Abstract
Dental enamel formation is coordinated by ameloblast differentiation, production of enamel matrix proteins, and crystal growth. The factors regulating ameloblast differentiation are not fully understood. Here we show that the high mobility group N (HMGN) nucleosomal binding proteins modulate the rate of ameloblast differentiation and enamel formation. We found that HMGN1 and HMGN2 proteins are downregulated during mouse ameloblast differentiation. Genetically altered mice lacking HMGN1 and HMGN2 proteins show faster ameloblast differentiation and a higher rate of enamel deposition in mice molars and incisors. In vitro differentiation of induced pluripotent stem cells to dental epithelium cells showed that HMGN proteins modulate the expression and chromatin accessibility of ameloblast-specific genes and affect the binding of transcription factors epiprofin and PITX2 to ameloblast-specific genes. Our results suggest that HMGN proteins regulate ameloblast differentiation and enamel mineralization by modulating lineage-specific chromatin accessibility and transcription factor binding to ameloblast regulatory sites.
Keywords
Introduction
As the hardest biological substance in the human body, dental enamel serves as the major protective layer of tooth crowns, insulating the dentin and pulp as well as being the site of most occlusal contacts during chewing. Dental enamel is formed from mature ameloblast cells during tooth development. The establishment of ameloblast cell identity is a stepwise process in which dental epithelium stem cells expressing the pluripotent marker SOX2 transdifferentiate into highly proliferative transit-amplifying cells (TACs), followed by terminal differentiation into functional ameloblasts that secrete the enamel matrix proteins amelogenin and ameloblastin (Sasaki et al. 2005; Juuri et al. 2012; Juuri et al. 2013; He et al. 2019).
The differentiation of stem cells into ameloblasts is regulated by multiple signaling cascades, transcription factors, microRNAs (miRNAs), and chromatin remodelers (Yoshizaki et al. 2020; Hermans et al. 2021). For example, miR-200 represses NOGGIN expression and modulates the BMP signaling pathway during ameloblast differentiation (Cao et al. 2013). Transcription factor AmeloD regulates early stage ameloblast migration, while transcription factor epiprofin regulates ameloblast terminal differentiation and transcription activation (Nakamura et al. 2008; Chiba et al. 2019; Miao et al. 2022). Additional transcription factors, including PITX2, LEF1, DLX3, and CIP2/BCL11b, also play significant roles during ameloblast differentiation (Amen et al. 2008; Golonzhka et al. 2009; Li et al. 2014; Duverger et al. 2017; Eliason et al. 2022). These studies greatly enhanced our knowledge of amelogenesis, yet the regulatory mechanisms that establish ameloblast-specific cell identity are still not fully understood.
Specific cell identity is established by lineage-specific transcription factors that bind to chromatin regulatory sites, thereby facilitating the recruitment of transactivation complexes, which leads to cell type–specific gene expression (Andersson and Sandelin 2020). Since cell type–specific regulatory sites are embedded into the 3-dimensional chromatin, the efficiency of transcription factors gaining access to regulatory sites plays a crucial role in gene transcription. The chromatin epigenetic landscape is known to play an important role in facilitating the binding of regulatory factors to regulatory sites. For example, transcriptionally active chromatin is enriched in H3K27ac-modified histones and DNase I hypersensitivity sites, 2 epigenetic marks known to be directly related to a higher level of gene transcription (Li et al. 2011; Arvey et al. 2012; Klemm et al. 2019). Therefore, deciphering the factors that modulate chromatin status and the efficiency of gene transcription is crucial to understanding how ameloblast cell identity is established.
Here we focus on the role of the high mobility group N (HMGN) proteins in ameloblast differentiation. HMGN are chromatin architectural proteins ubiquitously expressed in all vertebrate cells and are known to bind to chromatin without DNA sequence specificity. Previous studies revealed that HMGN proteins modulate chromatin accessibility, transcription factor binding, and promoter–enhancer interactions, and they stabilize cell identity (He et al. 2018; Garza-Manero et al. 2019; He et al. 2022; Zhang et al. 2022). We now use Hmgn1–/–; Hmgn2–/– double-knockout (DKO) mice to explore whether HMGN proteins regulate ameloblast differentiation. We found that HMGN1 and HMGN2 depletion accelerates ameloblast differentiation and dental enamel mineralization. By differentiation of wild-type (WT) and DKO induced pluripotent cells (iPSCs) into the ameloblast lineage, we found that HMGN proteins modulate chromatin accessibility, histone modification levels, and chromatin binding of lineage-specific transcription factor binding during ameloblast differentiation. Our study suggests that in mice, HMGN proteins act as epigenetic factors that modulate ameloblast differentiation and enamel mineralization. Since the HMGN amino acid sequence is highly conserved and HMGN proteins are expressed in all vertebrate cells, it is likely that these nucleosome binding proteins affect tooth development in other organisms than just mice.
Materials and Methods
Mice
All animal experiments complied with the protocols for animal use, treatment, and euthanasia approved by the National Cancer Institute (NCI) Animal Care and Use Committees (protocols: LMC-030 and LM-096). WT and DKO mice were from C57 BL/6 strain. Genotypes were verified by DNA sequencing and Western blot analysis. Procedures for generating Hmgn1–/–Hmgn2–/–DKO were as described (Deng et al. 2015).
