Abstract
Artificial Intelligence (AI) based ultrasound image processing is quite important for diagnosis of renal tumors. Traditional networks, however, struggle with accurate lesion segmentation and differentiation between benign and malignant cases in complex ultrasound images. This paper introduces an advanced multitask network model to overcome these limitations. Our model combines a deep separable Convolutional ResNet with a NestedUNet, enriched with a Convolutional Block Attention Module (CBAM) and residual connections, enhancing its precision in lesion segmentation and classification in intricate images. Compared with the existing methods, the model proposed in this study has the following innovative contributions: (1) combining depthwise separable convolution and NestedUNet, the computational efficiency and feature extraction ability of the model are significantly improved; (2) The Convolutional Block Attention Module (CBAM) is introduced to enhance the attention to key features, to effectively distinguish lesions in complex backgrounds; (3) The gradient flow is improved by the residual connection, which ensures the stable training of the deep network. Experimental results show that the IoU of the model on the kidney tumor dataset reaches 94.23%, which is significantly better than that of STUNet (92.31%). These results show that the model proposed in this study has significant advantages in processing complex ultrasound images, providing a more accurate basis for clinical diagnosis, and has a high application potential.
Keywords
Introduction
Currently, cancer continues to pose a significant threat to human well-being across the globe. The incidence of kidney cancer has been steadily rising, with over 400,000 new cases being reported annually worldwide.1,2 A majority of renal tumors exhibit subtle clinical manifestations. 3 Roughly 20%–30% of patients undergoing renal tumor resection are prone to preoperative misdiagnosis, leading to unnecessary surgical interventions that ultimately result in benign diagnoses post-surgery. 4 In such scenarios, the precision of tumor diagnosis for patients warrants enhancement, particularly with regards to accurate differentiation between benign solid tumors displaying low echogenicity and malignant tumors.
In the domain of medical imaging, ultrasound images are widely adopted for clinical diagnosis due to their non-invasive, real-time capabilities, and cost-effectiveness. However, the quality of ultrasound images is influenced by diverse factors including equipment parameters, procedural techniques, and patient physique, thereby engendering subjectivity and complexity in image interpretation. This complexity is particularly amplified in the context of renal tumor diagnosis, where distinguishing between benign and malignant tumors and precisely segmenting lesions is pivotal in formulating treatment strategies and predicting disease trajectories. Thus, leveraging advanced image processing techniques to heighten the diagnostic accuracy of ultrasound images of renal tumors constitutes a pivotal focal point of current research.
Over the past few years, deep learning has exhibited promising outcomes in the classification and diagnosis of kidney tumors.5–8 Unlike traditional machine learning, deep learning doesn’t necessitate the subjective definition of features, and it can capture comprehensive biological information from images.9–13 Literature underscores that deep learning algorithms surpass human experts in diagnosing a myriad of diseases such as liver ailments, breast conditions, lung pathologies, retinal diseases, and skin lesions.14–17 These investigations underscore the stability and universality of deep learning, bridging diagnostic discrepancies between medical professionals with varied levels of expertise. To the best of our knowledge, research dedicated to differentiating benign and malignant solid renal tumors through ultrasound-based radiomics is limited in scope, yielding results that are less than satisfactory.18,19
In line with our research objective, we developed a collaborative multitask network model for the analysis of grayscale ultrasound images of kidney tumors. The model incorporates deep separable convolutions from ResNet 20 and NestedUNet, 21 enhanced with the Convolutional Block Attention Module (CBAM) 22 and residual connections to improve feature extraction and maintain gradient flow. The workflow includes image preprocessing, lesion segmentation, and classifying lesions as benign or malignant, providing crucial diagnostic support. Experimental results show that the model achieves a classification accuracy of 81.8%, an IoU of 0.942, and an AUC of 0.88, demonstrating its effectiveness in lesion localization and classification, thus offering valuable support for kidney tumor diagnosis.
Main contributions
(1) Development of a Novel Multitask Network Model: Introduction of a new multitask network combining DSCResNet and AttentionNestedUNet. This model significantly improves the processing and analysis of grayscale ultrasound images of renal tumors. Our model outperforms StUNet 32 with an impressive IoU of 0.942, demonstrating advanced capability in accurately delineating tumor boundaries.
(2) Effective Classification of Renal Tumors: The model exhibits high accuracy in classifying renal tumors as benign or malignant, with an accuracy rate of 81.8%, providing reliable support for medical diagnosis. Utilization of accuracy (for classification tasks) and IoU (for segmentation tasks) as evaluation metrics, providing a thorough assessment of the model’s effectiveness in both aspects of renal tumor diagnosis. The ability of the model to precisely locate and segment lesions, coupled with its robust classification accuracy, significantly enhances clinical diagnostic processes and contributes to medical research in the field of renal tumors.
The subsequent chapters of this paper are structured as follows: Chapter 2 details our dataset and preprocessing methods. Chapter 3 describes our model architecture and implementation specifics. Chapter 4 presents our experimental results and performance evaluation. Chapter 5 discusses the implications of our findings, discussing the potential and limitations of our model in clinical applications. Finally, Chapter 6 concludes our study, summarizing our contributions and suggesting directions for future research. This organization aims to comprehensively showcase our research from data collection to model evaluation, concluding with prospects for further advancements.
Related work
The processing of renal ultrasound images has remained a critical focus of medical imaging research. In recent years, the surging popularity of machine learning and deep neural networks in diverse clinical applications has spurred heightened interest in the development of medical image processing systems rooted in deep learning. Presently, the Tumor Biological Characteristics Assessment (TBCA) stands as the primary approach for benign and malignant diagnoses of kidney tumors.23,24 Through TBCA, clinicians can evaluate the benign or malignant nature of kidney tumors without being reliant on specialized medical equipment. Nevertheless, TBCA-based diagnostic methods encounter certain limitations. To elaborate, the diagnosis heavily hinges on the observations, clinical expertise, and description of tumor biological attributes by physicians, which could be susceptible to subjective influences. Consequently, in the realm of benign and malignant diagnoses for renal tumors, numerous scholars have embarked on proactive exploration of computer-assisted technologies to develop more convenient and objective diagnostic methodologies.
This article focuses on two key areas: traditional approaches and deep learning based approaches. These trajectories of research hold the potential to furnish a fresh perspective and novel methodologies for enhancing the diagnostic accuracy of benign and malignant renal tumors.
Traditional approaches
In the realm of diagnosing benign and malignant renal tumors, traditional ultrasound image processing methods have played a significant role. These methods harness a sequence of image processing techniques, encompassing threshold segmentation, edge detection, and morphological operations, to extract pivotal features from kidney ultrasound images, thereby aiding the assessment of benign and malignant distinctions.
In a recent investigation, Xiong et al. 25 introduced a novel level set method for segmenting ultrasound images of kidney tumors. This approach leverages adaptive partitioning, segmenting the image into several subregions, and then executing level set evolution within each subregion, thereby enhancing segmentation accuracy and efficiency. The technique amalgamates localized region information, global information, and prior knowledge of shape, to amplify the robustness and precision of segmentation. This methodology adeptly tackles issues like noise, blurriness, and non-uniform illumination in ultrasound images, concurrently preserving the integrity and continuity of tumor boundaries. However, this method still necessitates manual initial contour selection, potentially impacting the consistency and outcome of segmentation. Additionally, it has not undergone comprehensive comparative experiments against alternative level set techniques to validate its superiority.
Pratondo et al. 26 proposed a medical image segmentation approach rooted in regional growth, which employed an enhanced multi-seed algorithm and was parallelized on a multi-core CPU machine. They introduced a seed selection strategy based on local gray mean and variance, effectively mitigating the phenomena of over-segmentation and under-segmentation. Simultaneously, a region merging criterion reliant on neighborhood similarity was incorporated, further enhancing segmentation precision and robustness. Furthermore, the parallelization scheme, powered by OpenMP, substantially augmented segmentation speed and efficiency. However, this method demands excessive manual parameter adjustment, potentially impacting the outcome and efficacy of segmentation. Moreover, it remains reliant on gray information, potentially presenting limitations and drawbacks when applied to images containing rich color or texture information.
