Abstract
A wide range of deep learning (DL) architectures with varying depths are available, with developers usually choosing one or a few of them for their specific task in a nonsystematic way. Benchmarking (i.e., the systematic comparison of state-of-the art architectures on a specific task) may provide guidance in the model development process and may allow developers to make better decisions. However, comprehensive benchmarking has not been performed in dentistry yet. We aimed to benchmark a range of architecture designs for 1 specific, exemplary case: tooth structure segmentation on dental bitewing radiographs. We built 72 models for tooth structure (enamel, dentin, pulp, fillings, crowns) segmentation by combining 6 different DL network architectures (U-Net, U-Net++, Feature Pyramid Networks, LinkNet, Pyramid Scene Parsing Network, Mask Attention Network) with 12 encoders from 3 different encoder families (ResNet, VGG, DenseNet) of varying depth (e.g., VGG13, VGG16, VGG19). On each model design, 3 initialization strategies (ImageNet, CheXpert, random initialization) were applied, resulting overall into 216 trained models, which were trained up to 200 epochs with the Adam optimizer (learning rate = 0.0001) and a batch size of 32. Our data set consisted of 1,625 human-annotated dental bitewing radiographs. We used a 5-fold cross-validation scheme and quantified model performances primarily by the F1-score. Initialization with ImageNet or CheXpert weights significantly outperformed random initialization (P < 0.05). Deeper and more complex models did not necessarily perform better than less complex alternatives. VGG-based models were more robust across model configurations, while more complex models (e.g., from the ResNet family) achieved peak performances. In conclusion, initializing models with pretrained weights may be recommended when training models for dental radiographic analysis. Less complex model architectures may be competitive alternatives if computational resources and training time are restricting factors. Models developed and found superior on nondental data sets may not show this behavior for dental domain-specific tasks.
Keywords
Introduction
Deep learning (DL) has been widely employed for image analytics in dermatology (skin photographs) (Jafari et al. 2016), ophthalmology (retina imagery) (Son et al. 2020), or pathology (histological specimens) (Kather et al. 2019). Also in dentistry, DL classification models have been employed to predict the modality of radiographs (Cejudo et al. 2021), the presence of caries lesions (Lee et al. 2018), periodontal bone loss (Krois et al. 2019), and apical lesions (Ekert et al. 2019) on dental radiographs. DL segmentation models, which perform a classification task at the pixel level, were used for the segmentation of anatomical structures in panoramic images (Cha et al. 2021), apical lesions on cone beam computed tomography scans (Orhan et al. 2020), periodontal bone loss on panoramic radiographs (Kim et al. 2019), and caries lesions on bitewings (Cantu et al. 2020).
Recent guidelines in the field call for rigorous and comprehensive planning, conducting, and reporting of DL studies in dentistry (Schwendicke et al. 2021). One key element in those guidelines is a hypothesis-driven selection of the DL model configuration, which includes, among others, its architecture, its complexity, and the initialization strategy for the model weights (e.g., via transfer learning). (1)
The sheer number of possible configurations of model architecture, including backbones, complexity, and initialization strategies, impedes systematic and comprehensive comparisons of existing study findings (Schwendicke et al. 2019). One strategy to overcome this issue is to perform benchmarking, which involves the systematic comparison of different model architectures and model configurations on an identical data set. Such benchmarking studies provide guidance for researchers in the model design process, which improves research efficiency by enabling the development of high-performing models in a shorter time at lower development costs. However, in the medical domain and, more so, dentistry, benchmarking initiatives are scarce, owing to limited data availability and high costs for establishing solid and accepted ground truth labels and annotations. To cope with these difficulties, the ITU/WHO Focus Group Artificial Intelligence for Health (FG-AI4H) is developing a standard evaluation process and benchmarking framework for artificial intelligence (AI) models in health. The present study will inform this initiative.
