Abstract
Chronic nonbacterial osteomyelitis is a rare bone disorder that can be found in the jaw. It is often associated with systemic conditions, including autoimmune deficiencies. However, little is known about how the genetic and immunologic background of patients influences the disease. Here, we focus on human leukocyte antigen (HLA), killer cell immunoglobulin-like receptors (KIRs), and their specific combinations that have been difficult to analyze owing to their high diversity. We employed a recently developed technology of simultaneous typing of HLA alleles and KIR haplotype and investigated alleles of the 35 HLA loci and KIR haplotypes composed of centromeric and telomeric motifs in 18 cases and 18 controls for discovery and 472 independent controls for validation. We identified an amino acid substitution of threonine at position 94 of HLA-C in combination with the telomeric KIR genotype of haplotype tA01/tB01 that had significantly higher frequency (>20%) in the case population than in both control populations. Multiple logistic regression analysis based on a dominant model with adjustments for age and sex revealed and validated its statistical significance and high predictive accuracy (C-statistic ≥0.85). Structure-based analysis revealed that the combination of the amino acid change in HLA-C and the telomeric genotype tA01/tB01 could be associated with lower stability of HLA-C. This is the first case-control study of a rare disease that employed the latest sequencing technology enabling simultaneous typing and investigated amino acid polymorphisms at HLA loci in combination with KIR haplotype.
Keywords
Introduction
Osteomyelitis of the jaw is a severe inflammatory disorder affecting the bones and is characterized by recurrent severe pain, swelling of the cheek, and trismus. The medical management of patients with osteomyelitis has been unsatisfactory due to the unknown etiology of the disease and difficulties in establishing a correct diagnosis. Chronic nonbacterial osteomyelitis (CNO) is the rare type of chronic osteomyelitis, with an incidence of 1 to 2 per million, and constitutes approximately 2% to 5% of all osteomyelitis cases (Buch et al. 2019). The infectious process in the development of CNO has yet to be elucidated, and some patients do not respond to conventional antibiotic therapy (Yoshiura et al. 1994); in such cases, the usual treatment is surgery. However, there are many intractable cases of postoperative acute exacerbation and recurrence, with symptoms persisting for several years (Montonen et al. 1993).
CNO, the most severe form of which is chronic recurrent multifocal osteomyelitis, is an inflammatory disorder that can occur in any bone, most commonly in children and with variable clinical manifestations. However, CNO of the jaw mostly occurs in adults; the symptoms are often more intense, and lesions in the jaw occur commonly with multifocal bone involvement or in isolation (Wipff et al. 2015). CNO of the jaw is associated with several systemic conditions, including autoimmune deficiencies as well as rheumatic arthritis, chronic inflammatory bowel disease, and palmoplantar pustulosis (Gudmundsson et al. 2017). The inflammatory nonsuppurative process suggests a possible autoimmune etiology (Gudmundsson et al. 2017). Importantly, bisphosphonates that negatively influence osteoclast activity are used in the treatment of CNO (Hofmann et al. 2017) when conventional therapies such as anti-inflammatory agents fail to control symptoms (Hatano et al. 2012). This suggests that local excess of osteoclasts could account for the bone destruction observed in some other autoimmune diseases—for example, in rheumatoid arthritis in which natural killer cells are abundant in inflamed joints (Soderstrom et al. 2010). However, little is known about the kinetics of inflammatory responses in CNO or how the genetic and immunologic background of patients influences the disease.
