Abstract
Background
Lactose intolerance is defined as the presence of gastrointestinal symptoms, such as bloating, abdominal pain or diarrhoea, after consumption of lactose in individuals with lactose malabsorption. Most cases involve primary lactose intolerance, caused by a loss of activity of the enzyme lactase, needed for digestion of lactose. A traditional method of establishing lactose intolerance is the hydrogen breath test (HBT), accompanied by a questionnaire to document complaints experienced by the patient during the test. Due to knowledge on lactase-persistent alleles, DNA genotyping has become available for the diagnostic work-up for lactose intolerance. Both methods are currently in use. The aim of this study is to provide a definite diagnostic approach for patients suspected of lactose intolerance in a Dutch population.
Methods
In this retrospective, observational study, patients aged 15 years or older were included after presenting to their treating physician with symptoms suggestive of lactose intolerance. HBT, including a questionnaire to document complaints and DNA genotyping of LCT-13,910 C/T was performed for each patient as part of a routine diagnostic work-up.
Results
1101 patients were included (29% men). Positive and negative predictive value, sensitivity and specificity of HBT versus DNA genotyping were 80% (CI 75–84), 97% (CI 96–98), 89% (CI 84–92) and 94% (92–96) respectively. The use of the questionnaire added little diagnostic value.
Conclusions
In a population with a high prevalence of lactase-persistent alleles, we advise to exclude HBT from the diagnostic route for suspected lactose intolerance, and replace it with genotyping of lactase-persistent alleles.
Introduction
Digestion of the disaccharide lactose takes place in the small intestine by the enzyme lactase, which is located in the intestinal brush border. About 65–70% of the world population will lose 80–90% of the capacity to digest lactose within a few years of weaning, due to a loss of lactase activity. After this loss, ingested lactose is no longer digested by lactase but by anaerobic bacteria present in the large intestine, producing carbon dioxide, hydrogen and short-chain fatty acids.1,2 A reduction of lactase activity, that is, lactase deficiency (LD), may thus lead to lactose intolerance (LI) with occurrence of gastrointestinal symptoms, such as abdominal pain, bloating or diarrhoea, upon consumption of lactose.2,3
Loss of activity, or non-persistence, of lactase combined with gastrointestinal symptoms is referred to as primary LI. LI can also be secondary to acquired conditions, such as short bowel syndrome, small bowel bacterial overgrowth and chronic inflammatory bowel disease. The loss of activity is normally reversible in secondary LI. Most cases however involve primary LI, where loss of lactase activity is due to a genetic disposition, and is irreversible. 3
LI is defined as the presence of gastrointestinal symptoms, such as bloating, abdominal pain or diarrhoea, after consumption of lactose in individuals with lactose malabsorption. However, such symptoms may be non-specific as self-reported milk intolerance showed sensitivities from 30 to 71% and specificities from 25 to 87%, with the hydrogen breath test (HBT) as reference method. 3 In fact, most individuals tolerate small doses of lactose, for example, 10–15 g, corresponding to 250 mL of cow’s milk. 4 Furthermore, not all individuals having lactose malabsorption (LM), which is defined as any cause of failure to digest or absorb lactose in the small intestine, suffer from symptoms matching with LI, while factors such as anxiety disorders, psychosocial stress and IBS are also related to LI. 4 Testing for both LM and LI can therefore be challenging. In literature, HBT is often still considered as reference method. 5 During HBT, the patient is orally challenged with a standardized dose of lactose and the production of H2 is measured in the end-expiratory air. This allows for a controlled exposure to lactose and registration of complaints experienced during the test.
Lactase persistence is thought to have evolved as a result of the rise of agriculture, as milk consumption gave better survival of ancient populations. 6 In 2002, Enattah et al. identified a C>T SNP at −13910 kb (rs4988235) upstream of the lactase transcriptional start site, that correlates strongly with lactase persistence in Caucasian individuals. 7 Genotype C/C corresponds with very low lactase activity (<10 U/g protein), while genotypes C/T and T/T are associated with higher activities. 8 Several other polymorphisms linked to lactase persistence have been described in African populations.9–17 These findings allow for genetic testing of lactase-persistent alleles and polymorphisms when LI is suspected.