Though traditional ultrasound image processing techniques can yield commendable results in simpler scenarios, their efficacy is often curtailed when confronted with intricate and ambiguous ultrasound images. These methodologies usually necessitate manual parameter tweaking and are sensitive to factors like image quality and noise, rendering them less adaptable to a diverse array of complex clinical contexts.
Deep learning
In recent years, significant strides have been made in the application of deep learning technology in medical image processing. Convolutional Neural Networks (CNNs), in particular, have showcased remarkable feature extraction capabilities, rendering them highly effective in tasks like image classification and segmentation.
In a recent study, Liu et al. 27 employed an MRI-based ResNet-50 convolutional neural network to predict the primary tumor site of spinal metastases. Utilizing ResNet-50 as the classifier, they fed a stack of five images and generated outputs across six categories: breast cancer, lung cancer, prostate cancer, kidney cancer, thyroid cancer, and other cancers. The training process involved the use of the cross-entropy loss function and the Adam optimizer. Their approach leveraged ResNet-50s strengths in deep residual learning, resulting in enhanced classifier performance and efficiency. However, the dataset labels they utilized might exhibit errors or inconsistencies due to primary tumor site diagnoses relying on clinical information and other examination methods. Additionally, a lack of comparison with alternative MRI-based classification methods limits the demonstration of the superiority or constraints of their approach.
Mahmud et al. 28 developed a machine learning-based model that harnessed CT scan images and clinical metadata to predict the pathological grade and surgical choices for patients with kidney cancer. The authors employed convolutional neural networks (CNNs) to extract features from CT images, combining them with clinical metadata. These combined inputs were fed into a multi-layer perceptron (MLP) to perform classification for pathological grading and surgical selection. The resultant predictions were then compared with those made by human experts. This pioneering study represents the first attempt to predict pathological grading and surgical choices in kidney cancer patients using machine learning in conjunction with CT images and clinical metadata. It demonstrates both innovation and practicality, showing that machine learning models can match or even exceed human experts in predicting pathological grading and surgical choices. Ultimately, this presents an effective auxiliary tool for clinical decision-making.
However, the deep learning approach also presents certain challenges. Firstly, deep learning models typically require a substantial amount of annotated data for training, which is often a challenge in the field of medical imaging. Secondly, deep learning models lack interpretability, limiting their applicability in clinical decision-making.
Although traditional ultrasound image processing methods perform well in simple scenes, these methods often show insufficient adaptability when dealing with complex ultrasound images containing a lot of noise and fuzzy boundaries. This is due to the fact that they often rely on manual parameter tuning and are very sensitive to image quality. In addition, traditional methods are difficult to effectively capture the complex features of renal tumors, especially in the case of overlapping lesion features and low signal-to-noise ratio.
Our proposed model addresses the challenges of traditional ultrasound image processing methods by introducing a joint multi-task network combining deep separable convolutional ResNet, NestedUNet, CBAM, and residual connections. This combination leverages deep separable convolutions to reduce computational complexity, NestedUNet for multi-scale feature fusion, CBAM to enhance focus on important regions, and residual connections to stabilize training and mitigate gradient vanishing. These innovations enhance the model’s ability to segment lesions and classify benign-malignant cases, improving its accuracy and robustness, and effectively aiding medical practitioners in diagnosing complex renal tumor ultrasound images.
The mixed domain attention mechanism, also known as the Convolutional Block Attention Module (CBAM), is an attention mechanism applied within convolutional neural networks to bolster the model’s focus on crucial features while downplaying insignificant ones. Originally designed for image classification tasks, CBAM’s concepts can be extended to other computer vision applications like object detection and image segmentation. By integrating both Channel Attention and Spatial Attention mechanisms, CBAM empowers the network to dynamically adjust the weighting distribution of feature maps, channeling heightened attention toward pivotal feature information.
Depthwise Separable Convolution represents an operation within convolutional neural networks that breaks down the standard convolutional operation into two phases: depthwise convolution and pointwise convolution. This partitioning assists in trimming the model’s parameter count and computational complexity, concurrently maintaining commendable performance in numerous instances. It augments feature extraction, diminishes overfitting risks, and thus strives to balance the equilibrium between model efficacy and computational efficiency.
ResNet, 20 as a deep CNN, introduces residual connections that tackle the vanishing gradient problem within deep networks by establishing direct connections between layers, thereby enabling the construction of deeper network structures. This structural refinement enhances training stability and network performance.
NestedUNet 21 (UNet+) constitutes an extended iteration of the U-Net architecture, intensifying the multi-scale representation of features through the progressive extension of nested structures. The amalgamation of feature maps from distinct depths empowers the network to adeptly capture intricate details and contextual cues, thereby lending itself well to tasks such as biomedical image segmentation.
In forthcoming chapters, we will meticulously expound upon our model architecture and implementation particulars, in addition to presenting experimental results and undertaking performance evaluations.
Dataset and model
Description of the dataset
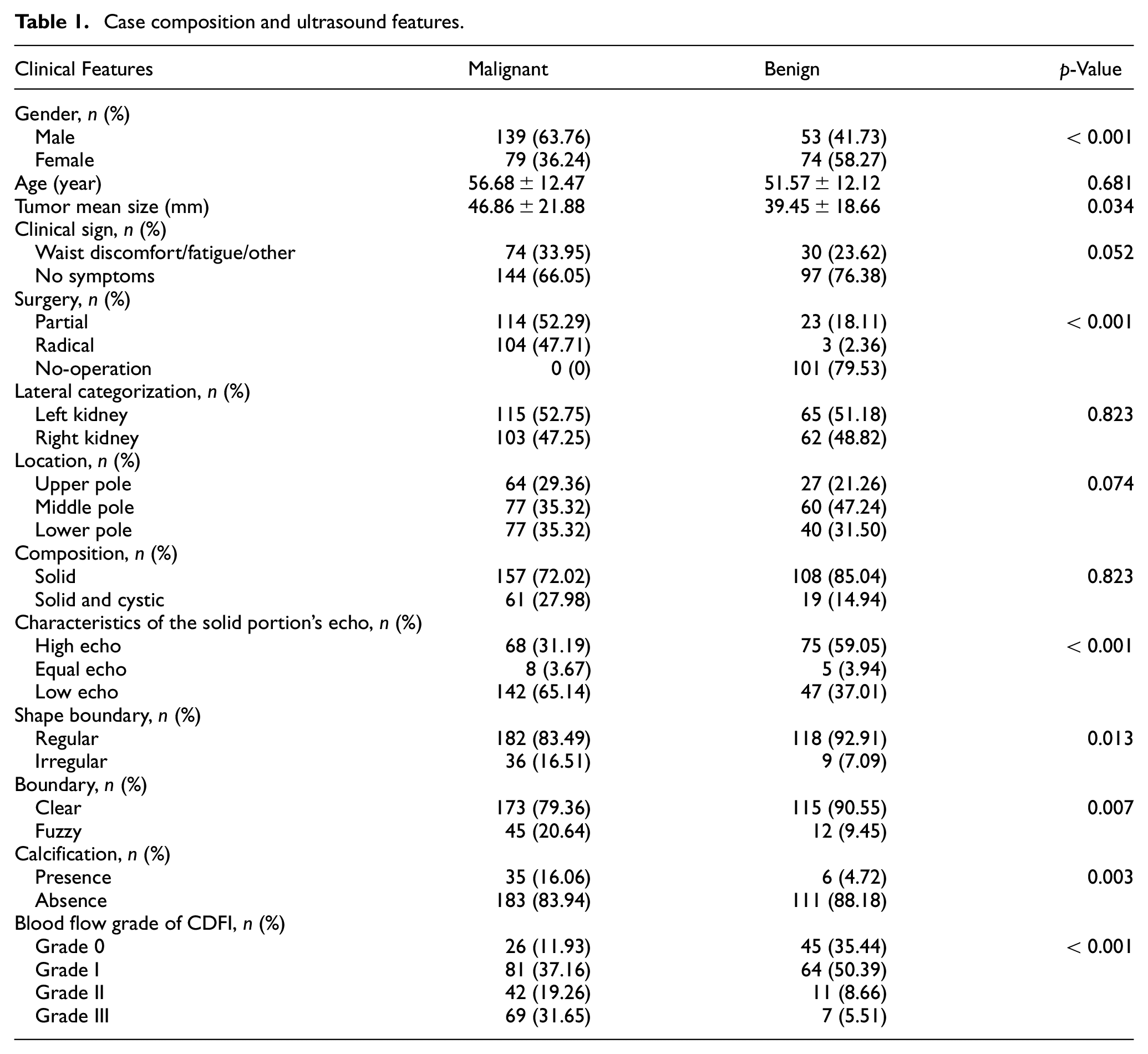
The dataset used in this study was gathered by specialized radiologists at the First Affiliated Hospital of Nanchang University. It involved a retrospective analysis of 345 cases of renal tumors treated and diagnosed at the hospital between January 2019 and May 2023. Among these cases, 218 were identified as malignant tumors, all confirmed via postoperative pathology. Additionally, 127 cases were benign tumors, all verified through postoperative pathology or confirmed by CT/MRI scans with no significant changes observed during a dynamic follow-up period of 3 months. Comprehensive case details and ultrasound features are outlined in Table 1.