Traditional typing of exons of a few loci in the human leukocyte antigen (HLA) gene region of patients with CNO showed that 4 of 12 patients had antigens B13 and B27 and, of these, 3 had the B13/B27 combination (Malmstrom et al. 1983). However, these studies had the following shortcomings. First, they had limited resolution and were not designed to identify amino acid polymorphisms associated with CNO. Second, until recent advances in next-generation sequencing, it was not possible to assess the contribution of all HLA genes spanning the 3.8-Mb segment of the human genome. Third, these studies did not evaluate possible interactions between killer cell immunoglobulin-like receptors (KIRs) and their HLA ligands. KIRs are members of the immunoglobulin family expressed on the surface of natural killer cells and some T cells (Rajagopalan and Long 2005). Recent studies have shown strong associations between KIR and HLA and susceptibility to autoimmune diseases such as rheumatoid arthritis (Fogel et al. 2013) and inflammatory bowel disease (Saito et al. 2018). It was reported that some KIRs can affect T cell–mediated immunity by controlling CD8+ T-cell survival (Boelen et al. 2018). These findings suggest that variable interactions between polymorphic variants of KIR and HLA influence immune cell education, which affects human health (Boudreau and Hsu 2018). Specific combinations of KIR and HLA were shown to be correlated with protection against or susceptibility to a range of infectious, reproductive, and autoimmune disorders (Rajagopalan and Long 2005; de Wit et al. 2016; Hilton and Parham 2017; Saito et al. 2018). However, analyzing the role of KIR and HLA genes in diseases has been difficult since it requires cutting-edge technology owing to the unusually high diversity of genes and their allelic variants at the KIR locus (Parham and Moffett 2013; Norman et al. 2016) and the remarkable polymorphism and complexity of the HLA region (Hosomichi et al. 2013).
We conducted a case-control study of CNO in the Japanese population and used the latest sequencing technologies to simultaneously determine the alleles of all 35 HLA loci and haplotype structures of the KIR region. We also used high-resolution analysis to explore how amino acid substitutions at HLA loci and KIR haplotype structure could contribute to CNO.
Materials and Methods
Diagnostic Criteria and Sample Collection
Although there is no international consensus on the definition of CNO, many researchers have agreed that the final diagnosis should be based on the following aspects: 1) clinical presentation and patient history, 2) imaging techniques, 3) culturing, and 4) histologic analysis (Coviello and Stevens 2007). In this study, we collected 7 mL of blood from 18 patients with the rare type of osteomyelitis and 18 healthy controls at several hospitals across Japan who met the following criteria: 1) recurrent pain and swelling; 2) radiographic appearance of a mixed pattern of sclerosis and osteolysis and uptake of scintigraphic agents such as technetium 99m in the region of the jaw bone; 3) little or no benefit from antibiotic treatment; and 4) increased bone resorption and deposition, and varying degrees of bone sclerosis and medullary fibrosis with no suspicion of malignancy.
Meanwhile, the healthy controls—aged 20 to 80 y with no history of chronic osteomyelitis and autoimmune diseases—were recruited from the same hospitals. We also recruited a substantially larger group of healthy controls (n = 472, aged 40 to 94 y with no history of chronic osteomyelitis and autoimmune diseases) from the Kanazawa prefecture, Japan, for validation. The age and sex of each subject were recorded, and DNA was extracted from blood samples.
Simultaneous Typing of HLA Alleles and KIR Haplotype
DNA libraries for sequencing HLA and KIR genes were amplified by ligation-mediated polymerase chain reaction and hybridized to SeqCap EZ Choice custom probes (Roche Sequencing Solutions). The captured target DNA libraries, with an average expected insert size of 600 bp, were subjected to paired-end sequencing (350 bp for read 1 and 250 bp for read 2) with the MiSeq system (Illumina).
Twelve classical HLA genes (HLA-A, -B, -C, -DRA, -DRB1, -DRB3, -DRB4, -DRB5, -DPA1, -DPB1, -DQA1, and -DQB1), 8 nonclassical genes (HLA-E, -F, -G, -DMA, -DMB, -DOA, -DOB, and MICA), 14 pseudogenes (HLA-H, -J, -K, -L, -T, -V, -W, -Y, -DRB2, -DRB6, -DRB7, -DRB8, -DRB9, and -DPB2), and the non-HLA gene TAP1 were selected for HLA typing according to the targeted capture method with SeqCap EZ Choice (Hirata et al. 2019), which was also used for targeted resequencing of 16 KIR genes (KIR2DL1, 2DL2, 2DL3, 2DL4, 2DL5, 2DP1, 2DS1, 2DS2, 2DS3, 2DS4, 2DS5, 3DL1, 3DL2, 3DL3, 3DP1, and 3DS1) to determine the KIR haplotype structure. KIR gene copy number was estimated with DepthOfCoverage in the Genome Analysis Toolkit to obtain the read depth for each gene in the sample interval summary file.
