Abstract
Objective
To evaluate the application effect of SPRi monoclonal antibody (mAb) chip in the detection of influenza virus antigen in complex mixtures.
Methods
A total of 115 strains of mAbs against different subtypes (H1N1, H5N1, A1, A3, B, H7N9, H9N2, and H3N2) of influenza virus were prepared. The chip of mAbs against influenza virus was prepared by surface plasmonic resonance imaging (SPRi) technology, which was used for the detection of influenza virus supernatant, and compared with the traditional antigen capture ELISA method.
Results
Comparative studies have shown that traditional antigen capture ELISA methods have a higher sensitivity (86.8% (46/53)
Conclusion
The SPRi mAb chip can be used for the detection of specific pathogenic microorganisms or viral proteins in complex mixtures such as influenza virus supernatant. It has significant advantages of label free, real-time, high-throughput, and good specificity, and can play an important role in disease diagnosis and infectious disease prevention and control.
Keywords
Introduction
The infection of pathogenic microorganisms and viruses poses a major threat to human life all over the world, and their constant mutation, evolution and spread have brought great difficulties to the disease prevention and control of infection. Therefore, how to sensitively and specifically detect and quantify specific pathogenic microorganisms and viral proteins in complex mixtures has become an important challenge for proteomics. 1
Among numerous biochemical detection technologies, microarray biochip technology, with its advantages of high throughput, high sensitivity, and low sample consumption, has been widely studied and applied by researchers to search for disease markers,2,3 protein interaction network, 4 disease diagnosis,5–7 glycosylation analysis, 8 and DNA interaction. 9 Surface plasmon resonance imaging (SPRi), as an optical biosensing technology, is based on the Surface Plasmon resonance phenomenon, 10 that is, the interaction of photoelectrons, and under special conditions, generating coupling resonance and the physical phenomenon of energy transfer. SPRi technology is ideal for label-free, high-throughput and real-time detection of biomolecules (including DNA/RNA,11,12 proteins or antibodies, 13 peptides,14,15 polysaccharides, and small molecule drugs16,17) interactions. The integration of SPRi technology with other biochemical technologies can provide an excellent solution to some of the most important biological problems currently being investigated.
The technology has perfect compatibility with microarray chip technology and can be used for real-time detection of molecular interactions at each sample point on the chip surface. Therefore, we developed a SPRi microarray chip and used it to detect and characterize interactions between influenza virus antigens and antibody proteins. The biological characteristics of influenza virus antigens were revealed by measuring the affinity difference of influenza virus monoclonal antibodies (mAbs) to capture virus antigens on the biochip surface. The combination of microarray biochip technology and SPRi technology has innovative significance for improving the research efficiency of protein and other biomolecular interaction, and accelerating the development of biodiagnostic products and other important fields.
Materials and methods
Antigens
The H1N1 influenza virus split vaccine (2009; SFDA approval no. S20090015) was obtained from Hualan Biological Bacterin Co., Ltd (Henan, China). H5N1 (Re-5+Re-4) avian influenza virus, seasonal influenza A1 (A/Brisbane/59/2007), A3 (A/Victoria/210/2009 NYMCX-187), Influenza Virus (B/Qingdao/102/91), H7N9 (A/Shanghai/1/2013), H9N2 (A/Hong Kong/1073/99), and H3N2 (A/New York/392/2004) virus strains were purchased from Qingdao Yibang Biotech Co. Ltd. (Qingdao, China). Influenza virus Ag308 (A/Hong Kong/308/2014 (H1N1)), Ag501 (A/Beijing/501/2009 (H1N1)), and PR8 (A/Puerto Rico/8/34(H1N1)) were all preserved in this laboratory.
Reagents
Myeloma SP 2/0 cells, DOME and RPMI 1640 medium were purchased from Hyclone, PEG 1500 from Roche, fetal bovine serum (FBS) from Gibco, and horseradish peroxidase-labeled sheep anti-mouse secondary antibody from Thermo. The antibody subtype identification kit was purchased from Southern Biotech, and BSA, ethanolamine, Tween-20, tetramethylbenzidine (TMB) and Freund adjuvants were purchased from Sigma. SPF BALB/c female mice aged 8 weeks, weighing 18–22g, were purchased from Laboratory Animal Center, Xi’an Jiaotong University (license No.: SCXK (Shaanxi) 2018-001).