Case composition and ultrasound features.
This study obtained the first affiliated hospital of Nanchang University ethics committee approval (approval number: IIT2023174), and from the requirement of informed consent. The retrospective analysis of this study ensured that all patient data had been anonymized to ensure privacy protection.
Additionally, all data processing and analysis are in accordance with relevant data protection laws and regulations to ensure the security and privacy of patient information.
In this study, we use distinct metrics to evaluate our network’s performance. For classification tasks, specifically distinguishing between benign and malignant renal tumors, we measure accuracy. In contrast, for segmentation tasks, like identifying tumor boundaries, we use Intersection over Union (IoU) as the primary metric.
For ultrasound data acquisition, various instruments were employed, including LOGIQ E9 from General Electric Healthcare (USA), Siemens ACUSON Sequoia from Siemens Medical Systems (USA), Resona R9 from Mindray Ultrasound Systems (China), and IU22 from Philips Medical Systems (Netherlands). A convex probe with a frequency range of 1.0–5.0 MHz was utilized. During the acquisition process, patients were positioned either flat or on their side, and the kidneys were scanned through a routine multi-sectional approach. Recorded information included parameters such as maximum diameter, lateral categorization (left kidney/right kidney), location (upper/middle/lower pole), composition (solid, cystic), characteristics of the solid portion’s echo (high echo/equal echo/low echo), shape (regular/irregular), boundary (clear/fuzzy), presence/absence of calcification, and Blood flow grade of CDFI (0/I/II/III grade).
The dataset used in this study includes images from multiple devices and features a range of kidney tumor types, providing a certain level of diversity. However, to further improve the model’s generalizability, it is important to consider expanding the dataset in future work. Since the current dataset is sourced from a single institution, this limited scope may affect the model’s ability to generalize to broader healthcare settings. Future studies should therefore aim to incorporate data from multiple institutions, which would help enhance the robustness and applicability of the model across diverse clinical environments.
Data annotation and preprocessing
Ultrasound images of renal tumors were obtained using four different types of ultrasound equipment mentioned in the previous subsection, with the proportion of images from each device being 1:2:2:1. These images were acquired through ultrasound scanning procedures performed on 345 patients, comprising 127 benign and 218 malignant cases. Each image is captured from the video at a certain frequency at the video frame rate, and from these obtained images are selected by a professional physician, focusing on capturing clear and representative views of the tumors.
During the preprocessing phase, we utilized OpenCV’s noise reduction functions, using bilateral filtering to enhance image clarity and reduce artifacts. Bilateral filtering was chosen for its effectiveness in reducing noise while preserving important details, such as the edges of anatomical structures in ultrasound images. This step was crucial in ensuring that the subsequent image processing and analysis could be conducted on high-quality data. Additionally, to address the imbalance between benign and malignant cases in the dataset, we employed a combination of oversampling and weight adjustment. We increased the number of benign images through oversampling and assigned higher weights to certain classes during model training. This approach ensured that more weight was given to errors in these classes in the loss function, thereby improving the model’s performance on less represented classes.
Contrary to the previous description, the images were not annotated using bounding boxes. Instead, we used a specialized radiomics annotation tool, allowing an experienced radiologist to mark the precise location of the tumors. These annotated regions were then processed using Python programing tools to generate label images, which accurately reflected the radiologist’s expert assessments.
The dataset was further enhanced through image processing techniques, including contrast enhancement, image flipping, and other augmentation procedures. Through these methods, we extracted and generated a total of 3504 images. These steps significantly improved the model’s ability to discern key image features, thereby enhancing its overall performance in segmenting and classifying renal tumors.
To rigorously evaluate the performance of our enhanced multitask network model, we employed a fivefold cross-validation technique. In this process, our renal tumor ultrasound dataset was divided into five equal parts, with each part being used as the test set while the remaining four parts served as the training set in a rotational manner. This method ensures a comprehensive assessment of the model by utilizing different subsets of the data for training and validation, thereby reducing bias and variance in the performance metrics.
Model design
We have devised a multi-task network model that incorporates a deep separable convolutional ResNet and NestedUNet equipped with a mixed domain attention mechanism. The DSCResNet component addresses the training challenge faced by deep neural networks through the introduction of residual connections, enabling the network to learn deeper features. Meanwhile, the AttentionNestedUNet employs a nested structure, deep supervision, and force mechanism to encourage the network to focus more on the local features of the image during image processing, thereby enhancing the model’s performance. We have enhanced the architecture by integrating residual connections, a critical element in deep learning models known for resolving the vanishing gradient problem in deep networks. These connections facilitate uninterrupted gradient flow, allowing for effective training of deeper network layers. Specifically, we have incorporated these residual connections within each block of our AttentionNestedUNet, ensuring a direct link between the input and output of each block. This modification is designed to improve the network’s capacity for feature extraction and recognition, crucial for the precise segmentation of renal tumor lesions and the accurate classification of their benign or malignant nature. Our model now takes advantage of both the attention mechanism provided by CBAM for refined feature focus and the enhanced gradient flow facilitated by the residual connections, aiming for superior performance in the complex task of ultrasound image analysis for renal tumor diagnosis.
To enhance both segmentation and classification tasks, our model employs deep separable convolutions (DSC) through ResNet specifically for the classification of renal tumors, while the AttentionNestedUNet is utilized for effective lesion segmentation. The AttentionNestedUNet combines the Convolutional Block Attention Module (CBAM) and residual connections, which significantly improves the model’s ability to focus on critical areas of the ultrasound images, enhancing the segmentation performance even in the presence of noise and blurred boundaries. CBAM enhances the model’s attention to important features, while residual connections help solve the vanishing gradient problem in deep networks and ensure stable training of the model.
The combination of DSCResNet for classification and AttentionNestedUNet for segmentation allows the segmentation and classification tasks to be conducted simultaneously. Importantly, the segmented lesions can be directly utilized for classification, improving the accuracy and efficiency of the model. This joint approach facilitates better feature learning for both tasks, providing a more comprehensive understanding of the complex characteristics present in renal ultrasound images, ultimately leading to superior performance compared to traditional standalone segmentation or classification models.
The DSCResNet plays a crucial role in refining the classification process. The integration of deep separable convolutions in ResNet architecture significantly reduces the model’s parameter count, enhancing computational efficiency without compromising the depth or performance. This adaptation allows for more efficient feature extraction and processing, especially in complex renal tumor classifications. The streamlined architecture of DSCResNet, combined with residual connections, ensures effective learning even at deeper levels, thereby optimizing the classification of ultrasound images into benign or malignant categories.
To further enhance the model’s segmentation capabilities, we have evolved the AttentionNestedUNet architecture. This advanced version now integrates not only the Convolutional Block Attention Module (CBAM) but also incorporates residual connections within each segmentation block. CBAM helps focus the network’s attention on the most informative features of ultrasound images, combining channel and spatial attention mechanisms for a nuanced understanding of image context and renal tumor intricacies. The addition of residual connections complements this by ensuring consistent and effective gradient flow, which is particularly beneficial for training deeper layers of the network. This dual enhancement with CBAM and residual connections significantly boosts segmentation accuracy, enabling more precise delineation of tumor boundaries. The AttentionNestedUNet, with its emphasis on detail and context, represents a substantial improvement in the field of biomedical image segmentation, particularly for challenging renal tumor cases.
