Abstract
Tick-borne spotted fever group (SFG) rickettsioses are emerging human diseases caused by obligate intracellular Gram-negative bacteria of the genus Rickettsia. Despite being important causes of systemic febrile illnesses in travelers returning from sub-Saharan Africa, little is known about the reservoir hosts of these pathogens. We conducted surveys for rickettsiae in domestic animals and ticks in a rural setting in western Kenya. Of the 100 serum specimens tested from each species of domestic ruminant 43% of goats, 23% of sheep, and 1% of cattle had immunoglobulin G (IgG) antibodies to the SFG rickettsiae. None of these sera were positive for IgG against typhus group rickettsiae. We detected Rickettsia africae–genotype DNA in 92.6% of adult Amblyomma variegatum ticks collected from domestic ruminants, but found no evidence of the pathogen in blood specimens from cattle, goats, or sheep. Sequencing of a subset of 21 rickettsia-positive ticks revealed R. africae variants in 95.2% (20/21) of ticks tested. Our findings show a high prevalence of R. africae variants in A. variegatum ticks in western Kenya, which may represent a low disease risk for humans. This may provide a possible explanation for the lack of African tick-bite fever cases among febrile patients in Kenya.
Introduction
T
ATBF is an acute febrile illness characterized by headaches, myalgia, regional lymphadenitis, and inoculation eschars typically present in multiple sites (Jensenius et al. 2003a). Although usually considered a benign condition, ATBF may be complicated by severe neuropathy (Jacquemard et al. 1998, Jensenius et al. 2006, Auce et al. 2011) or acute myocarditis (Bellini et al. 2005). Elderly patients seem to be prone to more severe clinical manifestations (Roch et al. 2008).
R. africae is transmitted by ticks of the genus Amblyomma (Kelly et al. 1994, Kelly et al. 1996). Two Amblyomma tick species, A. hebraeum in southern Africa and A. variegatum elsewhere in sub-Saharan Africa, are recognized as the principal reservoir hosts and vectors of R. africae (Jensenius et al. 2003a). R. africae has been detected in these ticks by PCR across much of their range (Mediannikov et al. 2010a), extending as far as the West Indies where A. variegatum was introduced in the 1800s (Kelly 2006). Infection is maintained through transovarial and transstadial transmission in both A. hebraeum (Kelly and Mason 1991) and A. variegatum (Socolovschi et al. 2009), and as a result all life stages of the ticks may transmit infection to humans. The rates of infection in these ticks in endemic areas are typically high and may reach 100% (Parola et al. 2001, Nakao et al. 2013).
Adults of A. hebraeum and A. variegatum prefer large hosts such as cattle and large wild ruminants, although they also infest sheep and goats. The immature stages (larvae and nymphs) have a wider host-range, feeding on the same hosts as adults do, as well as on smaller mammals, birds, and reptiles (Walker et al. 2003). These nidifugous (exophilic) ticks display active host-finding behavior, with nymphs and adults in particular attacking hosts in an aggressive manner (Kelly and Mason 1991, Sonenshine 1993). These ticks are known to attack humans, usually on the legs with attachment often behind the knee, in the groin, or in the axilla (Jensenius et al. 2003a).
Among domestic livestock, cattle serve as important hosts for both A. hebraeum and A. variegatum, and there is some evidence that they may play a role in the maintenance of human pathogenic rickettsiae (Kelly et al. 1991). A high proportion (80–100%) of cattle in endemic areas have serological evidence of exposure to SFG rickettsiae (Kelly et al. 1991, Parola et al. 1999, Kelly et al. 2010), and rickettsemia was detectable for at least 32 days following experimental infection of cattle (Kelly et al. 1991). High seroprevalence (87%) has also been reported in goats (Parola et al. 1999), although in another study seroprevalence rates in the three common domestic ruminants (cattle, goats, and sheep) reflected the host preference of the vector A. variegatum, with the highest prevalence in cattle and much lower prevalence in goats and sheep (Kelly et al. 2010).
In contrast to the number of reported cases of ATBF in travelers, reports of the disease in indigenous populations in affected areas of sub-Saharan Africa remain scarce. This is despite high seroprevalence to SFG rickettsiae in these populations: (16–55%) in eastern and southern Africa and (5–51%) in western and central Africa (Mediannikov et al. 2010b). In the coastal region of Cameroon, R. africae was identified in 6% of patients with acute febrile illness not caused by malaria or typhoid fever (Ndip et al. 2004). A study in rural Senegal found rickettsial DNA in a similar percentage (7%) of 134 febrile, nonmalarial patients (Mediannikov et al. 2010b); R. felis and R. conorii, and not R. africae, were identified.