Statistical Analysis of Disease-Associated HLA Amino Acid Polymorphisms and Their Combinations with the KIR Haplotype Structure
The univariate screening to identify polymorphisms with >20% allele frequency increase or decrease in the case population was performed with an in-house Perl script and JMP Pro 13 (SAS Institute). For analysis at the amino acid level, we downloaded amino acid sequences of HLA alleles found in at least 1 individual from the IPD-IMGT/HLA database (Robinson et al. 2015). Multiple logistic regression analysis adjusting for the effects of age and sex was carried out with R statistical software.
Amino Acid Sequence and 3-dimensional Structure Analyses
Amino acid sequences of KIR family members KIR2DS1, KIR2DS2, KIR2DL1, and KIR2DL2 were aligned and sequence identities calculated with ClustalW software (Thompson et al. 1994). The binding interfaces were identified by the “protein interfaces, surfaces, and assemblies” service of the European Bioinformatics Institute (http://www.ebi.ac.uk/pdbe/prot_int/pistart.html; Krissinel and Henrick 2007). Distances between residues were manually calculated with PyMOL software (https://pymol.org/2/).
Ethical Considerations
This study was approved by the ethics committees of the Research Institute National Center for Global Health and Medicine (NCGM-A-000271-01), National Institute of Infectious Diseases (700), Nagasaki University (16020820-3), Hokkaido University (015-0377), Tokyo Medical and Dental University (G2016-005), Hiroshima University (Hi-200), Shinshu University (2017-2), and Kurume University (258). This article was prepared in accordance with the STROBE guidelines.
Results
Screening of HLA Amino Acid Polymorphisms and Their Combination with the KIR Haplotype
We investigated alleles of the 35 HLA loci across the 3.8-Mb HLA region and KIR haplotypes composed of centromeric and telomeric motifs in 18 patients with osteomyelitis and 18 healthy controls sampled across several hospitals in Japan (Appendix Table 1, with information of sex, age, and phenotype). The typing of HLA alleles and KIR haplotype of each individual is available at https://doi.org/10.6084/m9.figshare.9724040.v1. The median number of alleles per locus was 3, and the interquartile range was 2 to 8; the following 8 loci harbored >10 alleles: HLA-B (24 alleles), HLA-DRB1 (21), HLA-C (14), HLA-DQB1 (14), MICA (14), HLA-A (13), HLA-DQA1 (13), and HLA-DPB1 (12). The KIR haplotypes were the commonly found centromeric (cA01, cB01, and cB02) and telomeric (tA01 and tB01) motifs, each differing in terms of gene content and arrangement (Vukcevic et al. 2015; Appendix Fig. 1).
Due to the small sample size, we first carried out a univariate screening procedure to identify a specific HLA allele or KIR haplotype motif that may be associated with the disease with a frequency that was >20% higher or lower in the case population than in the control. We identified only 2 HLA alleles (HLA-H*01:01 and HLA-T*01:01) with >20% higher frequency in the case population. However, since these are pseudogenes, we were unable to interpret this result. We did not find any KIR haplotypes that showed >20% higher or lower frequency in the case population.
Given the high allelic diversity of HLA loci, we explored disease-associated polymorphisms at the amino acid rather than allele level at 3 HLA loci corresponding to major histocompatibility complex class I (HLA-A, -B, and -C), since interactions between class I and KIR genes are a risk factor in autoimmune diseases (Rajagopalan and Long 2005). Given the alignment of amino acid sequences of the loci, we extracted polymorphic residues and used the univariate screening procedure to identify those that could be associated with the disease as showing >20% frequency increase or decrease in the case population. No such residues were found in HLA-B or -C. In HLA-A, 10 amino acid residues had a 22.2% higher frequency in the case population, although the frequency difference decreased to 13.1% when the large independent population (n = 472) was used as a control for validation.
We also investigated whether combinations of amino acid polymorphisms at the 3 HLA loci and KIR haplotype were associated with the disease. The univariate screening revealed that the threonine at position 94 (I94T, without the N-terminal 24-amino acid signal peptide) of HLA-C in combination with the telomeric KIR genotype of haplotype tA01/tB01 had a 22.2% higher frequency in the case population. The frequency difference increased to 26.3% when the large independent population (n = 472) was used as a control for validation.