In our multitask network model, we distinguish between the evaluation metrics for the two primary tasks: classification and segmentation. For classification tasks, we measure accuracy to assess the model’s efficacy in categorizing renal tumors. Conversely, for segmentation tasks aimed at precise tumor boundary delineation, we utilize IoU as our key performance metric.
To mitigate the risk of overfitting, which is common in deep neural network models due to their high complexity, we implemented two key strategies in our multitask network model. First, we applied data augmentation techniques such as rotation, scaling, and flipping, to artificially expand the diversity of our training dataset, ensuring that our model learns from a broader range of patterns and variations present in ultrasound images. Second, we incorporated dropout layers within the DSCResNet and AttentionNestedUNet architectures. By randomly deactivating a fraction of neurons during the training process, dropout prevents the network from becoming too dependent on any single node, thus encouraging a more generalized learning pattern. These measures have proven effective in enhancing the model’s generalization abilities and have shown to improve performance on unseen data, as evidenced by our rigorous validation process.
The convolutional block attention module (CBAM) plays a key role in our model. By combining channel attention and spatial attention mechanisms, CBAM enhances attention to important features in ultrasound images. In particular, CBAM first by channel module selectively focusing on the classification of the most important characteristics of the channel, and then through the space module on the feature map spatial dimension focusing on key areas. Figure 2 shows the CBAM in the application of image processing.
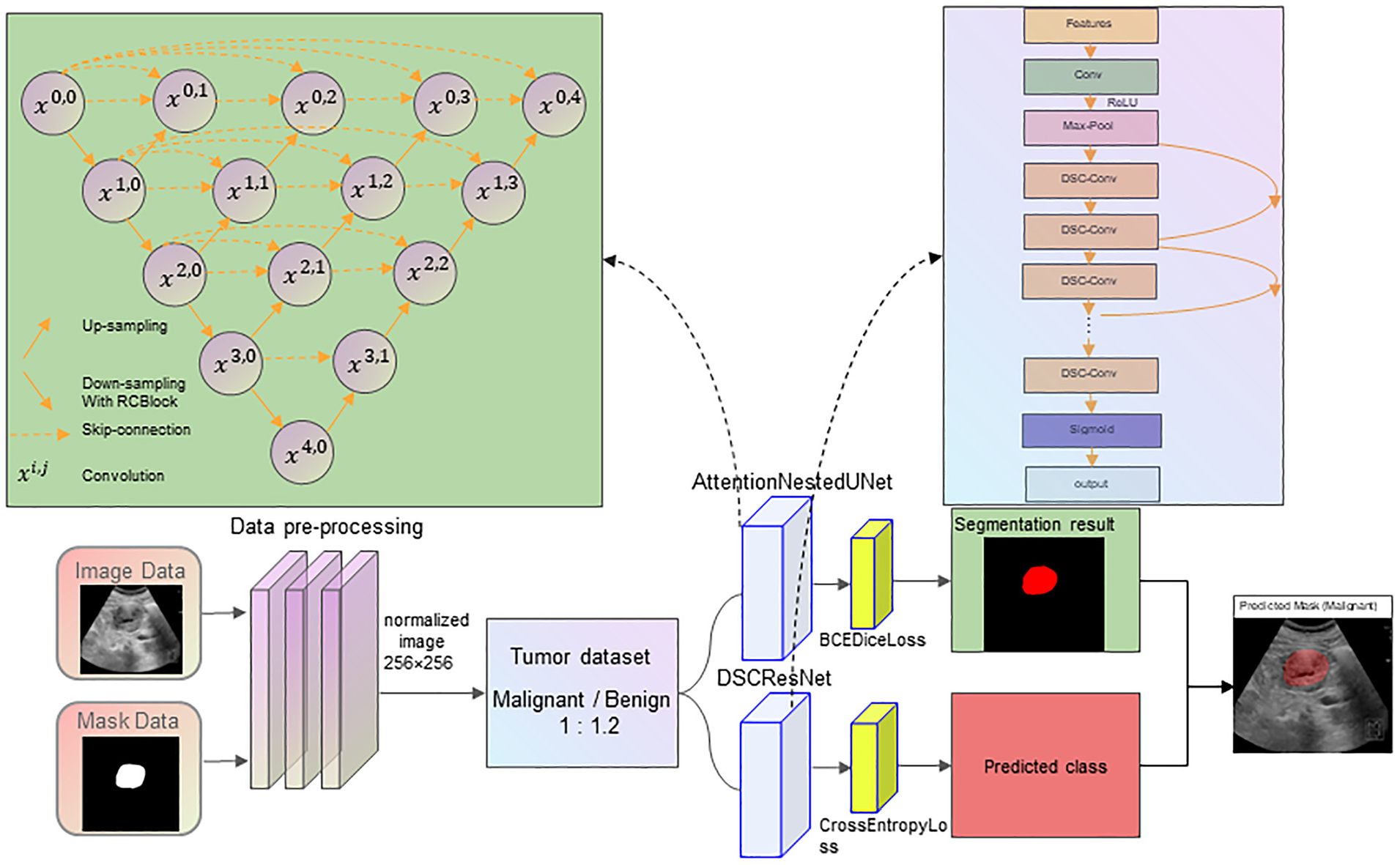
Model structure diagram.
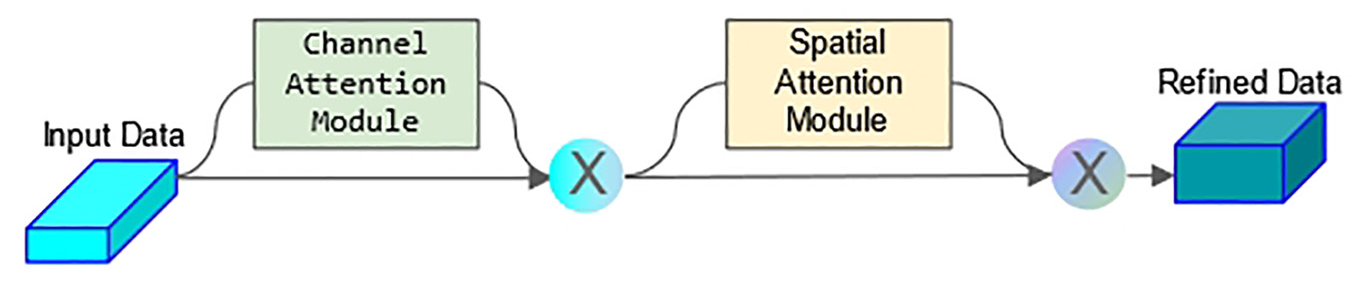
CBAM module architecture diagram.
To enhance the interpretability of the model, we integrated the Convolutional Block Attention Module (CBAM) to provide visual attention maps that highlight the key areas of focus during segmentation and classification. This improves the transparency of the model’s decision-making process, making it easier for clinicians to trust and use the model in practice.
