Abstract
Planning and designing a study that links content analysis and panel data is a complex endeavor and managing a collaborative research project across multiple organizations involves many hurdles and challenges. Both, the complexity of a linkage design and the challenging nature of a collaborative research project are enhanced by the COVID-19 pandemic, demanding creative solutions for many issues and a lot of planning by the researchers involved. Especially the challenges involved in gathering, storing, analyzing, and accessing data are amplified by the lack of face-to-face contact and standardized technical infrastructure for digital collaborative research projects. This article aims at giving an overview of the technical infrastructure involved in the content analysis part of a large-scale linkage study, providing researchers with a blueprint of the many moving parts involved in the study’s implementation. This overview will be discussed with a reflective eye toward the challenges encountered and solutions found in the process, the added layer of complexity of a global pandemic, and potential learnings for future projects like this.
Introduction
The COVID-19 pandemic has been a challenging time for many aspects of communication research. One of the consequences of the pandemic with the biggest impact is the lack of personal, face-to-face communication between collaborating researchers. While even direct colleagues of the same organization were—in most cases—not able to meet face-to-face, collaboration across organizations was met with even more hurdles. This becomes especially relevant when designing a new research project.
Designing a new research project generally has two distinct phases: The conceptual phase and the technical phase (Verschuren et al., 2010). In the conceptual phase, researchers need to set the goals of the project, determining what should—in the best-case scenario—be achieved by the end of the project and what methods will be utilized to achieve these goals. If the planned project involves many researchers across organizations or disciplines, during COVID-19 this phase might have involve online-meetings over videoconferencing services, where the most basic questions concerning the project were discussed: Which questions should be answered with which methods and to what end?
Following these considerations, the technical phase of the project design begins. When the conceptual design answers the what and why, the technical phase tackles the how, where, and when of the endeavor. In communication science, this might involve building a questionnaire, setting up the infrastructure for the automated retrieval of data, managing and training human coders for manual content analysis and so on. In the technical phase, researchers must also set up their research data management strategy, that is, how to store, analyze and access the data.
With more and more sources of (digital) data, research data management has gained more relevance across academic fields (Surkis & Read, 2015). Since there is little uniformity in the actual practices of research data management in academia (Barsky et al., 2017), agreeing on a shared strategy is a challenge for collaborative research projects, especially in an interdisciplinary context (Akers & Doty, 2013; Trimble et al., 2017). Particularly in distributed projects such as those accelerated by the COVID-19 pandemic, the technical phase also involves the usage of a (shared) digital infrastructure and the exchange of raw and processed data. Following best practices of open communication science (Dienlin et al., 2020), this phase should also include the inter-subjectively understandable documentation of these processes.
This article provides an overview of the technical infrastructure involved in the content analysis part of a large-scale linkage study conducted during the COVID-19 pandemic, providing a blueprint of the many moving parts involved in the study’s design, focusing on the development and usage of a research data management strategy. This overview will be discussed with a reflective eye toward the challenges encountered and solutions found in the process, the added layer of complexity of a global pandemic, and potential learnings for future projects.
Research Data Management
Originating mainly from information and library science, research data management refers to a sub-area of science management. Since there is little uniformity in the actual practices of research data management in academia (Akers & Doty, 2013; Barsky et al., 2017; Trimble et al., 2017), there is also no uniform consensus on the definition of the term (Wilms et al., 2016). Most often, it is used as in Whyte and Tedds: “Research data management concerns the organization of data, from its entry to the research cycle through to the dissemination and archiving of valuable results.” (Whyte & Tedds, 2011).
Research data thereby are all information and statistical findings gathered for later analysis. It includes, for example, tables, text, videos of experiments, biometrics data, or photographic documentation of media use situations. Ultimately, research data are everything that is used in the research process to produce scientific insights. Thus, research data are the basis of all scientific communication (Wilms et al., 2016).
According to Wilms et al. (2016), research data management is a means to collect, organize, validate, and preserve research data to make it available to the scientific community. Research data management must be seen as a process, as existing research data always have the potential to be used for new research. Especially when considering the need for transparent and open science practices, the relevance of a research data management strategy that enables other researchers to reproduce and validate prior findings becomes evident.
The process of research data management can be depicted in a circular fashion, as shown in Figure 1. The first three stages of the cycle can be understood analogously to classical social science research. Here, the research data are created, processed, and analyzed. Following FAIR (Findable, Accessible, Interoperable, Reusable) data principles (Wilkinson et al., 2016) the following stage, the storage of the research data, should not be understood as simply storing it on a single hard drive. Storage must also involve long-term considerations, such as choosing a format that will remain readable for as long as possible and safeguarding against data loss, for example by creating backups of the data. In the fifth step of the cycle, the data are made available to third parties (other scientists, laypersons, journalists, etc.). This step makes the research process more transparent and enables the reproducibility of research.