In East Africa, R. africae was previously detected in 16% of adult A. variegatum ticks in southwestern Kenya (Macaluso et al. 2003). A fatal case of spotted fever rickettsiosis in an expatriate in Kenya was attributed to R. conorii infection, the causative agent of Mediterranean spotted fever (MSF), on the basis of the severity of the disease (Rutherford et al. 2004), and a laboratory-confirmed case of MSF occurred in a Japanese traveler to Kenya (Yoshikawa et al. 2005). More recently, R. felis, the agent of flea-borne spotted fever, was detected in febrile patients in northeastern (Richards et al. 2010) and western (Maina et al. 2012) Kenya. We conducted serosurveillance for immunoglobulin G (IgG) antibodies to SFG and typhus group (TG) rickettsiae in domestic ruminants and surveyed for R. africae in Amblyomma ticks of domestic ruminants and ixodid ticks found on domestic dogs in a rural setting in western Kenya.
Materials and Methods
Study site
The study was conducted in the Asembo division of Rarieda (formerly Bondo) subcounty of Siaya County in western Kenya, in 2008–2010. This rural site on the eastern shore of Lake Victoria falls within a health and demographic surveillance system (HDSS) run by the Kenya Medical Research Institute (KEMRI) and US Centers for Disease Control and Prevention (CDC). The HDSS collects household demographic and socioeconomic data three times per year and includes information on the number of livestock owned (Adazu et al. 2005). The population undertakes agricultural production through a traditional mixed crop–livestock system and fishing. In this area, 44% of households own cattle (mean number owned was 1.84) and 43% own at least one sheep or goat (mean number owned was 2.12 [KEMRI/CDC HDSS data for 2008, unpublished]).
Tick collection and domestic animals sampling
The Kenyan International Emerging Infections Program of KEMRI/CDC has conducted human population-based infectious disease surveillance (PBIDS) in Asembo since late 2005, with between 23,500 and 25,000 people under surveillance in 33 villages within the HDSS (Feikin et al. 2011). As part of a study on zoonotic diseases in the site, 300 livestock-owning compounds (LOC) were randomly selected from all known livestock-owning compounds in the 33 PBIDS villages, with the sampling frame of LOCs compiled from livestock census data collected by the HDSS. Samples were collected within a 5-km radius of Lwak hospital, the designated referral health facility for Asembo site located at −0.171388 South and 34.391388 East. Domestic ruminants (cattle, goats, and sheep) and dogs were sampled from January through May of 2009. Where present, cattle, goats, sheep, and dogs were randomly selected (up to a maximum of three per LOC) and sampled for ticks, by examining specific body sites known to harbor ticks (ear, dewlap, shoulder, belly, groin, udder, and perineum) and collecting one to two nonengorged adults of different genera per site. Ticks, pooled by individual host animal (one to six ticks per pool), were preserved in 70% ethanol and stored at −80°C.
Ambylomma ticks from domestic ruminants and all ticks obtained from dogs were subsequently retrieved for identification and testing. Identification was based on entomological keys by Walker et al. (2003), after which ticks were pooled based on species and host animal. After identification, ticks were washed in molecular-grade water and mechanically disrupted using a bead mill (Qiagen TissueLyser LT). Genomic DNA was extracted using QIAamp blood and tissue kits (Qiagen, Valencia, CA) according to the manufacturer's instruction, using a final elution volume of 100 μL.
Blood specimens from cattle, goats, and sheep were collected by venipuncture of the jugular vein into plain vacutainers for serum preparation and EDTA vacutainers for anti-coagulated blood (VWR International, West Chester, PA). Blood collection tubes were kept upright at ambient temperature for 15–30 min and then transported to the field laboratory on ice. The EDTA and the plain vacutainers were centrifuged to obtain the white cell fraction (buffy coat) and serum, respectively. DNA was extracted from buffy coat specimens using Qiagen blood and tissue kit (Qiagen), as per the manufacturer's instructions. Testing of samples was conducted as summarized in Figure 1.