During the training of the segmentation networks, the original U-Net model employed the Softmax output results to compute cross-entropy, utilizing it as the overarching optimization function for the network. In the context of the segmentation experiments detailed in this paper, only the portion slated for segmentation and the background region were considered. Here, 0 denotes the background area, while 1 signifies the region designated for segmentation. The semantic segmentation of kidney parts essentially entails a pixel-level binary classification task. For this, the binary cross-entropy loss (BCELoss) function is typically employed. BCELoss represents a special instance of the cross-entropy loss function, tailored exclusively for binary classification problems. The expression of the loss function is as follows:
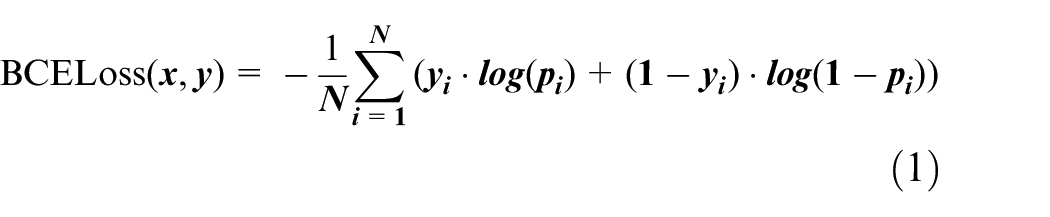
In the equation, N represents the total number of pixels; y denotes the ground truth;
In this study, the Dice loss function was adopted as a complement to the BCE loss, aiming to mitigate the impact of sample imbalance on the extraction of kidney regions. The Dice loss function is defined as:
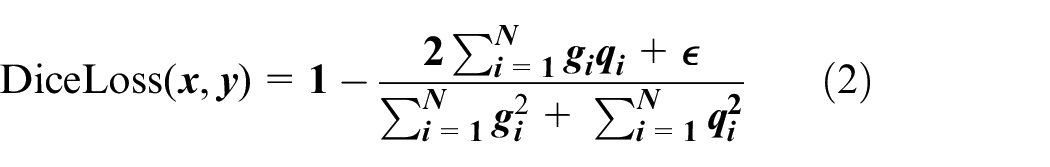
In the equation, g represents the binary mask of the ground truth; q is the binary mask of the predicted values;
In this study, the combination loss function Dice_BCEloss, which integrates Diceloss and BCEloss, is utilized to enhance the segmentation accuracy, particularly in scenarios involving small targets and sample imbalance. BCEDiceloss is defined as:
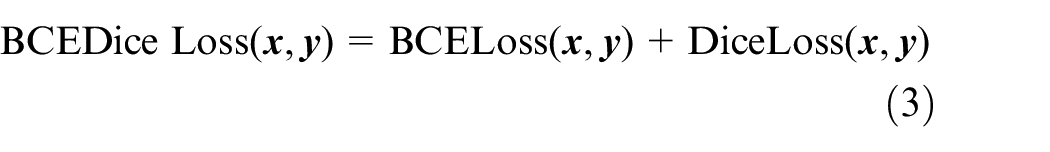
For image classification tasks, we opted to utilize the widely-used Cross-Entropy Loss, a common loss function in deep learning that’s particularly suitable for multi-classification problems. This loss function quantifies the difference between the predicted probability distribution of the model and the actual label distribution. It encourages the model to adjust its predicted probability distribution to closely match the true distribution, thus enhancing classification performance. The expression for Cross-Entropy Loss is:
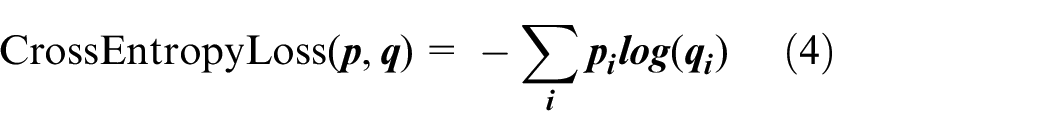
Our multitask network model is designed to balance performance and computational efficiency. The algorithm complexity of the model is mainly determined by its depth, number of layers, and number of nodes in each layer. In our model, we employ depth-wise separable convolutions and residual connections to reduce complexity while maintaining performance. Specifically, these techniques make the model more suitable for running in environments with limited computing resources by reducing the number of parameters and the amount of computation in the model, while maintaining good classification and segmentation performance.
Training and testing procedures
We conducted the model training on a server equipped with an A6000 GPU. The training set and test set were divided in a 7:3 ratio. Throughout the training process, we adjusted various parameters, including the learning rate and batch size, and conducted comparisons. The optimizer utilized was Adam (commonly used for optimization in classification and segmentation tasks).
We employed a range of evaluation metrics to assess the performance of our model, including IoU, accuracy, AUC, and others. These metrics comprehensively reflect the capabilities of our model in renal tumor ultrasound image classification and segmentation tasks. The model displayed promising performance across these indicators, attesting to its effectiveness.
Experimental setup
In this chapter, we will provide a detailed overview of the experimental setup, including model training parameters, dataset partitioning, and evaluation metrics.
Model training parameters
We set the learning rate of the model to 0.001 and the batch size to 16. To prevent overfitting, we implemented an early stopping strategy by monitoring the performance on the validation set. Training was halted when parameters such as loss and IoU on the validation set ceased to change.
Dataset partitioning
Our multitask network model is designed to balance performance and computational efficiency. To evaluate the model’s performance, we employed a fivefold cross-validation method. In each fold, the dataset was divided into a training set and a test set with a 4:1 ratio. Inside the cross-validation procedure, augment and balance only the training dataset while leaving the validation dataset intact. Such a division ensures that the model is effectively validated across different subsets of the data. The selection of the model for testing was based on the evaluation of specific validation metrics monitored during the training process, such as accuracy and loss.
Evaluation metrics
In this study, classification accuracy, sensitivity, specificity, IoU (Intersection over Union) and AUC (Area under the curve) were used to comprehensively evaluate the performance of the model. Accuracy measures the overall performance of the classification task, while sensitivity and specificity are used to assess the model’s ability to correctly identify positive and negative classes, respectively. The IoU was used to evaluate the accuracy of lesion segmentation, and the AUC was used to evaluate the overall performance of the classifier.
The Dice coefficient (also known as the Sørensen-Dice coefficient) is a similarity measure commonly used in the field of image segmentation, especially in medical image processing. It measures the similarity between two samples by calculating the size of the overlapping area between them. The DICE coefficient ranges from 0 to 1, with 1 indicating perfect overlap and 0 indicating no overlap. In this study, we use the DICE coefficient to measure the performance of our model in segmenting kidney tumor images.
Experiment and analysis
We introduced a combined multitasking network model comprising DSCResNet and AttentionNestedUNet. This mechanism’s incorporation enhances the model’s ability to capture global image dependencies, consequently boosting model performance.
Within our comparative experiments, we selected several prominent deep learning models for evaluation. These encompassed the classic U-Net, the U-Net pretrained with the VGG11 encoder (TernausNet), and the ResNeXt.
U-Net, proposed by Ronneberger et al., 29 is designed specifically for biomedical image segmentation. It features a symmetric encoder-decoder structure, where the encoder captures context information and the decoder enables precise localization. However, U-Net requires a substantial number of annotated samples during training, often challenging to procure for practical applications.
To address this limitation, Iglovikov and Shvets 30 introduced TernausNet in 2018, an improved U-Net model utilizing a VGG11 network pretrained on ImageNet for the encoder. Their experiments indicated TernausNet’s superior performance in image segmentation tasks compared to the traditional U-Net model.
ResNeXt 31 (Residual Next) is a deep convolutional neural network architecture that further extends and enhances ResNet. The key innovation of ResNeXt is its highly scalable modular structure, utilizing the concept of “cardinality” within the “block.” These branches execute parallel convolution operations and consolidate their outputs, enhancing the network’s capacity to represent data.
STUNet, 32 a variant of the traditional U-Net architecture, incorporates spatial attention mechanisms to enhance the network’s focus on relevant features within ultrasound images. This architecture optimizes feature extraction by selectively emphasizing spatial regions of interest, thus improving the segmentation accuracy. STUNet’s design is particularly adept at dealing with the complexities inherent in medical imaging, such as varying shapes and sizes of lesions. Its attention-driven approach allows for more precise segmentation, making it a valuable tool in medical image analysis, especially in scenarios where detailed delineation of anomalies is crucial.
Despite the impressive performance of these models across various tasks, they still face certain challenges when dealing with ultrasound images.
Firstly, ultrasound image quality is influenced by factors like equipment performance and operator skills, leading to considerable variability in image quality. This variability poses difficulties for a single model to handle all cases effectively. Secondly, ultrasound images exhibit low resolution and limited detail, making it challenging for models to precisely locate and segment lesions.
Experimental results
In this section, we will present the primary outcomes of the experiment, encompassing lesion object segmentation IoU, benign and malignant classification accuracy, AUC, and other metrics.
Object segmentation results
Our model effectively performed lesion object segmentation on kidney tumor ultrasound images. After evaluating the test set, our model achieved exceptional results in terms of IoU. The average IoU reached 0.942, signifying the model’s accurate localization and segmentation of kidney tumor lesions, which significantly aids clinicians in diagnosis.
Benign and malignant classification results
Our model successfully classified ultrasound images of kidney tumors as benign or malignant, facilitating lesion nature determination for medical practitioners. On the test set, the classification accuracy of the model in distinguishing benign and malignant lesions was 81.8%, the sensitivity was 82.2%, and the specificity was 80.6%. This demonstrates the model’s robust capability in assisting physicians by accurately distinguishing between benign and malignant renal tumors, thereby significantly improving the precision of clinical diagnoses.
Our experimental outcomes underscore the superiority of our model over U-Net, TernausNet, and STUNet models in terms of IoU.