Tissue Sections and Immunofluorescence Staining
Mouse embryonic mice mandibles were fixed in 4% Paraformaldehyde (PFA) overnight, followed by 15% and 30% sucrose treatment and embedding in Tissue-Tek O.C.T. Compound on dry ice. Postnatal mice mandibles were fixed in 4% PFA overnight, followed by embedding in O.C.T. Compound on dry ice. Procedures for immunofluorescence staining of mouse tooth frozen section were described in detail in our previous publications (He et al. 2019).
Data Availability
The RNA sequencing (RNA-seq) data in this article are available with the accession number GSE226942 (private token: kdijsuaobbgnzkp). The ATAC-seq and chromatin immunoprecipitation sequencing (ChIP-seq) data were downloaded from previously published data sets: PRJNA481982 and SRA: SRP154652.
Additional methods related to this study are in the Appendix.
Results
HMGN Proteins Regulate Ameloblast Differentiation
HMGN1 and HMGN2 are the 2 major variants of the HMGN protein family (Kugler et al. 2012). During ameloblast differentiation, HMGN1 and HMGN2 expression levels are slightly downregulated, as evidenced by immunofluorescence staining of mouse molars at various developmental stages, showing that HMGN1 and HMGN2 proteins are highly expressed at embryonic days 13.5 (E13.5) and E17.5 and gradually downregulated at postnatal day 1 (P1) and P3, while the enamel matrix protein amelogenin starts to express at P3 (Fig. 1A–C). Similarly, immunofluorescence staining of P3 mouse incisor sagittal sections showed higher HMGN protein expression in the cervical loop region and relatively lower HMGN protein expression in presecretory ameloblasts (Appendix Fig. 1A, B). In addition, transcription analysis of molar dental epithelium cells at developmental stages ranging from E15.5 to P7 revealed that Hmgn1 and Hmgn2 expression showed a reverse tendency with the expression of ameloblast differentiation markers Amelx and Ambn (Fig. 1D). These results suggest that HMGN expression is downregulated during ameloblast differentiation.
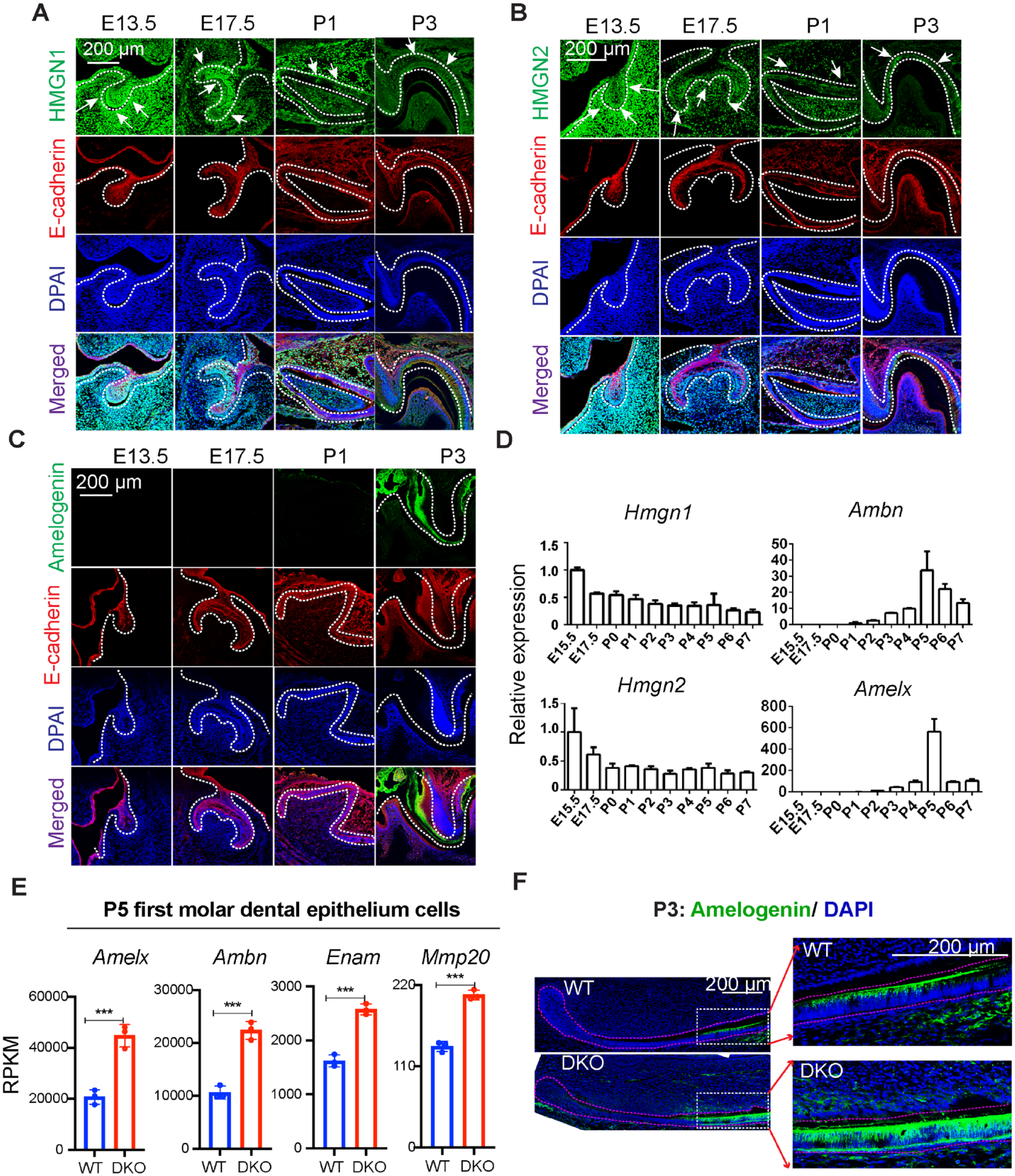
High mobility group N (HMGN) proteins regulate ameloblast differentiation. (
To study the role of HMGN proteins during ameloblast differentiation, we used Hmgn1–/–; Hmgn2–/– (DKO) mice as a model (Deng et al. 2015). We first analyzed enamel matrix gene expression in the first molar dental epithelium cells at P5, which is a developmental stage that Amelx and Ambn reach their peak levels, as indicated by our quantitative polymerase chain reaction (qPCR) data (Fig. 1D). RNA-seq analysis of P5 first molar dental epithelium cells showed significantly higher levels of Amelx, Ambn, Enam, and Mmp20 expression in the DKO mice as compared to the WT mice (Fig. 1E). Immunofluorescence staining of enamel matrix proteins amelogenin and ameloblastin in mice incisor and molar sections showed that indeed, the DKO group has a higher level of amelogenin and ameloblastin expression and a much thicker layer of amelogenin and ameloblastin protein deposition nearby the secretory ameloblast (Fig. 1F and Appendix Fig. 1C–E). Together, these results suggest that loss of HMGN proteins accelerate ameloblast differentiation.