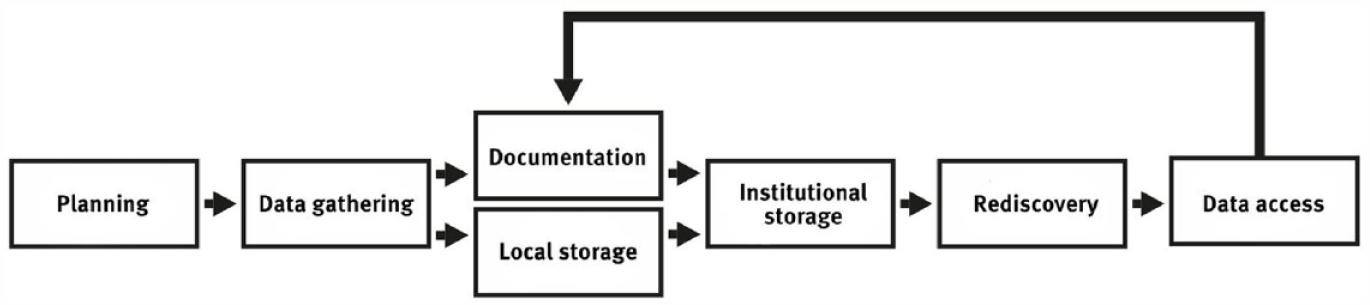
The research data management process.
Jarred by the so-called replication crisis, many researchers, including in communication science, have called for stronger open science practices, “which aim to increase the reproducibility, replicability, and generalizability of findings” (Dienlin et al., 2020, p. 2). Open science practices are discussed as a viable instrument for mitigating “questionable research practices,” such as the “undisclosed selective reporting of outcome variables, outlier removal, data imputation, hypothesizing after the results are known (HARKing)” (Bakker et al., 2021) and thus hoped to contribute to a more resilient and sustainable scientific process.
Yet, surveys of (quantitative) communication scholars show that many researchers do not yet fully embrace the open science agenda and doubt that their colleagues do so (Bakker et al., 2021; Bowman et al., 2022). An analysis by Gabelica et al. (2022) shows that while a large share of articles includes a data sharing statement, with the indication that data is available upon a reasonable request, few researchers (6.8%) are willing to grant access to requested data.
While some scholars have already noted the relevance of research data management (for an overview of the literature see Wilms et al., 2016), for others the hurdles and costs of research data management are still too high. Some researchers see the resources that sufficient research data management requires as a waste of time and an unnecessary burden (Surkis & Read, 2015). Others fear that by publishing their data, competition with their colleagues will increase or others will notice errors in their own work (Wilms et al., 2016). In a study by Barsky et al. (2017), more than 90% of the scientists surveyed said they were very interested in a structured research data management system but did not feel confident implementing one on their own. Support from the institutions of academia (research communities, libraries, administrations) thus seem necessary to enable structured research data management infrastructures.
Individual attitudes toward research data management vary greatly from researcher to researcher, from field to field, between countries. In some scientific fields, especially in the natural sciences, research data management has a long tradition and is also partly responsible for the most relevant achievements in these fields. In genetics for example, the human genome project (HGP) is an international research project in which scientists have worked together to completely decode the human genome. The HGP has been described as “one of the most revolutionary and captivating scientific endeavors ever conceived by mankind” (Sawicki et al., 1993) and would not have been conceivable without a strong system for structured research data management (Surkis & Read, 2015). Another current example are the global collaborations tackling the pandemic itself: “The pandemic could leave its mark on research collaborations for years to come. Many scientists [. . .] strengthened existing connections and forged new ones” (Maher & Van Noorden, 2021, para. 6).
One reason for the differences in research data management practices between academic fields is that research data itself varies greatly in mass, form, and content between the academic fields. For example, Akers and Doty (2013) asked respondents from the humanities (n = 54), social sciences (n = 78), medicine (n = 124), and natural/foundational research (n = 74) for their data storage needs and practices. Natural scientists reported the largest need for data storage with more than 30% of respondents reporting to need terabytes of data storage. In contrast, 80% of the respondents from the humanities did not even know about requirement for data storage and open access in research proposals. Particularly in multidisciplinary collaboration (e.g., between communication scholars and information science researchers), these differences between fields must be considered beforehand, as developing a shared and consistent research data management strategy must satisfy the requirements of all participants.
