Abstract
Density-based clustering for big data is critical for many modern applications ranging from Internet data processing to massive-scale moving object management. This paper proposes Cludoop algorithm, an efficient distributed density-based clustering for big data using Hadoop. First, we propose a serial clustering algorithm CluC by leveraging cell partition optimization and c-cluster to fast find clusters. CluC completes classification of the points using the relationships of connected cells around points instead of expensive completed neighbor query, which significantly reduce the number of distance calculations. Second, we propose the Cludoop, which can efficiently cluster very-large-scale data in parallel using already existing data partition on Map/Reduce platform. It employs the proposed serial clustering CluC as a plugged-in clustering on parallel mapper, along with a cell description instead of completed cell in transmission to reduce both network and I/O costs. Guided by proposed cell-based principles, we also design a Merging-Refinement-Merging 3-step framework to merge c-clusters on the overlay of assigned preclustering result on reducer. Finally, our comprehensive experimental evaluation on 10 network-connected commercial PCs, using both huge-volume real and synthetic data, demonstrates (1) the effectiveness of our algorithm in finding correct clusters with arbitrary shape and (2) the fact that our proposed algorithm exhibits better scalability and efficiency than state-of-the-art method.
1. Introduction
Clustering is to group data objects into different classes or clusters, and the objects within a cluster have high similarity, while objects in intercluster differ significantly with one another. Clustering has played a crucial role in numerous applications ranging from pattern recognition, mobile sensor networks, and moving object management to location-based service. With the rise of big data science, clustering analysis has attracted considerable interests in the big data mining, while clustering based on density is very useful to distance-based data mining in many applications with increasingly large-scale data owing to their capability to discover clusters with arbitrary shape.
Existing density-based algorithms such as DBSCAN [1], OPTICS [2], DENCLUE [3], and GDDSCAN [4] can obtain better groups of data points in the static large-scale and high-dimension databases, which were widely applied into many applications in the past decade. However applying the algorithms into current data-intensive applications is challenging due to rapidly increasing distributed stored big data. For example, the traffic data (GPS trajectories and infrared acquisition) from intelligent transportation system in Jiangsu Province reaches 6.94 billion records, and 4 TB sensor data were collected from older's healthcare monitoring in Shanghai for one week; the emerging wearable devices will further promote coming of big medical data era. The dataset released by Twitter is larger than 133 TB, increased data updated by Tencent's mobile applications reaches about 200 to 300 TB a day, and even the operational data of Yahoo reaches 5 PB. When the amount of data is this large, it is impossible to handle the data using the serial clustering methods on a single machine. Therefore, the best clustering algorithm is the one that (a) combines a scalable effective serial algorithm, (b) makes it run efficiently in distributed platform, and (c) does not need to preprocess all dataset.
For big data analysis, Map/Reduce and its open source equivalent Hadoop have attracted a lot of attentions due to its parallel way to handle massive-scale data. Hadoop is a desirable distributed computing platform based on a shared-nothing cluster architecture. It is designed to scale up from single servers to thousands of machines, each offering local computation and storage. Hadoop adopts HDFS (Hadoop Distributed File System) storage structure and simple Map/Reduce programming model. HDFS provides high-throughput access to application data and initially partitions data in multiple nodes, and data is represented as <key, value> pairs. In Map and Reduce phases, Map and Reduce functions take a key-value pair as input and then output key-value pairs. The Map function is first called on different partitions of input data on each slave node in parallel. The outputs by mappers are next grouped and merged by each distinct key. Then a Reduce function is invoked for each distinct key with the list of all values sharing the key. Finally the Map/Reduce framework executes the main function on a single master machine to postprocess the outputs of Reduce functions. Depending on the applications, a pair of Map and Reduce functions can be executed once or nested multiple times. Ever since Hadoop was first introduced in 2003, Map/Reduce received great success because it allows easy development of scalable parallel application to process big data on thousands of commodity nodes, tolerating the failure nodes in the process; some works also already reflected the fact in academia (see [5]). However, finding density-based clusters from big data is a very challenging problem today.