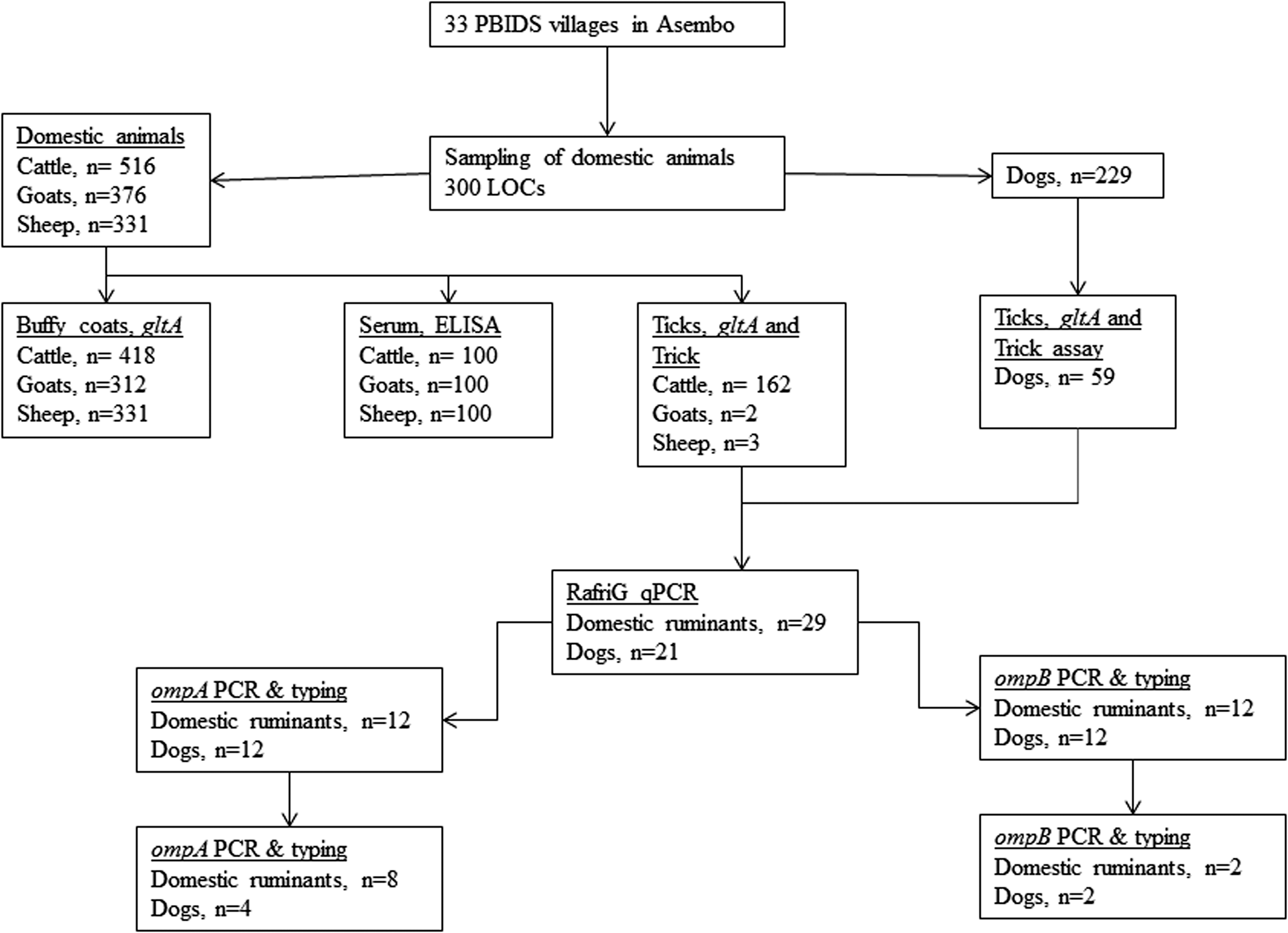
Framework for surveillance for rickettsiae in animals and ticks in Asembo collected in 2009 within a 5-km radius of Lwak hospital, located at −0.171388 South and 34.391388 East. PBIDS, population-based infectious disease surveillance; ELISA, enzyme-linked immunosorbent assay.
Serosurveillance for IgG antibodies
To assess previous exposure to rickettsiae, we examined a randomly selected subset of cattle, goat, and sheep sera (100 sera from each species) for SFG and TG rickettsiae-specific IgG. Serum specimens were screened at a dilution of 1:100 for IgG against SFG and TG rickettsiae, using an enzyme-linked immunosorbent assay (ELISA) as described by Graf et al. (2008). End point titers were determined for all specimens that were positive on screening.
Quantitative Real-time PCR assays
A quantitative real-time PCR (qPCR) assay that amplifies and detects a 74-bp segment of the citrate synthase (gltA) gene (Stenos et al. 2005) was used to screen nucleic acid preparations of blood samples from domestic ruminants and ticks for the presence of rickettsial DNA. Rickettsial DNA from ticks were further characterized using a tick-borne rickettsia-specific qPCR assay (Trick assay; Jiang et al. 2005a) that amplifies and detects a 128-bp fragment of the outer membrane protein B gene (ompB), and an R. africae (RafriG assay) genotype-specific qPCR assay using primers Raf1797F (TTGGAGCTAATAATAAAACTCTTGGAC), Raf1915R (GAATTGTACTGCACCGTTATTTCC), and Raf1879P (6-FAM-CGCGATGTTAATAGCAACATCACCGCCACTATCGCG-BHQ-1) that amplify and detect a 119-bp segment of the ompB gene.
