Abstract
Introduction:
Widespread transmission of Japanese encephalitis virus (JEV) genotype four (GIV) occurred across mainland Australia in 2022. This resulted in forty-five human cases, including seven deaths, and the identification of JEV infection in over 80 commercial piggeries.
Materials and Methods:
We collected mosquitoes which were trapped using CO2-baited light traps deployed near piggeries reporting disease or in regions linked to human cases in the Wide Bay region in the state of Queensland. Mosquitoes from four traps yielded JEV RNA by real-time RT-PCR. Pools containing RNA positive mosquitoes were inoculated onto mosquito cell monolayers.
Discussion:
A single isolate of JEV was obtained from a pool of mixed mosquito species. Near whole genome sequencing and phylogenetic analysis of the JEV isolate demonstrated its high genomic relatedness with JEV GIV pig sequences sampled from Queensland and the state of New South Wales in 2022.
Conclusion:
We report the first isolation of JEV GIV from mosquitoes collected in Australia. With only a few JEV GIV isolates available globally, the isolate we report will be essential for future research of JEV host interactions, evolution and disease markers, and development of effective therapies, vaccines, diagnostic assays, and mosquito control strategies.
Introduction
Japanese encephalitis virus (JEV) is an arthropod-borne virus (arbovirus) containing a positive-sense, single-stranded RNA genome. This encephalitogenic, zoonotic flavivirus causes an estimated 68,000 human clinical cases of Japanese encephalitis (JE) annually in Southeast Asia and the Indian subcontinent (Campbell et al., 2011). The virus exists in a zoonotic transmission cycle. Culex mosquitoes are primary vectors, wading birds and pigs are main amplifying hosts, and humans are among dead-end hosts.
Five JEV genotypes (GI-V) are classified according to genetic sequence phylogenetic analyses (Mackenzie et al., 2022).
Incursions of JEV into Australia have occurred on three recognized occasions, with GII and GI viruses isolated in 1995 and 2000, respectively (Mackenzie et al., 2022). In February–March 2022, JEV was identified as the cause of stillborn and mummified fetuses, and neurological disease in neonate piglets at commercial piggeries in three south-eastern states of Australia [World Organisation for Animal Health (WOAH) 2022]. The first human JE cases were diagnosed subsequently, and by the end of 2022, 45 human cases and 7 deaths had been reported and over 80 piggeries had been affected [Australian Government Department of Agriculture, Fisheries and Forestry (DAFF) 2023; Australian Government Department of Health and Aged Care (DoHAC) 2023].
In response to the emergence of JEV in the Wide Bay region of the state of Queensland (QLD), mosquitoes were collected in 2022 near commercial piggeries that had confirmed JEV pig infections and from locations where human cases may have been infected. We report the isolation and genomic characterization of a JEV genotype four (GIV) isolate from these collections, which is the first GIV virus isolated from mosquitoes collected in Australia. A comprehensive description of the entomological and epidemiological investigations into the emergence of JEV GIV in Queensland will be published elsewhere.
Materials and Methods
Mosquitoes were collected using CO2-baited Pacific Biologics light traps (Pacific Biologics, Scarborough, Australia) from the North Burnett local government area, Wide Bay region, QLD, during March–April 2022. Mosquito collections were sent frozen on dry ice to Brisbane, QLD. To determine species present in the collections, mosquitoes were morphologically identified on a chill table using the taxonomic key of Marks et al. (1982) before the contents of each trap collection were recombined for virus processing.
Mosquito collections were tested for virus in mixed-species pools of ≤50 mosquitoes which were prepared according to the protocol of Jansen et al. (2019). Nucleic acids were extracted using the Kingfisher Flex Purification System and MagMAX™ viral/pathogen II nucleic acid isolation kit (Thermofisher Scientific™, Australia) according to the manufacturer's instructions. Specific JEV RNA detection by reverse transcription, TaqMan™ real-time polymerase chain reaction (RT-rPCR), JEV isolation from a mosquito pool and next-generation sequencing of the JEV isolate were performed as previously described or a modification of published methods (Pyke et al., 2020). Further description of these methods is provided in the Supplementary Material. This work has received ethical clearance from the Forensic and Scientific Services Human Ethics Committee, approval reference HEC 24-01.