In this paper, we present data from a retrospective observational study of patients who presented with suspected lactose intolerance in a large Dutch non-academic hospital. Each patient underwent both HBT and genetic testing for −13910 C/T as part of routine diagnostic work-up. Despite numerous studies, detailing the merits of genetic testing over HBT, and spanning almost two decades of research, the latter is still the most widely used diagnostic test for primary lactose intolerance. The aim of this study is to provide a definite diagnostic approach for patients presenting with complaints suggestive of LI in a Dutch population with a high prevalence of lactase-persistent alleles. Based on genetic analysis of −13910 C/T, lactase non-persistence can be found in about 10% of the Dutch population. 18
Materials and methods
Ethics
Due to the observational nature of this retrospective study, the need for informed consent was waived. The study protocol was approved by the ethical committee of the Albert Schweitzer Hospital (Dordrecht, the Netherlands).
Patients
Patients were 15 years or older and presented to the clinical chemistry laboratory for diagnostic testing for presumably primary LI, after a referral by their general practitioner, paediatrician or gastroenterologist. The physician would refer a patient after taking a history of symptoms, which would generally include stomach pains, nausea and diarrhoea after ingesting food, and following relevant guidelines. No further selection criteria were set. Patients underwent both HBT and DNA genotyping. The study period lasted from October 2010 until June 2016.
Hydrogen breath test
Prior to HBT, patients followed the required diet, that is, no H2-producing vegetables 24 h prior to testing. Antibiotics needed to be halted 1 week prior. Patients abstained from eating from the evening before until the end of the test. Smoking during the test was not allowed. HBT started by collecting a baseline H2 reading (Gastrolyzer, Bedfont Scientific, Maidstone, United Kingdom), followed by consumption of 50 g of lactose, equivalent to 1 L of cow’s milk. The test lasted a minimum of 120 min (maximum 150 min), with measurement of H2 every 30 min. After each H2 measurement, the patient was asked to fill out a questionnaire regarding experienced symptoms. Symptoms included diarrhoea, abdominal cramping, nausea, flatulence and other. A positive result was defined as an increase of at least 20 ppm at t = 30 min, t = 60 min, t = 90 min, t = 120 min or t = 150 min, compared to the baseline value.19,20 Both registered symptoms and interpretation by a clinical chemist were reported to the requesting physician. Age and gender of each patient was registered for analysis.
Genotyping
DNA was extracted from ethylenediaminetetraacetic acid (EDTA) blood using QIAmp Blood kit (Qiagen, Hilden, Germany). Genotyping was performed by Sanger sequencing on a CEQ8000 (Beckman Coulter, Brea, United States) or, after it became available, realtime PCR (MRC Holland, Amsterdam, the Netherlands), using a CFX96 (Biorad, Hercules, United States). Primers used for Sanger sequencing were as follows: forward 5′ TGT AAC TGT TGA ATG CTC ATA CGA CCA TG 3′, reverse 5′ TTA CCT CGT TAA CCA CTG ACC TAT C 3’. Genotyping could give the following results: −13910 C/C (lactase non-persistent), −13910 C/T (lactase persistent) or −13910 T/T (lactase persistent).
Data analysis
All analyses were performed using Statistical Package for Social Sciences (SPSS) for Windows, version 24 (SPSS 24.0 for Windows, IBM Corp. New York, United States).
Results
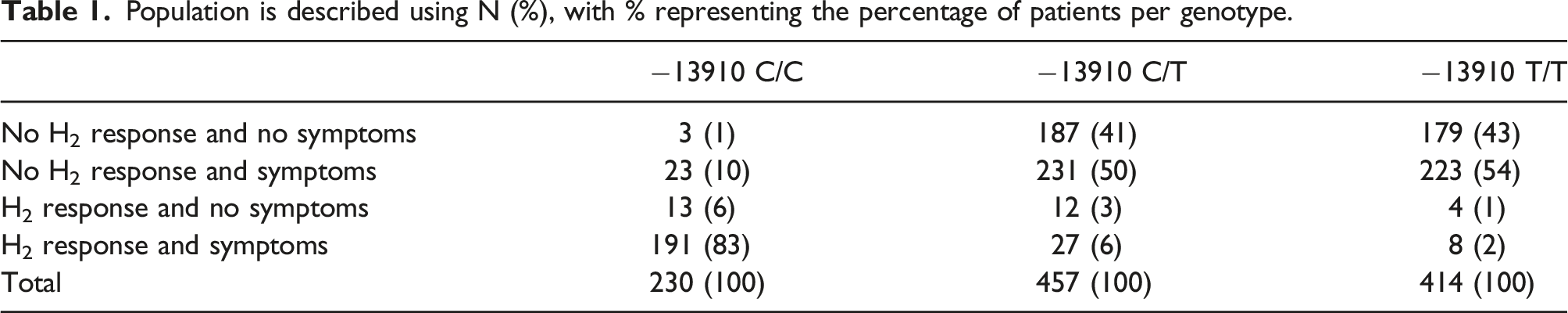
Population is described using N (%), with % representing the percentage of patients per genotype.

