Abstract
Background:
Anaplasma ovis are obligate intracellular bacteria that can endanger human and animal health, and they can be transmitted by arthropod vectors, such as Melophagus ovinus and ticks.
Materials and Methods:
In this study, 433 specimens, including 370 M. ovinus and 63 sheep blood samples, were collected from nine districts of South Xinjiang to investigate the distribution and molecular epidemiology of A. ovis in M. ovinus and small ruminant.
Results:
DNA of A. ovis was detected in 109 (25.2%, 109/433) of the 433 samples using PCR and sequencing. The analysis of A. ovis msp4 sequences revealed four different genotypes, including genotype III (47.7%; 52/109), GB3 (34.0%; 37/109), AoGOv3 (15.6%; 17/109), and XJ9 (2.8%; 3/109).
Conclusions:
To the best of our knowledge, A. ovis genotypes GB3, AoGOv3, and XJ9 detected in this study are the first to be reported in M. ovinus, and our data indicate that XJ9 is a novel A. ovis genotype presented herein for the first time. These findings provide important references for the new understanding and prevention of A. ovis in border counties in China.
Introduction
The family Hippoboscidae (Diptera) contains several hematophagous ectoparasites (Werszko et al, 2021). Melophagus ovinus (sheep ked) is the most widespread hematophagous ectoparasite that lives on many animals, including sheep, goats, rabbits, dogs, Tibetan antelope, European bison, and red foxes, it has also been detected in humans (Zhang et al, 2021), which is extensively distributed in North America, Oceania, Asia, Africa, and Europe (Werszko et al, 2021). Currently, M. ovinus has been reported in Tibet, Xinjiang, Liaoning, Qinghai, and Gansu and has been detected on imported sheep and sheep wool at quarantine ports in China (Liu et al, 2018).
The life cycle of M. ovinus consists of three stages: larva, pupa, and adult, and they often occur in the fleece of the host and can be carried from one animal to another by direct contact (Small, 2005). M. ovinus can cause damage of the skin, restlessness, anemia, weight loss, pruritus, anxiety, cutaneous myiasis, poor wool quality, and reduced wool growth (Liu et al, 2018; Zhang et al, 2021), and can transmit several microorganisms, some of which are zoonotic pathogens, such as Rickettsia spp., Anaplasma spp., Theileria spp., Bartonella spp., Borrelia burgdorferi, Trypanosoma spp., bluetongue virus, and border disease virus (Casco et al, 2021; Liu et al, 2019). In addition, it can also harbor abundant bacteria, including Wolbachia, Acinetobacter, Halomonas, Shewanella, Enterobacter, Bacillus, and Staphylococcus (Duan et al, 2020; Duan et al, 2017). Therefore, the livestock industry suffers significant economic losses due to M. ovinus.
Anaplasma species are small gram-negative, obligate intracellular bacteria belonged to the order Rickettsiales, family Anaplasmataceae (Stuen et al, 2013). Anaplasma spp. have become a global public health concern, among which Anaplasma ovis, Anaplasma phagocytophilum, and Anaplasma capra are the most widespread zoonotic pathogens. Anaplasma is responsible for causing anaplasmosis in a range of hosts, including humans (e.g., A. phagocytophilum), livestock (e.g., A. phagocytophilum, Anaplasma bovis, Anaplasma marginale, and A. ovis), and pets (e.g., Anaplasma platys) (Cabezas-Cruz et al, 2019; Sonenshine and Stewart, 2021). Ticks are the main carriers of these pathogens, however, these pathogens have also been reported to infect other arthropods (e.g., Culex quinquefasciatus, Ctenocephalides felis, Simulium damnosum, and M. ovinus) (Hornok et al, 2011; Sonenshine and Stewart, 2021).
A. ovis was first discovered in sheep in 1912, which is widely distributed in Asia, Europe, and Africa, and has been identified in goats, sheep, cattle, Tian Shan wapiti, and some wild ruminants (He et al, 2021; Hornok et al, 2007; Li et al, 2021; Yan et al, 2020; Yang et al, 2019). A. ovis is the etiologic agent of anaplasmosis in small ruminants and is the primary cause of host erythrocyte invasion (Cabezas-Cruz et al, 2019), which can frequently cause subclinical and persistent infection in small ruminants, particularly the older, malnourished, immunocompromised, and chronically debilitated animals (Cabezas-Cruz et al, 2019).
