewski et al. 2020); Het, heterozygous; HM, hypomaturation; Hom, homozygous; HP, hypoplastic; IC, isolated case; Phen, phenotype; smMIP, single-molecule molecular inversion probe; SP, sibling pair; WES, whole-exome sequencing.
Group 2 includes 2 cases of isolated AI from families G2-1 and G2-2, without any history of AI in the family. The probands in each family are heterozygous for the
Group 3 includes family G3-1, with an extensive family history of dominant AI but only a single affected individual recruited. This group also includes 2 additional families with possible AD-AI: G3-2, for which the father of the proband self-reported as unaffected but was not clinically examined, and G3-3, for which the proband’s mother is said to have AI but was not examined clinically. Probands from these families are all heterozygous carriers of the novel missense variant c.76G>A; p.(Ala26Thr). This variant was classified as likely pathogenic. No second
Founder Effect Screening
The presence of variant c.209C>G; p.(Ser70*) in 7 families (groups 1 and 2), c.295T>C; p.(Tyr99His) in 2 families (group 1), and c.76G>A; p.(Ala26Thr) in 3 families (group 3) suggests these variants may have been inherited from a common ancestor. To test this hypothesis, we examined the haplotype backgrounds of these variants at the
A 9,681-bp DNA segment spanning exons 4 to 13 of
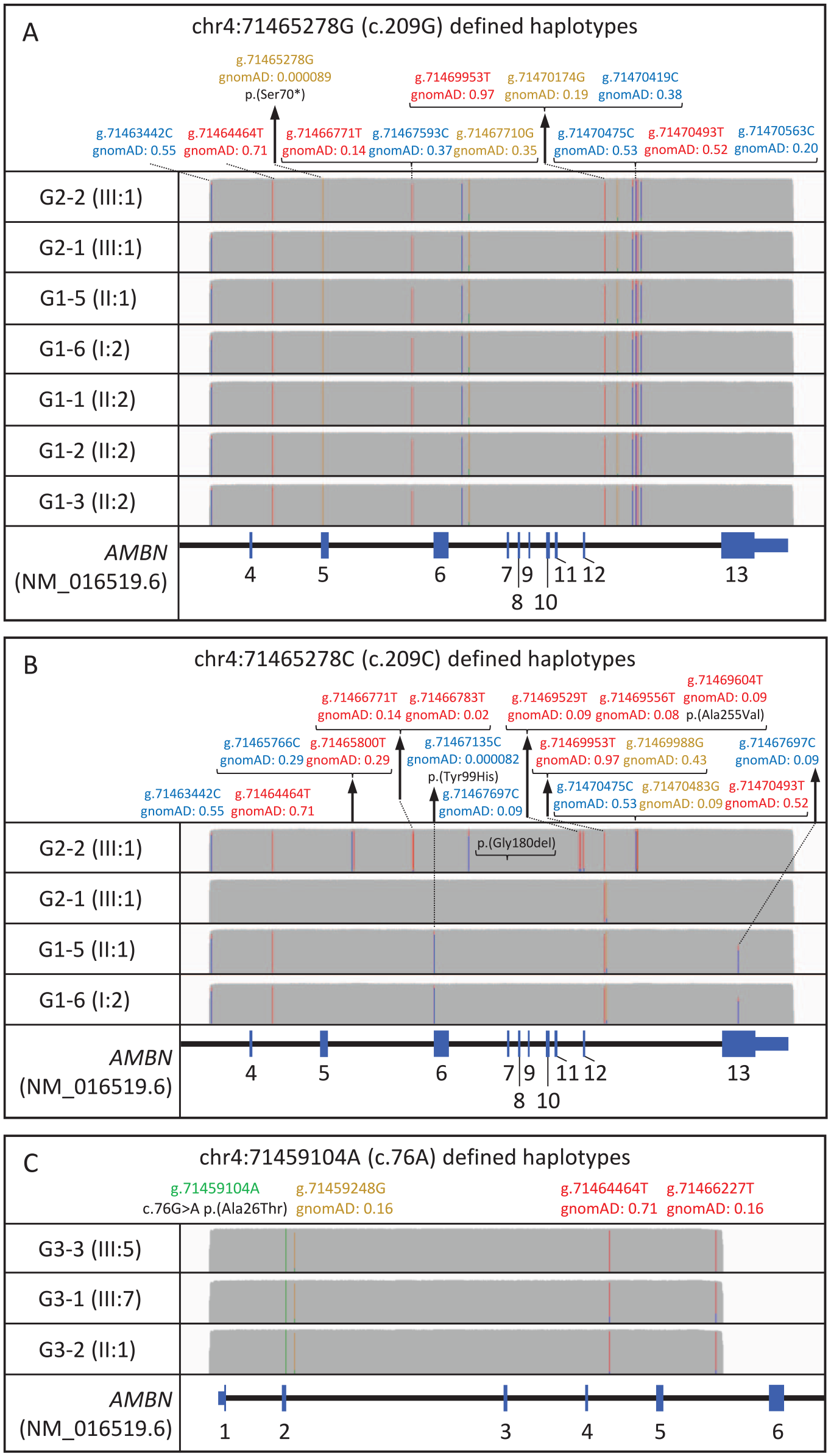
Long-read sequencing of a 9.7-kb amplification product from the