Enhanced rate of enamel mineralization in mice lacking HMGNs
Enamel matrix proteins play a crucial role in the proper assembly and growth of crystals during enamel mineralization (Fukumoto et al. 2014). The different levels of enamel matrix protein expression between the WT and DKO group prompted us to investigate whether HMGN proteins regulate dental enamel mineralization. We analyzed enamel mineralization levels in P10, P18, and adult mouse (2-mo-old) mandibles using micro–computed tomography (µCT). During normal development, at P10, mouse first molars and incisors are in the mineralization process, while the second and third molars are largely unmineralized and not yet erupted. Our µCT analysis showed that loss of HMGN proteins resulted in significantly higher mineralized enamel volume in the first molar and incisor at P10 (red in Fig. 2A, B, Appendix Fig. 2A). At P18, the third molar and incisor are in the mineralization process, and the first and second molar enamel formation is complete. Our µCT results showed the DKO group had significantly higher mineralized enamel volume in the third molar and incisor compared to the WT group, and there were no significant differences in mineralized enamel volume in the first molar or second molar between the WT and DKO groups (Fig. 2C, D, Appendix Fig. 2B). When mice reached adulthood, the WT and DKO groups did not show any differences in total mineralized enamel volume of first molar, second molar, third molar, or incisor (Fig. 2E, F, Appendix Fig. 2C). These results suggest that loss of HMGN proteins accelerates the rate of enamel mineralization but does not affect the final mineralization levels in adult mice.
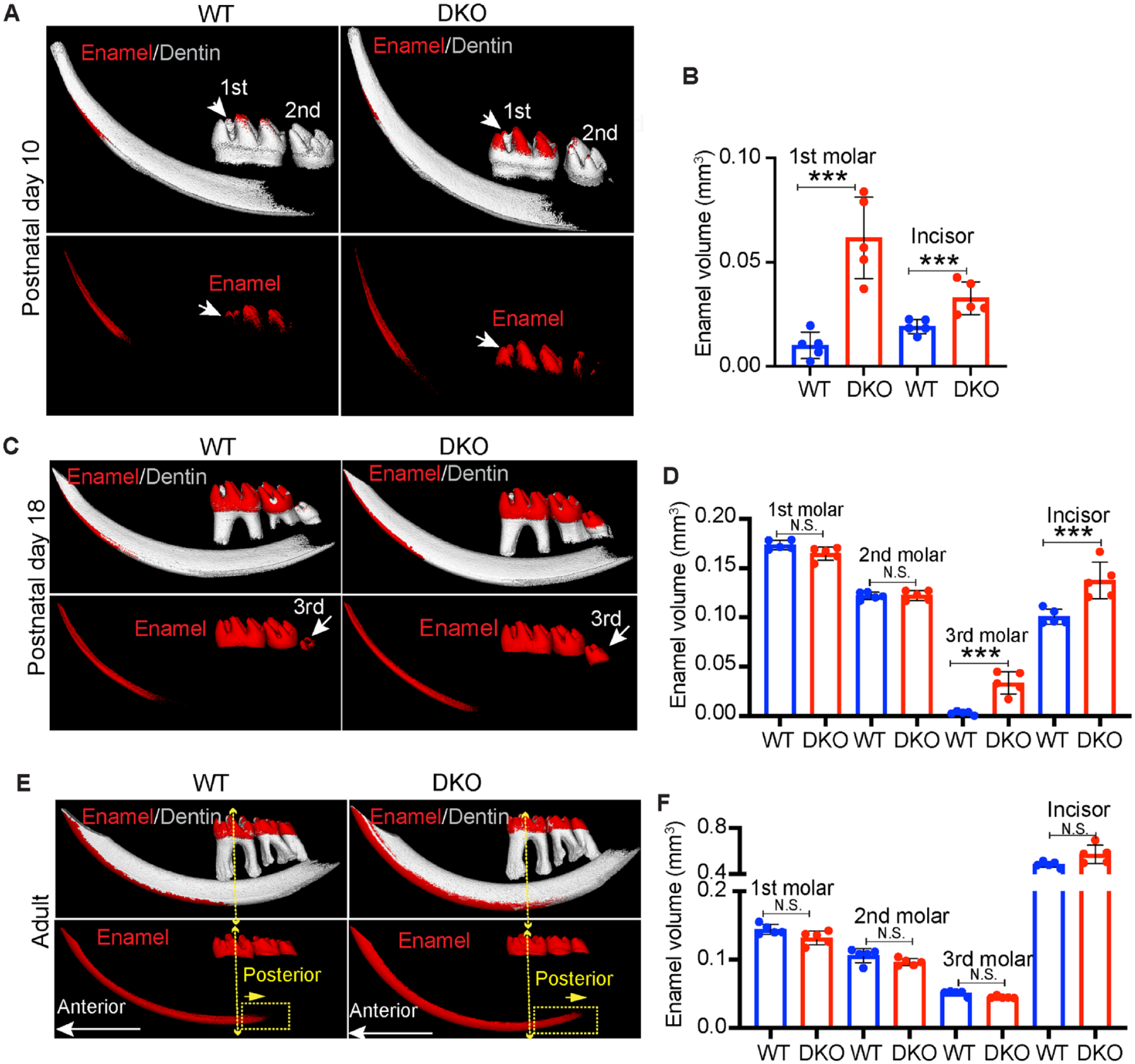
Enhanced rate of enamel mineralization in mice lacking high mobility group N (HMGN). (
We also detected differences between the WT and DKO mice in the mineralization patterns of adult incisors. The transition between immature and mature enamel extended more posterior in the DKO group, indicated by the dotted yellow rectangle (Fig. 2E, Appendix Fig. 2C). This observation was further confirmed by reconstructed µCT images of incisor coronal sections (Appendix