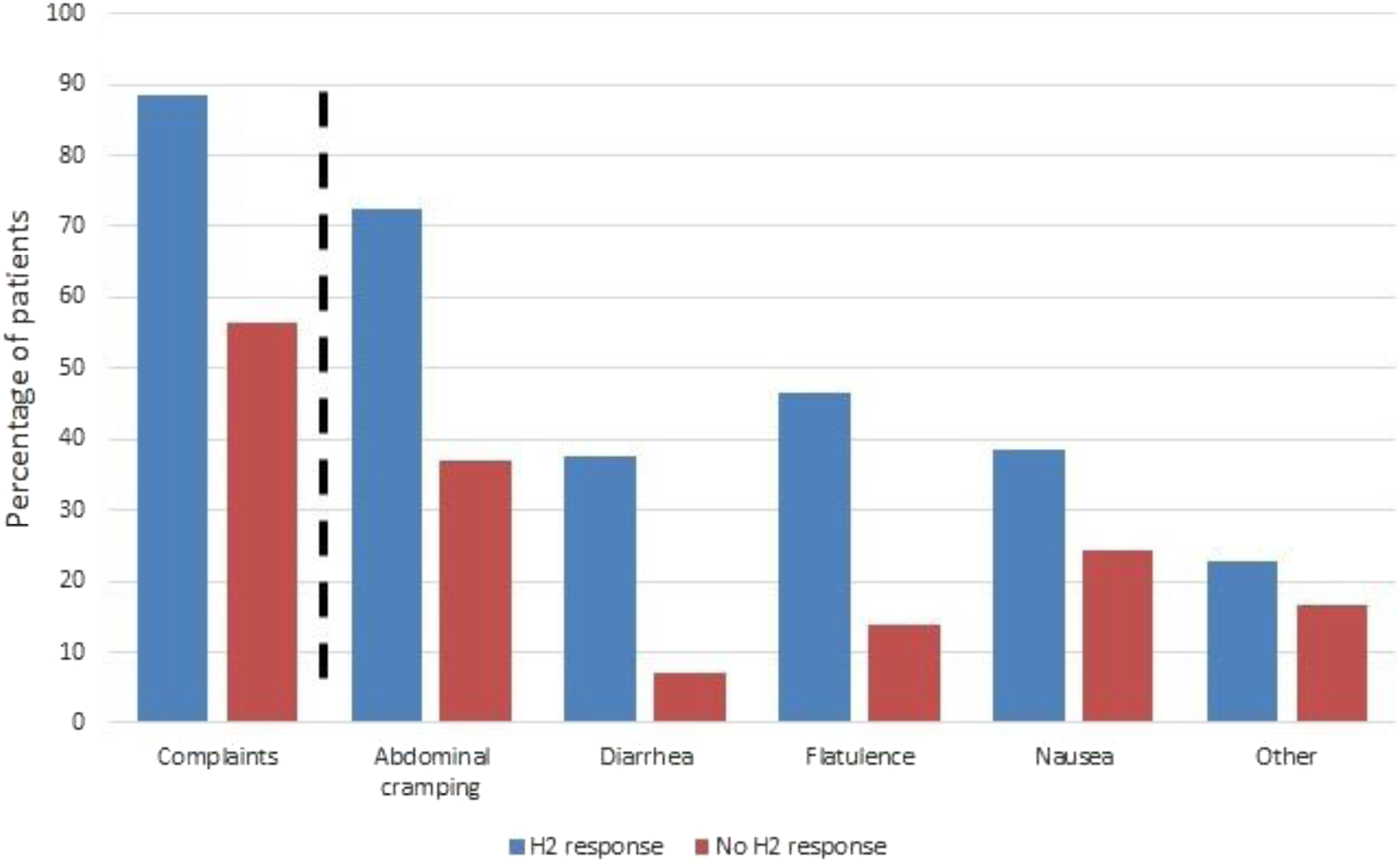
An overview of symptoms, as reported in the questionnaire during HBT, compared to H2 response. The most reported complaint in the category ‘other’ was headache (personal observation). N = 1101.