Clinical manifestations of the disease include fatigue, anorexia, progressive anemia, icterus, weight loss, milk yield decrease, and abortion; however, the most common symptom is fever (Belkahia et al, 2014; Cabezas-Cruz et al, 2019). Coinfection with A. ovis and other microbes or parasites can result in more severe outcomes than single infection (Renneker et al, 2013). In addition, A. ovis appears to be a potential zoonotic pathogen, as it has been detected previously in a patient from Cyprus (Chochlakis et al, 2010).
Currently, only limited studies have investigated the prevalence and molecular epidemiology of A. ovis in South Xinjiang, China (Song et al, 2018; Zhao et al, 2018). Analysis of the major surface protein 4 (msp4) gene indicated that it is highly conserved in Anaplasma spp. (Battilani et al, 2017; de la Fuente et al, 2007; de la Fuente et al, 2005a). A. ovis is a genetically diverse bacteria that can be classified into multiple genotypes based on msp4 gene using molecular evolutionary analyses (Belkahia et al, 2014; Ben Said et al, 2015; de la Fuente et al, 2007).
PCR amplification of A. ovis msp4 has high specificity and diagnostic accuracy (Belkahia et al, 2014; de la Fuente et al, 2007; Hornok et al, 2007; Torina et al, 2010; Zhao et al, 2018). Therefore, we aimed to determine the prevalence of A. ovis in M. ovinus and its geographical distribution in South Xinjiang, further to investigate the genetic variability of A. ovis in this region.
Materials and Methods
Sample collection
A total of 370 samples of M. ovinus were collected at 11 different sampling points of 9 counties in South Xinjiang from June to mid-October 2021 and during March 2022 (Fig. 1). Furthermore, blood samples were randomly collected from seven sheep via the jugular vein in each county, a total of 63 samples were collected into tubes coated with ethylenediaminetetraacetic acid (Table 1). M. ovinus and blood samples were transported to the Engineering Laboratory of Tarim Animal Diseases Diagnosis and Control, M. ovinus were stored in 75% ethanol at 4°C and blood samples at −20°C until analysis. Specimen information including geographic origin and host was recorded.