Comparative experiments
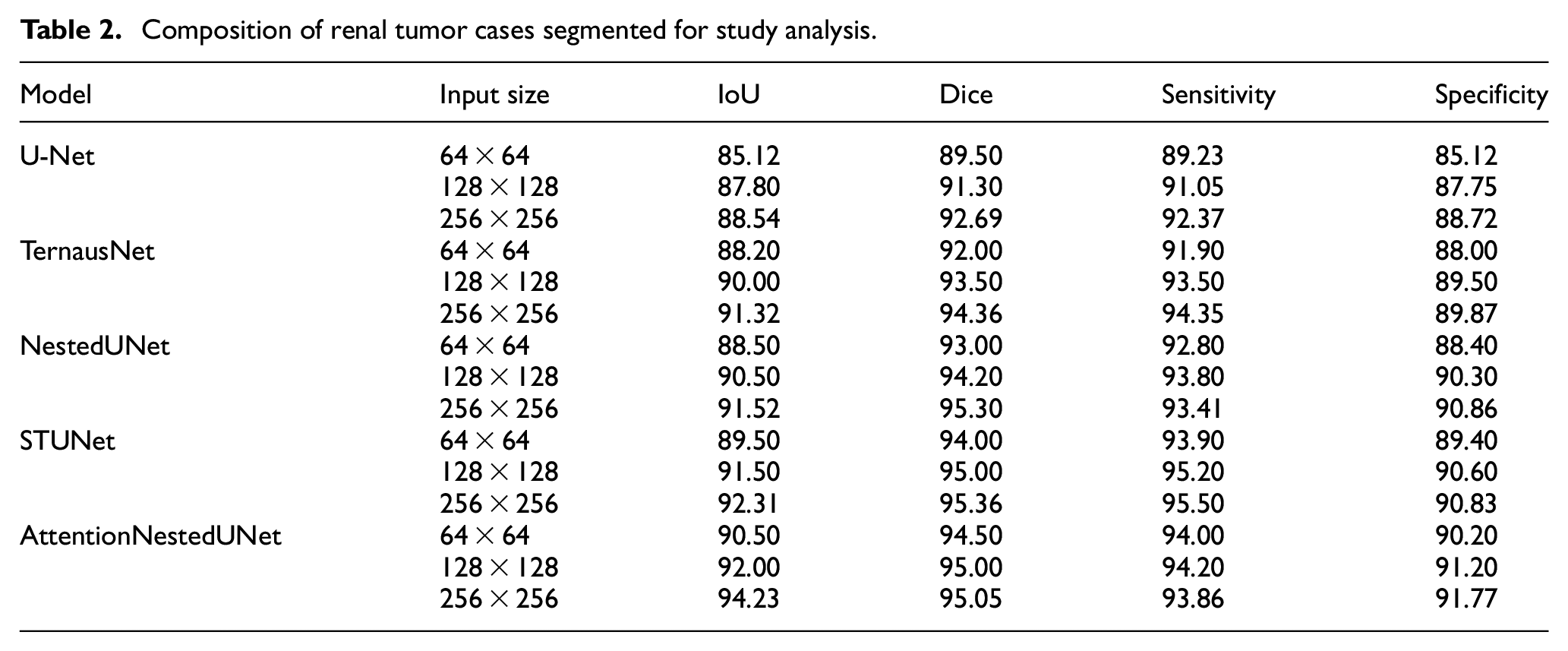
In order to validate the superiority of our proposed model, we conducted comparative experiments against traditional U-Net backbone networks, mainstream net-works like ResNet, SOTA network like STUNet and also compared our classification results with evaluations from radiologists. To establish a comprehensive and clinically relevant evaluation of our model, we incorporated the judgments of both junior and senior radiologists into our classification metrics. This approach allows us to bench-mark the model’s performance against human experts, providing a more nuanced understanding of its effectiveness and reliability in clinical settings. The results of these comparative experiments demonstrate that our model outperform1s in both lesion object segmentation and benign/malignant classification tasks, as shown in Table 2.
Composition of renal tumor cases segmented for study analysis.
The renal tumor ultrasound images were independently reviewed and annotated by a group of junior and senior radiologists. Their assessments included the identification of lesions and the classification of each as benign or malignant, based on their expertise and diagnostic criteria. The model’s performance was compared with the radiologists’ assessments to determine its relative accuracy and reliability. This comparison helps in identifying areas where the model excels or falls short compared to human judgment. Insights gained from this comparison are crucial in understanding the practical implications of deploying the model in a clinical setting. It also helps in identifying specific cases or conditions where the model might require further refinement or where it could potentially aid radiologists in making more accurate diagnoses.
We evaluated the model’s performance under different input sizes (64 × 64, 128 × 128, 256 × 256) to assess its performance under various input conditions. In addition, we compared our model’s performance with that of several existing models, including U-Net, TernausNet, ResNeXt, and STUNet, using a variety of evaluation metrics such as IoU, accuracy, and AUC. The results indicate that our proposed DSCResNet combined with AttentionNestedUNet outperforms across all evaluated metrics, particularly in lesion segmentation and benign/malignant classification tasks. These comparative experiments confirm the advantages of our model over traditional approaches, as shown in Table 3, highlighting its effectiveness in processing renal tumor ultrasound images for clinical diagnostic support.
Classification results of renal tumor cases with comparative analysis.
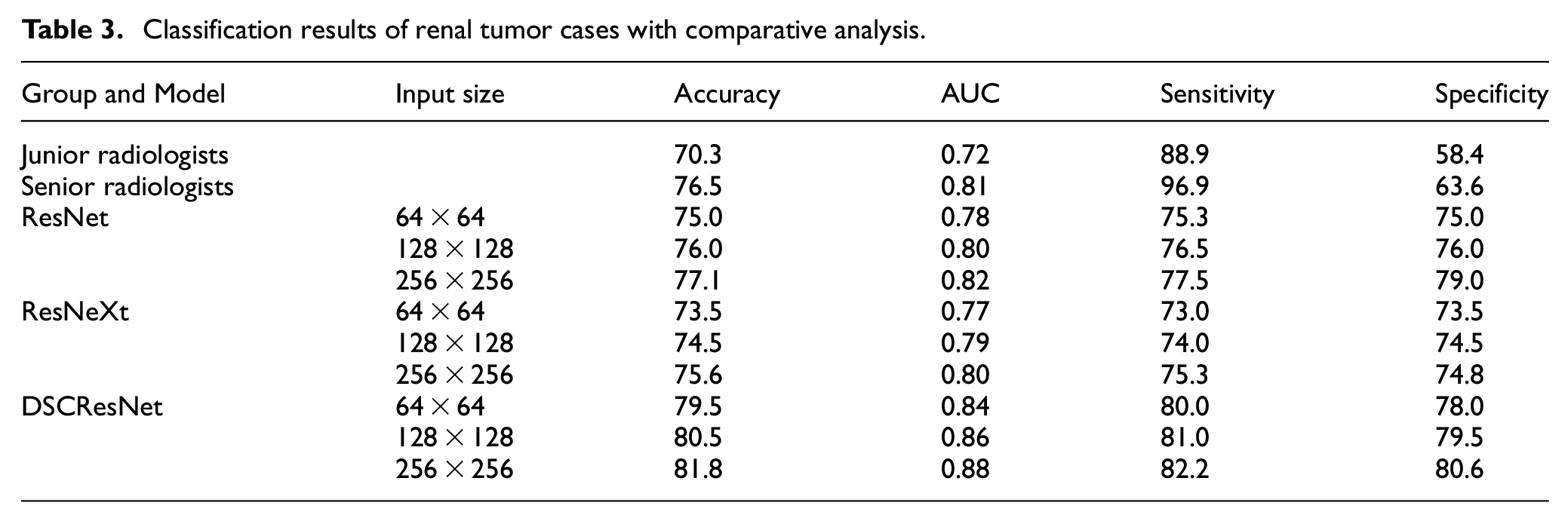
This detailed comparison helps to demonstrate more clearly the superiority of our proposed approach under various conditions and provides strong evidence to support it.
Result analysis
Our experimental results clearly demonstrate that the proposed joint multitask network model, which combines DSCResNet and AttentionNestedUNet, achieves significant performance improvements in the domain of renal tumor ultrasound image processing, as shown in Figures 3 and 4. The incorporation of DSCResNet allows for enhanced feature learning, reduced parameter count, improved training efficiency, faster inference speeds, and reduced overfitting. Additionally, the mixed domain attention mechanism facilitates the model in focusing on crucial regions within images, leading to improved accuracy in both image segmentation and classification tasks.