In a recent benchmarking study, Bressem et al. (2020) benchmarked 16 different model architectures for classification tasks on 2 openly available chest radiograph data sets: CheXpert (Irvin et al. 2019) and the COVID-19 Image Data Collection. They showed that complex and deep models do not necessary outperform simpler architectures. Similarly, Ke et al. (2021) addressed the assumption that model architectures that perform better on the ImageNet data set (Deng et al. 2009), a popular open-source benchmark data set containing millions of labeled images, also generally perform better on CheXpert. This assumption was not found to be valid based on the comparison of 16 convolutional architectures on 5 classification tasks.
In the present study, we aim to expand the studies of Bressem et al. (2020) and Ke et al. (2021) to a dental segmentation task. We benchmarked 216 DL models defined by their architecture, complexity, and initialization strategy. We evaluated these model configurations for a specific dental task: tooth structure (enamel, dentin, pulpal cavity, fillings, and crowns) segmentation on dental bitewing radiographs. We deliberately decided to use this application since first, there is evidence that segmentation models perform well on this task (Ronneberger et al. 2015a) and, second, there is less ambiguity about the establishment of the ground truth for this task, with tooth structures being easily discriminated even by nonsenior clinicians. We expect our results to inform dental researchers about suitable model configurations for their experiments and aim to contribute to evidence-guided DL model selection in dental research.
Materials and Methods
Benchmarking Tasks
This analysis is based on a segmentation task for tooth structures on dental bitewing radiographs. Several model development aspects were benchmarked. (1)
Ethics Statement
This study was ethically approved by the ethics committee of the Charité (EA4/102/14 and EA4/080/18).
Study Design
In the present study, 72 models were built from a combination of varying architectures and encoder backbones and were each trained with 3 different initialization strategies on a tooth structure segmentation task. Each model was trained with 5-fold cross-validation with varying train, validation, and test sets for each fold. Hence, for each model run, the data were randomly split into training, validation, and test data with proportions of 60% (3 folds), 20% (1 fold), and 20% (1 fold), respectively. We additionally applied a sensitivity analysis and assessed model performances on underrepresented classes (in our case, fillings and crowns), as in real life, medical data set class imbalance is likely the rule and not the exception. Reporting of this study follows the Standards for Reporting Diagnostic Accuracy guideline (STARD) (Bossuyt et al. 2015) and the Checklist for Artificial Intelligence in Dental Research (Schwendicke et al. 2021).
Performance Metrics
Model performances were primarily quantified by the F1-score, which captures the harmonic mean of recall (specificity) and precision (positive predictive value [PPV]). F1-scores are computed from the sum of true positives, false positives, and false negatives over all channels of segmentation masks and cross-validation folds. This method was described by Forman and Scholz (2010) and results in unbiased F-scores in cross-validation schemes. Secondary metrics were accuracy, sensitivity, precision, and intersection of union (IoU). Based on the distribution of the results, the median was chosen as a descriptive statistic.
Data Set, Sample Size, and Reference Test
The available data set consisted of 1,625 dental bitewing radiographs with a maximum of 8 to 9 teeth per image and is described in detail in the Appendix. Tooth structures visible on bitewing radiographs (namely, enamel, dentin, the pulp cavity, and nonnatural “structures” like fillings and crowns) were annotated in a pixel-wise fashion (as masks) by 1 dental expert. These masks represent the ground truth for each data sample. In a second iteration, those annotations were reviewed by another dental expert for validity and correctness. Each annotator independently assessed each image using an in-house custom-built annotation tool described in Ekert et al. (2019). All examiners were calibrated and advised on how to perform the segmentation. Images with implants, bridges, or root canal fillings were very rare (<1%) and therefore excluded.
Notably, enamel, dentin, and pulpal areas were present in every radiograph, while fillings and crowns were only available in 80% and 20% of images, respectively. Images and segmentation masks were resized to a resolution of 224 × 224 to provide a fixed input size of the images as required by the model architectures.