An 8,520-bp amplicon, spanning exons 1 to 5 and including residue 26, was PCR amplified and analyzed by long-read nanopore sequencing in probands from families G3-1, G3-2, and G3-3, all heterozygous for the
Phenotyping
Images of teeth and dental radiographs identified differences in the clinical phenotypes of the 3 groups recognized through genetic analyses (Fig. 3 and Appendix Fig. 1). Affected individuals in group 1 were characterized by hypoplastic AI with poor-quality enamel, with teeth having a yellow appearance following early posteruption loss of a thin layer of creamy, opaque mineralized tissue. Affected individuals in group 3 also had hypoplastic AI, but this differed from group 1 through the presence of a thin layer of more persistent enamel, which gives the teeth a whiter long-term appearance than group 1. Group 2 has a very different phenotype, characterized by hypomaturation AI with associated pits and minor morphological variations within a near-normal enamel volume that is more radio-dense than the supporting dentine on clinical radiography. No clear dentine abnormalities were evident on dental radiographs.
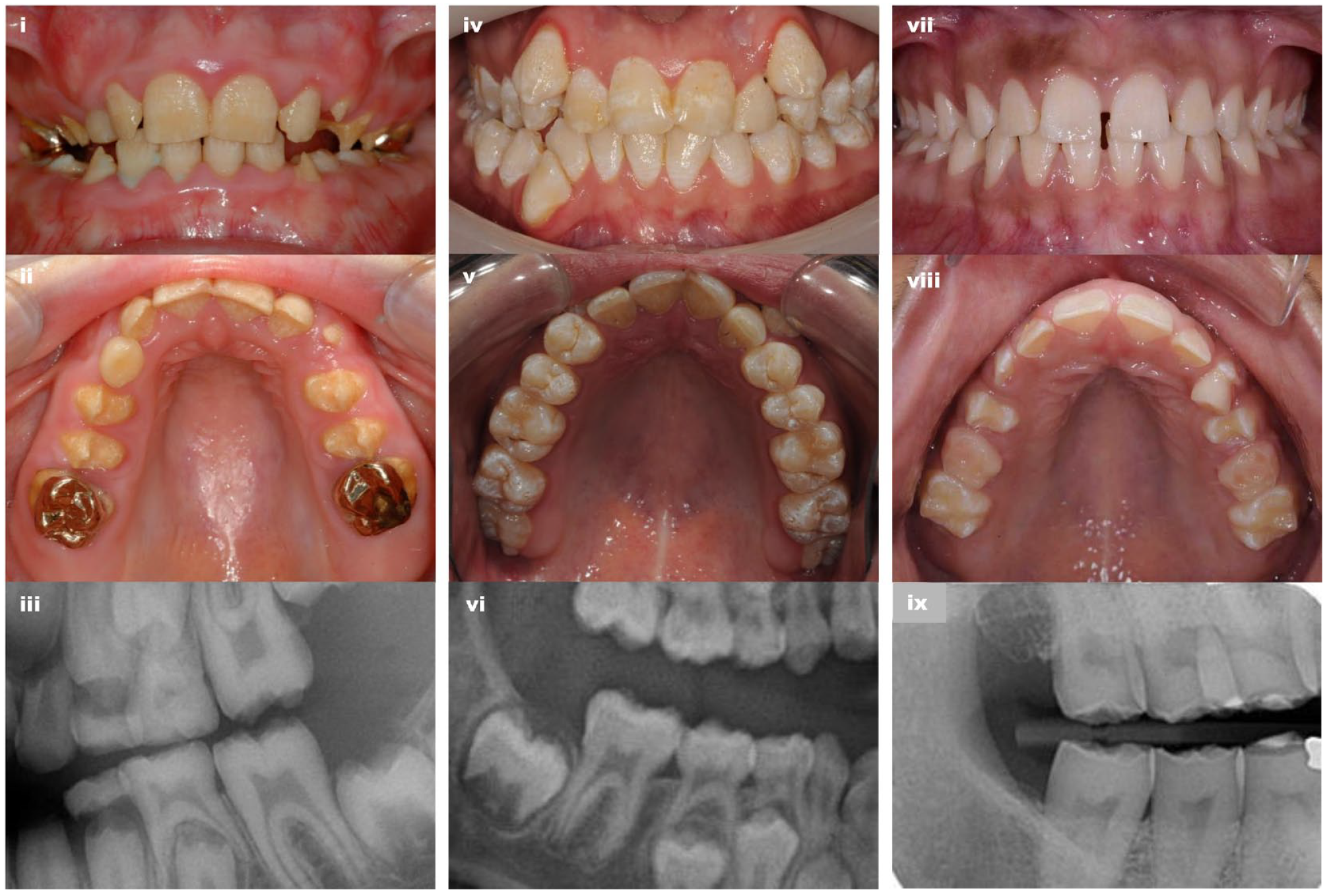
Clinical images and radiographs of the teeth capture the differences between the 3 groups. Group 1 (i–iii): Yellow hypoplastic amelogenesis imperfecta (AI) reflects the absence of any meaningful enamel on dental radiography (i and ii G1-2; iii G1-5 bitewing). Group 2 (iv–vi): Hypomaturation AI is characterized by variations in color with pits and other localized morphological changes that disrupt the normal