The RafriG assay was performed on a Smart Cycler (Cepheid, Sunnyvale, CA) with the following cycling conditions: First hold at 50°C for 2 min, second hold at 95°C for 2 min, followed by 45 cycles of denaturation at 95°C for 15 s and annealing and extension at 62°C for 30 s. Evaluation of the RafriG against a panel of closely related rickettsial (n=17) and nonrelated bacterial (n=12) DNA preparations (Jiang et al. 2012) showed the assay to be specific for the R. africae genotype (data not shown). Plasmid positive controls constructed with target sequences from Trick and RafriG assays were used at 103 copies per reaction. Molecular-grade water was included at every sixth sample as an extraction control, and two negative controls (molecular-grade water) were included in every qPCR run. All negative controls were consistently negative.
Molecular identification of rickettsiae
To confirm the identity of the rickettsiae present in the ticks, standard and nested PCR together with sequencing of the specific rickettsial outer membrane protein genes ompA and ompB were done. Two segments of the ompA gene were amplified. The first segment used primers RompA1F (Jiang et al. 2005b) and Rr190.5044R (Fournier et al. 1998b), and the second segment targeted the 5′ end of the ompA and used primers Rr190.70F and Rr190.701R, which amplify a 632-bp fragment (Fournier et al. 1998b). Molecular confirmation was also performed using the ompB gene with primers 120.607F (Roux and Raoult 2000) and ompB1902R (Jiang et al. 2005b). To further elucidate the extent of variability of the R. africae in the ticks, PCR and sequencing of a longer fragment of ompB was conducted in a subset of tick DNA preparations (four of 21) using previously reported primers (Table 1). The PCR products were detected using GelRed (Biotium, Inc, Hayward, CA) on 1% agarose gels.
Primers used for sequencing only.
Nested PCR products from the ompA and ompB assays were purified using QIAquick gel purification kit (Qiagen). A Big Dye Terminator v 3.1 Ready Reaction Cycle Sequencing Kit (Applied Biosystems) was used in subsequent sequencing reactions according to the manufacturer's instructions. Sequencing products were purified using Performa DTR Gel Filtration Cartridges (EdgeBiosystems, Gaithersburg, MD). Sequencing was done in ABI 3500 Genetic Analyzer (Applied Biosystems). The primers used for sequencing were the same as those for PCR. At least two reactions were done for each strand of DNA. Long contigs were generated using primers shown in Table 1 as well as RhoA4336F (Jiang et al. 2005b) for ompA to enable assemblage of a complete consensus. Sequences obtained in this study were aligned against Rickettsia sequences selected from different geographical regions, sourced from GenBank. Phylogenetic analyses were conducted using MEGA version 5 software, and BLAST searches were managed on the National Center for Biotechnology Information website (
GenBank accession numbers
The sequences of the R. africae variants detected in the A. variegatum ticks have been deposited in GenBank with accession numbers KF660530, KF660531, KF660532, KF660533, KF660534, and KF660535.
Ethical review
The collection of specimens from animals was approved by the KEMRI Ethical Review Committee and CDC Animal Care and Use Committees (KEMRI SSC 1191 and 1562BRETBDX, respectively).
Results
Overall, 516 cattle, 376 goats, 331 sheep, and 229 dogs were examined in the 300 LOCs for the presence of ticks. Of these, 418 cattle, 312 goats, and 331 sheep had a serum and buffy coat specimen collected to look for evidence of rickettsial infection. Of the sera collected, a subset of 100 specimens each from cattle, sheep, and goats were randomly selected and tested for the presence of SFG- and TG-specific IgG; 1% of cattle, 43% of goat and 23% of sheep sera had IgG antibodies against SFG rickettsiae with end point titers ranging from 400 to 6400 for goats, 400 to 1600 for sheep, and <100 for cattle. None of them were positive for antibodies against TG rickettsiae. In addition, none of the 418 cattle, 312 goats, and 331 sheep buffy coat specimens tested positive with the pan rickettsia qPCR screening assay.
For ticks collected from domestic ruminants, only A. variegatum ticks were assessed for the presence of rickettsiae. A total of 162 cattle (31.4%), two goats (0.5%), and three sheep (0.9%) carried A. variegatum ticks. In total, 263 adult A. variegatum were collected and pooled (one to five A. variegatum) by individual host (167 pools). DNA preparations were obtained from 162 tick pools of A. variegatum ticks from cattle, of which 79 (48.8%) consisted of a single individual tick. Of the 162 pools, 160 (98.8%) were positive for rickettsial DNA using the gltA pan-rickettsia qPCR assay whereas 150 (92.6%) were positive using the tick-borne rickettsiae-specific Trick assay. All of the Trick-positive pools selected for further testing were confirmed positive for the R. africae genotype using the RafriG qPCR assay. All A. variegatum ticks collected from goats and sheep (single tick per animal) were positive with the three assays. A total of 102 ticks were collected from 36 dogs, resulting in 59 pools of over five species of ticks summarized in Table 2.