Positive predictive value (PPV), negative predictive value (NPV), sensitivity and specificity of H2 response and reported symptoms.
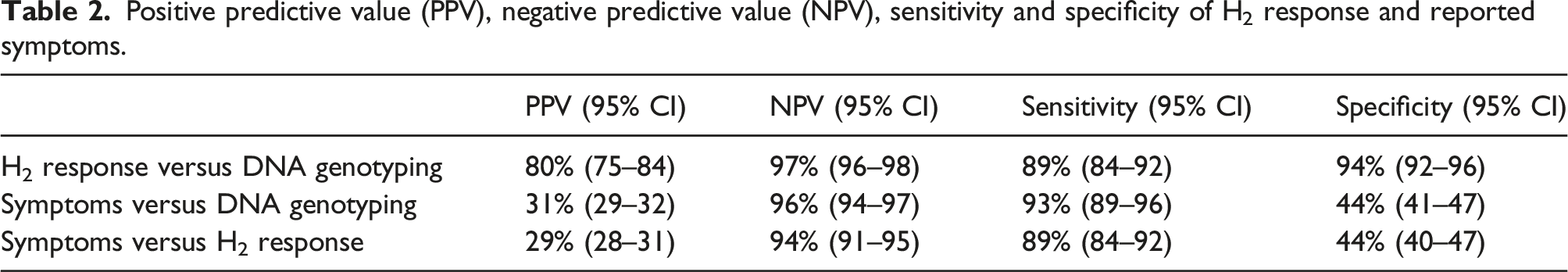
Discussion
The inability to digest lactose due to a genetic disposition to a loss of activity of the enzyme lactase, referred to as primary lactose intolerance, affects 65–70% of the world population. 1 Symptoms suggestive for lactose intolerance are quite non-specific, since gastrointestinal complaints may be caused by a great variety of disorders. Therefore, in a population with a high prevalence of lactase-persistent alleles, after taking a proper history of symptoms, further diagnostics tests are often necessary to determine if the complaints are due to an inability to digest lactose or a different underlying cause.
Traditionally, a hydrogen breath test (HBT) is performed to determine the diagnosis, but this test has considerable drawbacks. An alternative to HBT is genotyping of known lactase-persistent alleles. Both tests are currently in use as a diagnostic test for primary lactose intolerance. We reviewed both HBT and DNA genotyping in a cohort of patients presenting with symptoms suggestive of lactose intolerance. Prevalence of lactase-persistent allele −13910C>T was 79% in this cohort. The diagnostic value of both tests will be discussed here.
In our cohort, we found a sensitivity of 89% and a specificity of 94% for HBT (compared to genotyping of polymorphism −13910 C/T), which is comparable to the sensitivity and specificity found by a meta-analysis by Marton et al (1708 patients in 17 studies). 21 However, HBT does have considerable drawbacks. Prevalence of H2-non-producers range between 2 and 43% in populations. 3 In such individuals, the bowel flora is unable to produce hydrogen, leading to a potential for false-negative test results when performing HBT. While the test itself is relatively cheap to perform, HBT is time-consuming, requiring a minimum of 2 hours of the patient’s time, and may bring considerable discomfort. Performing this test means willingly exposing patients to a substance, which may cause them to suffer severe gastrointestinal complaints. The after-effects of the test may last for several days in severe cases.