This paper focuses on the problem of efficient, effective, and scalable density-based clustering for big data. Our method employs Hadoop as computing platform and incorporates cell partition and c-cluster optimizations into density-based clustering. Our major research works are (a) how to minimize the communication (network) cost among processing nodes, (b) how to avoid preprocessing the data, namely, reducing the I/O cost, and (c) how to extract exact density-based clusters. So this paper proposes the distributed density-based clustering using Hadoop-Cludoop method, which efficiently handles large-scale data without any preprocess.
The main contributions of this paper include the following. (1) We propose a c-cluster definition along with cell-connected observations to significantly reduce computational cost of neighbor range query. (2) We propose an efficient serial clustering algorithm CluC leveraging c-clusters and neighbor searching optimization. (3) We propose the distributed clustering framework for big data using Map/Reduce structure based on the proposed serial clustering algorithm as a plugged-in clustering on Map, along with a 3-step merging framework on Reduce. (4) We conduct comprehensive experiments on Hadoop platform deployed on 10 commodity machines to evaluate the performance of Cludoop using larger-scale real and synthetic data. The results show that our methods are both effective and efficient.
The paper is organized as follows: we first discuss related work in Section 2. In Section 3, we present preliminary notions and problem statement. Section 4 introduces theoretical ideas and presents the serial clustering algorithm. In Section 5, we then present the distributed Cludoop algorithm. In Section 6, we perform an experimental evaluation of the effectiveness, efficiency, and scalability of our algorithm. Section 7 concludes whole paper with a summary.
2. Related Work
In this section, we mainly review related work in the areas of density-based clustering, parallel variants of DBSCAN, and distributed clustering on Map/Reduce platform.
Density-Based Clustering. Density-based methods describe the clusters being the high density area of points separated from the low-density regions in the data space, so clusters exhibit arbitrary shapes in the high-density region. There are two types of density-based clustering method: algorithms based on local connectivity such as DBSCAN [1] and OPTICS [2] and algorithms based on density function such as DENCLUE [3]. DBSCAN determines a nonhierarchical, disjoint partitioning of the data into several clusters. Clusters are expanded starting at arbitrary seed points within dense areas. Objects in areas of low density are assigned to a separate noise partition. DBSCAN is robust against noise and the user does not need to specify the number of clusters in advance. OPTICS is proposed to solve parameters selection problem on density-based clustering algorithms. The paper proposes two concepts: core-distance and reachability-distance to organize points. The point objects would be ordered by reachability-distance and their core point to obtain the clustering structure, which contains hierarchical clusters under a broad range of parameter settings. OPTICS visualizes clearly the cluster structure via the ordered point list and could find arbitrary shaped clusters and overlapping clusters. DENCLUE formalizes the cluster notion by nonparametric kernel density estimation based on modelling the local influence of each data object on the feature space by a simple kernel function, for example, Gaussian. It defines a cluster as a local maximum of the probability density.
Parallel Variants of Density-Based Clustering. Xu et al. [6] propose a parallel clustering algorithm PDBSCAN for mining large distributed spatial databases. It uses the so-called shared-nothing cluster architecture, which has the main advantage that it can be scaled up to a high number of computers. However it is not fully paralleled while it still needs a single node to aggregate intermediate results. Januzaj et al. [7] depict a distributed clustering based on DBSCAN and DBDC and formed local and global two-level clustering. The local clustering is carried out independently on local data; then global clustering is done on a central server based on the transmitted representatives from local clustering. Subsequently, they design a density-based distributed clustering [8] which allows a user-defined tradeoff between clustering quality and the number of transmitted objects from the local sites to the global server site based on DBDC idea. They first order all objects located at a local site according to a quality criterion reflecting their suitability to serve as local representatives and then send the best of these representatives to a server site where they are clustered with a slightly enhanced DBSCAN algorithm to obtain high quality clusters. Dash et al. [9] propose a parallel hierarchical agglomerative clustering based on partially overlapping partitioning on shared memory multiprocessor architecture for handling nested data. The experiment shows the algorithm achieves near linear speedup. Brecheisen et al. [10] present a parallel DBSCAN on a workstation, which is parallelized by a conservative approximation of complex distance functions, based on the concept of filter merge points. The final result is derived from a global cluster connectivity graph. Böhm et al. [11] implement several data mining tasks under the highly parallel environment of GPUs, including similarity search and clustering. For density-based clustering, they design a parallel DBSCAN algorithm supported by their proposed similarity join on GPU. They then propose a massively parallel density-based clustering method [12] using GPUs by leveraging the high parallelism combined with a high bandwidth in memory transfer at low cost. Andrade et al. [13] also propose a GPU parallel version of DBSCAN, named G-DBSCAN, using graph to index point objects with less than a given distance threshold to each other. However the parallel density-based clustering can not straightforward be transferred on the Hadoop, because not only have GPU-capable parallel algorithms shared main memory but groups of the processors even share very fast memory units, while Hadoop uses a distributed file system.