Confirmation of rickettsiae in ticks by sequencing of ompA and ompB genes
Rickettsial DNA was amplified in 21/24 tick DNA preparations tested by PCR targeting ompA and ompB. The tick DNA preparations were from sheep (n=1), cattle (n=11), and dogs (n=12). A 1364-bp sequence was obtained after assembling the sequences of the ompA (Romp1F, 190-5044R, RhoA4336F, and RompA4433R). BLAST searches showed that this sequence, obtained from a tick collected from a sheep (tick 25) was 100% identical with that of R. africae ESF-5 (accession no. CP001612) and R. africae ESF 2500 (accession no. U83436), whereas the sequences of the rest of the rickettsiae from tick DNA samples were 99% identical with R. africae ESF-5. The ompA sequences in this fragment contained a deletion in 6/21 specimens and a 4-bp deletion in 1/21 (tick 214) DNA extracts. Sequencing of the 5′ end of the ompA gene that used primers Rr190.70F and Rr190.701R resulted in a 540- to 747-bp sequence summarized in Table 3. Rickettsia africae, Rickettsia sp. AvR4/Kenya, and some uncultured Rickettsia spp. were detected in the ticks. Unique among these sequences was a 173-bp insertion in four of 12 samples (tick 16, tick 56, tick 74, tick 226) assessed (Fig. 2). The insertion sequence was identical in all of the four ompA sequences except that the sequence of one tick extract (tick 74) had a 5-bp deletion.
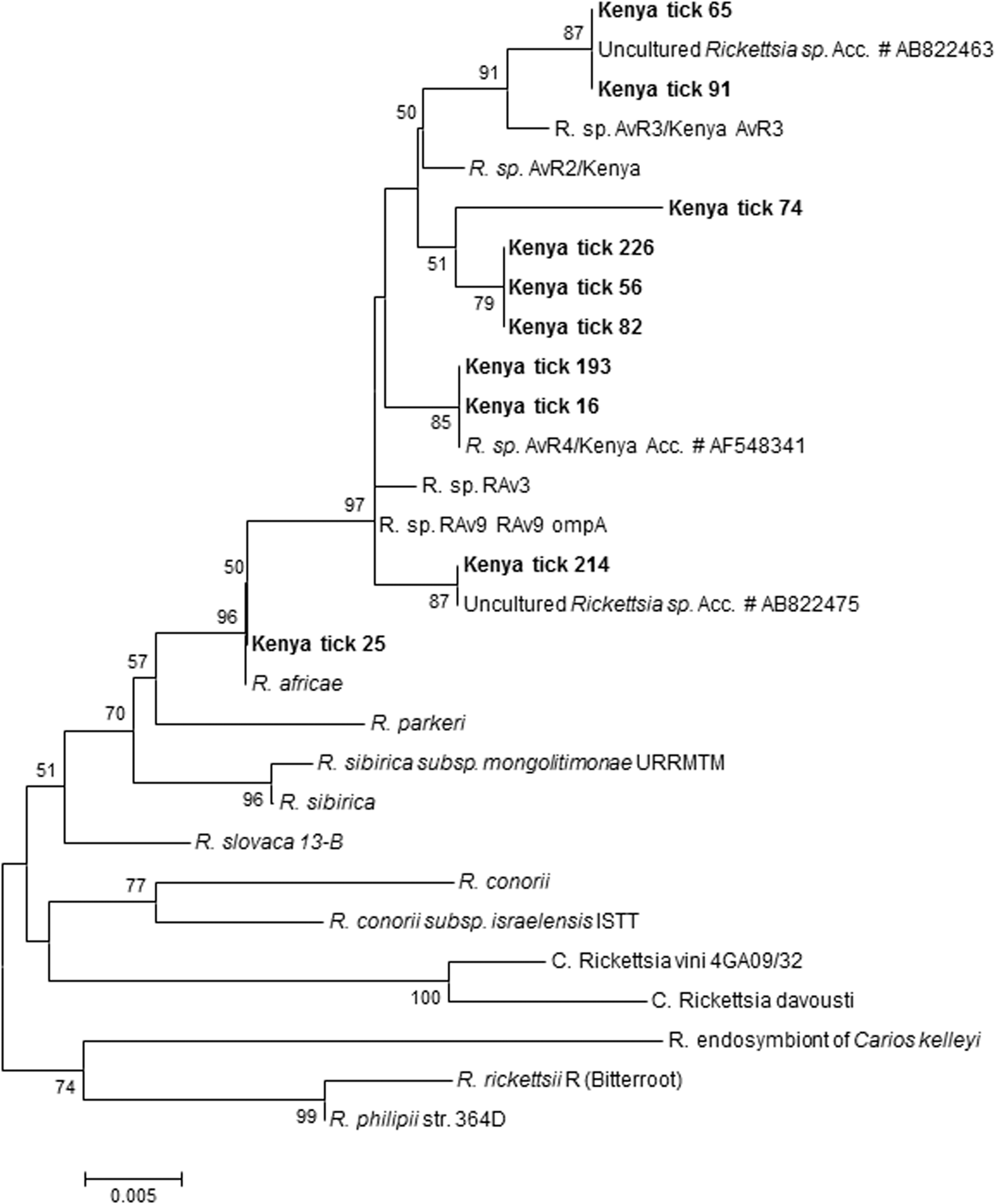
ompA partial gene sequences of rickettsiae detected in ticks obtained from domestic animals in western Kenya. The tree was inferred using the neighbor-joining method Bootstrap (1000 replicates) involving rickettsiae detected in this study and strains from GenBank.