Preparation and characterization of the monoclonal antibodies
Freund’s complete adjuvant was emulsified with H1N1, H5N1, A1, A3, B, H7N9, H9N2, and H3N2 subtypes of influenza viruses in a 1:1 ratio, respectively, and first immunized BALB/c mice by subcutaneous injection at a dosage of 25 μg/100 μL per mouse. Freund’s incomplete adjuvant was used for secondary immunization, the same dose and method as for primary immunization. Three days before fusion, additional intraperitoneal immunization was performed with influenza virus antigen. Myeloma SP2/0 cells in serum-free RPMI 1640 medium, reaching a logarithmic growth phase with a cell count of 109 cells; The spleen of mice immunized with influenza virus was extracted aseptically, ground and sieved, and culture them in serum-free RPMI 1640 medium until the number of splenic cells reaches 108. The prepared myeloma cell SP2/0 suspension was mixed with immune mouse spleen cell suspension at a ratio of 1:8–1:10 for cell fusion and screening of positive clones. Positive hybridoma cells were screened by indirect ELISA and clonized by limited dilution to establish stable hybridoma cell lines, 18 expanded culture and cryopreserved. BALB/c mice were intraperitoneally injected with 0.5 mL medical liquid paraffin, after 1 week, hybridoma cell lines with a number of 106 were intraperitoneally inoculated, and ascites could be collected and purified after another week. 19
Then, the ascites fluid was purified by the ammonium sulfate coprecipitation method. 20 Antibody subclasses were identified by SBA Clonotyping System/HRP kit. The hemagglutination inhibition activity of mAbs was determined by erythrocyte hemagglutination (HA) and hemagglutination inhibition (HI) tests. The indirect ELISA method was used to identify the cross-reactivity of purified ascites with other subtypes of influenza viruses, adenovirus, syncytial virus, parainfluenza virus, Echovirus, Coxsackie virus, enterovirus and other microorganisms outside of the corresponding antigen.
Construction of the SPRi microarrays chip surface
We use a novel block copolymer brush with a controllable layered structure to construct the chip surface. 21 The thickness, density, and composition of the polymer brush can be precisely regulated within the detection range. That is, the first layer of the polymer brush is prepared on a gold substrate by surface trigger electron activation regenerated atomic transfer radical polymerization. Based on this, the second layer is prepared by polymerization of glycidyl methacrylate (GMA) monomer.
Preparation of the SPRi microarray chips for influenza virus antibodies
The 115 different influenza virus antibodies and BSA (negative control) were diluted to 1 mg/mL with 10 mM PBS (pH 7.4), then 1 mg/mL antibody and 1 mg/mL BSA were spotted on the substrate of SPRi chip in the form of microarray using a contact mode spotting robot (AD3210, BioDot, USA) and dried overnight in a vacuum drying oven. The next day, them were washed in turn with phosphate buffered solution (PBST) supplemented with Tween-20 at 0.05% volume ratio, PBS, and deionized water three times, respectively, and blown dried with nitrogen. Subsequently, the surface was closed with ethanolamine. Finally, them was washed using PBST with 0.05% Tween-20, PBS, and deionized water three times, respectively, then dried with nitrogen and reserved. Each antibody of the microarray chip had five replicate spots to reduce the error. Three chips were prepared to contrast the repeatability between the different chips.
Detection of influenza virus binding to antibody by SPRi microarray
To obtain a typical binding curve (A curve plotted based on the strength (response value/RU) and time (s) of each antigen binding to antibody), the diluted protein samples were circulated and bound at 2 μL/s flow rate for 300 seconds, then dissociated with PBS at 2 μL/s for 300 seconds, and regenerated at 3 μL/s with a volume ratio of 1:100 phosphoric acid for 200 seconds to remove specific adsorption or weak binding. For the detection of influenza virus supernatants, the culture supernatants of three influenza viruses (Ag308, Ag501 and PR8) with a hemagglutinant titer of 1:320 were diluted 5 times in DMEM medium and detected in SPRi instrument (PlexArray HT 100, Suzhou, China). Sensitivity correction and refractive index unit conversion were performed for each sample point on the surface by using PBS solution and PBS solution of glycerol with a mass to volume ratio of 1% with known refractive index coefficient variation (1200 RU), and all the resulting SPRi binding signals (arbitrary unit, AU) were converted to the standard unit (resonance unit, RU) of refractive index according to the formula (1AU = 200RU).