Models and Training
As represented in Figure 1, models were built by combining different model architectures (U-Net, U-Net++, FPN, LinkNet, PSPNet, MAnet) with backbones from 3 different families (ResNet, VGG, DenseNet) of different depths (ResNet18, ResNet34, ResNet50, ResNet101, ResNet152, VGG13, VGG16, VGG19, DenseNet121, DenseNet161, DenseNet169, DenseNet201). This led to a total of 72 model designs, which were each initialized with 3 different strategies (random, ImageNet, CheXpert), resulting into 216 trained models in total. All models were trained under a 5-fold cross-validation scheme, where the combination of samples in training, validation, and test set was varied for each fold to achieve a reasonable estimate of the model performance independent from the data split. Details on training are described in the Appendix.
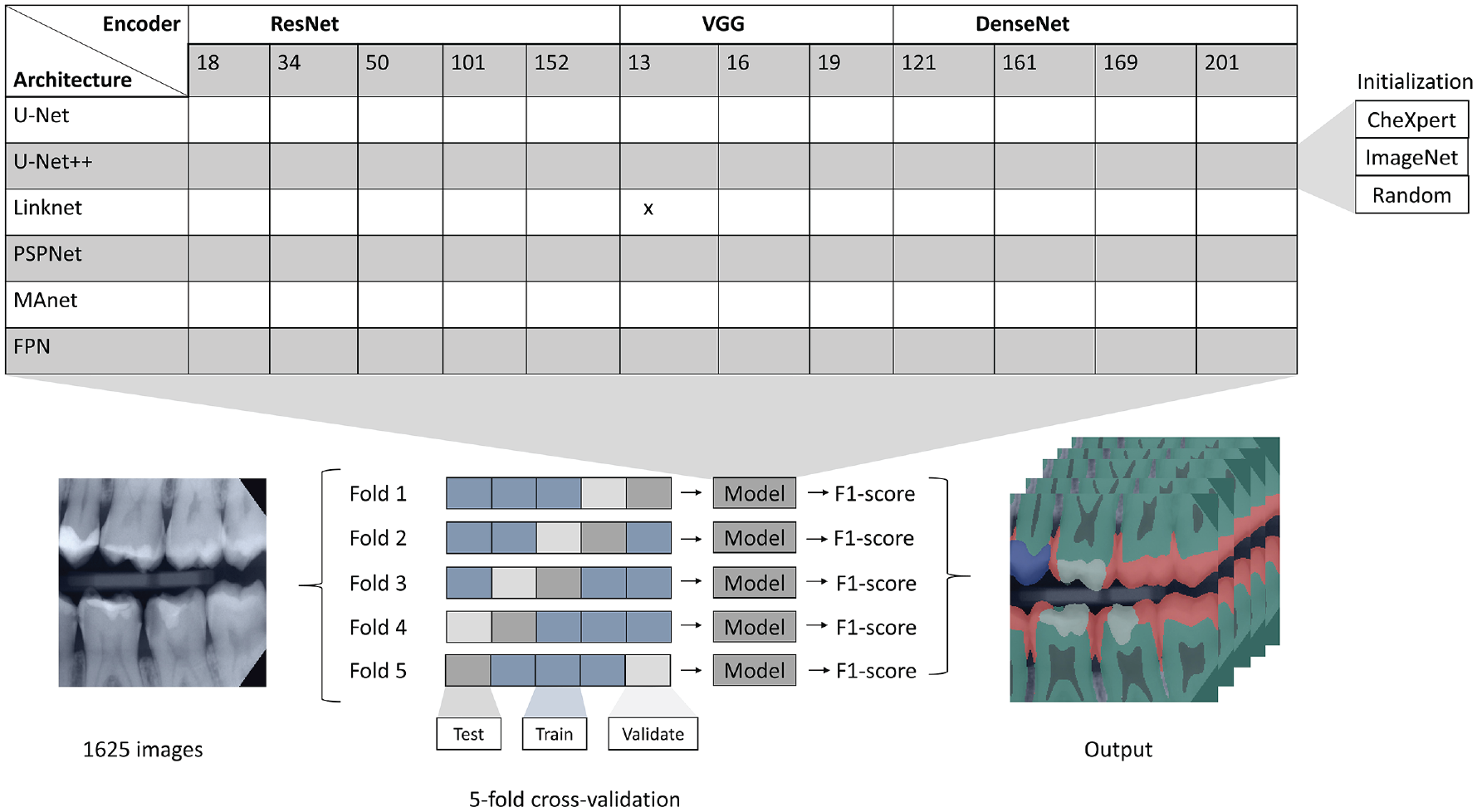
Illustration of the study design. Model setups were based on different architectures, encoder backbones, and initialization strategies (top) and 5-fold cross-validation with varying train, validation, and test sets for each fold (bottom). Exemplary bitewing radiograph (left) and tooth structure components overlaid on an input image (right).