For the combination of threonine at position 94 (I94T) of HLA-C and the telomeric KIR genotype of haplotype tA01/tB01, we carried out multiple logistic regression analysis with a dominant model with adjustments for age and sex. There was a bias between the case and control populations for age (median, 58.5 vs. 46.5 y; P = 0.034, Wilcoxon’s rank sum test) but not sex (12% difference in the proportion of males; P = 0.5, chi-square test). There was no bias between the case and large independent control populations for age (median, 58.5 vs. 62.0 y; P = 0.21) and sex (3.7% difference in the proportion of males; P = 0.67). The analysis revealed that the disease association of the combination had a P value of 0.036 despite the small sample size. The C-statistic, a measure of predictive accuracy ranging from 0.5 to 1.0, was 0.86. When the large independent control population was used, the C-statistic was almost the same (0.85), and the P value decreased to 0.0005 (odds ratio, 5.5; 95% CI, 2.1 to 14.5).
Threonine at position 94 (I94T) of HLA-C was shared by HLA-C*01:02:01, HLA-C*04:01:01, HLA-C*04:09N, HLA-C*05:01:01, HLA-C*06:02:01, HLA-C*07:02:01, HLA-C*08:01:01, HLA-C*12:02:02, HLA-C*14:02:01, and HLA-C*14:03 alleles.
Structure-Based Modeling of the HLA-C Polymorphism Identified in This Study
The telomeric KIR genotype tA01/tB01 was dominant in the case population (10 of 18 vs. 8 of 18 and 159 of 472 in the control populations), while tA01/tA01 was dominant in the control populations (10 of 18 and 291 of 472 vs. 6 of 18 in the case population). This differed from the centromeric KIR genotype cA01/cA01, which was almost fixed irrespective of phenotype (14 of 18, 16 of 18, and 397 of 472 in the case and 2 control populations, respectively). The telomeric KIR haplotype tB01 was characterized by the presence of 3 activating KIR loci (3DS1, 2DS5, and 2DS1) and 1 inhibitory KIR gene (2DL5; Appendix Fig. 1). Although the binding mode of class I major histocompatibility complex HLA-C to these KIRs remains elusive, the structures of complexes formed by HLA-C and 2DL1/2DL2 have been reported (PDB IDs: 1IM9/1EFX; Fan et al. 2001; Boyington and Sun 2002). Given the strictly conserved amino acid sequence identity between 2DS1 and 2DL1/2DL2 (Fig. 1), we investigated how the identified HLA-C polymorphism (I94T) could affect binding by structure-based modeling with the structure of the HLA-C/2DL2 complex (Fig. 2), which is predicted to be similar to that of 2DS1 characteristic of KIR haplotype tB01.

Amino acid sequence alignment of KIR2DS1 and human homologs KIR2DS2/2DL1/2DL2. Black dots are shown above the alignment to indicate every 10 residues.