Detection of influenza virus binding to antibody by indirect ELISA
Three influenza viruses (Ag308, Ag501, and PR8) were coated with 2 μg/mL coating solution, washed with PBST for three times, then added 100 μL sealed solution, and incubated in 37 °C water bath for 1 h. The plates were washed with PBST for three times, and 115 kinds of mAb supernatants were added to each well, respectively, and 100 μL culture supernatant of Sp2/0 cells was added to the corresponding well as the negative control, and incubated at 37°C for 1 h. The plates were washed with PBST for 3 times, and 100 μL of sheep anti-mouse enzyme-labeled secondary antibody in 2500 times dilution was added, and continued to incubate at 37 °C for 1 h. After washing the plate with PBST for 5 times, add color developing liquid A and B. After 5 min, color development was terminated with 2 mol/L sulfuric acid, and the OD value was read at 450 nm using a microplate reader.
Data analysis
The SPRi test data was collected and analyzed using commercial data processing software (SPR Data Analysis Model, Plexera, USA). The
Results
Preparation and characterization of monoclonal antibodies
Monoclonal antibody cell lines against influenza virus antigens of different subtypes were obtained by conventional hybridoma technology, and the antibodies could still be secreted stably after 3 months of passage. A total of 115 strains of mAbs were selected for the preparation of high-throughput influenza virus antibody microarray chips based on SPRi technology by naming, testing subclass, hemagglutination activity, specificity and cross-reactivity, among which 25 strains derived from H1N1 influenza virus, 35 strains from H5N1 influenza virus, 4 mAbs from seasonal A1 influenza virus, 8 mAbs from A3 influenza virus, 17 mAbs from influenza B virus, 9 mAbs from H1N1 influenza virus (PR8), 15 mAbs from H7N9 influenza virus, and 2 mAbs from H9N2 influenza virus.
Preparation of high-throughput antibody microarray chips based on SPRi
After identification and screening of the prepared mAbs against influenza virus, 115 antibodies against different influenza virus epitopes and BSA (negative control) were prepared into microarray on the surface of SPRi chip (Figure 1) and applied in the detection of influenza virus. Printed image of SPRi influenza virus mAb microarray. It was confirmed that influenza virus mAbs immobilize to the surface of SPRi chip.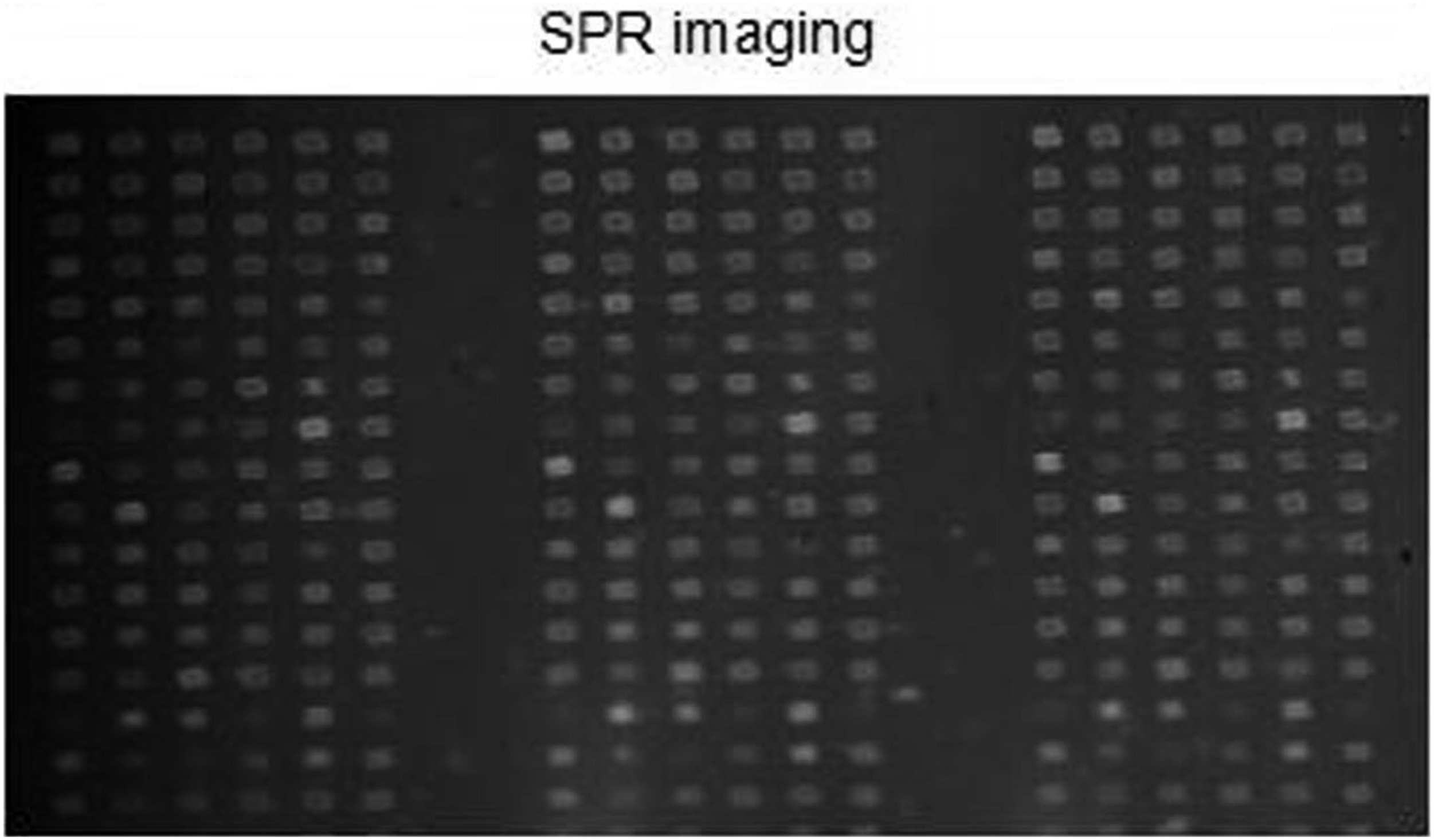
Interaction of SPRi antibody chips with different influenza virus antigens