A 1265-bp amplicon was produced using ompB nested PCR in 21 of 24 of the gltA-, Trick-, and RafriG-positive samples. A 1200-bp sequence was obtained after assembling the sequences of the ompB (Romp607F, RompB1007F, RompB1452R, and RompB1902R). BLAST searches showed that this sequence had 100% and 99% homology with R. africae ESF-5 (accession no. CP001612) in one of 21 (tick 25) and 20 of 21 DNA preparations, respectively (Fig. 3).
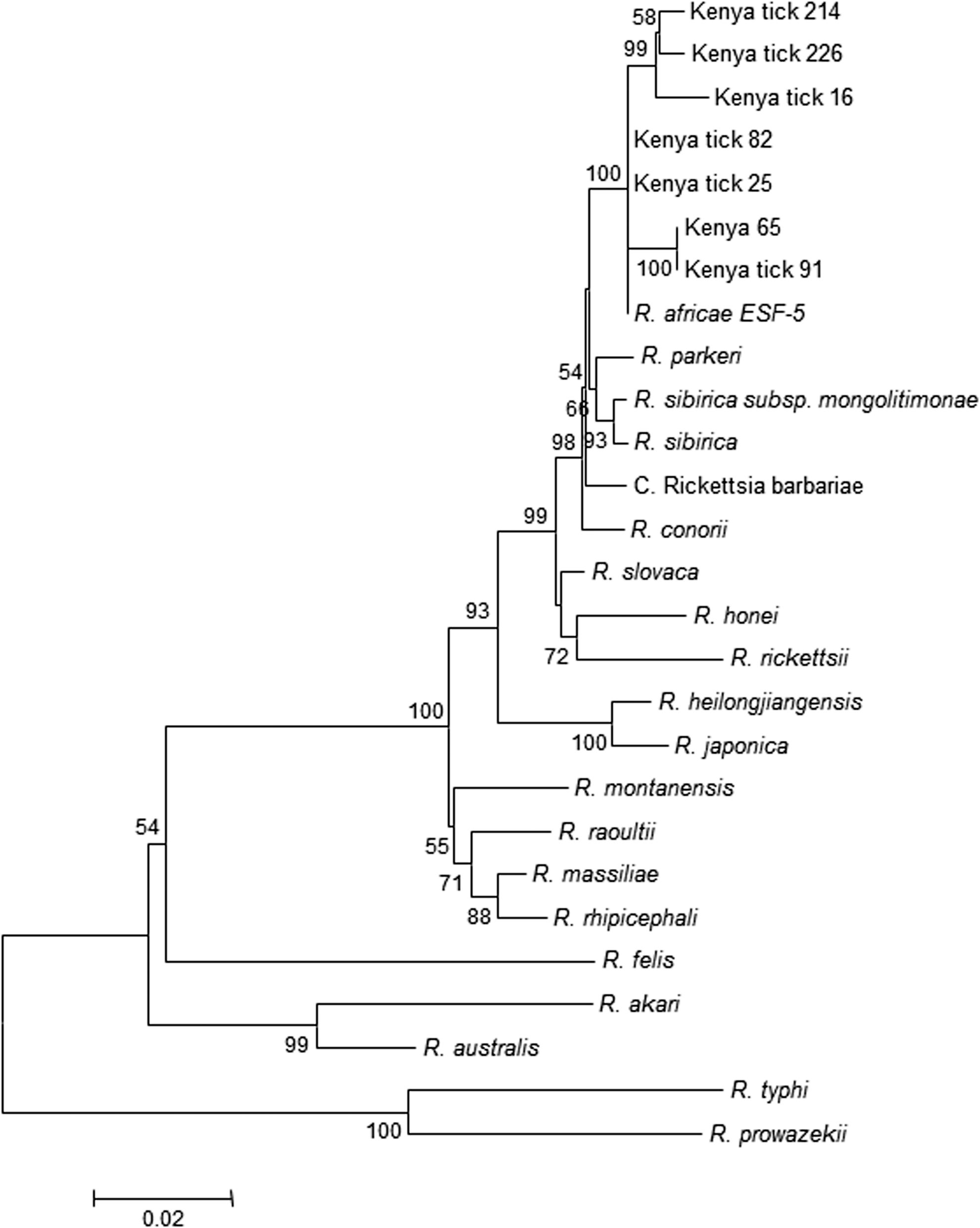
ompB partial gene sequences of rickettsiae detected in ticks obtained from domestic animals in western Kenya. The tree was inferred using the neighbor-joining method Bootstrap (1000 replicates) involving rickettsiae detected in this study and 20 historical strains.