Fig. 2D), showing a larger area of mineralized dental enamel in the DKO group. We also sectioned the mouse mandible vertically near the mesial root canal of the first molar, followed by scanning electron microscope (SEM) analysis. Our results showed that the coronal section of the DKO incisor has higher mineralized levels than in the WT incisor, as indicated by the brighter signal (Appendix Fig. 2E, upper panel); however, magnified SEM images showed that loss of HMGN proteins does not change the overall enamel rod structure and arrangement pattern (Appendix Fig. 2E, lower panel).
In summary, HMGN proteins regulate the rate of dental enamel mineralization without affecting the mineralization levels of adult, fully developed teeth.
HMGN Proteins Regulate Transcription Dynamics during Mouse Ameloblast Differentiation
During mouse tooth development, E17 is a transition stage between dental epithelium stem cells and preameloblast cells, P0 is a transition stage between preameloblasts and secretory ameloblasts, and at P5, ameloblasts start to mature (Caton and Tucker 2009). To study whether HMGN proteins modulate gene expression dynamics during ameloblast differentiation, we performed RNA-seq analyses of dental epithelium cells prepared from the first molars of E17, P0, and P5 WT and DKO mice. IGV snapshots at the Hmgn1 and Hmgn2 loci in the genome verified the genotype of WT and DKO cells (Appendix Fig. 3A–C). Pearson correlation analysis among replicates showed high R2 values, indicating high replicate reproducibility and high quality of the RNA-seq libraries. Comparison of averaged WT to averaged DKO transcription profile showed a broader scatterplot with lower R2 values, suggesting that loss of HMGN proteins alters the overall gene expression levels (Appendix Fig. 4A–C). Volcano plot comparison between the WT and DKO transcripts indicates that loss of HMGN proteins leads to a large number of differentially expressed genes (DEGs) at each developmental stage: 299 genes upregulated and 91 genes downregulated at E17, 484 genes upregulated and 290 genes downregulated at P0, and 194 genes upregulated and 783 genes downregulated at P5 (Fig. 3A, C, E).