Noteworthy, albeit half of the participants in Akers and Doty’s (2013) study indicated their interest in sharing data in the future, only one-fifth of them did already engage in such routines (one-third in the natural sciences). Medical scientists and social scientists in particular state that they do not share data in order not to restrict the privacy of their subjects and therefore need tools for data release management, whereas natural and social scientists report being afraid that their research results will be published in advance by third parties. In the natural sciences and medicine, there are additional concerns about possible patents and the commercial usability of their results. Slightly less than one-third of scientists across academic fields were concerned that sharing data would be too much effort, and that data could be misinterpreted. Another 15% across fields thought that data might be worthless for others or might violate privacy or legal rights. Thus, to build effective research data management systems, scholars’ specific practices must be understood and then mapped into the systems (Cragin et al., 2010).
Researchers are often caught between public and private interests when it comes to research data management. Many scholarly journals require that research data be made public (Piwowar et al., 2007; Zenk-Möltgen & Lepthien, 2014), and the German Research Foundation (DFG), the German Academic Exchange Service (DAAD), the Fraunhofer-Gesellschaft, and the Helmholtz Association, as well as research societies from various other countries, have strict guidelines on how to handle collected data (Erway & Rinehart, 2016; Jones, 2012). Vines et al. (2013) found that a requirement to retain research data alone was already sufficient to greatly increase the availability of data to other researchers although the practical availability of research data which is not published in public databases is relatively poor (Alsheikh-Ali et al., 2011; Savage & Vickers, 2009). Research data management systems and structured plans for storage and publication can remedy this situation. Particularly in the context of third-party funders from the private sector, data sharing might also conflict with intellectual property interests (David, 2004; Murray-Rust, 2008).
Especially when cloud storage services are used to store and manage research data management, legal problems add to the challenge. The legal conditions for using cloud storage in Germany are still unclear and results in “a high degree of legal uncertainty” (Hilber & Reintzsch, 2014, p. 702) for researchers. This is especially true for services that operate under foreign laws.
Against this background, the requirements for a modern research data management system come from two sides: the specifications of institutional sponsors and funders and the research data management practices of the researchers themselves.
Particularly public founders increasingly pronounce the value of open data, as the research data management requirements of the largest science funders from Germany (German Research Foundation, DFG), the United Kingdom (Research Councils UK, RCUK), and the United States (National Science Foundation, NSF) show. Each of these three institutions has several billion euros available annually to distribute to research projects in their respective countries and provides specific rules to the funded scientists on how to handle the research data collected. Based on Wilms et al., (2016), the following seven categories can be identified (see Table 1).
Research Data Management Requirements Across Institutions.
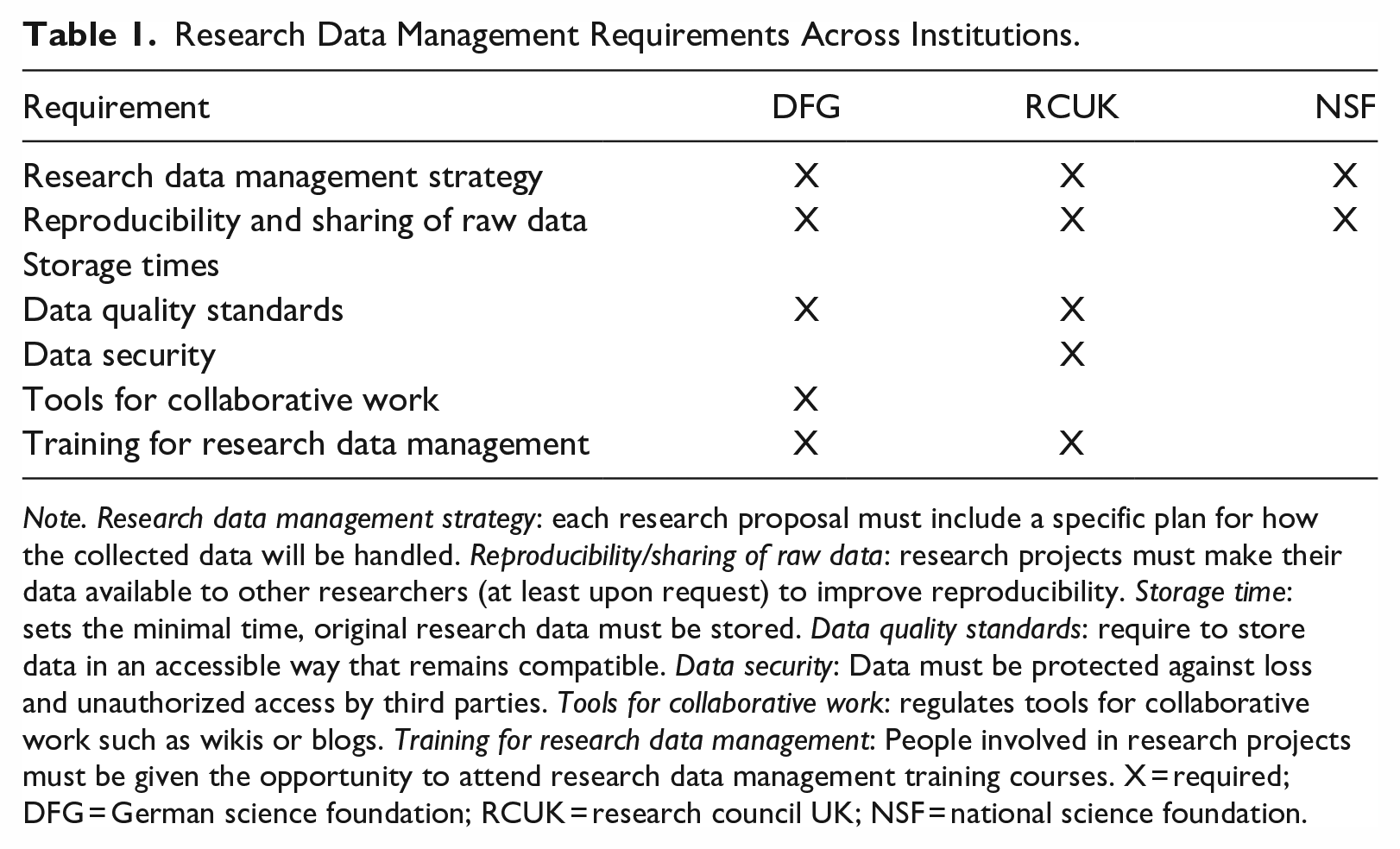
Note. Research data management strategy: each research proposal must include a specific plan for how the collected data will be handled. Reproducibility/sharing of raw data: research projects must make their data available to other researchers (at least upon request) to improve reproducibility. Storage time: sets the minimal time, original research data must be stored. Data quality standards: require to store data in an accessible way that remains compatible. Data security: Data must be protected against loss and unauthorized access by third parties. Tools for collaborative work: regulates tools for collaborative work such as wikis or blogs. Training for research data management: People involved in research projects must be given the opportunity to attend research data management training courses. X = required; DFG = German science foundation; RCUK = research council UK; NSF = national science foundation.