Results and Discussion
A total of 4,010 mosquitoes collected in 34 traps were processed in 90 pools. JEV RNA was detected in 15 pools of mosquitoes from 4 traps (Supplementary Table S1). Virus isolation was attempted from 9 pools, and a virus isolate (QLD_S46716_M2022) was obtained from a single pool of mosquitoes collected from the Mundubbera region on March 15, 2022 (Fig. 1A). Limited isolate recovery from JEV RT-rPCR positive pools is likely owing to low viable virus loads in most samples as indicated by high cycle threshold values ≥27. In three of the four JEV RT-rPCR positive traps, recognized JE vectors (Mackenzie et al., 2022) comprised >95% of mosquitoes. However, known JEV vectors constituted only 7% of the 418 mosquitoes in the trap collection which yielded the isolate, with the dominant species being Aedes vittiger (89%). QLD_S46716_M2022 is the first JEV GIV isolate derived from Australian mosquitoes and to our knowledge, only the second GIV isolate that has been recovered from the 2022 transmission of JEV GIV on mainland Australia, the other being derived from a pig in New South Wales (NSW).
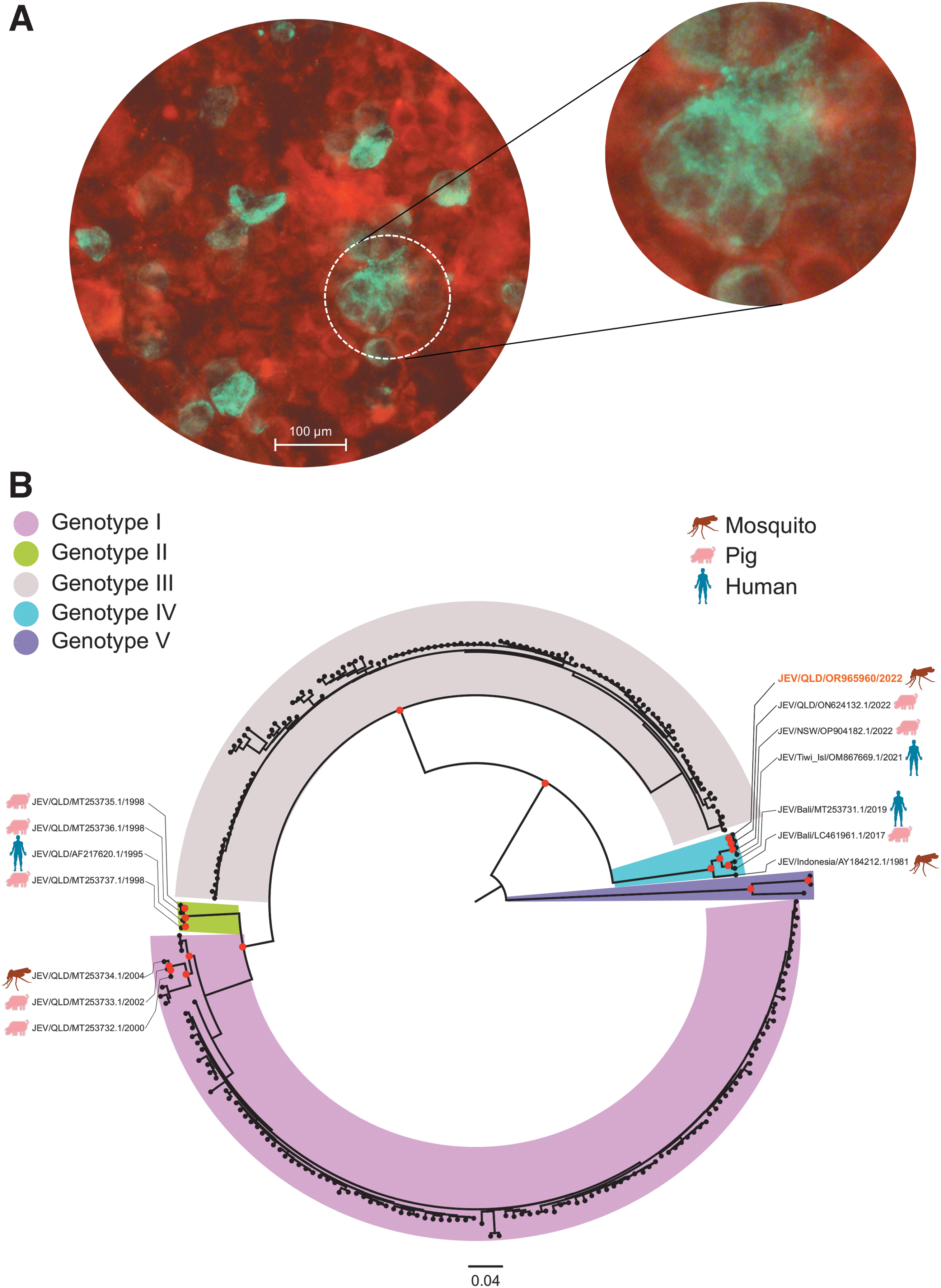
Isolation of JEV and phylogenetic characterization.