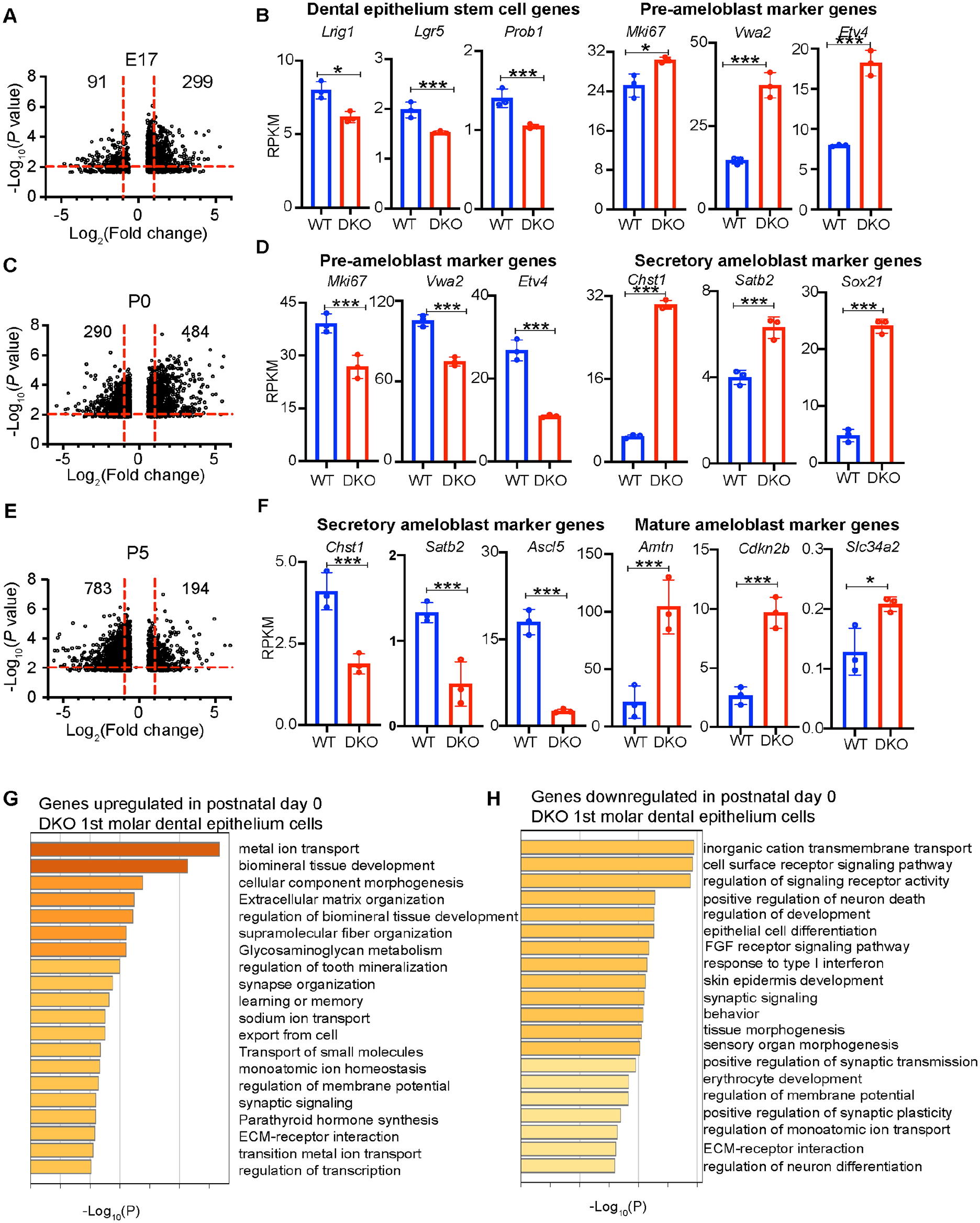
High mobility group N (HMGN) proteins regulate transcription dynamics during mouse ameloblast differentiation. (
Examination of the expression levels of select genes that mark unique ameloblast developmental stages (Krivanek et al. 2020) provided further support that loss of HMGNs leads to acceleration of ameloblast differentiation. Thus, at E17, loss of HMGN proteins led to decreased expression of dental epithelium stem cell marker genes but increased expression of preameloblast marker genes, including marker genes of both the highly proliferative transit-amplifying (Sfrp5, Mki67) and the preameloblast (Vwde) populations (Fig. 3B, Appendix Fig. 5A), suggesting that in the DKO group, a higher fraction of dental epithelium stem cells are committed to preameloblasts. At P0, loss of HMGNs led to decreased expression levels of preameloblast marker genes and increased expression of secretory ameloblast marker genes (Fig. 3D and Appendix Fig. 5B, C), suggesting that a larger fraction of the ameloblast cells transited into the secretory stage in the DKO group. At P5, loss of HMGN proteins led to decreased expression levels of the secretory ameloblast marker genes but increased expression of Amtn, Cdkn2b, and Slc34a2, which are marker genes for ameloblast maturation (Krivanek et al. 2020) (Fig. 3F), suggesting a higher efficiency of ameloblast maturation in the DKO group. The differentially expressed genes at the 3 developmental stages are enriched in the developmental process, growth, and other signaling pathways that are important for tooth development. For example, at P0, genes upregulated in the DKO group are enriched in categories of biomineral tissue development, extracellular matrix organization, and metal ion transport, while the downregulated genes are enriched in categories involved in cell–cell surface signaling pathways and epithelial cell differentiation (Fig. 3G, H, Appendix Fig. 6). In summary, the RNA-seq data analysis indicates that HMGN protein depletion enhances ameloblast differentiation and maturation.
HMGN Depletion Enhances the Conversion Efficiency of iPSCs into Dental Epithelium Cells
To gain insights into the mechanisms whereby HMGN proteins modulate ameloblast differentiation, we used WT and DKO iPSCs as in vitro models. The WT and DKO iPSCs colonies are morphologically indistinguishable and show comparable levels of SOX2, SSEA1, OCT4, and NANOG immunofluorescence staining signal (Fig. 4A, B) and similar expression levels of pluripotent marker genes (Fig. 4C). These results further confirm and agree without previous studies showing that HMGN depletion does not compromise the iPSCs’ ground state pluripotency (He et al. 2018).