Research Data Management in Communication Science
Beyond these general requirements, communication science has also specific needs. A report by the German communication association (Peter et al., 2019) identified several questions communication scientists have to answer for themselves when planning their data sharing: These can be summarized broadly as what should be stored for how long, to whom should it be accessible, and what risks must be mitigated?
First, researchers must clarify, what data should be stored. In most cases, text (~80%), numeric data (~40%), and multimedia data (~40%) should be stored in research data management systems (Barsky et al., 2017; Trimble et al., 2017). Software and statistical models account for an average of 15% of the data stored. Thus, a research data management system must be able to handle at least these types of data. At best, the research data management system’s functions must also be able to be customized for specific users; 10% of the stored data falls into this category. Depending on the data, the storage space can be planned. Studies show that about half of researchers need between 5 and 20 GB of storage volume, with about one-tenth of researchers needing more than 50 GB (Akers & Doty, 2013; Barsky et al., 2017; Trimble et al., 2017). Noteworthy, the demand for storage sizes increases exponentially over time, analogous to the computational power of computers. Thus, these numbers represent only a snapshot; scalability of research data management system will become extremely important in the future. Related to this aspect is the question for how long data should be stored. Ideally, there should be no restrictions on the storage length of their research data (Murray-Rust, 2008; Trimble et al., 2017; Wilms et al., 2016) as it is unclear, when a specific data point might become relevant for future studies. Data storage and sharing are therein closely connected. Most researchers require a research data management system to be able to share research data online via access control tools (Akers & Doty, 2013; Barsky et al., 2017; Trimble et al., 2017). Often, research data should also be able to be managed, created, and analyzed collaboratively (Trimble et al., 2017). At the same time, data protection against both external threats (e.g., hacking) and data loss must be considered from the beginning.
As outlined above, research data management is in general an important puzzle piece in the larger goal of open, transparent, and reproducible (communication) scholarship. The pandemic and its repercussions have only underscored its relevance. The pandemic heightened the need for remote work solutions, even in projects at a single institution or university. Without physical infrastructures, accessing, working with, and documenting collected research data can become chaotic and can lead to errors (like not being able to keep track of the most current state of datasets). Furthermore, the principles of sustainable research data management outlined above can also help to structure the way researchers think about their data. Well-structured and documented research data management practices can mitigate the effects of disruptive events like the pandemic, especially when they are formulated and implemented early on. By structuring a part of the research process that is often seen as trivial and developing adequate routines, makes the process more resilient against sudden changes and disruptions. For example, storing completed survey responses on the computers of individual researchers working on the data might be feasible when the researchers can coordinate themselves in a face-to-face setting. However, switching to remote work is much easier when the raw data and the survey are stored in a digitally accessible shared space.