The culture supernatants of three influenza viruses (Ag308, Ag501, and PR8) were injected into the constructed SPRi antibody microarray chip, and high-throughput interaction curves between 115 antibodies and three inactivated viruses were obtained. As described in Figure 2, the surface of the chip has a good anti-nonspecific adsorption capacity for such complex system sample. The binding signals of inactivated viruses and antibodies were sorted according to the signal-to-noise ratio, and it was found that different viruses recognize the antibody microarrays in different distribution patterns (Figure 3), indicating that the three viruses carried different epitopes of antigens on their surfaces. The binding curve of the reaction between influenza virus and SPRi antibody chip. Three types of influenza virus, (a) Ag508, (b) Ag501, and (c) PR8 show different binding curves after reacting with influenza virus mAbs on SPRi chips. The different colored curves represent the time variation curves of different influenza virus binding to different mAbs, detected by SPRi chips. RU, resonance unit; Time/s, time in seconds. Heatmaps of specific binding of three influenza viruses interacting with mAbs on SPRi chips. The left figure (a) lists the influenza virus mAbs immobilized on the surface of the SPRi chip; The right (b) is the heatmaps of three influenza viruses corresponding to the left image, with the distribution of different mAbs as rows and columns, and the SPRi signal-to-noise ratio of three influenza viruses reacting with different mAbs as color functions. S/N, signal/noise ratio. Note: S/N>3 the binding signal is significant.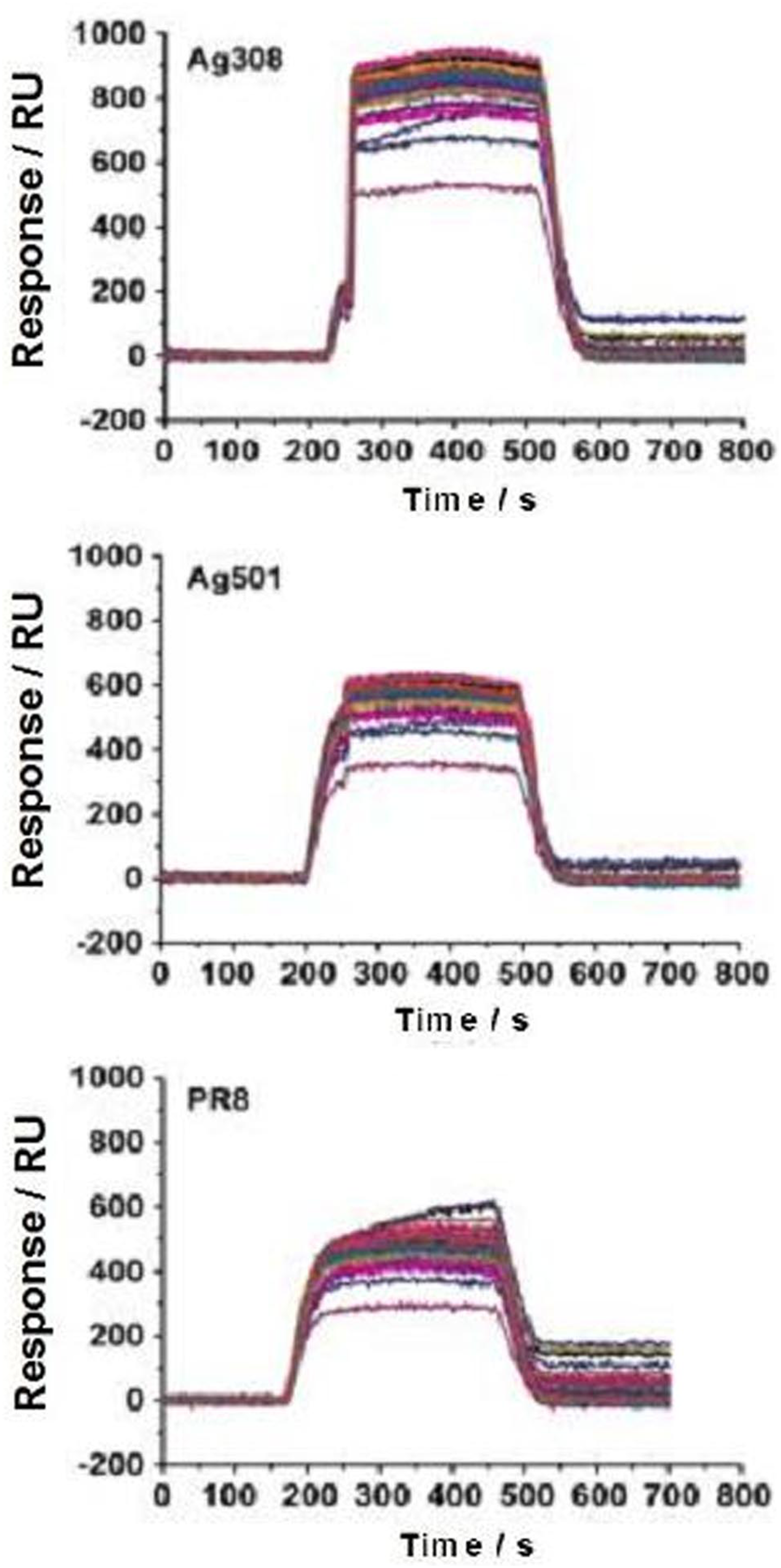