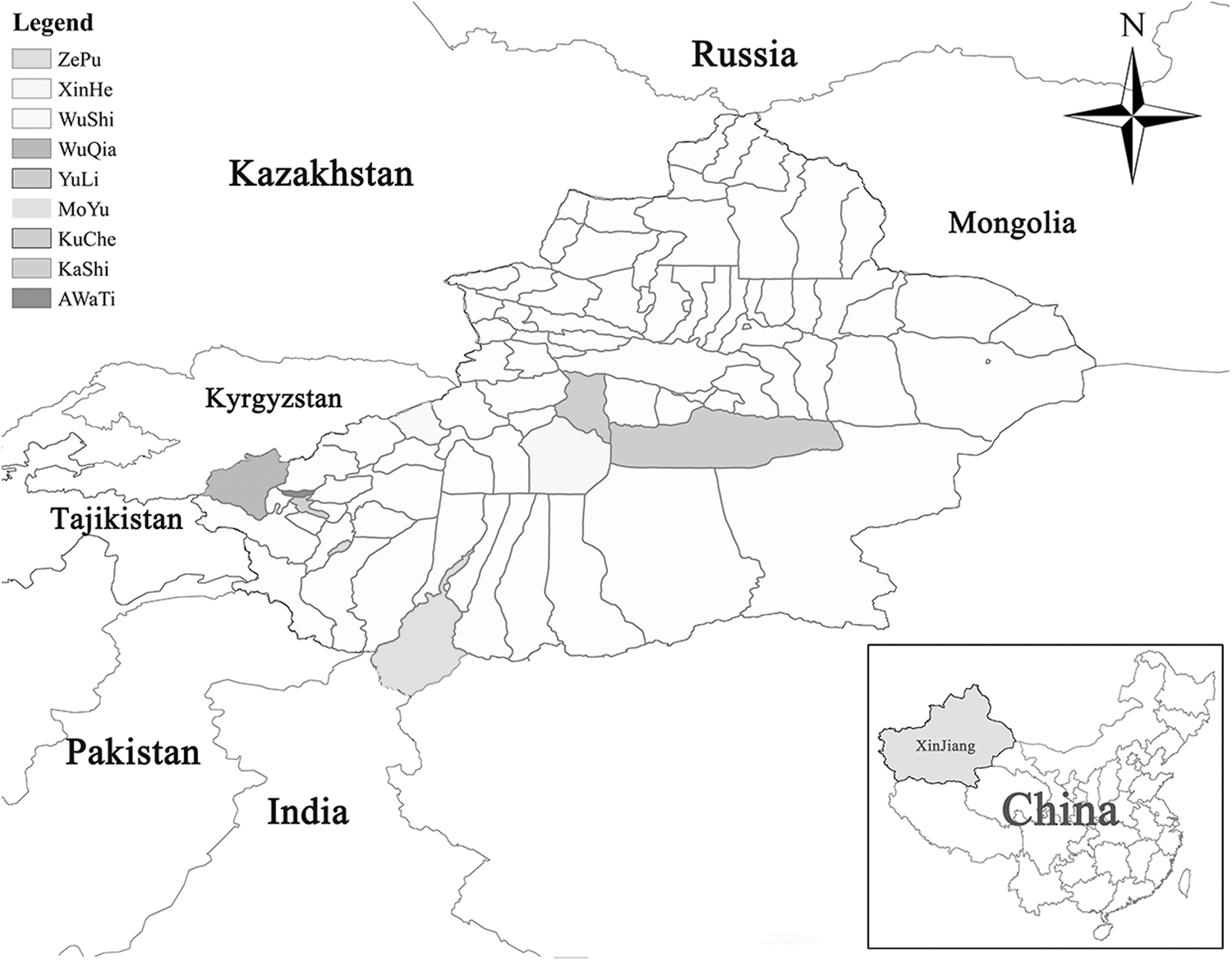
Distribution map of sampling districts of nine counties and cities in southern Xinjiang of China.
The Prevalences of Anaplasma Ovis Different Genotypes in Melophagus Ovinus and Sheep in Border Areas of South Xinjiang
Collected in 2021.
Collected in 2022.
Ⅲ, Anaplasma ovis genotypes Ⅲ; GB3, Anaplasma ovis genotypes GB3; AoGOv3, Anaplasma ovis genotypes AoGOv3; XJ9, Anaplasma ovis genotypes XJ9, “—”, no pathogen infection; CI = 95% confidence interval.
Demographic information of the study population
A total of 433 specimens were studied, among which 140 (32.3%) samples were collected from border region, 172 (39.7%) from nonborder, and 121 (27.9%) from central region. Most animals are free-ranging, there are 137 (31.6%) and 296 (68.4%) samples from animals that maintained in captivity and grazed year-round, respectively. Prevention-based sampling analysis indicated that the samples from animals without anthelmintic treatment were 358 (82.7%), and 75 (17.3%) from animals that received anthelmintic treatment.
M. ovinus morphological identification and total DNA extraction
The sex of the adult M. ovinus was identified according to the morphological features visualized under a stereomicroscope (LEICA M165 C; Solms, Germany), as described by Duan et al (2017). All M. ovinus were sterilized by washing with a graded ethanol series (70%, 50%, 30%, and 10%) under a constant temperature oscillator for 30 min at 37°C and 180 rpm. Subsequently, the samples were washed twice with sterile deionized water to remove potential nucleic acid contaminants, dried on sterile filter paper, cut into pieces with a sterilized scalpel, and placed into 2 mL sterile microcentrifuge tubes. DNA was extracted from the processed M. ovinus samples and 500 μL of whole blood using the TIANamp Genomic DNA Kit (TIANGEN Corporation, Beijing, China) according to the manufacturer's instructions. Extracted DNA was stored at −20℃ until PCR was performed.