Four tick DNA preparations were selected for sequencing of a longer fragment of the ompB gene. A 4400- to 4867-bp sequence was generated in which the sequence from one of the four ticks (tick 25) was confirmed to be 100% identical to R. africae ESF-5 (accession no. CP001612). The other sequences (from tick DNA preparations 74, 214, and 226) had 99% homology with R. africae ESF-5. These sequences had a 4-base deletion at exactly the same site (Table 4) and 32–36 nucleotide substitutions. One sequence (tick 74) had an extra nucleotide deletion in a different site of its ompB gene.
Note the 4-base deletion mutation in some R. africae–genotypes that is absent in Kenya tick 25. The alignment also shows substitutions in several positions.
Discussion
This report corroborates that of Macaluso et al. (2003, who found R. africae variants in ticks from Kenya. Diversity has also been reported among variants in Amblyomma laculosum ticks obtained from humans and a bird in New Caledonia (Eldin et al. 2011). Although the present study was conducted to determine the prevalence of R. africae in A. variegatum ticks of domestic animals, all tick species obtained from dogs were tested owing to their small numbers. R. africae variants have previously been detected in A. variegatum ticks collected in various countries in Africa, namely Niger, Mali, Sudan, Burundi, and Mauritania (Parola et al. 2001). On the basis of the nucleotide sequences of the ompA gene, two of 10 sequences were identical to Rickettsia sp. AvR/4 Kenya (GenBank accession no. AF548341) that was detected in A. variegatum from Maasai Mara, Kenya (Macaluso et al. 2003). Uncultured Rickettsia spp. (GenBank accession nos. AB822463 and AB822475) (Nakao et al. 2013) were detected in three of 10 samples analyzed. However, ompA sequences of four of 10 tick DNA preparations were unique in that they had a 173-bp nucleotide insertion and a 5-nucleotide deletion in one of the sequences. In addition to nucleotide substitution, we report for the first time existence of a deletion mutation in the ompA and ompB genes of some R. africae variants. The percent divergence of the ompA and ompB sequences in the R. africae variants to the next closest accepted Rickettsia species, R. africae, was less than that required to define a new species (Fournier et al. 2003). Thus, we refer to these molecular isolates as R. africae variants as previously reported (Parola et al. 2001).
The variants described in this study have not been reported to be associated with ATBF, and we hypothesize that, A. variegatum infected with these variants represent a low risk for R. africae transmission to humans in Kenya. The above discussion suggests that R. africae variants may have reduced virulence due to host adaptation or possibly competitive exclusion that might account for the lack of ATBF in the indigenous populations. This kind of exclusion may be mediated by the host immune system (concomitant immunity) or by direct interference as shown for R. peacockii, where it has been shown to interfere with multiplication of the pathogenic R. rickettsii in Dermacentor andersoni ticks (Baldridge et al. 2004). Additionally, this is plausible given that no mixed results were found when assessing pooled samples in the present study with such a highly prevalent organism.
Supporting this notion, a study conducted in rural western Kenya, the region where these ticks were collected, found no evidence of R. africae infections in febrile patients and afebrile controls (Maina et al. 2012). R. felis was detected to be a major cause of rickettsial infections. It is unknown whether R. felis infections alone are responsible for the high (56%) seroprevalence to SFG rickettsiae. Furthermore, ATBF was not detected in indigenous patients presenting with fever in northeastern Kenya, a region that is predominantly inhabited by pastoralist communities (Richards et al. 2010). Similarly, other investigators noted lack of clinical cases due to ATBF in native Africans (Cohen et al. 1996, Frean et al. 2007, Mediannikov et al. 2010b).
