Abstract
In recent years, hepatitis B virus (HBV) has been detected in some species of animals. In this study, we found HBV-like nucleotide sequences in murine rodents and Asian house shrews (Suncus murinus) collected in China. A total of 801 animals were trapped. We found that 0.48% (3/624) of the murine rodents and 1.69% (3/177) of Asian house shrews were positive for HBV-like DNA. Detection of HBV-like DNA in brown rats (Rattus norvegicus), rice-field rat (Rattus losea), and Asian house shrews indicated that these species of animals might be hosts for HBV. However, none of the HBV-like DNA-positive animals was additionally positive for anti-HBV antibodies or hepatitis B surface antigen. A 585 bp nucleic acid sequence, mapping to a hepadnavirus, was extracted from rice-field rat, and bores strong resemblance to human HBV genotype B sequences. Further research is required to investigate the hepadnaviruses within the murine rodent and Asian house shrew populations to uncover the origin and zoonotic potential of HBV.
Introduction
Hepatitis B
HBV has been found in some species of animals; however, the host range of HBV remains elusive (Locarnini et al. 2013). With the discovery of the tent-making bat HBV, HBV is now viewed as a potentially zoonotic pathogen (Drexler et al. 2013).
To date, HBV or HBV-like viruses have not been found in murine rodents or shrews (Van Nguyen et al. 2018). In this study, we investigated HBV in these species of animals.
Methods
Samples
Between 2014 and 2017, small animals were trapped in China (Supplementary Table S1 and Supplementary Fig. S1). Serum and liver tissue samples were collected and stored at −80°C until testing. The species of animals were identified by sequencing of cytochrome B (cytB) gene (Arai et al. 2008).
Extraction of nucleic acid and detection of HBV
The MiniBEST Viral RNA/DNA Extraction Kit (TaKaRa, Japan) was used to extract total RNA and DNA from liver tissue samples. A broadly reactive and highly sensitive nested PCR assay was used to detect HBV DNA (Drexler et al. 2013).
The positive samples were sent to the Beijing Genomics Institute for sequencing.
Amplification of HBV genome
Based on six human HBV sequences (GenBank accession numbers: KM392074, FJ562311, AY217369, AF121250, KC774399, DQ993706), we designed primers to amplify the viral genome (Supplementary Table S2). Lasergene SeqMan software (DNASTAR, Madison) was used to assemble the sequences.
Phylogenetic analysis
Sequences obtained in this study were aligned using the ClustalW multiple sequence alignment program in MEGA (version 7.0; Oxford Molecular Ltd., UK). Phylogenetic tree was constructed using the neighbor-joining method, with 1000 bootstrap replicates.
The sequences identity was calculated using Sequence Identity Matrix program in BioEdit (Version 7.2.5).
Detection of anti-HBV antibodies and HBV surface antigen
Serum samples and supernatants of liver tissue samples from HBV-like DNA-positive animals were assayed for the presence of HBV e antibody, HBV surface antibody, HBV core antibody, and HBV surface antigen (HBsAg) by using enzyme-linked immunosorbent assay (ELISA) kits (CUSABIO, China).
Ethics guidelines
The protocol for this study was approved by the ethics committee of the Institutional Animal Care and Use Committee of Southern Medical University.
Results and Discussion
A total of 801 animals were trapped. HBV-like DNA was detected in 0.48% of murine rodents (3/624) and 1.69% of the Asian house shrews (Suncus murinus) (3/177) (Supplementary Table S1). The HBV-like DNA-positive Asian house shrews were trapped in the same site in Guangzhou, but not at the same time. Discovery of HBV-like DNA in brown rats (Rattus norvegicus), rice-field rat (Rattus losea), and Asian house shrews indicated that these species of animals might be hosts for HBV, and the host range of HBV could be wider than previously thought. The low detection rates indicated either a spillover infection or a low prevalence of HBV in these populations. More sensitive detection techniques are required in the future investigation.
The sequences, about 100 bp in length from HBV-like DNA-positive animals, were obtained from the nested PCR assay, and showed high degree of similarity with human HBV sequence (Supplementary Table S3). We only obtained a 585 bp sequence (XM24, GenBank accession number: MK158228), which is closely matched to human HBV sequence at both nucleotide and amino acid levels (Supplementary Table S4), from a rice-field rat with one pair of the designed primers (HBV-F25, HBV-R414, and HBV-R421; Supplementary Table S2) and the primers reported in a previous study (Van Nguyen et al. 2018). Amplification was failed in the other five HBV-like DNA-positive samples. The remaining designed primers failed in amplifying full-length genome, this may be attributed to the diversity of nucleic acid regions that we had not amplified.
HBV is diverse between different species of animals and it was traditionally considered to be a host-specific virus (Drexler et al. 2013). However, the discovery of human HBV-like DNA in murine rodents and Asian house shrews highlights the zoonotic potential of HBV. To ensure the credibility, phylogenetic analysis was only performed based on the 585 bp sequence. The sequence clustered with human HBV genotype B (Fig. 1). HBV genotype B is common in Asia (Rasche et al. 2016). If there are novel hepadnaviruses from murine rodents and Asian house shrews, and the HBV-like viruses shared similarities with the human HBV prevalent in the same geographical region, one may speculate that they might share a common viral ancestor. However, without full-length HBV genome, we cannot conclude whether the hepadnaviruses in these animals were of human origin due to pathogen spillover or derived from rats or shrews.