clinical enamel surface. Dental radiography confirms near-normal enamel volumes with a clear difference between enamel and dentine radiodensity (iv and v G2-2; vi G2-2 detail from panoramic radiograph). Group 3 (vii–ix): White hypoplastic AI reflects the presence of a thin layer of enamel on dental radiography (vii and viii G3-3; ix G3-1 detail from panoramic radiograph). Further clinical images are included in Appendix Figure 1.
Teeth were available from a primary upper lateral incisor from the G2-1 proband and a permanent canine from the G2-2 proband for laboratory analyses. µCT of these teeth revealed normal enamel volume (Fig. 4i –iv). No significant differences were observed in average enamel mineral density (EMD) between affected and control teeth of the same type obtained from unrelated unaffected individuals. The EMD in G2-1 and its respective control were 2.561 g.cm–3 and 2.546 g.cm–3, and in G2-2 and its respective control, they were 2.569 g.cm–3 and 2.721 g.cm–3. An outer layer of particularly high mineral density seen in the control was missing in the G2-1, while the enamel of the G2-2 appeared pitted (Fig. 4i, iii) with pits extending through the enamel layer to the dentine–enamel junction (DEJ) (Fig. 4vii –ix, Appendix video). SEM analysis of these teeth showed disrupted, poorly formed prismatic microstructure with little demarcation between rod and interrod regions (Fig. 4xiii–xv). Hunter–Schreger banding was also absent in the affected tooth (Fig. 4xvi) as opposed to the control (Fig. 4xvii).