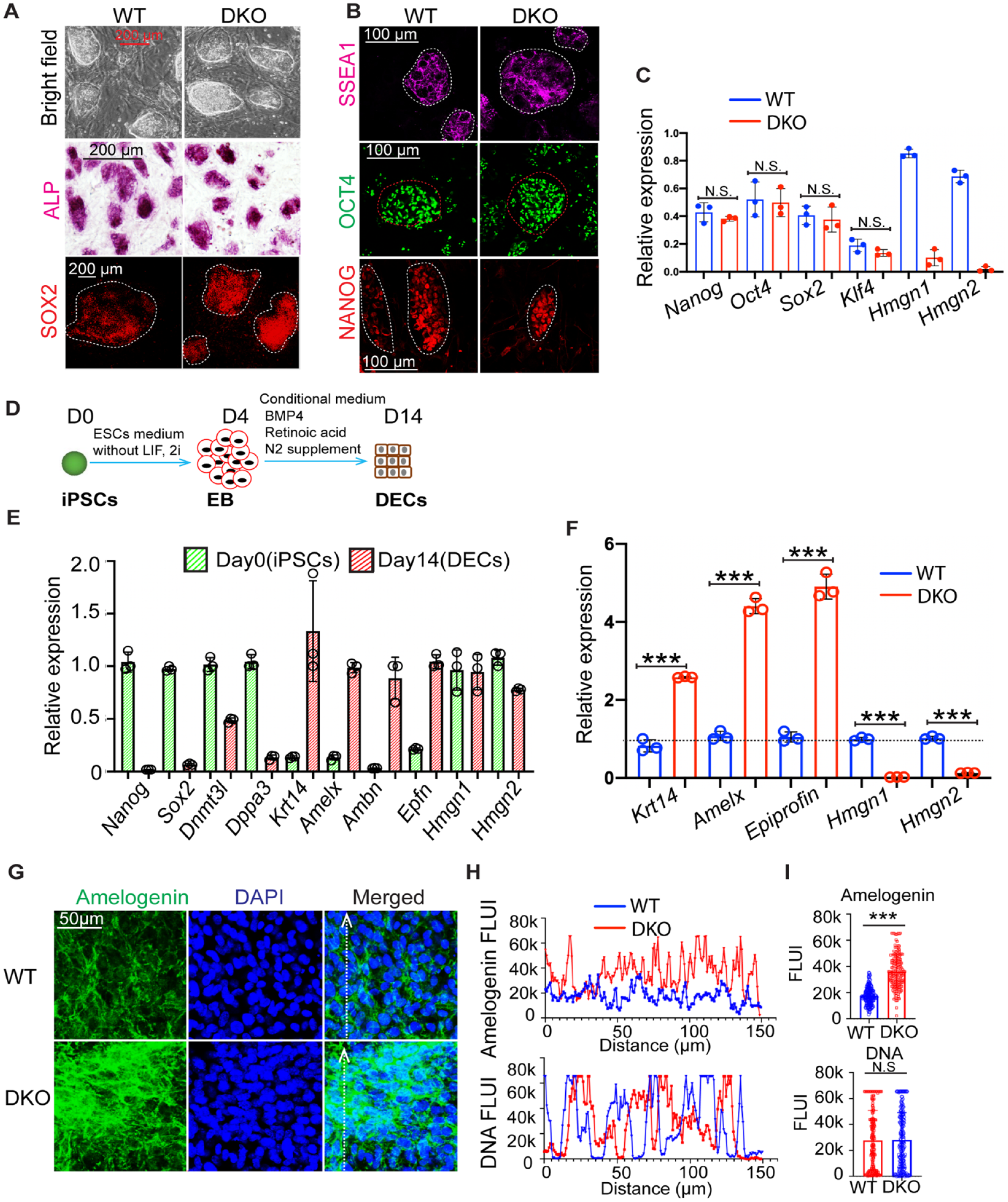
High mobility group N (HMGN) depletion enhances the conversion efficiency of induced pluripotent stem cells (iPSCs) into dental epithelium cells (DECs). (
We differentiated WT and DKO iPSCs into dental epithelium cells (DECs) (Fig. 4D) by following an established protocol (Kim et al. 2019). The genotype of WT and DKO iPSCs and of DECs was confirmed by immunofluorescence staining of HMGN proteins (Appendix Fig. 7A, B). The DECs derived from iPSCs have high levels of ameloblast differentiation marker gene expression, including Krt14, Ambn, Amelx, and Epfn, and very low levels of pluripotent markers such as, Sox2, Dnmt3l, and Dppa3, suggesting a successful transition from iPSCs to ameloblast lineage (Fig. 4E). Interestingly, gene expression analysis of WT and DKO DECs revealed that the DKO group had significantly higher levels of ameloblast differentiation marker gene expression, such as Krt14, Amelx, and Epfn (Fig. 4F). Likewise, immunofluorescence staining (Fig. 4G) and quantitative analysis of fluorescence intensity (Fig. 4H, I) showed significantly higher amelogenin protein expression in DKO DECs. Together, these results indicate that loss of HMGNs enhances the differentiation efficiency of iPSCs to DECs.
To further verify the roles of HMGN proteins in dental epithelium cell differentiation, we transfected Cervical Loop Derived Dental Epithelial (CLDE) cells (Yoshizaki et al. 2014) with plasmids expressing the transcription factor epiprofin together with plasmids expressing either Hmgn1/2 DNA or Hmgn1/2 small interfering RNAs (siRNAs). qPCR analyses showed that Hmgn1/2 overexpression significantly decreased the epiprofin-induced Amelx gene expression level while transfection with Hmgn siRNAs enhanced the Amelx gene expression level (Appendix Fig. 7C). These results support the notion that depletion of HMGN protein enhances ameloblast differentiation.