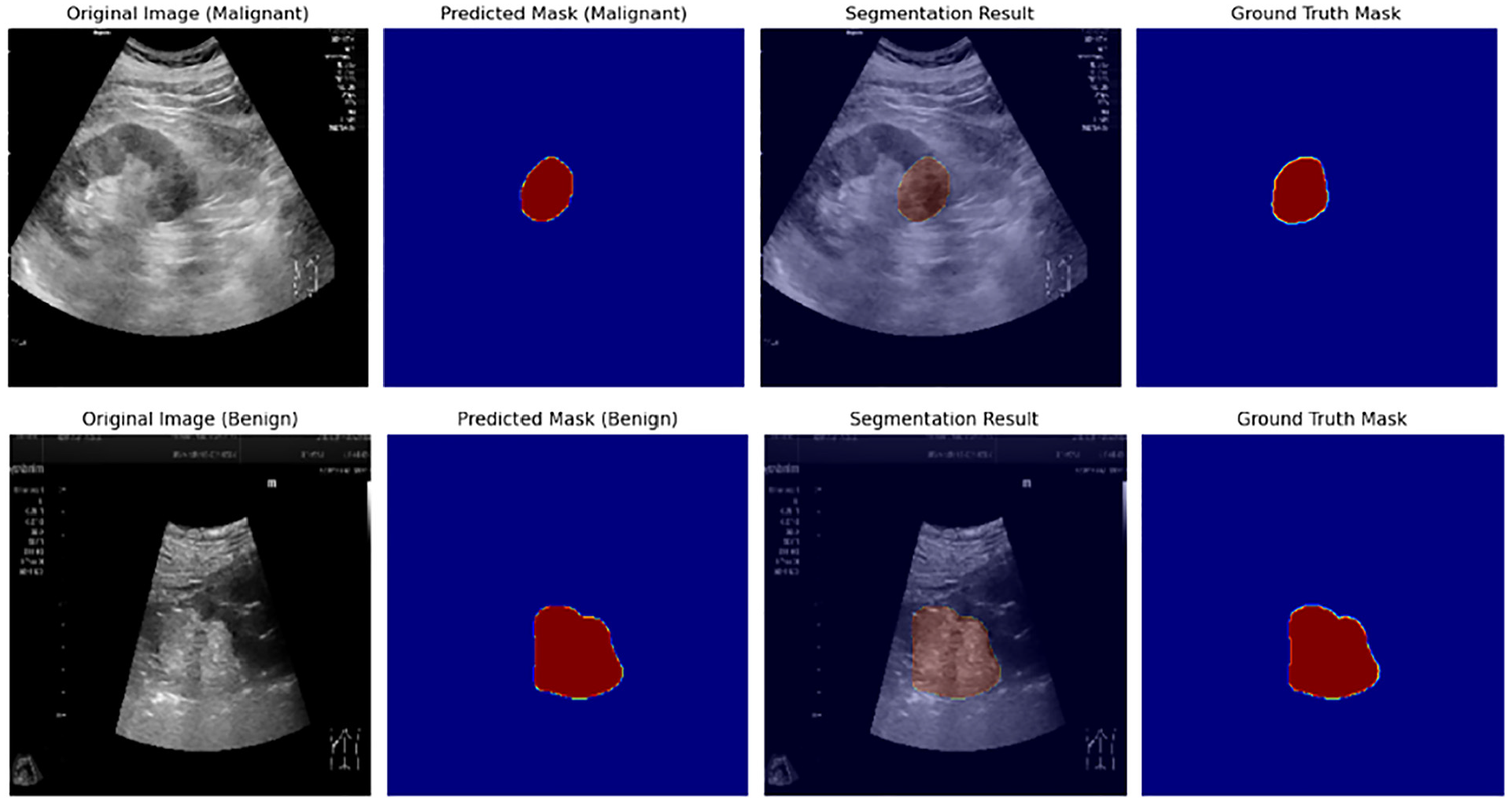
Model output results (classification and segmentation results).
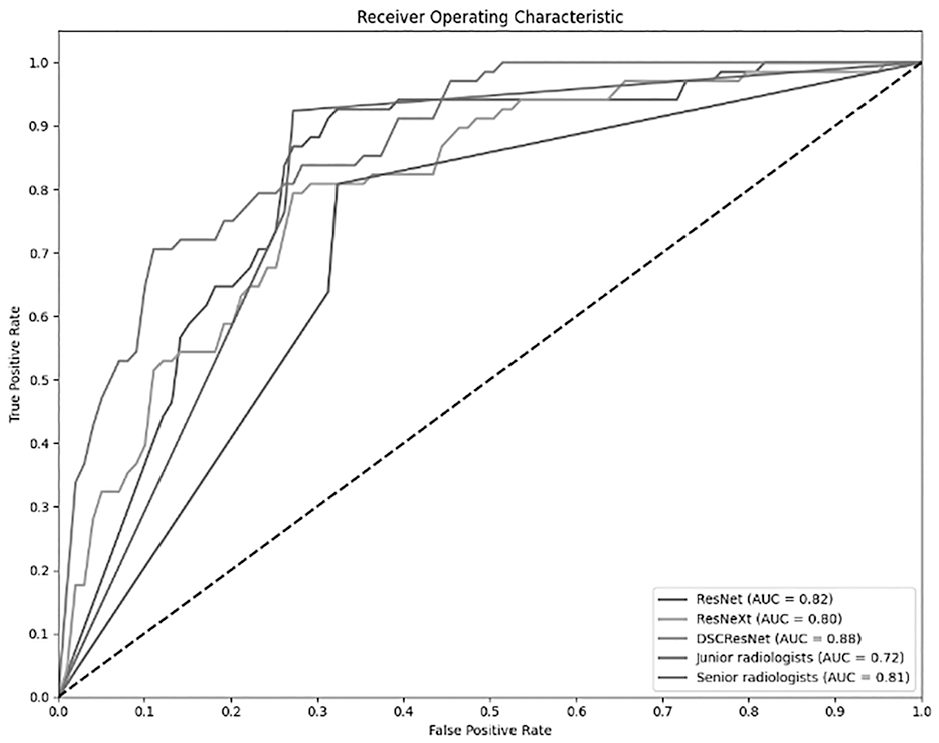
Model output results (classification results).
Computation of complexity
In order to fully evaluate the utility and efficiency of AttentionNestedUNet networks, we performed a detailed algorithm complexity analysis as well as a probability-based time and effort calculation analysis
Algorithmic complexity
The AttentionNestedUNet network consists of multiple convolutional layers, pooling layers, batch normalization layers, and the Convolutional Block Attention Module (CBAM). The computational complexity of each layer is determined by its parameters and the dimensions of the input feature maps. Table 4 summarizes the computational complexity of key layers in the network:
Composition of algorithm complexity.
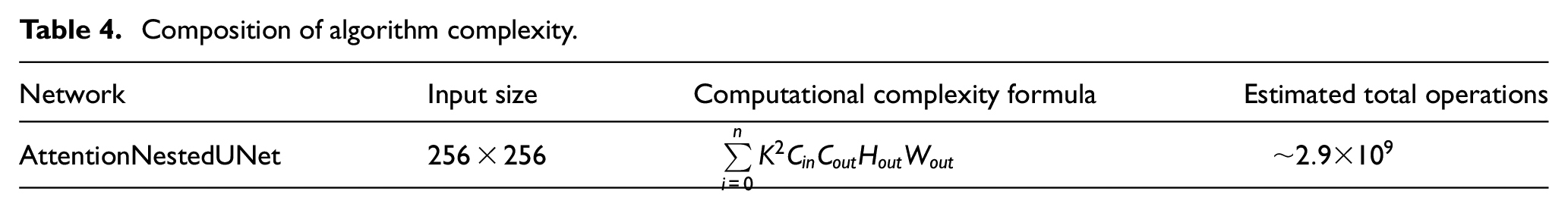
Where:
i, n: The number of layers
Probability-based time and computational analysis
The AttentionNestedUNet network was trained on a server equipped with an NVIDIA A6000 GPU, using a dataset of 3600 images with a batch size of 8. Experimental findings revealed that one epoch took approximately 2 min and 40 s. Based on this data, the average processing time per image is estimated to be around 0.044 s. This indicates that the AttentionNestedUNet exhibits relatively high efficiency in processing large volumes of data, which is critical for real-time or near-real-time medical image processing applications.