Amplification and sequencing of A. ovis msp4 gene
PCR targeting msp4 gene was performed to determine A. ovis infection in sheep herds. Amplification of the 867 bp fragment was carried out using previously designed primers (MSP4-F:5′-GGG AGC TCC TAT GAA TTA CAG AGA ATT GTT TAC-3′/MSP4-R:5′-CCG GAT CCT TAG CTG AAC AGG AAT CTT GC-3′) (de la Fuente et al, 2007). The PCR mixture, which had a total volume of 25 μL, contained 13 μL of 2 × Taq PCR Master Mix (Tiangen), 1 μL of the relevant primers (10 μM final concentration), 9 μL nuclease-free deionized water, 1 μL template DNA, and was performed under the following amplification conditions: denaturation at 95℃ for 5 min; 35 cycles of 94°C for 30 s, 54°C for 40 s, and 72°C for 1 min, followed by final elongation at 72°C for 10 min.
For quality control, genotype III DNA of A. ovis stored in our laboratory was used as positive control, nuclease-free sterile distilled water as a negative control to monitor contamination. Gel electrophoresis of the PCR product was performed by loading 5 μL onto 1.5% agarose gel containing GelStain (Beijing TransGen Biotech Co., Ltd., Beijing, China) and was subsequently visualized using a gel documentation system (FluorChem, ProteinSimple, CA, USA). Amplicons were submitted for sequencing at GENEWIZ, Inc. (Suzhou, China), genomic sequences were analyzed and compared using BLAST (
The A. ovis amino acid sequence was deduced from the corresponding genomic sequences using DNAStar, EditSeq software (DNASTAR, Inc.) and amino acid sequence alignment was performed using MEGA 7.0 (
Statistical analysis
All statistical analyses in this study were carried out using SPSS 22.0 statistical software (IBM SPSS, Chicago, USA). The potential association and statistical significance between the pathogen prevalence and other independent variables are evaluated with multivariable logistic regression models.
Ethical approval
The study was conducted in compliance with animal care guidelines of the Ethics Committee of Tarim University, Xinjiang, China. Sheep blood samples were obtained in accordance with good animal practices required by the Animal Ethics Procedures and Guidelines of the People's Republic of China.
Results
From June to mid-October 2021 and in March 2022, a total of 433 samples, including 370 adult M. ovinus and 63 sheep blood samples, were collected from 11 sampling points of nine counties in South Xinjiang (Table 1 and Fig. 1). During sample collection, we observed that most of the M. ovinus were mainly attached to the neck of sheep and often existed alone, the biting site exhibited erythema and inflammatory tumefaction on the skin surface. Among these M. ovinus, 198 (53.5%) were morphologically identified as male and 172 (46.5%) as female.
Detection of A. ovis
A. ovis in 63 blood samples collected from sheep and 370 M. ovinus samples from South Xinjiang were detected by targeting msp4 gene. Among the 370 M. ovinus, A. ovis was detected in 88 (23.8%) of the samples, including 32/198 (16.2%) from males and 56/172 (32.6%) from females, A. ovis were also detected in 21/63 (33.3%) sheep blood samples. Further Amino acid sequence alignment revealed that nucleotide changes resulted in amino acid variations (Table 2). The sequences detected in our study had 99.7–100% nucleotide similarity, and four different genotypes based on four distinct nucleotide positions were confirmed (Table 2), including genotype III (n = 52), GB3 (n = 37), XJ9 (n = 3), and AoGOv3 (n = 17) (Table 1).
Differences in the msp4 Nucleotide and Amino Acid Sequences Among Anaplasma ovis Strains
Asterisks indicate nucleotides identical to the reference Idaho (GenBank acc. no. AF393742) isolate sequence; Amino acid changes are indicated between parentheses with single letter code. (amino acid: R, Arginine; I, Isoleucine; V, Valine; A, Alanine; L, Leucine; Nucleotide: T, Thymine; C, Cytosine; G, Guanine; A, Adenine).
Idaho, Italy 147, Italy 20, B3, and Al2 samples were collected from sheep. MD samples were collected from mule deer. BS samples were collected from bighorn sheep. XJ3, XJGB3, XJ8Gov3, and XJ9 samples were collected from M. ovinus.
GenBank accession number of the variant.
Numbers represent the nucleotide position starting at translation initiation codon Adenine.