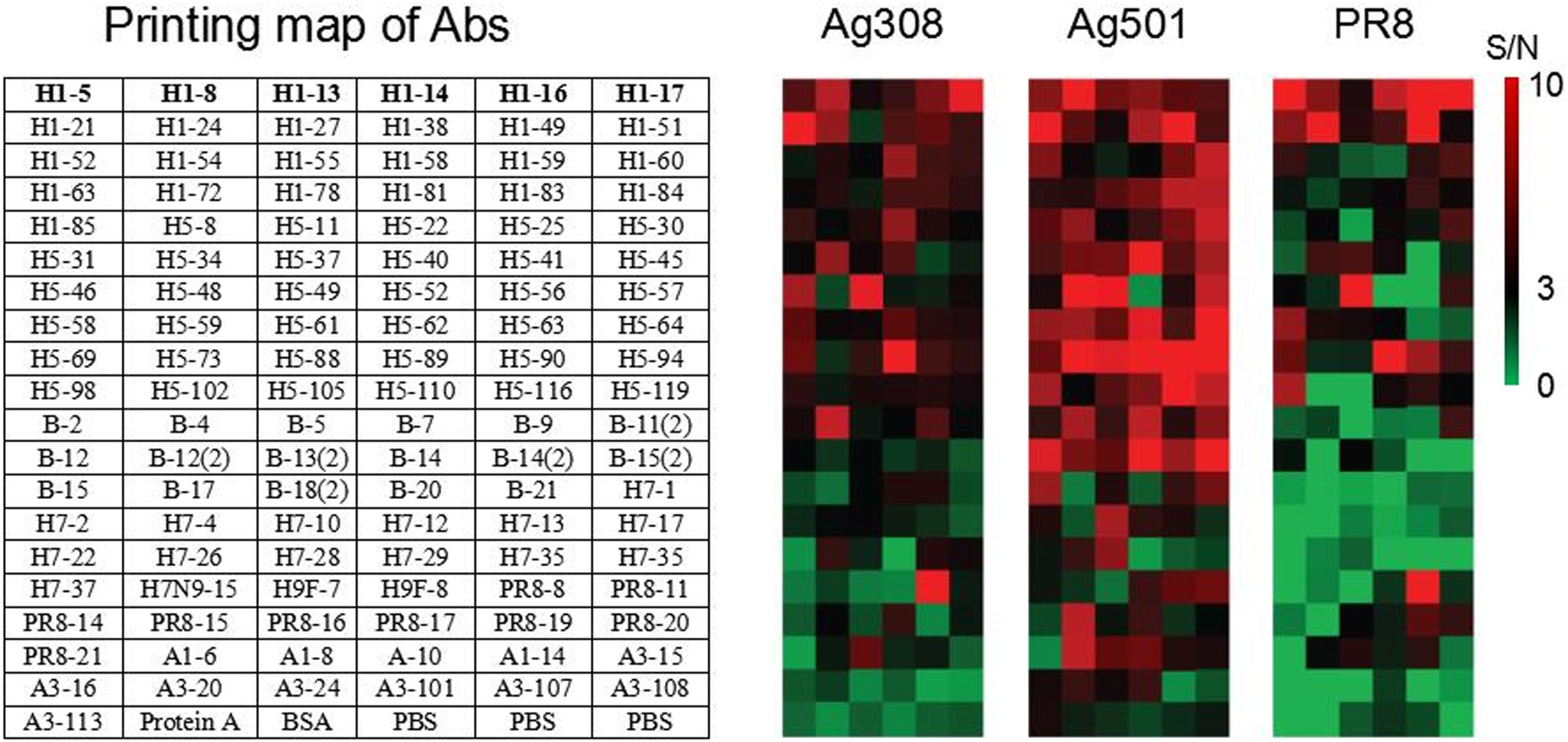
Comparison between the results of SPRi influenza virus antibody chip and ELISA
DMEM culture medium and 1:5 dilution of influenza virus PR8 culture supernatant was injected into antibody microarray chips, respectively, and the SPRi array was compared with ELISA. The results showed that among the total 115 strains of antibodies, 53 strains could be identified by SPRi detected virus supernatant, 99 strains could be identified by ELISA, and 46 strains could be identified by both methods (Figure 4). Although ELISA methods have a higher sensitivity (86.8% (46/53) Comparison of results between SPRi chip and ELISA method for detection of influenza virus PR8. The detection results of SPRi chip (a) are expressed in S/N, with a range of −20∼20. When S/N>3, it is considered positive; The ELISA test results (b) are expressed as OD values, ranging from 0 to 2. When the detected OD value is more than 2.5 times the control OD value, it is considered positive; The results of comparison (c) between SPRi chip and ELISA detection is profiled with a Venn Diagram. PR8, 1:320 dilution of influenza virus PR8 culture supernatant; PR8 Super, 1:80 dilution of influenza virus PR8 culture supernatant; DMEM, DMEM culture medium; S/N, the signal-to-noise ratio of three influenza viruses reacting with each mAbs; OD, optical density value.