Statistical Analysis
Model configurations with respect to initialization strategies and architectures were ranked according to their median F1-score and formally tested for differences between configurations with the nonparametric Wilcoxon rank-sum test. The nonparametric Spearman’s rank-order correlation was estimated to determine the relationship between complexity and model performance (F1-score). To account for multiple comparisons, we adjusted the P values using the Benjamini–Hochberg method (Benjamini and Hochberg 1995). P values below 0.05 were considered statistically significant. The number of pairwise comparisons C of conditions k was computed via equation (1).
Results
Figure 2 presents an overview of segmentation outputs generated by different model architectures in comparison to the ground truth. Figure 3 shows the F1-scores of different model configurations grouped by architecture, backbone family, and initialization strategy.
(1)
(2)
(3)
(4)
(4.3)
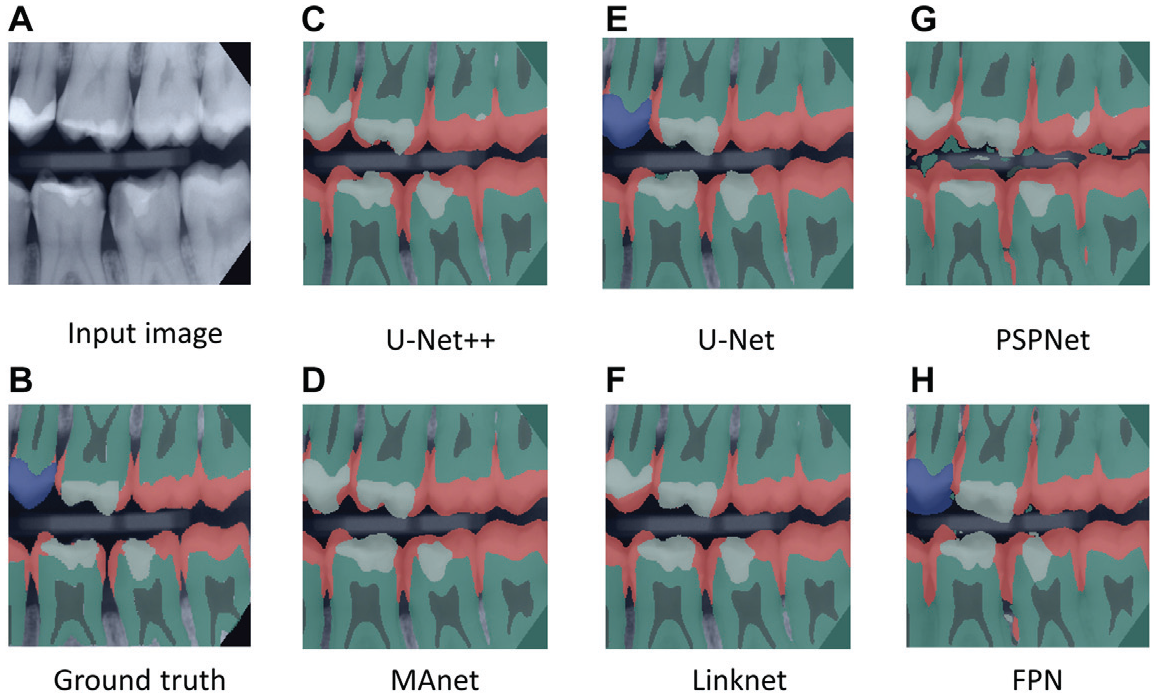
Examples of segmented bitewing radiographs. (
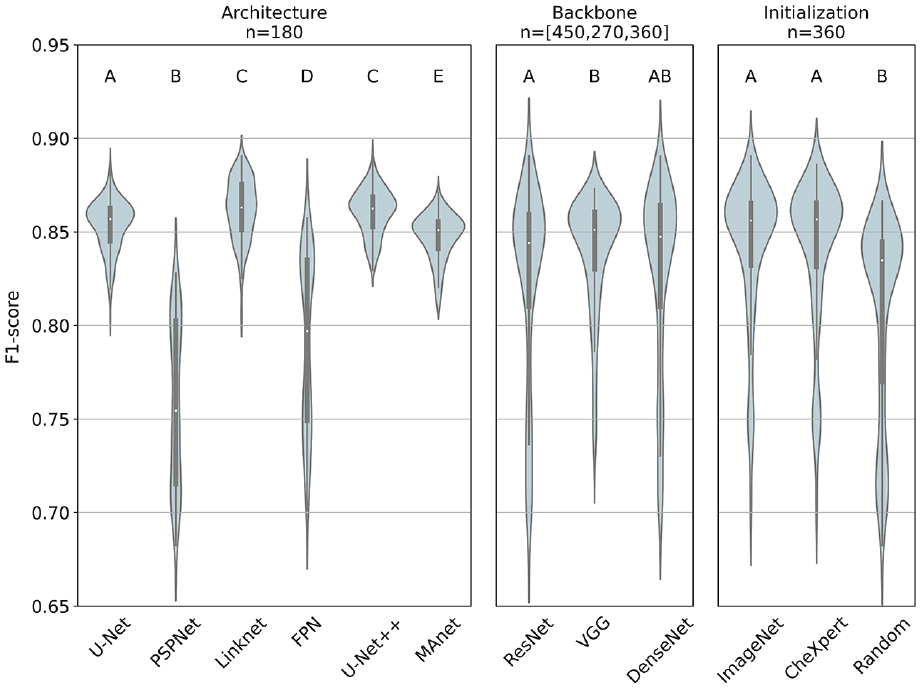
F1-scores stratified by initialization strategy, architecture, and backbone family based on sample sizes n. Median, interquartile range, and 95% confidence interval are represented by the white dot, the black box, and the black line, respectively. Different superscript letters indicate statistically significant difference (e.g., between U-Net and LinkNet), while the same superscript letters represent no significant difference (e.g., between LinkNet and U-Net++) (see Appendix for more details).