Distributed Clustering on Map/Reduce. Ene et al. [14] develop partition-based clustering algorithms, k-center and k-median, running on Map/Reduce, which use sampling to decrease the data size and run in a time constant number of Map/Reduce rounds. For density-based clustering, He et al. [15] propose a parallel DBSCAN using Map/Reduce framework, MR-DBSCAN, which partitions all spatial data into different Maps by the space location in the preprocessing stage and then performs DBSCAN in each mapper and merges the bordering spaces in Reduce step. Similarly, Dai and Lin [16] also propose a Map/Reduce-based DBSCAN with a data partition, partition with Reduce boundary points (PRBP), selecting partition boundaries based on the distribution of data points. However the data partition based on data space easily causes load unbalancing due to the sparse data. Subsequently, He et al. [17] also propose a load balancing mechanism based on a cost-based spatial partitioning for heavily skewed data. However above methods expend I/O cost due to preprocessing, especially for the distributed stored big data. This is also one of the key points which we resolve in this paper. Cordeiro et al. [18] present a clustering solution to find subspace for high-dimensional data using Map/Reduce; however they focus on the tradeoff problem between the I/O cost and network cost and aim to dynamically choose the best strategy. These techniques did not explore the optimization opportunities enabled by the nature insights of serial density-based clusters; this is exactly what our work does for delivering highly scalable distributed solutions along with an efficient optimized serial density-based clustering.
3. Preliminaries and Problem Statement
3.1. Definitions and Theorems
This section introduces the definitions of related terms based on the notion of connected density in [1].
Definition 1 (
-range).
The
Definition 2 (
-neighbor).
All points included in
Therefore,
Definition 3 (core point).
A point p is called a core point if
Definition 4 (border point).
For a point p, if
Definition 5 (isolated point).
A point p is classified as an isolated point if it is neither a core point nor a border point.
Note that isolated points can be considered as either anomalous points or noise.
Definition 6 (directly density reachable).
A point p is directly density reachable from another point q, if the point p is one of
The
Definition 7 (density reachable).
A point p is density reachable from a point q, if there is a chain of points
Definition 8 (density connected).
A point p is density connected to a point q, if there exists a point o such that both p and q are density reachable from o.
Definition 9 (density-based cluster (d-cluster)).
Let D be points set. A density-based cluster is a nonempty subset of D including at least a core point and all points which are density reachable from the core point.
Note that all core points would be classified in density-based clusters.
Lemma 10.
If a core point p belongs to two d-clusters
Proof.
Since point
Figure 1 shows an example demonstrating Lemma 10; p is a core point and belongs to both blue and green d-clusters, so the two d-clusters would be merged.

An example of Lemma 10; p is a core point.
Definition 11 (cell).
According to the
Let
Lemma 12.
Given a point p in the cell
Proof.
Lemma 12 can be easily proven using the definition of
Thus all points in the cells
Corollary 13.
Given a cell
Lemma 14.
Given a point p in the cell
Proof.