The Current Example
Planning a Collaborative Linkage Study During a Pandemic
This article aims at providing insights into the planning, technical realization, and data management infrastructure of a linkage study, designed and conducted by three research groups at three distinct universities. A linkage study describes a methodological design within media effects research that combines survey data with observational data about the media environment of the survey participants. The advantage of a linkage design is that while content analyses can, for example, reveal certain patterns in the coverage of political actors, but cannot measure their effects, while surveys can be used to measure the attitudes and usage patterns of participants, “but these attitudes are hard to meaningfully relate to the media without knowing about the contents of the media” (De Vreese et al., 2017, p. 221).
The linkage study covered here combines a (semi-) automated content analyses of news coverage and user comments on three popular social media platforms (Facebook, Twitter, and YouTube) with a three-wave user panel and aims at better understanding the processes of and interplay between media use and political attitudes. The study was planned and conducted between 2020 and 2021, during the “hot period” of the COVID-19 pandemic. Linkage studies often involve relatively complex designs (De Vreese et al., 2017), and this study was no exception. To ease understanding of the use case, we thus provide an overview of the data collected for the study.
For the survey component of the study, a commercial panel host (Respondi) collected a three-wave panel study of German social media users, aged 18–69, that use Facebook, Twitter, and/or YouTube at least once per week, stratified for gender and age. A total of N = 1277 individuals completed all three waves of the panel (wave 1 was collected in February 2021; wave 2, 6 weeks later in April; and wave 3, another 6 weeks later in May). Participants were on average 44.6 years old (SD = 13.7, range 18–69) and more likely male (57.09%) than female (42.52%), and no one was non-binary. More than two-thirds used Facebook (77.53%) and YouTube (69.03%), more than a quarter used Twitter (28.07%).
For the content analysis, we crawled the output of five selected German legacy news on Facebook, Twitter, and YouTube. The selected outlets represented the broadness of the German legacy news media landscape (public broadcaster news Tagesschau, private broadcaster NTV, left-center-leaning newspaper Spiegel, right-center-leaning newspaper Welt, boulevard magazine Bild). We also included three alternative news brands (Russian-based Russia Today, short RT, left-leaning Junge Welt, and right-leaning Junge Freiheit). Overall, we collected N = 122,865 news posts uploaded by either of the eight brands on either of the three platforms between January 1, 2021 and May 17, 2021. Most were tweets (n = 78,042), followed by Facebook posts (n = 35,110) and YouTube videos (n = 9713). We also crawled all N = 4,287,048 comments/replies to all tweets, posts, or videos. To ensure access to the full coverage, we archived the content from all URLs included in the post’s texts. The data gathering was split across the three involved research groups, with each group being responsible for a specific social media platform.
Challenges Regarding the Research Data Management Infrastructure
There were multiple challenges to be addressed by the technical infrastructure in our study as multiple datasets had to be homogenized, merged, and documented. Overall, we worked with ten unique datasets: (1) three datasets for the output of all media brands, one per social networking site, (2) three datasets for the comments and replies to these posts, one per social networking site, (3) a dataset containing the shared webpages, combining all three social networking sites, and (4) three datasets for the three panel waves. During the data gathering process, these datasets had to be interlinked, as the posts by the media outlets were the basis for the other two datasets and the usage data from the survey panels had to be linked to the observed outlets. Finally, as the data gathering was conducted by multiple universities, the datasets had to be transferred to one shared space, that conformed with the standards of a modern research data management system (e.g., providing a fine-grained access-management-solution for all involved researchers including student coders).
As the pandemic rendered in-person meetings impossible, coordinating a shared research data management strategy for this project was an even higher challenge than usual (and requested several video conferencing debates). Yet, relying on the standardized principles drawn from funding organizations and the open data principles developed by other researchers also made our process less error prone. By stating common goals informed by the best practices of research data management and by respecting specific requirements of the researchers involved, we were able to find a joint solution matching our requirements regarding data crawling and storage.
Data Crawling
First, we had to develop a process to gather the social media posts across three quite different social networking sites. As the project had to be conducted remotely due to the pandemic, a mostly automated solation had to be developed that could be managed and maintained by multiple researchers at different locations. We opted for a server-based solution that used tools like git to manage the codebase.