The registering of symptoms experienced during HBT, using a questionnaire, is considered an advantage of this test and may explain its enduring popularity. In our cohort, the presence of symptoms was found to have low predictive value, compared to both HBT and DNA genotyping, and adds little diagnostic value for the physician. The used dose of 50 g of lactose, the equivalent of a litre of cow’s milk, may not reflect the general daily exposure in a normal diet. Patients may be able to tolerate lower amounts of lactose and the complaints triggered by such a high dose may therefore be incorrectly labelled as caused by lactose intolerance.
While primary LI is irreversible, secondary LI is often transitory and can be reversed after treatment of the underlying cause, for example, infectious gastroenteritis. The essential distinction between the two forms cannot be made by HBT. Furthermore, other diagnoses such as coeliac disease or systemic sclerosis may also give positive HBT results. 4 An incorrect differentiation may lead to unnecessary exclusion of dairy and diagnostic delay. DNA genotyping will definitely determine or exclude the presence of LD and thus a strong predictor for primary LI, allowing for further diagnostic testing into secondary causes if necessary. Of course, as mentioned before, a diagnosis of primary LI may not always completely explain a patient’s symptoms. LI is generally treated by switching to a diet low in lactose, since generally some ability to digest lactose remains. 4 If this switch does not (completely) alleviate reported symptoms, further diagnostic evaluation may be necessary.
A drawback of DNA genotyping may be the need for a blood draw. This invasive procedure may be daunting, especially for children. However, DNA can also be obtained from swabs of the mucosal lining of the mouth. In our laboratory, mucosal swabs have been verified as a source for DNA genotyping lactase-persistent alleles. This allows for a quick, patient-friendly method to obtain DNA (manuscript in preparation).
Strengths and limitations
The main strength of this study is the large and representative cohort of patients referred to our non-academic hospital after presenting with complaints suggestive of lactose intolerance. Limitations of our study include its retrospective nature. The outcomes could not be correlated with the success of a potential treatment and secondary causes of lactose intolerance were not registered. Moreover, as stated above, the used dose of 50 g may not reflect a regular daily intake of lactose. Indeed, doses of 25 g and even 12,5 g have been advised.22,23 However, at the time of testing, 50 g was the regular dose in the Netherlands for HBT and is in accordance with the routine diagnostic process. 20
The study population included patients from three separate specialisms. We did not register per patient which specialism requested the test and it is unclear whether a difference in prevalence of lactase persistence can be expected between patients from the different specialisms. However, the combination of patients referred from both GPs and in-house specialists is common for non-academic hospitals in the Netherlands. Since our study population included all patients referred for the test, we feel it is an accurate representation of the regular diagnostic process. It should also be noted that the prevalence of lactase-persistent alleles in our study population was high. Our results and the accompanying diagnostic approach may not be applicable to populations with a lower prevalence of these alleles.
Conclusion
In a population with a high prevalence of lactase-persistent alleles, it is strongly advisable to start with DNA genotyping when lactose intolerance is suspected. DNA can be quickly obtained, either through a blood draw or mucosal swabs, and can establish or rule out the presence of primary lactose intolerance. Once primary LI has been ruled out, further diagnostic tests into other potential causes of abdominal complaints may be considered. Genotyping is well optimized, patient-friendly, and relatively cheap to perform (in the Netherlands, the cost lies between 50 and 100 euros). HBT on the other hand is time-consuming, patient-unfriendly and unable to make the distinction between primary and secondary LI. A questionnaire of complaints during the test does not add to its diagnostic efficacy. We therefore strongly advice to exclude HBT from the diagnostic route for suspected LI, when the population is comparable to the one described here, and replace it with genotyping of lactase-persistent alleles.
Footnotes
Acknowledgements
The authors thank the study participants, their relatives and the clinical staff at the participating hospital for their support and contribution to this project.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Ethical approval
Due to the observational nature of this retrospective study, the need for informed consent was waived. The study protocol was approved by the ethical committee of the Albert Schweitzer Hospital (Dordrecht, the Netherlands) Study 2020–08.
Guarantor
HJV.
Contributorship
KS, RC and HJV researched literature, conceived the study, performed data analysis and wrote the first draft of the manuscript. MW and JK performed all DNA tests. FW and FV were involved in data analysis and subsequent drafts of the manuscript. All authors reviewed and edited the manuscript and approved the final version of the manuscript.