Lemma 14 is intuitive by Lemma 12 and definition of
According to the d-cluster definition (Definition 9) and Lemma 12, we observe that if a core point p in the cell
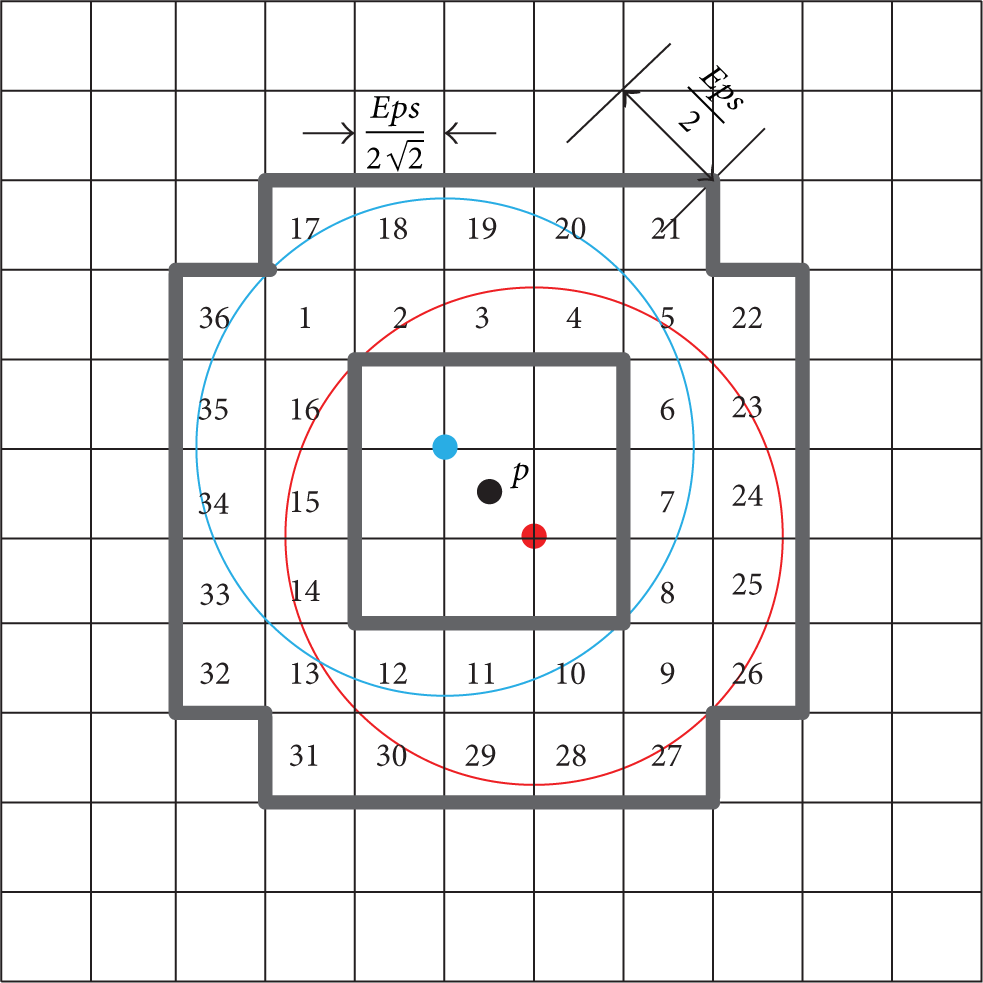
Demonstration of Lemma 14.
Definition 15 (cell-cluster (c-cluster)).
Given a d-cluster C, a cell-cluster is 2-tuple
c-Cluster simplifies the d-cluster by leveraging inclusive cells instead of large amounts of density-connected points. One no longer checks the status of every point in the clustering; instead one only changes the cluster identifier of the inclusive cells. Therefore, one uses c-cluster instead of d-cluster in the subsequent clustering process.
Definition 16 (exclusive cell).
Given a cell
Obviously, an exclusive cell must be an inclusive cell of the c-cluster, while an inclusive cell may not be an exclusive cell.