We collected YouTube videos and corresponding comments via custom crawlers written in Python using YouTubes’ application programming interface (API) developed by Röchert et al. (2022). For the Facebook data, we combined two data sources. The posts by the media outlets were automatically crawled using the CrowdTangle API. CrowdTangle is a public insights tool owned and operated by Meta. As CrowdTangle allows for historic data retrieval, posts were exported on a weekly basis in batches. Among others, CrowdTangle provides unique IDs for Facebook posts. We used these IDs to crawl the corresponding comments using Facepager (Jünger & Keyling, 2019). Access to Twitter data was provided through a custom crawler written in Python that queried the “search/recent” endpoint of the Twitter API to gather all Tweets, Retweets, Quote Tweets, and Replies posted by any of the eight media outlets (similarly to Schatto-Eckrodt et al., 2020). These tweets were then stored in a Postgres Structured Query Language (SQL) database. Using a SQL database for this step had the benefit of being able to quickly query this database for the tweets’ IDs, which were needed to extract the replies to the original tweets and to extract the posted URLs. Furthermore, this allowed us to fully automate the process. This was necessary as the replies and comments could not be crawled immediately after a media outlet published a tweet. Users needed time to comment on the original tweet. To provide an automated solution to this problem, the SQL database of the original media outlet posts was checked daily for posts that were older than 3 days and that had not previously been checked for potential replies. Using the “conversation_id” provided by version 2 of the Twitter API, all replies and quote tweets (and replies to these posts, and so on), were queried using self-developed Python scripts by the first author.
The challenge of gathering the webpages of the news articles posted by the media outlets for (automated) content analysis, without having to manually save the content of the webpages, was also solved through SQL databases. Once per hour we queried the database for new tweets and checked if they included a weblink (URL). The URLs were then passed to a Python script that utilized a standalone version of the “readability library” used for the Firefox Reader View to extract both a clean (i.e., containing no HTML markup) version of the webpages and their full HTML source for later analysis. By using the readability library, there was no need for custom web-scrapers for each media outlet; a process that otherwise can become time-consuming and resource intensive. Figure 2 summarizes the project’s technical infrastructure.
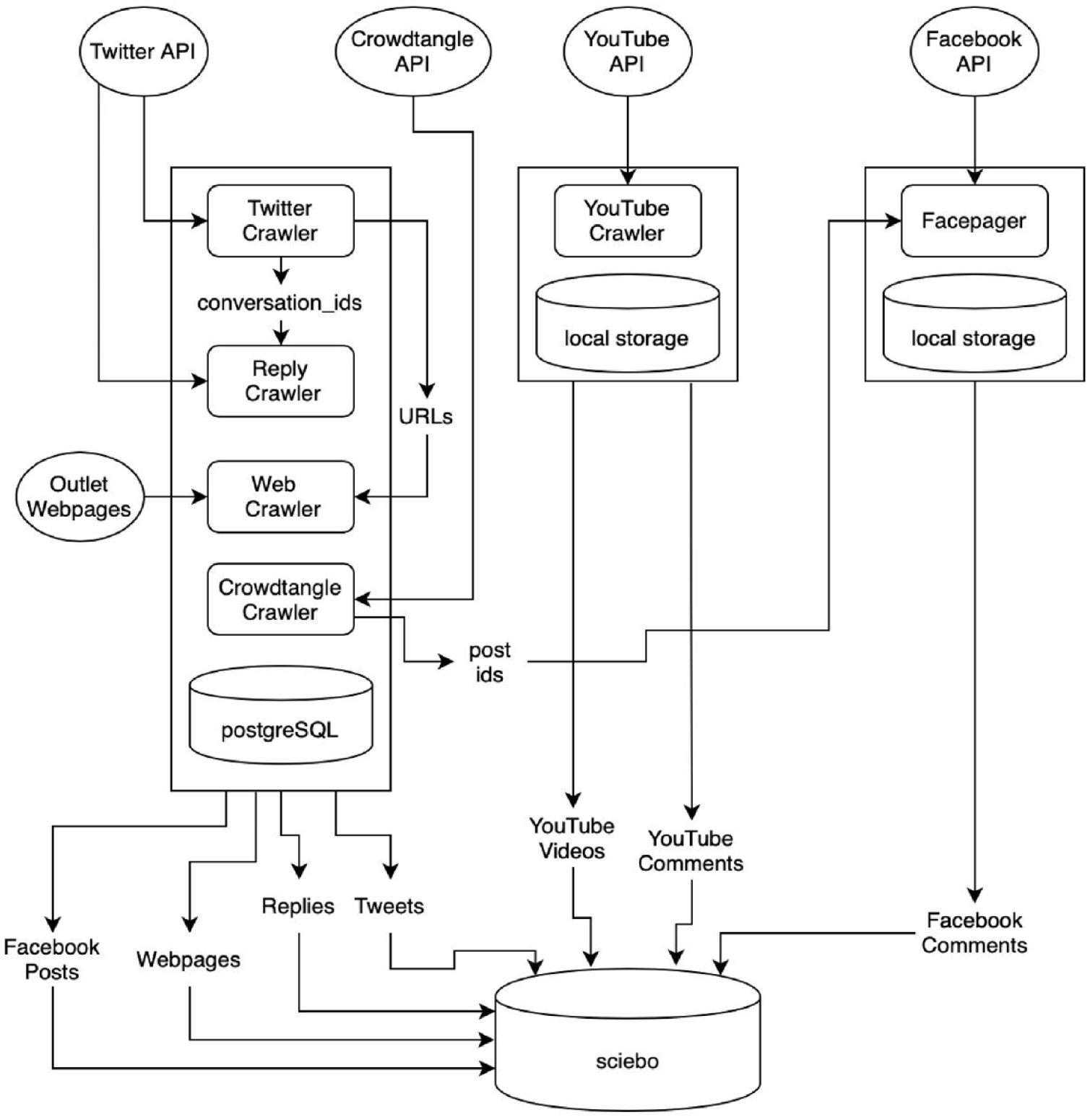
Schematic overview of the data gathering and storage infrastructure.