Although high seroprevalence of SFG antibodies has been reported in native Africans, the actual cause of this high prevalence has not yet been fully elucidated. We speculate that because R. africae shares common antigenic determinants with other SFG rickettsia, the high reactivity reported in previous studies could be due to not only R. africae, but also to other rickettsia species. Members of the Rickettsia genus have surface cell antigens (sca) genes that are annotated as sca0 (ompA), sca1, sca2, sca3, sca4, sca5 (ompB), and so on. These genes are often characterized by frequent degradation resulting in genome reduction (Blanc et al. 2005). Deletions and mutations have been observed in several rickettsial genes, including ompA in R. peacockii and are said to be responsible for its lack of pathogenicity (Feisheim et al. 2009). In the present study, the rickettsia ompB gene had both deletion and substitution mutations in three out of four ompB sequences analyzed, and only one of the four sequences conformed with the type strain (R. africae ESF-5). This suggests a high infection rate of the ticks with the variants, which might have reduced pathogenic potential for causing ATBF. Comparative analysis of clinical isolates with those from ticks warrants further investigation.
In the present survey, antibodies against SFG rickettsiae were detected in domestic ruminants—goats (43%), sheep (23%), and cattle (1%). The findings of this study confirm previous reports of the occurrence of SFG antibodies to rickettsiae in domestic ruminants (Philip et al. 1966, Kelly et al. 1991, Parola et al. 1999, Kelly et al. 2010). In contrast, however, the prevalence of SFG antibodies in cattle in our study were lower (1%) compared to what was reported in Zimbabwe (26–100%) (Kelly et al. 1991) and Guadeloupe (≥38.4%) (Parola et al. 1999). It is intriguing, however, that A. variegatum ticks containing the R. africae genotype were predominantly found on cattle, but only a small proportion of the cattle sera tested had antibodies to SFG rickettsiae.
Although antibodies reactive to Rickettsia typhi had been demonstrated using a complement fixation test in domestic ruminants in Egypt (Schmatz et al. 1978), our study did not find antibodies to TG rickettsia in these animals. This corroborates previous studies conducted in humans in eastern Africa where prevalence of TG rickettsia was found to be low (Prabhu et al. 2011, Maina et al. 2012). Likewise, there was no evidence of rickettsiemia in the ruminants from which these ticks were collected. This is likely to be associated with the high prevalence of an R. africae variant that may be an endosymbiont of A. variegatum. Insufficient numbers of rickettsiae in the blood of these animals coupled with the prevalent use of tetracyclines/antimicrobial drugs in these animals (Mitema et al. 2001) may explain the lack of detection by PCR method and the low prevalence of seroreactivity to SFG rickettsial antigens by ELISA. As preferred hosts of the tick vectors, they likely still play a pivotal role in the ecology of the pathogen through maintenance of tick populations, particularly in areas devoid of other large mammal species. Although this study recorded higher prevalence of IgG antibodies to SFG in goats (43%) and sheep (23%) than cattle (1%), only a small proportion of goats and sheep had A. variegatum ticks (2.9% and 1.5% in goats and sheep, respectively). These findings are consistent with those found in Ethiopia, where antibodies to SFG rickettsiae were significantly higher in goats and sheep than in cattle (Philip et al. 1966).
Whether these variants reported in this and other previous studies can cause human disease remains to be studied. A more comprehensive study is needed to elucidate whether translation of ompA and ompB to functional OmpA and OmpB do occur in the variants that have nucleotide deletions in the ompA and ompB genes. This study provides three important findings: First, antibody levels reported among livestock in this region appear unrelated to exposure to ticks infected with R. africae; second, a high tick infection rate was evident in the absence of R. africae infection; and finally, the variability observed among strains underscores the need to undertake comparative assessment of strains associated with clinical ATBF and those circulating within the larger tick population to decipher potential virulence markers.
Footnotes
Acknowledgments
We thank the IEIP-Z field team (Samuel Asembo, Michael Otieno, James Oigo, and Pauline Otieno) for the collection of blood and tick specimens. We also thank Dr. John Waitumbi of USAMRU-K for reviewing the manuscript, and the Director, KEMRI for approving publication of this manuscript. This work was supported by the Wellcome Trust, UK (grant no. 081828/B/06/Z).
Author Disclosure Statement
No competing financial interests exist.
The views expressed in this article are those of the authors and do not necessarily reflect the official policy or position of the Department of the Navy, Department of Defense (DoD), nor the US Government. This work, partially funded by DoD AFHSC-GEIS work unit no. 188M.0931.001.A0074 and as an employee of the US Government (A.L.R.) under Title 17 U.S.C. 107, this article is not available for copyright protection.