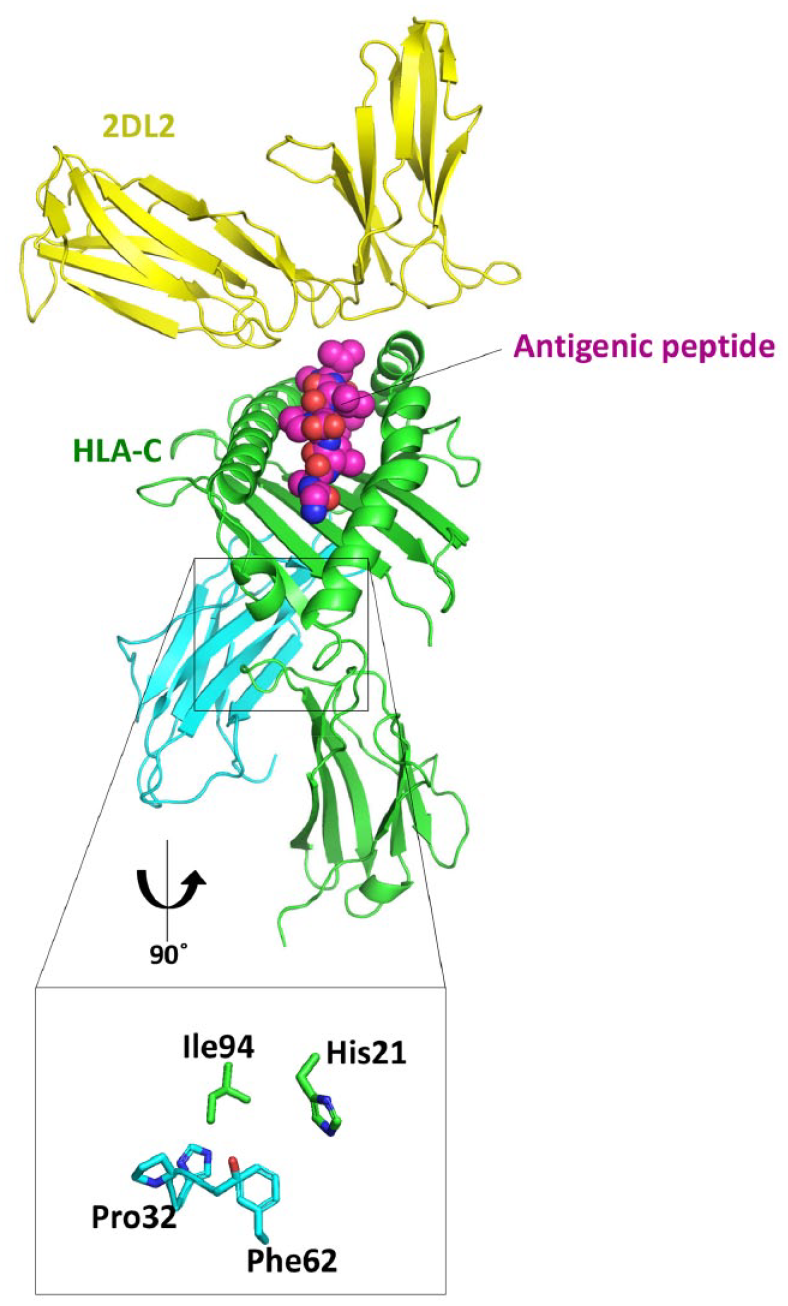
Structural modeling and mapping of disease-associated amino acids. β2-Microglobulin is shown in light blue. The human HLA-C/2DL2 complex (PDB: 1EFX) molecule shown in this figure does not include the N-terminal 24-amino acid signal peptide, and the positions shown are representative of the molecule after subtraction of the signal peptide. Close-up views of the interfaces are indicated by black rectangles with the same color coding. Surface representations of the binding interfaces are shown as transparent.
In the 3-dimensional structure, the identified polymorphism I94T (Ile94 in Fig. 2) is located on the side opposite to that for presentation of antigenic peptides to KIR family members, indicating that Ile94 is not directly involved in the interaction with KIR2DS1. However, structural analyses have demonstrated that Ile94 is located proximal to the binding interface between the HLA-C α chain (consisting of α1, α2, and α3 proteins) and β2-microglobulin in 2DL1/2DL2-bound structures. Notably, Ile94 engages in a hydrophobic interaction with Pro32 and Phe62 in the β2-microglobulin domain, suggesting that replacing the hydrophobic isoleucine with a hydrophilic threonine residue at this site could lower the binding affinity for β2-microglobulin.
Discussion
The KIR haplotypes found in this study were the cA01, cB01, and cB02 centromeric and tA01 and tB01 telomeric motifs. As a convention, KIR haplotypes formed by centromeric- and telomeric-A are defined as KIR group A haplotypes, whereas the other 3 combinations are defined as KIR group B haplotypes (Colucci 2017). Group A haplotypes outnumber group B haplotypes in the Japanese population, with individuals harboring 2 group A haplotypes accounting for 56% of the population (Yawata et al. 2002). In contrast, although the centromeric-A haplotype (cA01) was almost fixed irrespective of phenotype in the present study, the telomeric-B haplotype (cB01) was more frequently detected in the case population: the frequencies of the telomeric KIR genotypes tA01/tA01, tA01/tB01, and tB01/tB01 were 6, 10, and 2, respectively, in the case population (n = 18) and 10, 8, and 0 in the control population (n = 18). Although the frequency of the tB01 haplotype itself was not >20%, the combination of telomeric genotype tA01/tB01 with the amino acid change I94T (Ile94 in Fig. 2) in HLA-C showed a statistically significant >20% higher frequency and high predictive accuracy (C-statistic ≥ 0.85). Exploring such combinations between the usually minor KIR group B haplotype and specific amino acid changes in HLA may be an effective approach in association studies of various diseases, including CNO.