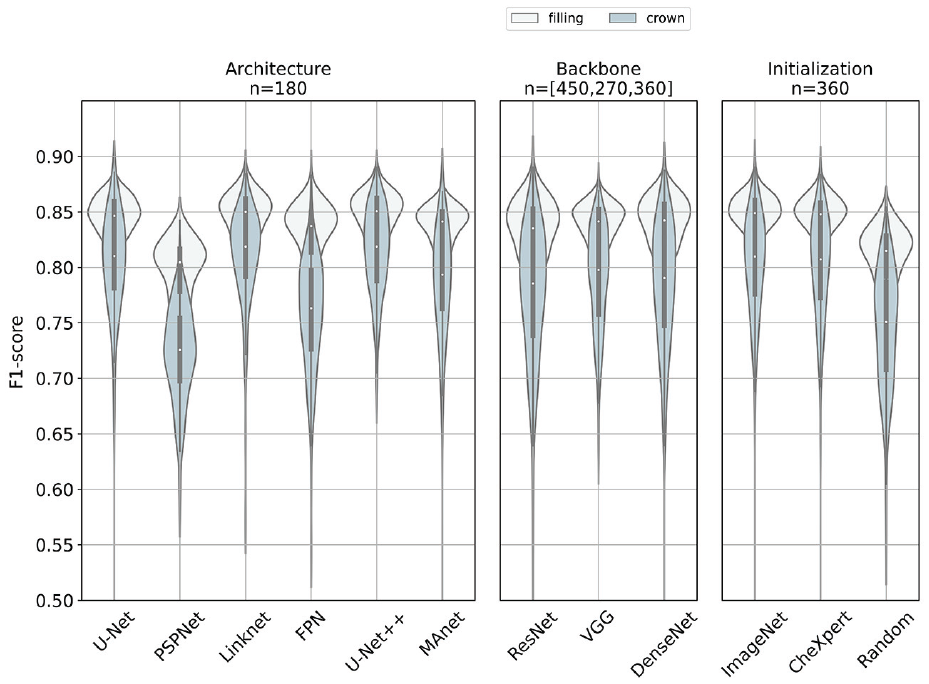
F1-scores of different models in the minority classes, filling (white) and crown (steel blue), respectively. We stratified the analyses by initialization strategy, architecture, and backbone family. Median, interquartile range, and 95% confidence interval are represented by the white dot, the black box, and the black line, respectively. Results are based on a sample size n. This figure is available in color online.
Discussion
We benchmarked 216 models defined by their architecture, complexity, and initialization strategy on a tooth structure segmentation task of dental bitewing radiographs. Several findings require a more detailed discussion.
First, we aimed to evaluate whether there are superior model architectures for the tooth segmentation task at hand. We discovered a performance advantage of models with backbones from the VGG family over models with backbones from the ResNet family. Our findings are consistent with those from Ke et al. (2021), who reported that architecture improvements reported on ImageNet may not always be translated to performances on medical imaging tasks. New model architectures and model improvements seem to be prone to overfitting on ImageNet data sets. Hence, transferability of newest AI research results into other domains, here the dental domain, may not be guaranteed.
The statistically significant performance advantage of models with VGG encoder backbones plead for the usage of VGG encoders, when solid baseline models are required, which perform reasonably well across different model configurations and settings. This may be relevant for the implementation of proof of concepts, for example. The top 10 performing models on the tooth structure segmentation task were built with backbones from the ResNet and DenseNet family. Consequently, if the focus is on model performance, it seems warranted to invest time to find an optimal model configuration based on more complex models (e.g., from the ResNet family). If, however, the validation of general concepts or benchmarking is the focus of the study, VGG-based models seem a reasonable choice as they are more robust across model configurations.
Second, one of our objectives evolved around the effect of the model complexity on the model performance. One of the key findings was a weak positive relationship between model depth and model performance. Therefore, we accept our hypothesis. Notably, however, the number of parameters increased in large steps, with only incremental improvements of model performance. Hence, the performance improvement was oftentimes disproportionate to the increasing demands for computational resources, training time, or the need to reduce image resolutions. The largest network in the present study was MAnet combined with a ResNet152 backbone, which reached an F1-score of 0.85 (0.85, 0.85) over all folds (ImageNet initialization). LinkNet in combination with a ResNet50 backbone was 5 times smaller but reached an F-score of 0.88 (0.88, 0.88) in comparison. It should be highlighted that lower computational costs allow for input imagery of higher resolution, which may be relevant for many dental applications.
Our third objective, aimed to give insights whether initializing with ImageNet or CheXpert, is consistently superior even when there is a difference in performance between both initialization strategies. We found statistically significant performance boosts for models initialized with ImageNet or CheXpert weights in comparison to a random initialization. These findings are consistent with those from Ke et al. (2021), who reported that 12 of 16 architectures benefited from an initialization with ImageNet weights for a classification task of chest radiographs. The comparison of ImageNet and CheXpert initialization showed no significant differences.