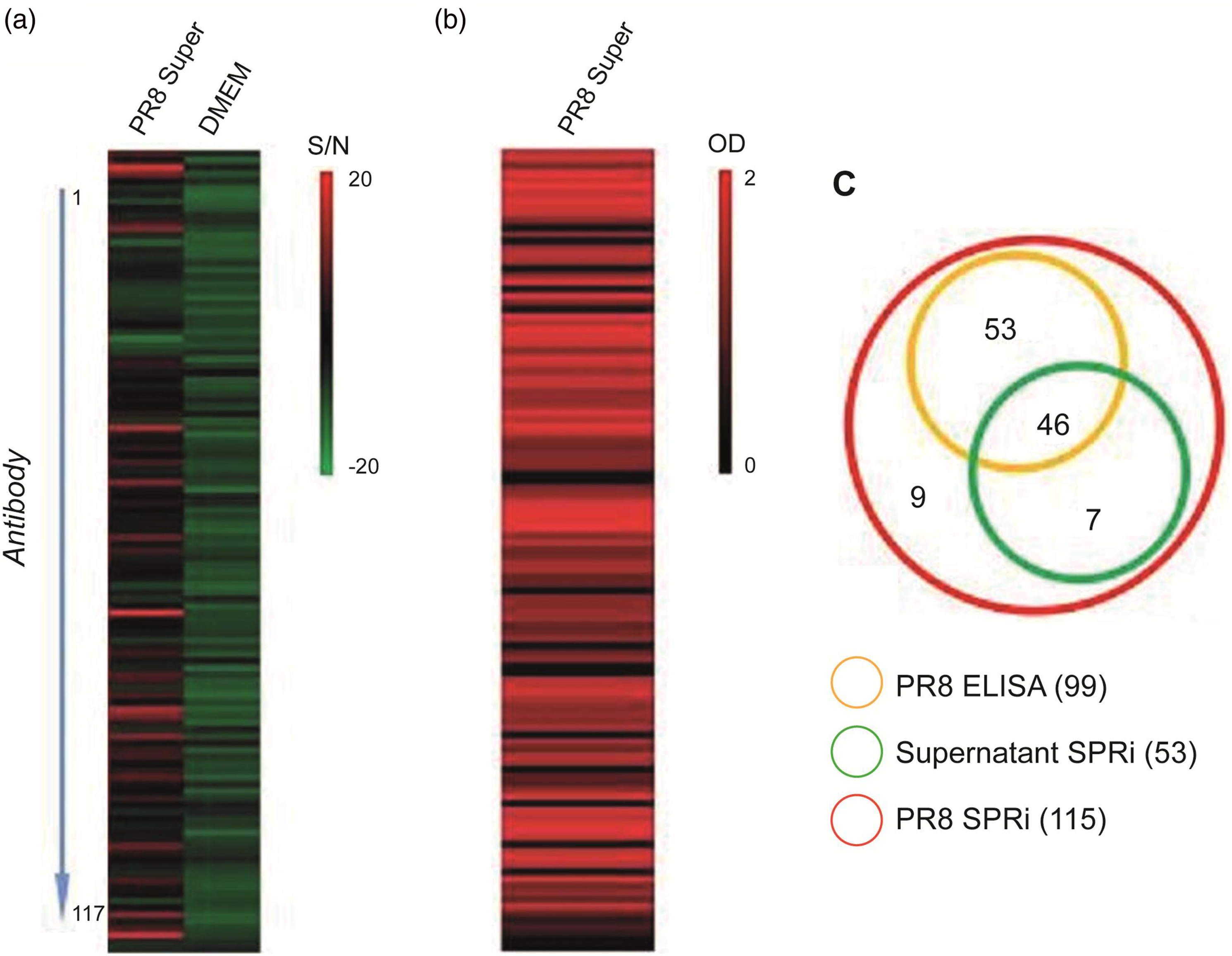
Discussion
In modern biomedical research, serum, body fluid, sputum, alveolar lavage fluid, ascites, allantoic fluid, and virus culture supernatant are commonly used as biological samples for clinical detection and disease diagnosis. However, these mixtures have complex components and different properties. It has always been one of the difficulties in biomedical laboratory technology to accurately identify and high-throughput detect target substances sensitively and specifically.
Microarray biochips are widely used in scientific research and industrial fields, for the acquisition of life information provides an efficient and cheap analysis means. In particular, protein microarray chip has been widely used in biological and medical research, including the discovery of biomarkers, disease early diagnosis, clinical detection, new drug development, and other important fields.22–24 The emergence and development of chip technology has improved the efficiency of obtaining information about biomolecular interaction, and greatly promoted the understanding of life activities and the application of biological sample analysis.