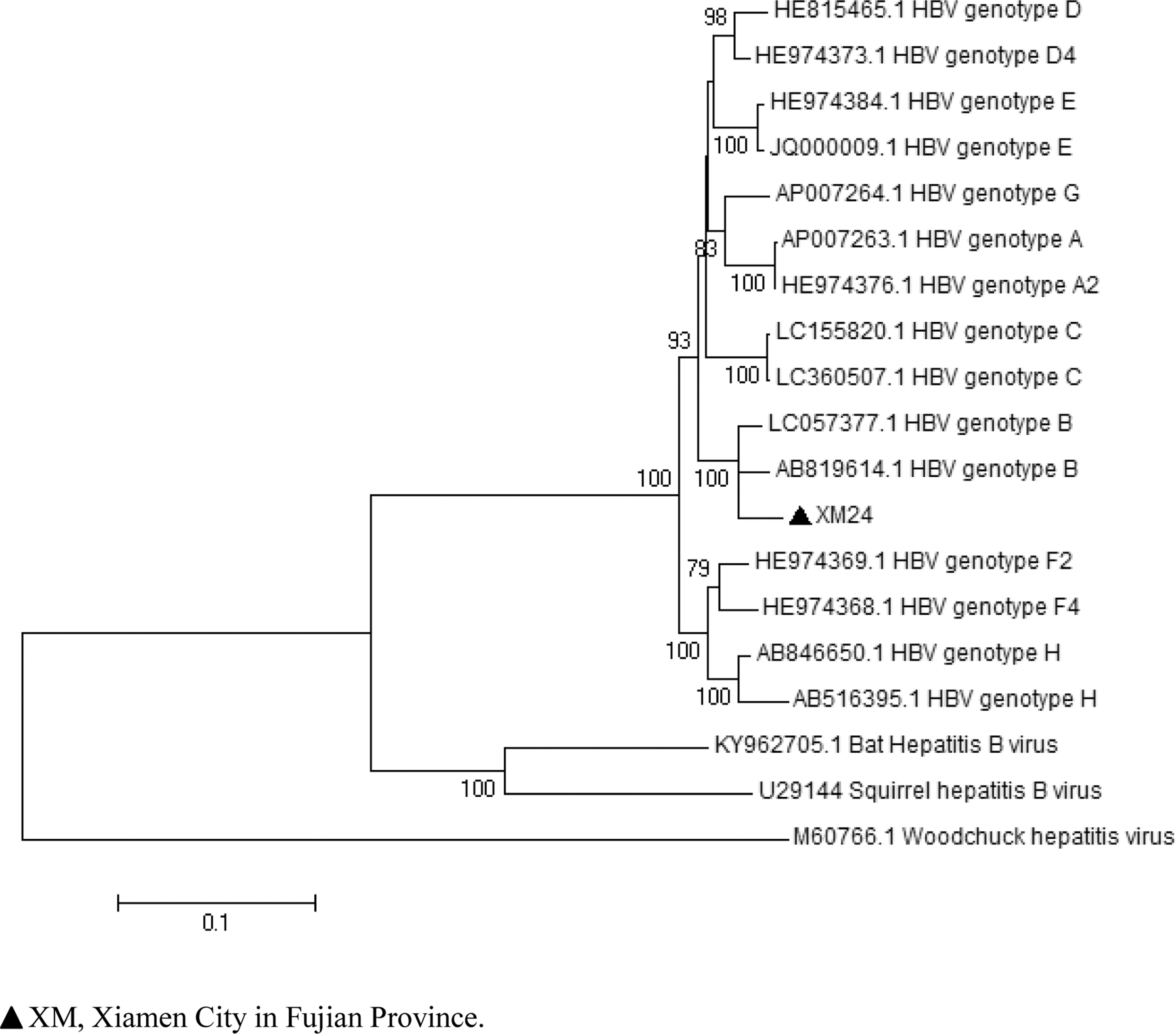
Phylogenetic tree constructed using the neighbor-joining method, based on partial nucleotide sequence (585 bp) alignment of 19 HBV sequences. Eighteen representative HBV sequences derived from humans, bat, woodchuck, and squirrel. HBV, hepatitis B virus.
None of the HBV-like DNA-positive animals was positive for anti-HBV antibodies or HBsAg, it might be explained by the seronegative occult HBV infection (Makvandi 2016).
Although this study provides new insights into the host range and origin of HBV, the authors acknowledge its limitations. For instance, attempts at the full-length hepadnaviral genome amplification failed. Furthermore, without any suitable HBV cell culture models, we did not perform cell culture experiment. We endeavor to conduct further experiments, with the aim of obtaining the full-length sequence of the virus, in the future.
Conclusion
HBV-like DNA was detected in brown rats, rice-field rat, and Asian house shrews. A 585 bp sequence, sourced from rice-field rat, showed high similarity to human HBV sequences. More research is needed to investigate the nature of hepadnaviruses in these animals, be they of rodent or human origin. The zoonotic potential of HBV also merits further investigation.
Footnotes
Acknowledgments
We are grateful to Xue-Shan Zhong, Shao-Wei Chen, Xue-Yan Zheng, Shu-Juan Ma, Li-Na Jiang, Jin Ge, Ming Qiu, Shu-Ting Huo, Xue-Mei Ke, Wen Zhou, Xing Li, Yun Mo, Yi-Quan Xiong, Ming-Ji Cheng, Yan-Xia Chen, Min-Yi Zhang, and Fang-fei You for their participation in sample collection. This study was supported by the National Natural Science Foundation of China (grant no. 81373051).
Author Disclosure Statement
No conflicting financial interests exist.
Supplementary Material
Supplementary Figure S1
Supplementary Table S1
Supplementary Table S2
Supplementary Table S3
Supplementary Table S4
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