A. ovis XJ3 (GenBank acc. no. OP169076) exhibited 100% identity with genotype Ⅲ, which is mostly represented by “Italy20” A. ovis strains isolated from Sicilian sheep (GenBank acc. no. AY702923). A. ovis XJGB3 (GenBank acc. no. OP169077) revealed 100% sequence identity with genotype GB3 that is mostly represented by “GB3” A. ovis strains from Tunisian sheep (GenBank acc. no. KC432643). In contrast, A. ovis XJ8Gov3 (GenBank acc. no. OP169078) demonstrated 100% nucleotide identity with “A. ovis A-54” (GenBank acc. no. MT311201) from Pakistani sheep and the AoGOv3 genotype represented by the “A. ovis AoGOv3” strain (GenBank acc. no. KM285220) from Tunisian sheep. In addition, a novel A. ovis msp4 genotype (XJ9, GenBank acc. no. OP169079) was identified, phylogenetic analysis revealed that XJ9 was classified into a separate cluster and shared high similarity (99.9%) with A. ovis genotype III (GenBank acc. no. AY702923) (Table 2; Fig. 2).
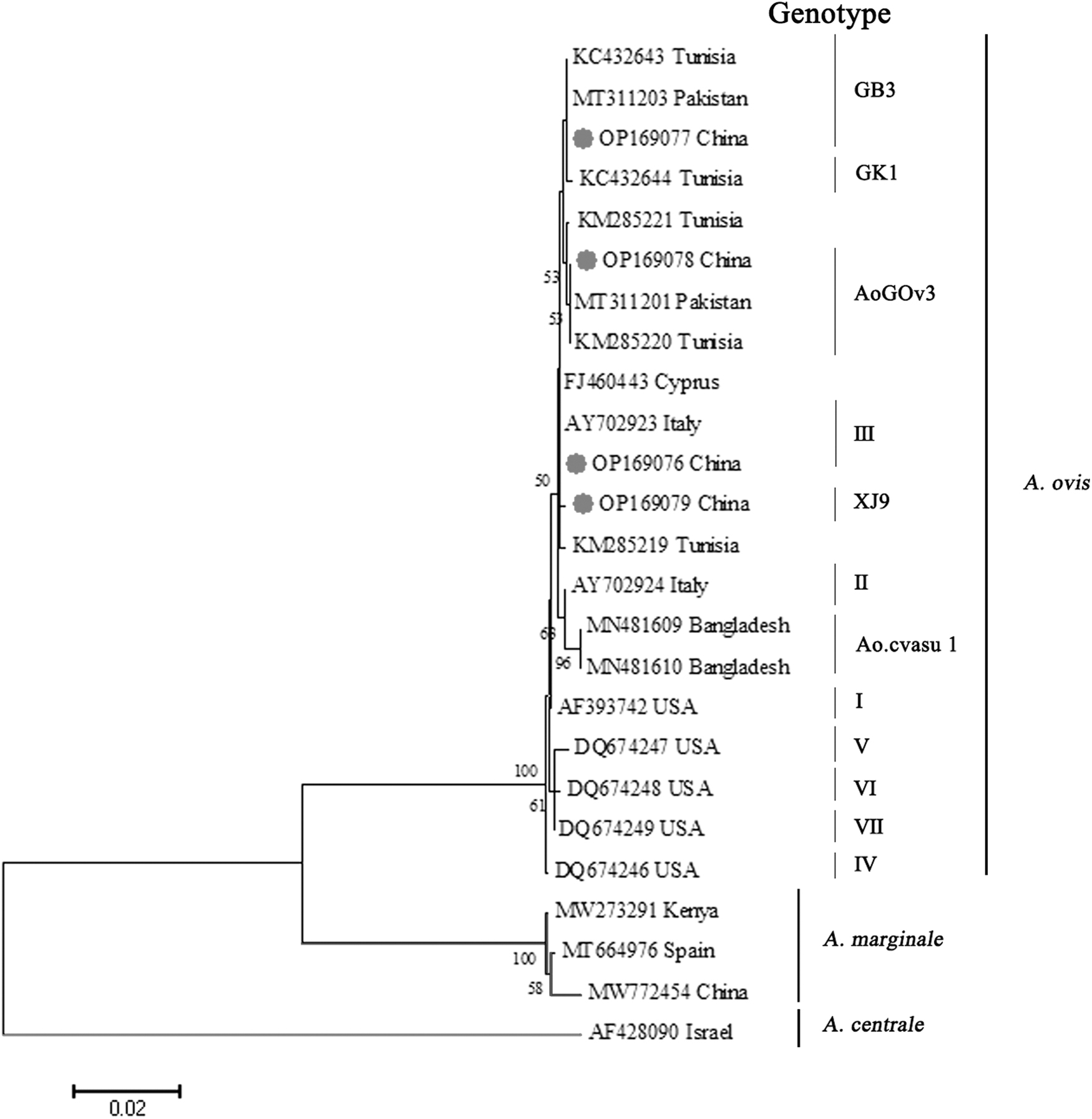

Phylogenetic tree of Anaplasma ovis based on the msp4 partial gene sequence. The tree was constructed with the ML (bootstrap replicates: 1000) phylogenetic analysis on the basis of MEGA10. Sequences of our specimens are marked with red circles. ML, maximum likelihood.