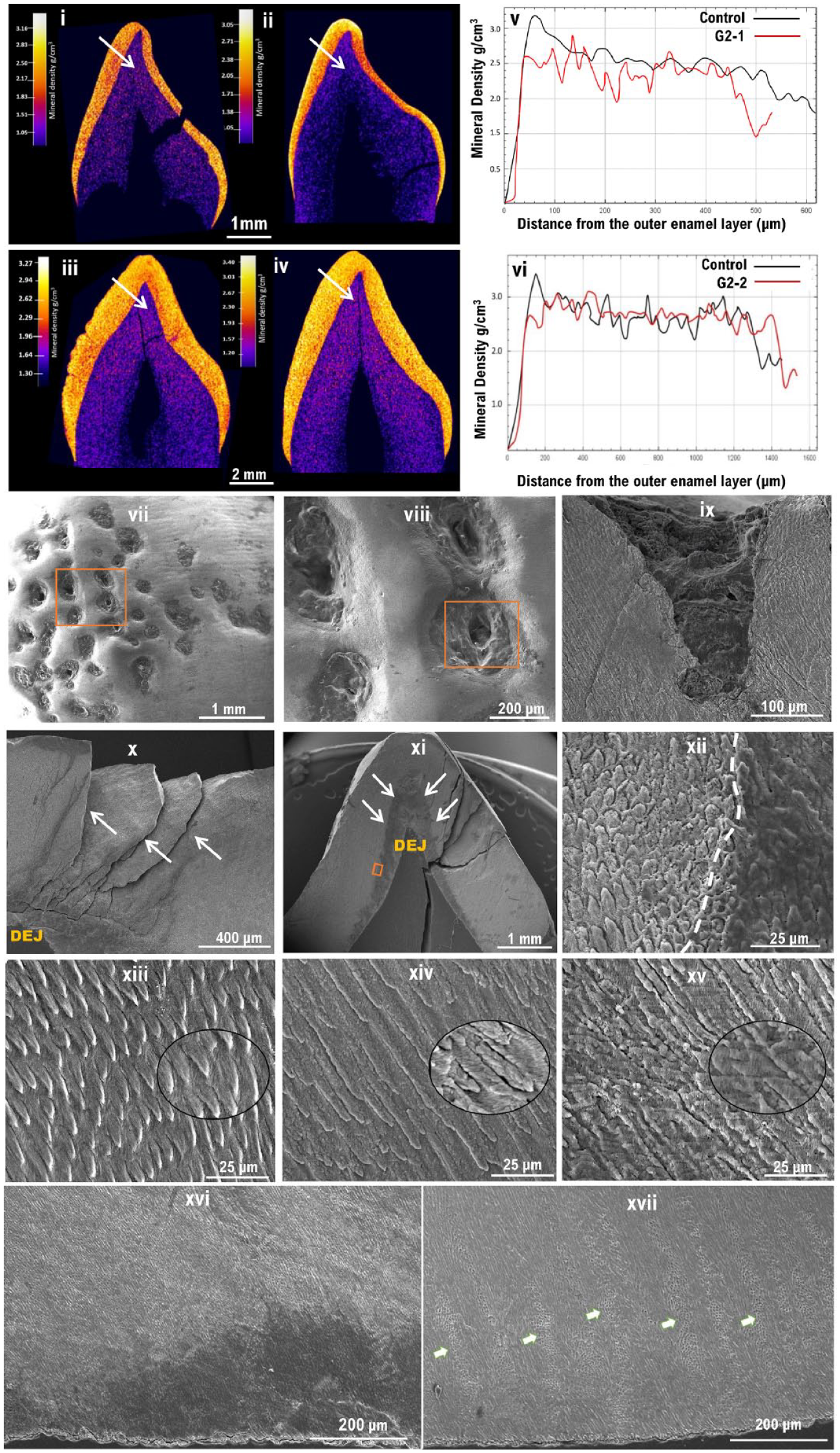
Phenotypic characterization of the teeth of group 2 families. (i–iv) Micro–computed tomography (µCT): False-colored calibrated heatmaps. (i) A primary upper lateral incisor from the affected individual from G2-1 and (iii) a permanent canine from G2-2. Teeth of the same types obtained from unrelated unaffected individuals were used as controls for comparison and are shown in (ii) and (iv), respectively. (i, iv) No significant differences in enamel volume was observed between affected and healthy controls, but the mineral density distribution was found to be disturbed. (v, vi) Mineral density line scans showing the distribution of enamel density form the enamel surface to the dentine–enamel junction (DEJ) in G2-1 and G2-2, respectively, as shown by the arrows in i–iv. In both controls, note the initial high peaks present at the surface and the gradual decrease of mineral density toward the DEJ, which are absent in the amelogenesis imperfecta–affected teeth confirming the disturbed mineral density observed in the µCT images. (iii) G2-2 tooth shows an uneven, pitted surface that is absent from the control tooth. (vii–xvii) Scanning electron microscope images: (vii–viii) Surface topography features in G2-2 showing numerous clear pitting of the labial enamel surface, extending through the enamel (ix) and to the DEJ (x). (xi) Labiolingual section of G2-2, a more disrupted inner enamel below the cuspal area is shown by the white arrows. (xii) Higher magnification of the inset in (xi) showing a clear demarcation (white dotted line) between a less dense enamel located near the DEJ and the rest of the enamel. (xiii–xv) Higher magnification of the enamel microstructure at ×1k, with ×3.5k insets to illustrate prismatic structure as well as crystallite orientation. (xiii) Healthy enamel. (xiv–xv) Affected teeth from the family G2-1 and G2-2, respectively. (xiv–xv) Note the poorly formed enamel rods that appear fused at many areas, making it difficult to distinguish the boundaries between rod and interrod regions. (xvi) Low magnification of G2-2 showing missing Hunter–Schreger banding as opposed to the healthy enamel (arrows) in (xvii).