Structure-based analysis of the combination of the telomeric genotype tA01/tB01 and the amino acid change I94T (Ile94 in Fig. 2) in HLA-C suggested that this substitution weakens the interaction between the HLA-C α chain and β2-microglobulin domain of KIR, possibly resulting in self-oligomerization leading to the unfolded protein response and/or destabilization of HLA-C itself. It has been suggested that the variation in HLA stability among alleles is associated with autoimmunity (Miyadera et al. 2015). Further studies are needed to determine how this weakened interaction is related to CNO at the molecular level, since some HLA molecules are known to “misbehave”—for example, by misfolding inside cells—and cause diseases (McHugh and Bowness 2012; Colbert et al. 2014).
The precise ligands and functions of the KIR3DS1 receptor specifically encoded by the tB01 haplotype remain unclear (Chapel et al. 2017). In general, activating members of the KIR family—although closely related to inhibitory KIRs—weakly interact with HLA class I, and KIR2DS1 was reported in some assays to weakly interact with HLA-C group 2 (HLA-C2), which is characterized by a lysine at position 80 (without the N-terminal 24-amino acid signal peptide; van der Ploeg et al. 2017). It was recently reported that KIR2DS1 binding was narrowly restricted to HLA-C2 (Chapel et al. 2017). However, the Japanese population has the lowest HLA-C2 and the highest KIR group A haplotype frequencies (Arima et al. 2018). Indeed, most individuals in our study were homozygous for HLA-C1 characterized by an asparagine at position 80, and only 2 patients and 1 healthy control were heterozygous for HLA-C2 and HLA-C1. Further studies are needed to determine the expression levels of KIR2DS1 in patients with CNO with the KIR group B haplotype. If, as predicted, KIR2DS1 is indeed differentially expressed in patients with CNO, clarifying the influence of the amino acid change I94T on the binding affinity of HLA-C1 for KIR2DS1 could provide molecular-level insight into the disease.
Although the sample size was small, this is the first multicenter case-control study of CNO that employed the latest sequencing technology to identify alleles of all 35 HLA loci with the haplotype structure of the KIR region to investigate how amino acid polymorphisms at HLA loci in combination with KIR haplotype could influence the disease. Our study provides a basis for genetic association studies of rare diseases that focus on HLA, KIR, and their interaction.
Author Contributions
H. Yahara, contributed to conception, design, data acquisition, analysis, and interpretation, drafted and critically revised the manuscript; S. Horita, K. Hosomichi, contributed to analysis and interpretation, drafted and critically revised the manuscript; S. Yanamoto, Y. Kitagawa, T. Asaka, T. Yoda, K. Morita, Y. Michi, M. Takechi, H. Shimasue, Y. Maruoka, E. Kondo, J. Kusukawa, H. Tsujiguchi, T. Sato, T. Kannon, H. Nakamura, contributed to data acquisition, critically revised the manuscript; A. Tajima, contributed to data interpretation, critically revised the manuscript; K. Yahara, contributed to design, data analysis, and interpretation, drafted and critically revised the manuscript. All authors gave final approval and agree to be accountable for all aspects of the work.
Supplemental Material
DS_10.1177_0022034520901519 – Supplemental material for A Targeted Genetic Association Study of the Rare Type of Osteomyelitis
Supplemental material, DS_10.1177_0022034520901519 for A Targeted Genetic Association Study of the Rare Type of Osteomyelitis by H. Yahara, S. Horita, S. Yanamoto, Y. Kitagawa, T. Asaka, T. Yoda, K. Morita, Y. Michi, M. Takechi, H. Shimasue, Y. Maruoka, E. Kondo, J. Kusukawa, H. Tsujiguchi, T. Sato, T. Kannon, H. Nakamura, A. Tajima, K. Hosomichi and K. Yahara in Journal of Dental Research
Footnotes
Acknowledgements
We thank Dr. Simon Kollnberger (Cardiff Institute of Infection and Immunity) and Dr. Noriko Sorimachi (National Center for Global Health and Medicine in Japan) for helpful discussions.
A supplemental appendix to this article is available online.
This research was supported by JSPS Research Fellowships for Young Scientists to H.Y. and Grants-in-Aid for Scientific Research of Education, Culture, Science, Sports, and Technology (MEXT) of Japan (No. 15H06794 to H.Y. and No. 16H06502 and 19H05344 to K.H).
The authors declare no potential conflicts of interest with respect to the authorship and/or publication of this article.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