Lemma 17.
Given a cell
Proof.
Lemma 17 is intuitive. Since there exists one core point p in
Lemma 18.
Given a cell
Proof.
Lemma 18 is similar to Lemma 10. Since
3.2. Problem Statement
In this work, we focus on the efficient parallel solution of finding density-based clusters from big data on Hadoop platform. Next, we define the distributed clustering problem based on the above definitions, which this paper just aims to resolve.
Problem (distributed clustering with c-cluster). Given a massive-scale dataset D, parameter setting (
4. Proposed Serial Clustering with c-Cluster
Our proposed distributed clustering solution is designed based on one optimized serial clustering method that we propose next: clustering with c-cluster—CluC. CluC aims to find out c-clusters from a large-scale dataset. In the following, we present a basic version of CluC omitting details of data structure and additional information for understanding, as shown in Algorithm 1.
(1) (2) Get SetOfCells from D; (3) (4) (5) Get Cell (6) (7) (8) Create c-cluster (9) (10) Create c-cluster (11) (12) (13) (14) (15) (16) (17) (18) (19) (20) (21) (22) (23) (24) (25) (26) (27) (28) (29) (30) (31) (32) (33) (34) (35) (36) (37) (38) (39) (40) (41) (42) (43) (44) (45) (46) (47) (48) (49) (50) (51) (52) (53)
The
The
CluC first expands the inclusive cells recursively to reduce a large amount of distance calculations; then it can fast searches the border points in the pruned space that eliminate the inclusive cells. Therefore, the CluC achieves an efficient and also effective improvement compared with the state-of-the-art serial methods.
5. Cludoop: Distributed Density-Based Clustering on Hadoop
5.1. Cludoop Framework
The overall architecture of our Cludoop framework is depicted in Figure 3. Cludoop uses the existing data partition and does the parallel CluC algorithm in mappers and then does the merging work based on intermediate c-clusters in reducers. The final c-clusters that consist of inclusive cell descriptions and a tiny amount of border points are obtained on one reducer/machine.

Overall architecture of Cludoop clustering.
5.2. Preclustering on Mapper
As shown in Figure 3, Cludoop starts with m mappers reading the data in parallel from the HDFS and employs CluC as a plugged-in clustering on each mapper. We call the phase preclustering to distinguish whole clustering. In this phase, we first build the cell index according to the received dataset, mapping the points into the cells. Then each mapper performs CluC algorithm on the cell-structure data in parallel, aiming to output the c-clusters and noises as preclustering result. However, the preclustering results need to be normalised and shipped to the appropriate reducers over the network, to get final clusters. Thus the network traffic and normalization would cost much CPU resources when shipping all c-clusters and noises to reducers. How can we reduce the network cost for this large-scale dataset?
Our main idea is to only ship the c-clusters with inclusive cells’ descriptions and the border points to reducers, rather than send all points which have already been classified into c-clusters. Thus, we use a simple description (
To efficiently merge the c-clusters located in near area on different mappers, meanwhile to avoid the computation unbalancing for skewed data, we combine the spatial distribution and uniform division in shuffle mechanism. We set
5.3. Merging-Refinement-Merging Framework on Reducer
In Reduce phase, each reducer receives the normalized c-clusters and noises, denoted by
First, we observe that the c-clusters should be merged when they share certain cells or connected cells. To characterize the merging process we formalize two merging rules.
Merging 1.
Given an exclusive cell
Merging 1 is intuitive. First, since
Merging 2.
Given an exclusive cell
Merging 2 implies that two c-clusters covering one of two connected exclusive cells, respectively, should be merged into one cluster. This is obvious, because any core points in these two cells are neighbors; thus the density reachable core points would be classified into one cluster. Therefore, they should be merged.
Actually, Merging 1 already contains the same cases of Merging 2. For example, given two connected cells
Next, we analyse the refinement process for the nonexclusive inclusive cells, border points, and noises on the overlay of
Refinement 1.