Risk factor analysis of A. ovis occurrence
The highest and lowest prevalence of A. ovis were in WuShi (24/37; 64.9%) and KuChe districts (5/67; 7.5%), respectively. The risk factor analysis for A. ovis showed a higher prevalence in the border region (50/140; 35.7%) than in the nonborder region (37/172; 21.5%) and central region (22/121; 18.2%) (p < 0.001). Moreover, A. ovis was more common among animals that grazed year-round (86/296; 29.1%) compared to those that did not graze at all (23/137; 16.8%) (p < 0.001). Using vitro acaricide treatment (11/75; 14.7%) significantly reduced A. ovis prevalence compared to those without acaricide treatment (98/358; 27.4%) (p < 0.001).
Multivariable logistic regression analysis revealed that there was a potential association between the pathogen prevalence as a dependent variable and other independent variables, including acaricide treatment (odds ratio [OR] = 1.64; [confidence interval; CI 0.75 − 3.59]; p < 0.001), rearing paradigms (OR = 2.08 [CI 1.07 − 4.07]; p < 0.001), and geographic location (OR = 0.66 [CI 0.48 − 0.89]; p < 0.05).
Discussion
Previous studies have confirmed the presence of A. ovis in M. ovinus (Zhang et al, 2021; Zhao et al, 2018), however, this is the first report of molecular detection and phylogenetic analysis, in which we describe the overall prevalence of A. ovis and respective genotypes in distinct geographic regions across South Xinjiang, China. Notably, our investigation demonstrated that A. ovis is highly prevalent in the blood and M. ovinus from small ruminants in South Xinjiang, this result is consistent with previous reports that confirmed the widespread prevalence of this pathogen (Li et al, 2021; Li et al, 2020; Song et al, 2018; Zhao et al, 2018).
In China, A. ovis has mainly been reported in Xinjiang Uygur Autonomous Region, Ningxia Hui Autonomous Region, Qinghai Tibet Plateau, Inner Mongolia Autonomous Region, Gansu provinces, Shaanxi Province, and Anhui Province (Lu et al, 1997; Song et al, 2018; Wang et al, 2021; Yang et al, 2019; Zhang et al, 2021). The overall infection rate of our study was 25.2% (109/433), which is higher than that previously reported in Senegal (11.5%) (Djiba et al, 2013) but similar to that previously reported in China (25.6%) (Chi et al, 2013), and lower than that reported in Spain (91.1%) (Lacasta et al, 2021), Tunisia (65.3%) (Ben Said et al, 2015), and Turkey (46.6%) (Benedicto et al, 2020).
The prevalence of A. ovis genotype III was 47.7% (52/109), which was higher than previous report in China (32.1%) (Yang et al, 2022). Genotype III A. ovis has also been detected in Turkey (Benedicto et al, 2020), Pakistan (Niaz et al, 2021), Tunisia (Ben Said et al, 2015), and Italy (de la Fuente et al, 2005b), this indicate that A. ovis genotype III was common in most countries. The prevalence of A. ovis genotype GB3 was 34.0% (37/109), which was higher than previous report in Tunisia (32.6%) (Belkahia et al, 2014) and Pakistan (24.7%) (Niaz et al, 2021).