Epigenetic Remodeling during Differentiationof iPSCs into DECs
HMGNs are epigenetic regulators that affect gene expression by binding to chromatin regulatory sites (Deng et al. 2015). To gain insight into the epigenetic role of HMGNs during iPSC differentiation into DECs, we queried our ChIP-seq data of HMGN proteins and of several histone modifications at iPSC-specific and ameloblast-specific genomic loci (He et al. 2018). We found that in iPSCs, HMGN proteins are preferentially enriched at iPSC-specific sites such as Nanog, Dppa3, and Dnmt3l and colocalize with H3K27ac, H3K4me1, and H3K4me3, epigenetic markers of active chromatin, but not with H3K9me3, a marker of inactive chromatin (Fig. 5A and Appendix Fig. 8, left panel). In contrast, iPSCs do not show HMGN enrichment or histone marks of active chromatin at ameloblast-specific loci such as Amelx, Ambn, and Amtn (Fig. 5B and Appendix Fig. 8, right panel). ChIP-qPCR verified HMGN protein binding and the presence of H3K27ac modifications at the Nanog locus in the iPSCs (Fig. 5C) and at the Amelx locus in DECs (Fig. 5D). Thus, HMGN protein occupancy displays a cell type–specific pattern: in iPSCs, HMGN occupancy levels are relatively high at the Nanog locus and low at the Amelx locus, while in DECs, HMGN occupancy is relatively low at the Nanog locus but high at the Amelx locus (Fig. 5C, D).
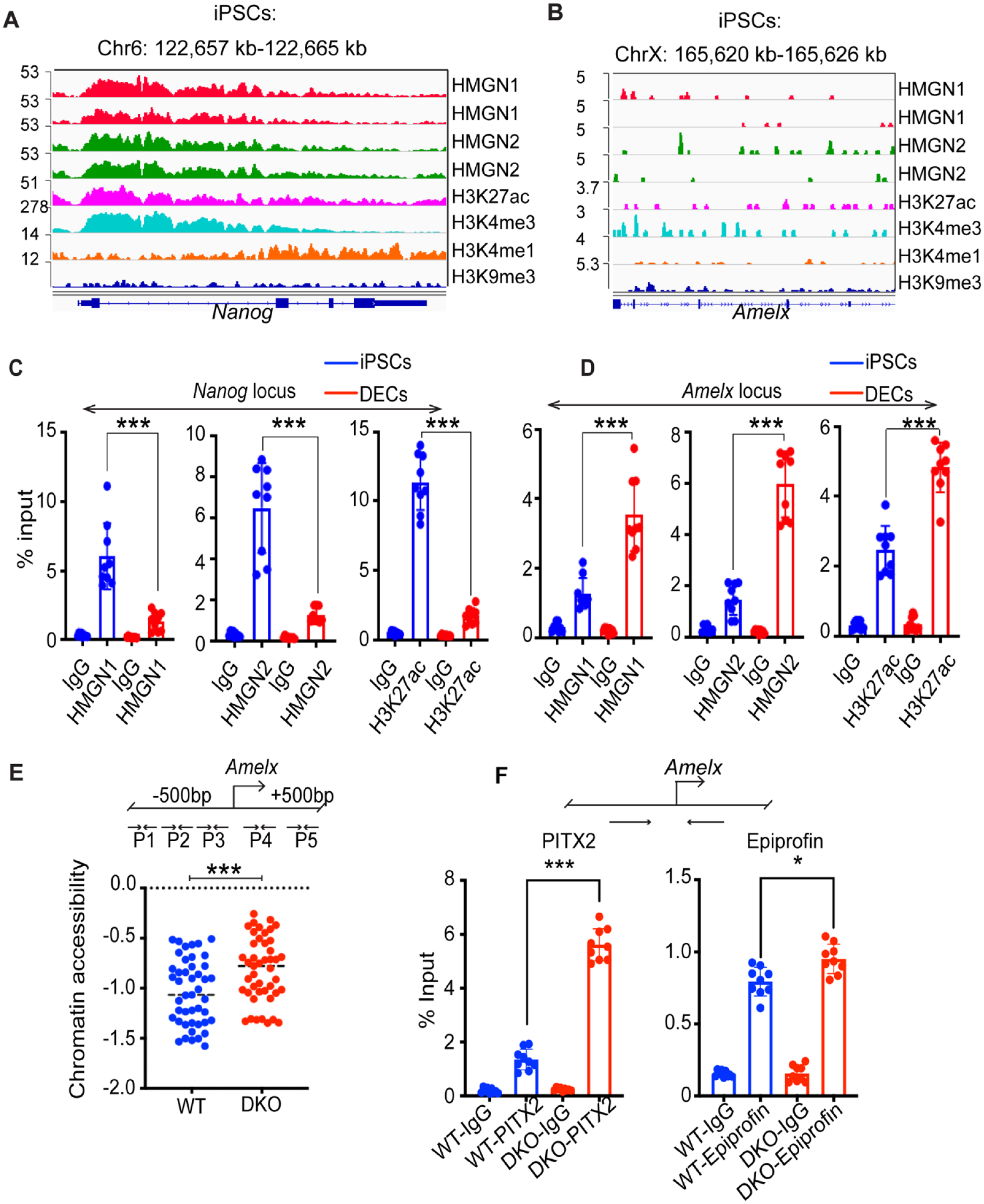
Epigenetic remodeling during differentiation of wild-type (WT) and double-knockout (DKO) induced pluripotent stem cells (iPSCs) into dental epithelium cells (DECs). (
Chromatin accessibility is a major epigenetic mark of active chromatin (Klemm et al. 2019). We used Tn5-mediated chromatin tagmentation (Buenrostro et al. 2015) followed by a qPCR assay of chromatin accessibility at a gene-specific locus to compare the chromatin accessibility at the Amelx locus in the WT and DKO DECs. Using 5 pairs of primers, which cover the ±500 bp of the Amelx transcription start site, we find that the Amelx locus in DKO DECs is significantly more accessible than that in WT DECs, suggesting more efficient differentiation in the DKO cultures (Fig. 5E).