Data Storage
To ensure sustainability of the project, we aimed for a long-term storage solution for the data. While being a helpful tool during the data gathering, SQL databases are not suitable for this step, as accessing them requires a certain level of technical know-how and are not easily extendable to include analytical routines later one. Thus, for the final storage, all seven relevant datasets (three post datasets, three comment datasets, and one webpage dataset) were exported from the databases to a common format (i.e., csv) and given a unique ID that provided a way to reference each post in the following analysis. The data were then stored on a “Sciebo Projektbox,” a special form of account on sciebo, “a non-commercial cloud storage by [German] universities for universities” (sciebo, 2021). Sciebo Projectboxes are particularly suitable for long-term projects with changing project management and satisfy all the requirements for a modern research data management solution. Sciebo is based in Germany and is founded and lead by a non-commercial consortium, thus satisfying all the strict data protection and privacy regulations required by German and European law. The “sciebo Projektbox” can store up to 2 TB, which is especially relevant here, as YouTube videos takes up large amounts of space. Sciebo also offers tools to share data externally and internally (across universities), through a sophisticated access management system, where administrators can grant read and write access to the stored data. This can be especially useful when giving student assistants access to the data for manual coding. The tools offered by sciebo for collaborative writing are similar to for example, Google Docs and were used for the collaborative documentation of the data.
Learnings from Complex Data Gathering Processes
Due to its design, the infrastructure of the linkage study involved many technical, legal, and practical challenges, and we hope that some of our solutions are helpful for future projects involving the collaborative analysis of social media data across platforms and universities. We thus summarize our key recommendations in the following.
Developing a Full Research Data Management Strategy at the Start of the Project
In the technical phase of designing a research project (especially when multiple organizations are involved), the research data management strategy should be laid out. Early deliberations on the plan for long-term storage crucially ease later steps in the process. Researchers must not only consider where the research data will be stored (e.g., using Sciebo, open-science specific platforms such as the open science framework, or specific scientific resources such as the German GESIS), but also in what format and who should be able to access the stored data. For communication scientists, the question of data storage can be informed from the report of the German Communication Association (Peter et al., 2019) and the requirements for a modern research data management system outlined above. The (technical) format of data storage should also be discussed early in the process. Finding the smallest common denominator (e.g., csv files that can be imported to Python, R, SPSS, and Excel) between the collaborators will inform many decisions related to long-term data storage and the data gathering process in general. Data formats commonly used by one scientific field might not be easily accessible for researchers from another field (e.g., data formats for specific analysis tools) and data formats that require proprietary software should be avoided, as the access to the data would be tied to access to these tools. In the described linkage study, we discussed data sharing in the project planning phase with all collaborating research teams and then declared one person responsible for setting up the technical infrastructure.
This practice of taking the time within in project to develop a shared strategy was particularly helpful as the pandemic disrupted communication even between researchers at the same institution. As the pandemic often resulted in researchers having temporarily to cover for other colleges or their tasks having to covered by others, being able to refer to the agreed upon strategy when building the data management solution made it possible for other researchers and assistants to step up with little friction. The gained flexibility by these well-documented strategies can also be useful in the context of other disruptive events.