For an inclusive cell
Refinement 1 can be easily shown. On the overlay of
Refinement 2.
Given a c-cluster C, for a border point
Similarly Refinements 1 and 2 depict the update case for the border points not in inclusive cells on the overlay of
Refinement 3.
Given a noise
Refinement 3 indicates the update process for noise on the overlay of assigned preclustering results after first-round merging.
Based on the above merging and refinement rules, we describe the Merging-Refinement-Merging framework to merge the preclustering in Reduce phase. The pseudocode of the framework is shown in Algorithm 2. First, we execute directly the first-round merging using the proposed two merging rules (lines 2–10). This step would merge the overlapping c-clusters and noise in
(1) //First-round Merging (2) (3) (4) (5) Merge all c-clusters by Merging 1; (6) //update the number of point in sharing inclusive cells and border points (7) Merge all c-clusters by Merging 2; (8) (9) (10) (11) //Refinement (12) (13) (14) (15) (16) (17) (18) (19) (20) (21) (22) (23) (24) (25) (26) (27) (28) (29) get cell (30) (31) (32) (33) (34) (35) create new c-cluster C; (36) (37) (38) update the cell; delete p from (39) (40) (41) //Second-round Merging (42) (43) Merge all c-clusters by Merging 1; (44) Merge all c-clusters by Merging 2; (45)
In the phase, r reducers execute the Merging-Refinement-Merging process in parallel. However we still need to perform one reducer to return final clusters on one machine in final phase as shown in Figure 3.
6. Experimental Study
6.1. Experimental Setup and Methodologies
A comprehensive performance study has been conducted on 10 network-connected commodity computers that deployed Hadoop platform to evaluate the effectiveness and efficiency of the proposed Cludoop algorithm. Each PC/node is equipped with an Intel Core 2 Duo P8600 processor with 8 GB memory and runs a Ubuntu 11.4 operating system. One node is configured as both NameNode and JobTracker. Other nodes are configured as DataNode and TaskTracker.
Real Datasets. We use two real datasets in our experiments. The first real dataset is a trajectory database from
Synthetic Datasets. We also generate two synthetic datasets by our developed data generator using Matlab. The first synthetic dataset, denoted by
The second large-scale dataset
Alternative Algorithms. Our experiments focus on evaluating the effectiveness and efficiency of proposed Cludoop. For the effectiveness evaluation, we compare directly clustering results of our algorithm against the most wildly used method DBSCAN. For the efficiency evaluation, we compare the CPU time utilized by our algorithm against the state-of-the-art method MR-DBSCAN as introduced in Section 2.
Metrics and Methodology. We measure the quality of clusters for effectiveness by Precision and Recall as follows:
We also evaluate the performance of Cludoop method by varying the most important parameters. In particular, we measure the scalability of methods by varying the volume of dataset and the number of work nodes. Moreover, we also measure sensitivity of our Cludoop algorithm on efficiency by varying the input parameter
6.2. Effectiveness Evaluation
First, we evaluate the effectiveness of our proposed algorithm compared to DBSCAN using two evaluation metrics (visualization comparison, Precision/Recall) on Ssyn and GeoLife data.