Phylogenetic analyses comparing 247 JEV complete open reading frames (ORFs) demonstrated the JEV QLD_S46716_M2022 isolate (GenBank accession number OR965960) grouped with GIV (Fig. 1B). Although this virus was passaged in both mosquito and vertebrate cells, it shared 99.9% nucleotide identity with Australian JEV GIV 2022 pig sequences ON624132.1 (QLD) and OP904182.1 (NSW), and JEV GIV 2021 sequence OM867669.1 obtained from a fatal human JE case from the Tiwi Islands (Waller et al., 2022). In comparison, these four sequences which clustered within GIV were less closely related to other previous JEV sequences obtained in QLD. This included the sequence from the first reported fatal human case of JE caused by GIV (MT253731) obtained from a traveler returning to Australia from Bali, 2019 (Pyke et al., 2020) which shared 96.7% nucleotide identity with JEV QLD_S46716_M2022. The remaining QLD human, mosquito, and pig JEV GI and GII sequences collected between 1995 and 2004 (Fig. 1B) all shared >85.0% nucleotide identity with QLD_S46716_M2022.
Further scrutiny of amino acid (aa) coding sequences demonstrated QLD_S46716_M2022 was most closely related to the QLD GIV 2022 pig sequence (ON624132.1) with which it shared 99.97% aa identity and differed from it by only one nonconserved aa change, R493K in the NS3 gene. However, other recent GIV NSW (OP904182.1), Tiwi Islands (OM867669.1), and Bali 2019 (MT253731.1) sequences all contained a K residue at NS3 position 493 identical to QLD_S46716_M2022. Interestingly, the cluster of 4 GIV sequences containing QLD_S46716_M2022 (Fig. 1B) all contained at least 23 aa changes within the JEV ORF and were distinctly different to the recent Bali 2019 (MT253731.1) sequence obtained from nearby Indonesia.
The transmission dynamics and disease potential of JEV GIV is largely uncharacterized but before 2022, infections of this emerging genotype had caused two known human fatalities (Pyke et al., 2020; Waller et al., 2022). Notably, it was the only genotype detected during the Australian 2022 mainland JEV outbreak [Australian Government Department of Health and Aged Care (DoHAC) 2023]. The recovery of representative GIV isolates, such as the one reported herein, is essential for viral phenotypic experiments including assessment of mosquito vector competence, and in vitro and in vivo pathogenicity studies which will assist future research of JEV host interactions, evolution, and potential disease markers, and the development of effective therapies, vaccines, diagnostic assays, and mosquito control strategies.
Footnotes
Acknowledgments
We are sincerely grateful to Public Health Virology staff at Forensic and Scientific Services, Queensland Health for their assistance in mosquito sample preparation, and we acknowledge the effort of Public Health Units, particularly the Darling Downs Public Health Unit, and local government personnel, who performed additional mosquito collections in response to JEV emergence in 2022.
Author Disclosure Statement
No conflicting financial interests exist.
Funding Information
This project was funded by the Department of Health, Queensland Government.
Supplementary Material
Supplementary Data
Supplementary Table S1
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