Discussion
The objective of this work is to apply advanced deep learning techniques to the processing and analysis of grayscale ultrasound images of renal tumors. We introduced a joint multitask network combining DSCResNet and AttentionNestedUNet, aiming for effective lesion segmentation and benign/malignant classification. Our model significantly advances current state-of-the-art techniques, particularly outperforming STUNet, with AttentionNestedUNet achieving an IoU of 94.23%, compared to STUNet’s 92.31%. This improvement is mainly due to the integration of the Convolutional Block Attention Module (CBAM), which enhances focus on important regions, resulting in more accurate segmentation. The combination of DSCResNet, with its deep separable convolutions to reduce computational complexity, and AttentionNestedUNet allows simultaneous segmentation and classification, enabling superior lesion localization and analysis.
Comparative experiments demonstrate that our model outperforms traditional STUNet and ResNet networks in both segmentation and classification tasks, showing consistent improvements across multiple evaluation metrics. The residual connections integrated into the AttentionNestedUNet effectively mitigate gradient vanishing, ensuring deeper features are propagated and stabilizing the training process. This joint approach also addresses challenges inherent to ultrasound images, such as blurred boundaries and noise, thereby providing robust performance even under low-resolution and variable image quality conditions. These innovations not only contribute to advancing renal ultrasound imaging research but also offer valuable support for clinical diagnostic applications.
In segmentation and classification tasks, the main challenges lie in the noise in ultrasound images and blurred lesion boundaries. We effectively overcome these challenges by introducing CBAM and residual connections to enhance the extraction of key features and ensure gradient stability.The results obtained from the fivefold cross-validation underline the robustness and consistency of our model. Across all fivefolds, the model demonstrated a high degree of accuracy in lesion segmentation and tumor classification, with the Intersection over Union (IoU) and classification accuracy metrics remaining consistently high. This consistency across different data subsets underscores the model’s reliability and effectiveness in diagnosing renal tumors from ultrasound images. The use of cross-validation has also provided insights into the generalizability of our model across varied data samples.
While our model outperforms existing solutions in several areas, it is important to recognize its limitations. Its performance, particularly in classifying renal tumors, relies on the quality and diversity of the training dataset. Additionally, our experiments were based on data from a single organ, which may limit the model’s generalizability to other contexts. Future work will focus on expanding the dataset to include data from different devices and imaging conditions, and incorporating multicenter data to improve robustness and broader applicability. Interpretability is another challenge, particularly in clinical settings where understanding model decisions is essential. The use of CBAM and visual attention maps can help clarify the features driving the model’s predictions, enhancing its transparency and clinical usefulness. While the current focus is on renal tumors, the model’s architecture is adaptable to other organs and tumor types. Expanding its scope could further increase its clinical impact and applicability across a wider range of medical diagnoses.
While the dataset used in this study is diverse, it still underrepresents certain scenarios, such as rare tumor types and images with specific quality issues. Future work could address this by using synthetic data augmentation, such as Generative Adversarial Networks (GANs), to simulate these conditions and improve the model’s robustness. This could also help adapt the model to different imaging devices and settings, enhancing its generalizability in clinical practice. Additionally, integrating multimodal data, such as ultrasound, CT, MRI, and contrast-enhanced ultrasound (CEUS), could significantly boost the model’s accuracy, particularly in distinguishing benign from malignant tumors, by leveraging complementary information from various imaging modalities. Combining CEUS with conventional ultrasound provides functional insights, further enhancing tumor detection and characterization.
Moreover, incorporating additional types of attention mechanisms or exploring other advanced deep learning architectures could further enhance the model’s performance. Future studies could also delve into the interpretability of the model’s decisions, providing insights into the features most relevant for classification and segmentation tasks. Although our experimental results show that the proposed model has significant advantages in renal tumor ultrasound image diagnosis, we realize that the model still needs further improvement in multicenter validation. At present, we have not yet multicenter data validation, but in the future study, we plan to collect data from different clinical centers, in order to further verify the universality and robustness of the model. This will help to ensure the robustness of the model under different equipment and operating conditions, enhancing its application value in real clinical settings.
Recently, robot-assisted partial nephrectomy (RAPN) has shown good results in the treatment of complex renal hilar masses. 33 These advances highlight the importance of precise surgical planning and execution to preserve renal function without compromising the outcome of cancer treatment.
Our AI-driven model has the potential to enhance surgical procedures by providing detailed and accurate image analysis. In the operating room, such models could assist surgeons in better identifying tumor boundaries, optimizing surgical decisions, reducing intraoperative risks, and improving both postoperative functional and oncological outcomes. While the model shows high accuracy in controlled testing environments, its real-time performance in clinical settings has yet to be tested. Future studies should focus on validating the model in live clinical workflows to evaluate its effectiveness under real-world conditions, including varying image quality and clinical scenarios. We hope that, in the future, our model will play a significant role in enhancing RAPN surgeries and improving patient outcomes.
In our discussion of contemporary methods in medical image processing, it is pertinent to highlight the recent development of MedSAM, 34 a novel adaptation of the Segment Anything Model (SAM) 35 for medical imaging. MedSAM represents a significant leap in medical image segmentation, offering a universal tool for segmenting a variety of medical targets. This model, which extends SAM’s capabilities to medical images, was developed through the curation of a large-scale dataset encompassing over 200,000 masks across 11 different modalities.
Medical image segmentation, a fundamental task in medical imaging analysis, involves identifying and delineating regions of interest (ROI) in various medical images, such as organs, lesions, and tissues. MedSAM addresses the limitations of current models that are often tailored to specific imaging modalities. Through comprehensive experiments on 21 3D segmentation tasks and 9 2D segmentation tasks, MedSAM demonstrated superior performance with an average Dice Similarity Coefficient (DSC) improvement of 22.5% and 17.6% on 3D and 2D segmentation tasks, respectively, compared to the default SAM model.
The inclusion of MedSAM in our discussion provides a broader perspective on the state of the art technologies in medical image processing. It highlights the potential of leveraging large-scale supervised training and advanced network architectures to improve segmentation performance in medical imaging, which is a key aspect of our research as well.
While our model has demonstrated promising performance in controlled testing environments, its effectiveness in real-time clinical workflows remains to be validated. To assess its practical utility, future work will focus on conducting prospective clinical studies that involve deploying the model in live clinical settings. Such validation will allow us to evaluate the model’s performance under real-world conditions, where variables such as image quality, diverse patient profiles, and varied imaging settings can significantly impact outcomes.
Conclusion
This study introduces a joint multitask network model that combines DSCResNet with AttentionNestedUNet. The model is designed to address the tasks of lesion object segmentation and benign/malignant classification in renal tumor ultrasound grayscale images. Through the presentation and analysis of experimental results, we have confirmed the remarkable performance enhancements achieved by this model in the realm of renal tumor ultrasound image processing. The model excels at precisely locating and segmenting lesions while also demonstrating impressive capabilities in the classification of benign and malignant cases. We have engaged in discussions concerning the model’s strengths and limitations, as well as contemplated future research avenues. We propose future research directions, including expanding the dataset to improve the robustness and accuracy of the model, exploring more attention mechanisms, further improving model performance, and validation in real-world clinical Settings. These improvements will further promote the development of medical image processing field, and provide more accurate and effective support for the clinical diagnosis.
Footnotes
Authors’ contributions
Weiping Zhang and Long Gao contributed equally to this paper. Weiping Zhang and Long Gao were responsible for the article writing and ultrasound image processing; Weiping Zhang and Li Chen were in charge of cases collection; All authors have read and agreed to the published version of the manuscript.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was funded by Science and Technology Project of Jiangxi Provincial Administration of Traditional Chinese Medicine (no. 2021A060), and Teaching Reform and Research Project of Jiangxi Province (no. JXJG-22-1-60), and Health Commission Science and Technology Plan of Jiangxi Province (no. 202510031).
Data availability statement
The data used to support the findings of this study are available from the corresponding author upon request.