The prevalence of A. ovis genotype AoGOv3 was 15.6% (17/109), which was lower than that previously reported in Pakistan (24.7%) (Niaz et al, 2021), this genotype was first identified in Tunisia in 2015 (Niaz et al, 2021). We also identified a novel genotype, XJ9, with 2.8% (3/109) prevalence. The variation of prevalence among these studies may be attributed to different climatic condition, seasonal change, selective pressure, feeding pattern, and the number of study samples, as previous report (Belkahia et al, 2014; Cabezas-Cruz et al, 2019; Torina et al, 2008).
Human anaplasmosis caused by A. ovis is extremely rare, the only case with fever, lymphadenopathy, and hepatosplenomegaly was reported in Cyprus in 2007 (Chochlakis et al, 2010). In the present study, the msp4 sequence of A. ovis XJ3 (acc. no. OP169076) detected in M. ovinus and sheep achieved 99.8% similarities with the human strain (FJ460443), and were clustered into the same branch on the phylogenetic trees (Fig. 2), this indicates that the human strain from Cyprus is similar to genotype III. Therefore, the high prevalence of genotype III strains detected in M. ovinus and sheep may pose a threat to human health.
In recent years, several genotypes of A. ovis have been reported in small ruminants from Pakistan (acc. nos.: MT311203 and MT311201) (Niaz et al, 2021), these strains shared 100% sequence similarity with genotypes XJGB3 (acc. no. OP169077) and XJ8Gov3 (acc. no. OP169078), respectively, which were distinctly clustered into two branches in our study (Fig. 2), belonged to genotype GB3 and AoGOv3, respectively. China is contiguous with Pakistan, however, if the genotypes XJGB3 and XJ8Gov3 of A. ovis were originated from Pakistan or other contiguous countries, a systemic epidemiological study should be carried out. In our study, genotype XJ9 (acc. no. OP169079) of A. ovis was discovered for the first time from M. ovinus and sheep (originated from KaShi and ZePu County), this genotype has never been reported at one time, it maybe a mutant generated under specified environment.
A. ovis has been proved to be a vertically communicable pathogen, and the prevalence of A. ovis was higher in female sheep (Niaz et al, 2021; Zhao et al, 2018), once the animal has an A. ovis infection, without adequate treatment, the pathogen can be carried for several years (Jiménez et al, 2019; Yasini et al, 2012).
In the present study, a higher prevalence of A. ovis was observed in females M. ovinus than in males, which poses a serious threat not only to animals but also to human, the difference may be due to several factors, including the reproductive characteristics of M. ovinus, the effect of different sexes on M. ovinus life cycle (Small, 2005), and the vertically communicable trait of A. ovis in M. ovinus (Zhao et al, 2018). In addition, our risk factor analyses suggested that a combination of acaricide treatment, rearing paradigms, and geographic location can influence the prevalence of A. ovis in animals, therefore, the persistent improvement of feeding pattern and regular anthelmintic treatment will help decrease the prevalence of A. ovis in domestic livestock.
Conclusions
This study reported a high prevalence of A. ovis in sheep and M. ovinus from nine border counties in southern Xinjiang, China. XJ9 was identified as a novel genotype in our study, and A. ovis genotypes GB3 and AoGOv3 were first reported in M. ovinus. The result of this study provided a critical epidemiological information of A. ovis, which can give a remarkable improvement to prevent and control A. ovis in this region. Further study should be carried out to evaluate the pathogenicity of different genotypic A. ovis, a unified nomenclature for A. ovis genotype is required to facilitate global communication and epidemiological reference.
Footnotes
Acknowledgment
We thank Professor YuanzhiWang (Department of Basic Medicine, School of Medicine, Shihezi University, Shihezi, Xinjiang, Uygur Autonomous Region, China) for his critical suggestion of the article.
Authors' Contributions
S.L.: Data curation, formal analysis, investigation, and writing original draft. L.Z.: Investigation, resources, software, and writing review and editing. Z.L.: Investigation, software, and formal analysis. H.S.: Investigation, methodology, and formal analysis. Z.Q., S.Z., Y.L., and Y.G.: Investigation and resources. J.W.: Conceptualization, funding acquisition, project administration, and supervision.
Author Disclosure Statement
No conflicting financial interests exist.
Funding Information
This research was supported, in part, by grants from the National Natural Science Foundation of China (Grant Nos. 31960705 and 32160841).
