Abstract
Background
The foetal anomaly screening (FAS) programme in England screens for trisomies 21, 18 and 13 using a combination of maternal age and biochemical and ultrasound markers in a multivariate Gaussian approach to calculate the chance of a foetal trisomy. The screened population is divided into higher and lower chance categories using a 1 in 150 cut-off. A truncation of 1 in 5000 is used for the lowest chance results.
A series of postal strikes resulted in delay to specimen delivery and necessitated rebleeding of hundreds of affected clients across England. The effect of delay in processing specimens was investigated based on the final chance reported, rather than the effect on the individual biomarkers used in the chance calculation.
Methods
The FAS chances from samples delayed in postal strikes were compared against those from repeat samples from the same patients to determine the absolute effect on the final chance reported rather than the individual biomarkers.
Results
A total of 119 of 120 chances reported remained in the same high/low chance category.
Conclusion
Marker concentrations are affected by delayed separation, but the effect on calculated chance is only significant in those close to the 1 in 150 cut-off. There is little benefit in rebleeding clients at very low chance or at very high chance as this leads to a delayed result and the possibility of missed screening for some affected pregnancies.
Introduction
Screening for trisomies 21, 18 and 13 has been offered by the NHS in England since 2015 as part of the foetal anomaly screening programme (FASP). Birmingham Women’s Hospital is the largest foetal anomaly screening laboratory in England, providing screening for around 100,000 pregnancies each year.
The tests used to screen depend on the gestation at which the sample is collected. Between 10 and 14 weeks gestation (combined test), the biomarkers used are pregnancy-associated plasma protein A (PAPP-A) and free beta subunit of human chorionic gonadotrophin (fbhCG) and an ultrasound marker, nuchal translucency (NT). Chance is calculated for trisomy 21 and a joint chance for trisomies 13 and 18.
Between 14 and 20 weeks gestation (quadruple test), the biomarkers measured are alpha feto-protein (AFP), human chorionic gonadotrophin (hCG), Inhibin A (InhA) and unconjugated estriol (uE3). Chance is calculated only for trisomy 21.
To standardise concentrations between individuals, biomarkers are corrected for gestational age, maternal weight, maternal family origin, diabetes and smoking and reported as multiples of a median concentration (MoM). Biomarker MoM values are combined using a multivariate Gaussian approach to provide a chance based on the markers and maternal age. 1
Combined testing is expected to detect 86% of all trisomy 21 cases and 80% of trisomy 18 and 13 cases and quadruple testing 82% of trisomy 21 cases. 2 Approximately 85% of clients have the combined test.
If the calculated chance is 1 in 150 or greater, further testing is offered.
Background
Serum analytes can be affected by pre-analytical time (PAT). Prolonged PAT can lead to individual biomarker variation. The accuracy of FAS chance calculation will depend on the cumulative effect of the PAT on all biomarkers. 3
FASP states the acceptable PAT for unseparated whole blood specimens is 48 h at 20°C, or 72 h at 20°C for separated serum specimens for trisomy chance calculation to be valid. 4
Postal strikes caused increased PAT for significant numbers of specimens which were delayed in transit beyond 48 h, leading to 194 clients requiring rebleed. This delayed results and many clients became too late for first trimester screening. Nine did not re-attend and so were not screened.
Methods
The 194 rejected specimens were all analysed for FAS biomarkers, a chance calculated but not reported and the results compared to those from the repeat specimen. Of the 194 samples, 65 were repeated using the same combination of biomarkers (10 quadruple tests and 55 combined tests). The remainder were either too late for screening or returned too late for the same test combination to be performed and were therefore not included in the study. Analysis of AFP, hCG, fbhCG and PAPP-A was performed on the Roche Cobas 6000 platform. Analysis of InhA and uE3 was performed on the Beckman Access II platform.
For the 55 combined test repeats a chance was calculated for T21 and T18/13 (110 chances) and for the 10 quad tests only a chance for T21 was calculated (10 chances), meaning overall 120 chance results were recalculated and compared. Chance calculation was performed using SsdwLab software (v6.3).
Results
Specimen delay (PAT)
The original specimens were between 3 and 10 days old on receipt. 71% (39) were 4 days old and 20% (11) 7 days old. All repeat specimens were received within 48 h of collection.