The transcription factors PITX2 and epiprofin are important regulators of ameloblast differentiation (Nakamura et al. 2008; Li et al. 2014). By ChIP-qPCR analysis, we now find that the occupancy levels of PITX2 and epiprofin at the Amelx promoter are significantly higher in DKO DECs (Fig. 5F). Thus, our chromatin accessibility assay and ChIP-qPCR results suggest that HMGN depletion in iPSCs leads to increased chromatin accessibility and transcription factor binding at ameloblast-specific sites, as well as to elevated DEC-specific gene expression. Together, these changes enhance the efficiency of differentiation of iPSCs into DECs.
Discussion
Mouse tooth development is initiated by the reciprocal interactions between dental epithelium cells and the underlying mesenchymal cells (Thesleff 2003). Elucidating the molecular mechanisms that regulate embryonic mouse tooth development could provide valuable insights into mammalian odontogenesis and organogenesis. We found that loss of HMGN enhances ameloblast differentiation and dental enamel mineralization, a finding that agrees with previous results showing that overexpression of HMGN2 protein in the dental epithelium cells decreases ameloblast differentiation (Li et al. 2014). By analyzing the phenotype of HMGN1/2 double mutant mice and the in vitro differentiation of iPSCs derived from WT and DKO mice, we now find that HMGN proteins regulate ameloblast differentiation and enamel formation by modulating chromatin accessibility and transcription factor binding at lineage-specific sites.
Our in vitro differentiation analyses also showed that HMGN proteins relocate from iPSC-specific regulatory sites to ameloblast-specific sites during iPSC differentiation. The relocation of HMGNs is accompanied by the deposition of active histone marks such as H3K27ac at ameloblast regulatory sites (Fig. 4). These studies indicate that HMGNs affect the epigenetic chromatin remodeling during iPSC differentiation into DECs.
Cell fate conversion involves an interplay between regulatory factors that maintain the original cell identity and the regulatory factors that establish the new cell identity. At the epigenetic level, this process involves the erasing of the existing epigenetic memory followed by the establishment of a new cell type–specific epigenetic signature (Nashun et al. 2015). HMGN proteins have been shown to affect the rate of cell fate conversions in several biological systems (Furusawa et al. 2006; Zhang et al. 2016; Deng et al. 2017; He et al. 2018). Most likely, HMGNs modulate cellular differentiation by preferentially binding to nucleosomes containing histone marks of active chromatin such as H3K27ac and H3K4me3, thereby modulating the binding of transcription factors to regulatory chromatin (Amen et al. 2008; Eliason et al. 2022; Zhang et al. 2022). We postulate that the presence of HMGN at cell type–specific regulatory chromatin sites stabilizes the cell identity (He et al. 2018; Garza-Manero et al. 2019). Consequently, HMGN-depleted cells, devoid of the “stabilizing effect” exerted by HMGNs, display higher efficiency of cell fate conversion, a possibility that agrees with our present finding indicating that HMGN-depleted cells show a higher efficiency of ameloblast differentiation. Thus, our results showing that HMGNs modulate ameloblast differentiation provide further support for the possibility that the ubiquitous HMGN proteins fine-tune the rate of cell fate conversion, a process that may be of particular importance at early stages of embryogenesis and development.
Author Contributions
B. He, contributed to conception, design, data acquisition, drafted and critically revised the manuscript; V. Kram, O. Duverger, R. Nanduri, contributed to data acquisition and analysis, critically revised the manuscript; T. Furusawa, E.Y. Chu, M. Ishikawa, contributed to data acquisition, critically revised the manuscript; P. Zhang, J.S. Lee, contributed to data analysis, critically revised the manuscript; B.A. Amendt, contributed to conception, data interpretation, critically revised the manuscript; M. Bustin, contributed to conception, design, data analysis and interpretation, drafted and critically revised the manuscript. All authors gave final approval and agree to be accountable for all aspects of the work.
Supplemental Material
sj-docx-1-jdr-10.1177_00220345231202468 – Supplemental material for Epigenetic Regulation of Ameloblast Differentiation by HMGN Proteins
Supplemental material, sj-docx-1-jdr-10.1177_00220345231202468 for Epigenetic Regulation of Ameloblast Differentiation by HMGN Proteins by B. He, V. Kram, T. Furusawa, O. Duverger, E.Y. Chu, R. Nanduri, M. Ishikawa, P. Zhang, B.A. Amendt, J.S. Lee and M. Bustin in Journal of Dental Research
Footnotes
Acknowledgements
We thank Dr. Alexei Lobanov, Center for Cancer Research (CCR) Collaborative Bioinformatics Resource, CCR, NCI, National Institutes of Health (NIH), for help with data deposition; Dr. Marian Young (National Institute of Dental and Craniofacial Research [NIDCR]/NIH) and Dr. Pamela Robey (NIDCR/NIH) for their help with µCT analysis; and Dr. Yoshihiko Yamada NIDCR/NIH (deceased) for significant contribution to conceptual aspects of this manuscript.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Center for Cancer Research, Intramural Research Program of the National Cancer Institute of the National Institutes of Health.
A supplemental appendix to this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