Fourth, we additionally found predictions on the minority class of filling (80%) to be generally more stable over different model configurations than predictions on class crowns (20%). Our results showed that there are superior architectures for segmenting minority classes (e.g., U-Net, U-Net++, LinkNet), but choosing a reasonable architecture may not be sufficient to overcome class imbalance. Hence, it could be recommended to address this problem with weighted loss functions (Guerrero-Penã et al. 2018) or oversampling (Buda et al. 2018).
This study comes with several limitations. First, our results were based on 1 specific DL task, a tooth structure segmentation on bitewing radiographs, and are limited to the examined model architectures. Hence, we do not claim generalizability of our findings across other segmentation tasks or over all existing model architectures. Second, images of our data set originate from varying machines, which may lead to different behavior of the models. Furthermore, radiographs with bridges, implants, and root canal fillings were not considered in the present study as they were very rare. We accept this as our aim was to benchmark models and not to build clinically useful ones in this study. In line with this, we were only aiming at a model comparison instead of proposing a high-precision model. Hence, we did not take any actions against the existing class imbalance and did not perform an extensive hyperparameter search. Finally, we based our analysis of the relationship between model performances and model complexity exclusively on the number of model parameters. It may be the case that model architectures with more parameters require less computational power through more efficient structures of layers. Furthermore, we did not evaluate the effect of minor differences in performance within the dental environment or how computational resources are affected by differences in the number of parameters of the models.
Conclusion
We benchmarked different configurations of DL models based on their architecture, backbone, and initialization strategy regarding their performance on a tooth structure segmentation task of dental bitewing radiographs to provide guidance for researchers in their DL model selection process. Regarding the superiority of certain model architectures, we found that VGG backbones provided solid baseline models across different model configurations, while peak performances were reached through combinations of U-Net++, LinkNet, and ResNet or DenseNet encoders. Superior architectures did not overcome class imbalance. Models known to perform better than others on a nondental data set like ImageNet did not demonstrate such superiority on our dental imaging task. The analysis of the relationship between model complexity and performance showed that deeper models did not necessarily perform better than shallow alternatives with lower demands in computational resources. Finally, we found that transfer learning boosts model performance, independent of the origin of transferred knowledge.
Author Contributions
L. Schneider, contributed to conception, design, data analysis, and interpretation, drafted and critically revised the manuscript; L. Arsiwala-Scheppach, contributed to analysis, critically revised the manuscript; J. Krois, contributed to conception, design, and data analysis, drafted and critically revised the manuscript; H. Meyer-Lueckel, contributed to interpretation, critically revised the manuscript; K.K. Bressem, contributed to acquisition and interpretation, critically revised the manuscript; S.M. Niehues, contributed to acquisition, critically revised the manuscript; F. Schwendicke, contributed to conception, design, data acquisition, and interpretation, drafted and critically revised the manuscript. All authors gave final approval and agree to be accountable for all aspects of the work.
Supplemental Material
sj-docx-1-jdr-10.1177_00220345221100169 – Supplemental material for Benchmarking Deep Learning Models for Tooth Structure Segmentation
Supplemental material, sj-docx-1-jdr-10.1177_00220345221100169 for Benchmarking Deep Learning Models for Tooth Structure Segmentation by L. Schneider, L. Arsiwala-Scheppach, J. Krois, H. Meyer-Lueckel, K.K. Bressem, S.M. Niehues and F. Schwendicke in Journal of Dental Research
Footnotes
A supplemental appendix to this article is available online.
Declaration of Conflicting Interests
The authors declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: F. Schwendicke and J. Krois are cofounders of the dentalXrai Ltd., a startup. dentalXrai Ltd. did not have any role in conceiving, conducting, or reporting this study.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.