Documenting the Data Gathering and Storage Solutions
To ensure the usability of the stored data for future access it is vital to fully document how the data was gathered, stored, and what types of analyses were planned. A detailed and clear documentation enables reproducing the data gathering process utilized in the original study and can reduce errors, especially when the researcher responsible for analyzing the data is not the same as the one gathering the data. Beside the documentation of the data gathering process (e.g., what API was used at what time?), the documentation of the structure of the resulting datasets is also important (e.g., for tabular data describing what each column and row represents). While the specifics of this documentation process depend on the gathered data and goals of the study at hand, the involved researchers should agree upon a shared strategy early on, so no information is lost in the process. In the research project described above, we used a collaborative document where we documented how we crawled, stored, and processed the data? Other examples include README-files within project folders, which are inspired by software development and include important background information about the data as well as the people and processes responsible for the data collection and processing.
Furthermore, version control systems, like Git or Mercurial, can be helpful. Combined with public repositories (e.g., on GitHub or GitLab), these systems ensure a fine-grained image of the collected data, the instruments used to collect the data (e.g., code or coding schemes), and any changes to them. They are especially helpful for collaborative projects (Hepp et al., 2021).
Logging Automated and Manual Data Gathering Helps to Find Errors and Report Potential Holes in the Data
In computer science and computing in general, a log file records events that occur in during the usage of certain software. When using automated methods to gather data (e.g., crawling social media posts over a long period), it can be useful to use log files to make potential errors visible and to ensure complete data coverage. Logging not only errors, but all events makes even fully automated processes transparent and gives researchers the ability to find potential lacks in the data coverage (e.g., caused by downtime of a queried API) and thus avoid biases introduced by missing data being not at random. When developing software for the automated retrieval of data (e.g., crawlers that connect to an API), researchers should utilize log files to report errors, messages of successful runs, and warnings raised by the research software. These logs should also be stored alongside the gathered data.
Discussion
The COVID-19 pandemic that hit the world in spring 2020 challenged many preconceived notions about scientific collaborations (Maher & Van Noorden, 2021). With the lack of face-to-face communication, new ways to organize research projects had to be developed and implemented. One of the most challenging facets of scientific collaborations that had to be addressed in new ways in this context is research data management. Against the backdrop of the many calls for open data, transparent academic practices, and reproducible results (Dienlin et al., 2020) the systems and infrastructure developed for this purpose must satisfy a number of different needs, especially for projects spanning different institutions or countries. The linkage study described above can be seen as a case study of cross-institutional collaboration, where the involved researchers had to find solutions to the challenges posed by the complexity of project and the hurdles set by the ongoing pandemic.
Situated in the technical phase of designing and planning a research project (Verschuren et al., 2010), research data management can be a challenging requirement for (collaborative) research projects. The many different requirements on the institutional, legal, and quality control level can be difficult to address in a satisfying manner. The COVID-19 pandemic highlighted how difficult it can be for collaborating researchers to find data management solutions that do not impede scientific work, while still making relevant data available for internal and external use, in a consistent and well-documented manner. From the challenges faced when designing the described linkage study, we found three main learnings that have the potential to support researchers in planning their own projects. First, our project benefited greatly from developing a full research data management strategy early on, considering the needs of all collaborating researchers. Second, documenting the data gathering and analysis processes involved ensured that our findings remain replicable, even when the project is completed. Collaborating on this (continuous) documentation can be aided by modern research data management tools. Finally, projects like ours, that use some form of computer-assisted data gathering benefit greatly from utilizing log files to automatically document every step of the (automated) data gathering. While automated data collection scales well, both in terms of the quantity and the covered time of the data, these methods can often be hard to track (e.g., loss of data through program errors). Logging can help close this gab and make automatized processes that run in the background more transparent.
The pandemic was, for many, a unique disruptive event, that has challenged many assumptions on how research projects should be structured and made traditional data storage on individual researchers’ computers finally useless. In response to these challenges, we and many other researchers have implemented creative solutions in their planning and day-to-day research. Many of the steps necessary during the pandemic (e.g., extensively documenting datasets to mitigate the many sudden changes in personnel) will remain beneficial even after the pandemic. Thus, we argue that establishing and implementing research data management strategies early on, and a general mindset of building processes to be more resilient against disruptive events should become standard in (communication) research in related areas well beyond the current COVID-19 pandemic. Without doubt, the development of research data management solutions and their implementation in (collaborative) research project is an ongoing challenge that researchers have to face. The COVID-19 pandemic, with its constraints on personal communication, has forced the academic community to be more vigilant and precise in the documentation of our methods and the transparency of our processes. Overcoming these challenges can aid to build a more transparent and open communication science.
Footnotes
Acknowledgements
We would like to thank the members of the research groups responsible for the planning and execution of the linkage study described in this article: Manuel Cargnino, Dominique Heinbach, German Neubaum, Daniel Röchert, Anke Stoll, and Marc Ziegele. Without their expertise and collaboration, this whole project would not have been possible.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