Discussion
The data presented support
Six of the families described here have genotypes and inheritance patterns consistent with autosomal recessive hypoplastic AI with poor-quality enamel. Clinical images were consistent with rapid failure of a thin creamy mineralized tissue after eruption, leaving a predominantly yellow appearance. These group 1 families fit with previous reports of recessive AI due to biallelic pathogenic variants in
Intriguingly, group 2 included isolated AI cases heterozygous for
Group 3 includes 3 families with the same heterozygous variant, c.76G>A; p.(Ala26Thr), which is absent from gnomAD. Of these, G3-1 has a clear family history of dominantly inherited AI. However, only a single family member was recruited, so it was not possible to confirm cosegregation of the variant with AI. Observation of 2 other apparently unrelated patients with AI (G3-2 and G3-3) with the same variant on the same founder haplotype implies descent from a common ancestor, which is further evidence of cosegregation of this variant with AI as a dominantly inherited trait. No potentially pathogenic second
The spectrum of pathogenic variants in
For missense variants, the disease mechanism is less clear, but they may also be functional knockouts. The p.(Tyr99His) substitution changes an aromatic tyrosine to a basic histidine in the first of 15 highly conserved amino acid residues in the proline-rich region of AMBN. This region is retained in AMBN isoform I (ISOI) but removed in isoform II (ISOII) due to alternative splicing, and both isoforms are highly conserved and are coexpressed in vitro, suggesting they might perform different functions during enamel development (MacDougall et al. 2000; Vetyskova et al. 2020). Probands in families G1-5 and G1-6, who are compound heterozygotes for p.(Ser70*) and p.(Tyr99His), had thin poor-quality enamel, similar to families homozygous for p.(Ser70*). It is therefore likely that p.(Tyr99His) is also a functional knockout.