The advent of SPRi provides a label-free and real-time detection readout mode for microarray biochip applications, which has been applied to the detection of various types of biomolecular interactions. Due to the excellent compatibility between SPRi technology and microarray chip technology, almost all kinds of biomolecules (including DNA/RNA, proteins or antibodies, peptides, polysaccharides, molecular drugs, etc.) can be immobilized on SPRi chips. In the detection of biological samples, SPRi chips can detect small molecular compounds, 17 biological macromolecules such as nucleic acids11,12 and proteins,1,25 and even detect cell adhesion and migration on the surface. 26 However, in the face of complex mixtures such as serum, alveolar lavage fluid, ascites, allantoic fluid, and virus culture supernatant, the detection and quantification of specific proteins becomes a challenging test because many proteins and other components span dynamic ranges of many orders of magnitude.
Therefore, SPRi microarray chips were prepared with 115 mAbs against different influenza virus epitopes and BSA (negative controls) using the constructed double-layer epoxy surface 21 and applied to the detection of influenza virus antigens (Figure 1). The experiment showed that the culture supernatant of three influenza viruses (Ag308, Ag501, and PR8) (with a blood coagulation titer of 1:320) was diluted 5 times with DMEM medium and then injected into the SPRi influenza virus antibody microarray chip for high-throughput detection. The surface of this complex system sample has good anti-nonspecific adsorption ability, and its antibody can recognize influenza virus timely and effectively in light of its high frequency mutation characteristics (Figure 2). Due to the different epitopes of the surface antigens of three influenza viruses, as indicated by the signal-noise ratio of their binding signals, these different viruses recognize antibody microarrays in difference distribution patterns. Figure 3 also shows that some antibodies in the SPRi microarray only have specific binding or significant differences in binding signals with a certain virus, indicating that these antibodies detected by the SPRi microarray can be used for rapid typing of influenza virus, thus meeting the needs of clinical treatment and disease control in response to the spread of influenza. The above results fully confirm that the SPRi antibody microarray prepared by double-layer epoxy surface can cope with the detection of biological samples of complex systems such as influenza virus culture supernatant. The surface can not only provide abundant binding sites to immobilize proteins or antibodies effectively but also has good anti-nonspecific adsorption and biomolecular interaction detection performance, which is suitable for the detection of antigen-antibody interaction in complex biological samples.
The comparative study with traditional ELISA detection methods also demonstrates the advantages and future development prospects of SPRi microarray chip method. Although ELISA has been widely used for the detection of target molecules, it is well known that this method is susceptible to interference from autoantibodies and heterologous antibodies, and is prone to false positives. In addition, for protein microarray methods, it usually needs to label the samples by secondary antibodies, added additional cost and measurement error. In our control verification, SPRi microarray chip method and traditional ELISA method were applied to test the binding of influenza virus PR8 to 115 influenza mAbs (Figure 4). The results showed that although the ELISA method showed high sensitivity in the recognition of influenza virus by monoclonal antibodies (46/53), the specificity of SPRi antibody microarray detection (9/16) was much higher than that of ELISA, which is consistent with previous research reports. 27 This indicates that SPRi microarray chip, as a new biochemical detection technology, can construct label-free and high-throughput biomolecular probes, and obtain accurate information about the interaction between surface molecules and the objects to be tested in complex biological samples in real time, with little sample consumption and high detection specificity.
Conclusion
In short, although preliminary results have been obtained in the detection of influenza virus using SPRi antibody microarray, and optimization of surface and detection sensitivity is still necessary. The above studies show that SPRi antibody microarrays with the advantages of real-time, label-free and high throughput, and can be used for the detection of biological samples of complex systems such as influenza virus culture supernatant.
Footnotes
Acknowledgements
We sincerely thank Professor Jinsong Zhu and his team of Suzhou Puxin Life Science and Technology Co., Ltd. for their great assistance in this study.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Key Research and Development Project of Shaanxi (grant number 2021ZDLSF01-03); the Natural Science Basic Research Program of Shaanxi Province (grant number 2023-JC-QN-0844); and the Talent Support Program of Shaanxi Provincial People’s Hospital (grant number 2023JY-27).
Ethical approval
The study was approved by the Ethics Committee of Shaanxi Provincial People’s Hospital in China.
Guarantor
Lijun Sun.
Contributorship
Data availability statement
The data are available from the corresponding author on reasonable request.