Combined testing
T21 chance
None changed from low to higher chance or vice versa. 41/55 chances reported did not change numerically as both the original and repeat chance were below the truncation of 1 in 5000 (Figure 1). Comparison of chance for specimen delayed in transit (circle) and repeat specimen received within acceptable time frame (cross).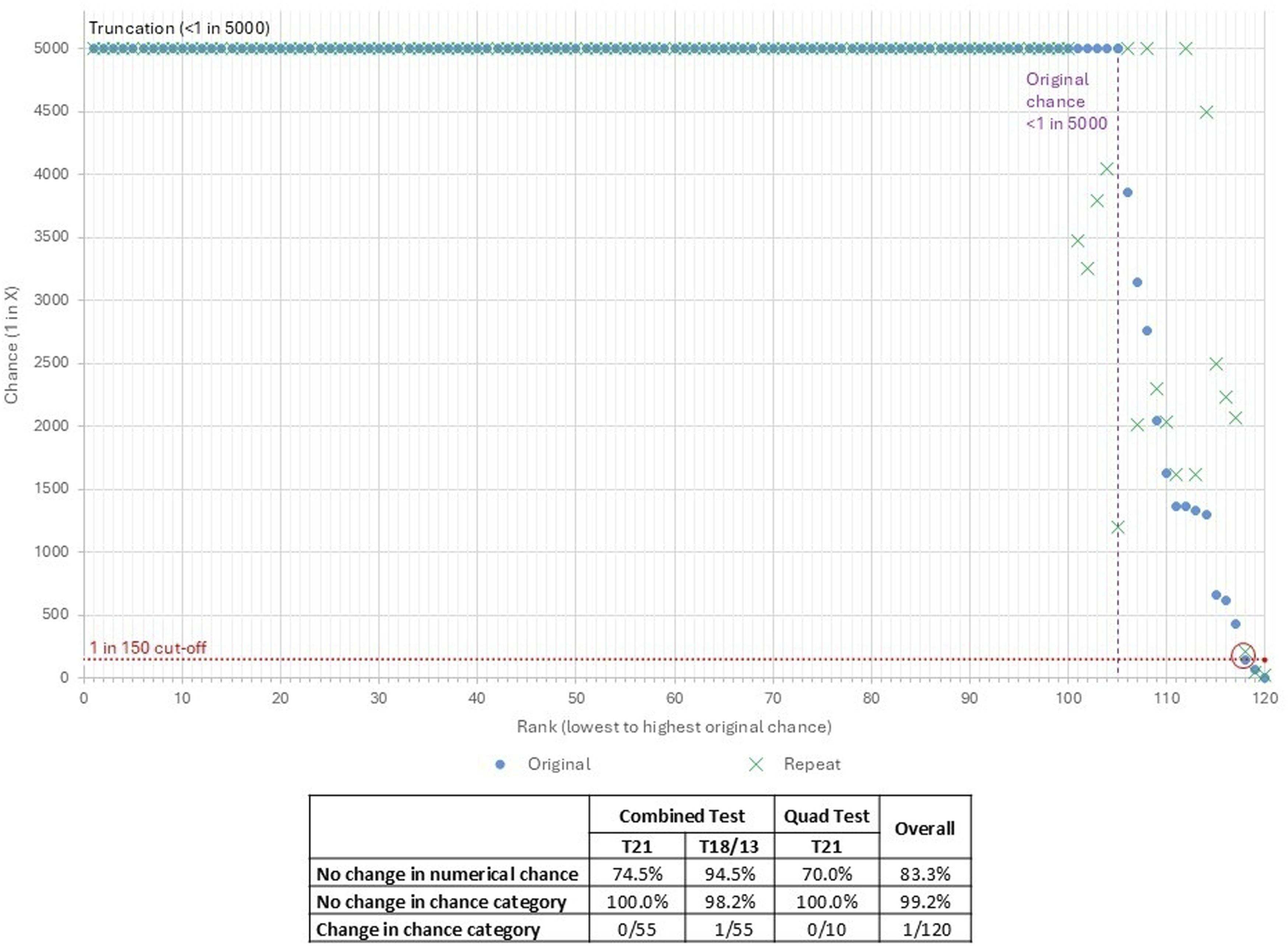
T18/13 chance
95% of chances reported did not change numerically as both the original and repeat chances were below the truncation of 1 in 5000.
One T18/13 chance changed from 1 in 145 (increased chance) on the original specimen to 1 in 208 (low chance) on the repeat specimen (circled in Figure 1). The T21 chance was increased for both specimens and the client was therefore offered further testing. The pregnancy was not affected.
Quadruple testing
None changed from low to higher chance or vice versa. 70% of chances reported did not change numerically as both the original and repeat were below the truncation of 1 in 5000 (Figure 1).
Discussion
A total of 119 of the 120 chances calculated did not change classification (i.e. remained lower or higher chance). A total of 48 of the 65 clients (73.8%) had every chance remain below 1 in 5000 (the lower limit of chance reported) in both samples. The single case where the category changed from higher to lower chance (circled in Figure 1) was close to the clinical cut-off, and the other factors contributing to the imprecision of the chance calculation should be considered. There were no affected pregnancies in this cohort.
Changes in chance are not only partly due to the pre-analytical conditions but also partly due to other factors including the relatively high imprecision of a chance caused by combining several different assays. The discriminatory powers of the biomarkers change with gestational age.5,6 This means that the chance calculated will change even if both samples are analysed under the same conditions, as the biomarkers may not remain stable relative to the median value through pregnancy.
Conclusion
Chance category was not impacted by PAT in most cases. There was no benefit in obtaining a repeat specimen where the chance was less than 1 in 5000 and the original sample less than 7 days old (left of the dashed line in Figure 1). Further work should be undertaken to determine if PAT limits can be extended for the combined test where the most impactful marker (NT) is not biochemical. Many repeat specimens were collected too late to use the NT measurement and so had the quad test, which has poorer performance. Extending the acceptable PAT for the combined test would enable more clients to have this better performing test in the event of delays, such as postal strikes.
Footnotes
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Ethical approval
Ethical Committee approval was not required for this retrospective review of anonymised data.
Guarantor
IM.
Contributorship
IM and SH conceived the study. PLB was involved in data analysis. IM wrote the first draft of the manuscript. All authors reviewed and edited the manuscript. All authors approved the final version of the manuscript.