On clinical examination, group 1 enamel is yellow, thin, and of poor quality without normal microstructure (Poulter et al. 2014). By contrast, group 3 enamel associated with c.76G>A; p.(Ala26Thr) is thin on radiography, yet the white appearance of the teeth is consistent with little posteruption breakdown. Without laboratory analyses, it is unknown if this has a normal enamel microstructure. This substitution changes a nonpolar hydrophobic alanine (conserved in all mammals except the toothless platypus) at the C-terminal amino acid of the AMBN secretory signal peptide, immediately adjacent to the cleavage site, to a polar hydrophilic threonine (Delsuc et al. 2015). Proteins such as AMBN that are destined for the extracellular environment are transported to the endoplasmic reticulum (ER) under the direction of the signal peptide, which is cleaved from the protein before secretion. Many human diseases, including AI, are caused by ER stress, resulting from the misfolding of newly synthesized proteins as they are trafficked through the ER (Brookes et al. 2017; Morikawa and Urano 2022). It is therefore plausible that the apparently dominant hypoplastic AI phenotype associated with the p.(Ala26Thr) signal peptide variant arises by a different disease mechanism compared to null variants causing recessive AI. A dominant negative effect may lead to impairment of the normal ameloblast secretory pathway, ER stress, and ultimately ameloblast apoptosis.
Accordingly, these data contribute to the debate as to whether mutations in
These data provide new insight into how null
Author Contributions
U. Hany, C.M. Watson, C.F. Inglehearn, A.J. Mighell, contributed to conception, design, data acquisition, analysis, and interpretation, drafted and critically revised the manuscript; L. Liu, G. Nikolopoulos, C.J. Brown, A. Patel, H.D. Rodd, R. Balmer, A. Harfoush, M. Al-Jawad, contributed to data acquisition and interpretation, critically revised the manuscript; C.E.L. Smith, J.A. Poulter, contributed to conception, design, data acquisition and interpretation, critically revised the manuscript. All authors gave final approval and agree to be accountable for all aspects of the work.
Supplemental Material
sj-docx-1-jdr-10.1177_00220345231203694 – Supplemental material for Novel Ameloblastin Variants, Contrasting Amelogenesis Imperfecta Phenotypes
Supplemental material, sj-docx-1-jdr-10.1177_00220345231203694 for Novel Ameloblastin Variants, Contrasting Amelogenesis Imperfecta Phenotypes by U. Hany, C.M. Watson, L. Liu, G. Nikolopoulos, C.E.L. Smith, J.A. Poulter, C.J. Brown, A. Patel, H.D. Rodd, R. Balmer, A. Harfoush, M. Al-Jawad, C.F. Inglehearn and A.J. Mighell in Journal of Dental Research
Supplemental Material
sj-pptx-1-jdr-10.1177_00220345231203694 – Supplemental material for Novel Ameloblastin Variants, Contrasting Amelogenesis Imperfecta Phenotypes
Supplemental material, sj-pptx-1-jdr-10.1177_00220345231203694 for Novel Ameloblastin Variants, Contrasting Amelogenesis Imperfecta Phenotypes by U. Hany, C.M. Watson, L. Liu, G. Nikolopoulos, C.E.L. Smith, J.A. Poulter, C.J. Brown, A. Patel, H.D. Rodd, R. Balmer, A. Harfoush, M. Al-Jawad, C.F. Inglehearn and A.J. Mighell in Journal of Dental Research
Footnotes
Acknowledgements
The authors thank the families involved for their support for this study.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by Rosetrees Trust Grant PGS19-2/10111, Wellcome Trust Grant WT093113MA, and a Leeds Doctoral Scholarship awarded to U. Hany.
A supplemental appendix to this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