Figure 4 shows the clustering result comparison of two algorithms on Ssyn data when
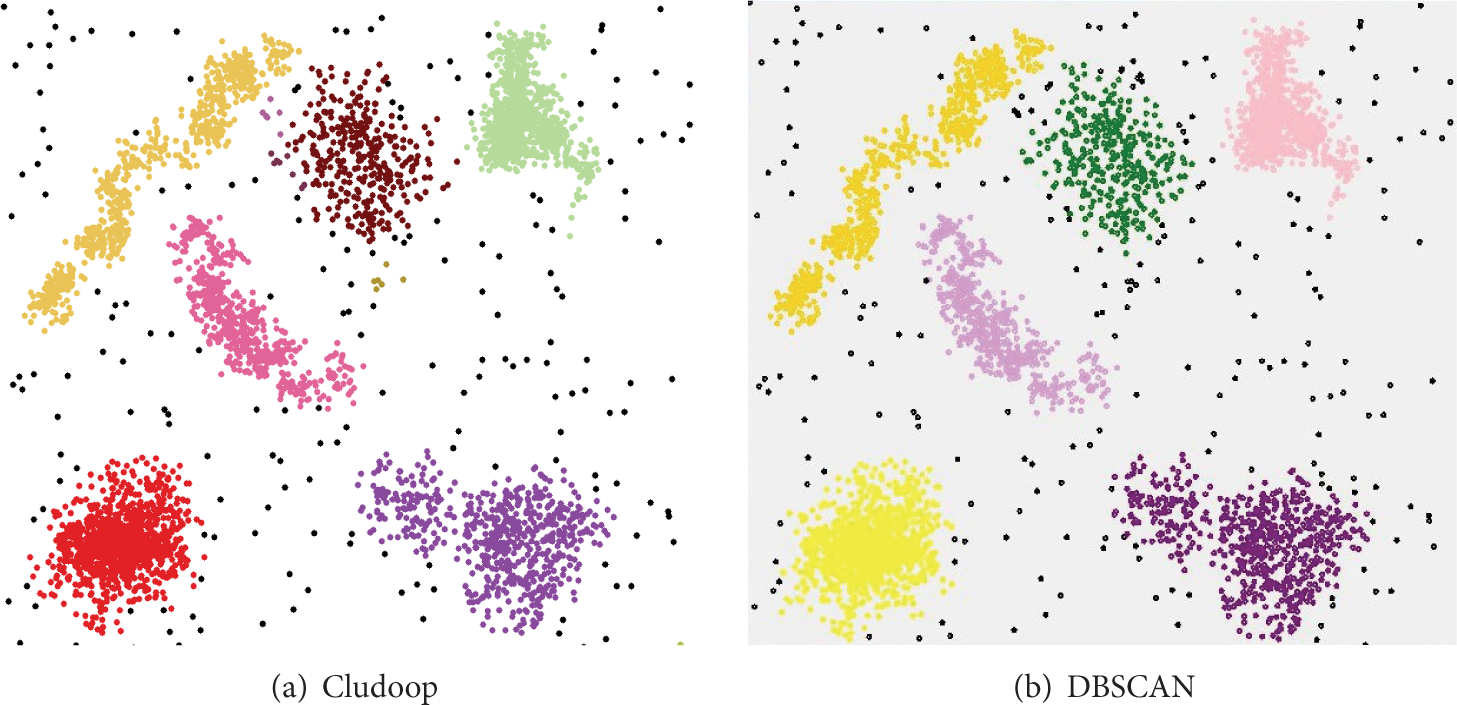
Visualization comparison on Ssyn data.

Visualization comparison on GeoLife data.
The Precision and Recall of Cludoop on Ssyn data are shown in Figure 6. For Ssyn data, the Precision and Recall are computed based on labelled points. From Figure 6(a), the Precision is nearly 100% once the parameter

Effectiveness evaluation on Ssyn data.

Effectiveness evaluation on GeoLife data.
In summary the effectiveness study confirms that our distributed algorithm can obtain the correct clusters with noise under a loose parameter setting.
6.3. Efficiency Evaluation
Next we evaluate the efficiency of our algorithm compared to MR-DBSCAN algorithm using the Taxi and Lsyn data. We vary the most important parameters, to (1) assess the scalability of Cludoop versus MR-DBSCAN in terms of efficiency and (2) evaluate sensitivity of parameter variation on our method.
6.3.1. Varying Volume of Dataset D
We first evaluate the scalability of our algorithm in terms of the volume of dataset using Lsyn data. In this experiment we randomly extract four subsets from the Lsyn data from 20% to 80%. We fix the
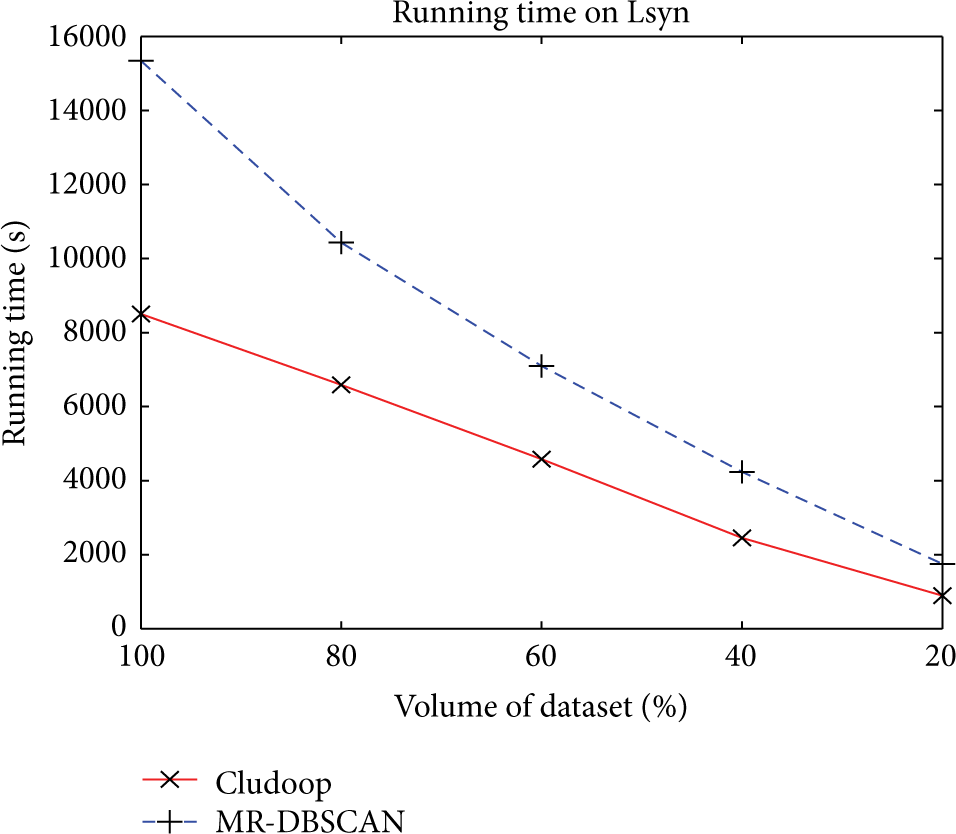
Varying volume of dataset D on Lsyn data.
6.3.2. Varying Number of Nodes
Next, we valuate the speedup of our algorithm as the number of nodes increases by varying work node from 2 to 9 on the Taxi data when

Varying number of nodes
6.3.3. Varying Neighbor Range Threshold
Finally, we analyze the sensitivity of our algorithm with respect to the important parameter

Varying neighbor range

Varying neighbor range
7. Conclusion
Density-based clustering for increasing big data applications is very important yet difficult task. This paper proposes an efficiency and load-balanced distributed density-based clustering for big data using existing data partition on Hadoop platform. Our algorithm incorporates a proposed serial clustering with c-cluster as plug-in on mapper and a Merging-Refinement-Merging 3-step framework to fast merge c-cluster on reducer. The empirical study on large-scale real and synthetic data shows that our Cludoop algorithm effectively finds the correct clusters and exhibits better scalability and efficiency than the state-of-the-art. An interesting direction for future work is to leverage optimized shuffle mechanism for further improving the workload balancing problem of clustering on Hadoop platform.
Footnotes
Conflict of Interests
The authors declare that there is no conflict of interests regarding the publication of this paper.
Acknowledgments
This work is partially supported by the National Natural Science Foundation of China (nos. 61403328, 61403329, 61302065, and 61172049), the open project program of Key Laboratory of Symbolic Computation and Knowledge Engineering of Ministry of Education, Jilin University (no. 93K172014K13), the Shandong Provincial Natural Science Foundation (no. ZR2013FM011), and the project of Shandong Province higher educational science and technology program (no. J14LN24).
