Abstract
Choroideremia (CHM) is a rare X-linked chorioretinal dystrophy causing progressive vision loss due to mutations in the
Introduction
Choroideremia (CHM) is a chorioretinal dystrophy marked by progressive degeneration of the choroid, retinal pigment epithelium (RPE), and photoreceptors (PR). Symptoms begin in the first decade of life, with male patients experiencing nyctalopia, followed by peripheral vision loss and complete blindness in late adulthood. This X-linked recessive disease is caused by
Loss of REP1 can be compensated in most tissues by its isoform REP2 (encoded by CHM-like gene,
The RPE is a monolayer of polarized cells that supports the PR, performing several essential functions such as providing nutrients, growth factors, and ions to the PR, while eliminating waste products from retinal metabolism. 9 –11 Furthermore, RPE cells perform daily phagocytosis of POS and contribute to inflammatory signaling. 10
Multiple retinal dystrophies impair RPE function, resulting in vision loss 11 ; thus, several RPE models are currently routinely used. ARPE-19 (Adult Retinal Pigment Epithelial cell line-19) is one of the most used cell lines to study retinal diseases because of its simplicity and availability. Although this cell line has some advantages, such as being readily available commercially and having good viability and proliferation properties in culture, it is now clear that it has some limitations regarding its relevance and data reproducibility. 12 Because ARPE-19 immortalization occurred spontaneously, these cells exhibit replicative exhaustion and senescence, as well as chromosomal aberrations, as the number of passages increases. 12,13 In addition, ARPE-19 cells produce some RPE-specific mRNA transcripts without the corresponding protein expression, such as RPE65, BEST1, and CLDN19, leading to an incompletely differentiated phenotype. Regarding pigmentation, most studies report absent or heterogeneous melanin accumulation in ARPE-19, which, on top of an incomplete protein network supporting the activity of junctional complexes, do not represent accurately the physiological functionality of a complete monolayer. 12
In a study by Markert and colleagues, the authors compared the transcriptional profile between various RPE sources, including adult human primary RPE (ahRPE), human induced pluripotent stem cell (hiPSC)-derived RPE, and ARPE-19, to investigate differences in aging, maturation, and plasticity. 14 They observed that ahRPE and hiPSC-RPE shared a similar RPE gene expression profile, while ARPE-19 had a significantly lower RPE signature gene expression profile, consistent with a reduced transepithelial electrical resistance (TEER). 14
Given the limitations of ARPE-19 usage, it is of high interest to develop more relevant and robust models to study retinal diseases. 12 The use of hiPSC-derived RPE (hiPSC-RPE) models may help study and understand various retinal disorders as they can offer an unlimited supply of RPE for cell therapies and are already used in multiple clinical trials. 11
The development of cellular disease models from individual patients was made possible with hiPSC technology, enabling the study of genetic alterations in various cell types. This, coupled with advances in genome editing techniques such as the clustered regularly interspaced short palindromic repeat (CRISPR) system, allows the repair of causative alleles in patient cell lines or the introduction of disease alleles into healthy cell lines. Together, this permits the investigation of isogenic cell pairs varying in a single genetic change, enabling research into the molecular and cellular phenotypes underlying specific mutations. 15
In this study, we developed two isogenic cell lines using CRISPR/Cas9 to model CHM disease, from two different sources: a healthy donor cell line (RBi001-A) and a patient-derived hiPSC line (CRFi001-A). Moreover, we demonstrate that these cells can differentiate into RPE and show typical RPE morphological and molecular characteristics.
The development of these isogenic lines is a valuable contribution for CHM research since it provides a reliable platform for further investigating the pathological mechanisms underlying this degenerative disease.
Materials and Methods
hiPSC culture
hiPSC lines were cultured according to the StemFlex™ medium (#A3349401, Gibco™) user guide, on 6-well plates (#3506, Corning®) precoated with 20 µg cm−2 of Growth Factor Reduced Matrigel (GFR, #354230, Corning®). When 80% confluence was reached, the hiPSCs were subcultured for expansion or experimental use, using Versene Solution (#15040066, Gibco™) at a ratio between 1:2 and 1:4 in StemFlex™ medium and maintained at 37°C in a humidified atmosphere containing 5% CO2.
Gene editing of hiPSCs
In this study, we generated two cell lines by genome editing. First, we used a healthy hiPSC line, RBi001-A (#66540025, ECACC), to produce a
We designed single guide RNAs (sgRNAs) for gene editing using the CRISPOR (http://crispor.tefor.net/) online tool. On both strategies, the sgRNAs target CHM exon 6 (5′-TGCATTTCGAGAAGGACGAGTGG-3′; 5′-TGCATTTCGAGAAGGATGAGTGG-3′—Strategy 1 and 2, respectively). However, on Strategy 2, in addition to the sgRNA, whic targets the mutation, a single-stranded oligodeoxynucleotide (ssODN) 5’-AATCAAATCTAATGTTAGTCGATATGCAGAGTTTAAAAATATTACCAGGATTCTTGCATTTCGAGAAGGACGAGTGGAACAGGTTCCGTGTTCCAGAGCAGATGTCTTTAATAGCAAACAACTTACTATGGTAGAAAAGC-3′ was used to induce the homology-directed repair (HDR) mechanism and correct the CHM mutation on CRFi001-A cells.
The two hiPSC lines were generated using CRISPR/Cas9 components following the LipofectamineTM Stem Transfection Reagent (#STEM00001, Invitrogen™) protocol. Briefly, one day before transfection, cultures with <85% confluence were seeded as single cells, using approximately 75 000 cells/well, in 24-well plates (#3524, Corning®) precoated with 2 μg cm−2 of Biolaminin 521 LN (#LN521-05, BioLamina). The next day, wells with ∼30% confluence were transfected using LipofectamineTM Stem Transfection Reagent (#STEM00001, Invitrogen™), as per manufacturer’s instructions. For
hiPSC-RPE differentiation
hiPSCs were grown as described previously, and differentiation into RPE was induced using Foltz LP et al.’s 11 method, by sequential addition of growth factors for 14d in Retinal Differentiation Medium (RDM). On d14, immature RPE was collected and plated at ∼1x105 cells cm−2 on laminin-coated 6-well plates (passage 0, P0) and were allowed to mature in X-VIVO (TheraPEAK™ X-VIVO™−10, #BP04-743Q, Lonza) containing 1% Antibiotic-Antimycotic (AB/AM, #15240062, Gibco™), which was changed twice per week. On d30, when pigmentation was visible, the cells were further expanded to P1 and matured roughly 30d later, at which point, they were used for experiments (from P1 to P4).
Retinal organoid differentiation from hiPSCs
Retinal organoid (RO) differentiation was carried out according to Capowski EE et al.’s
16
protocol with slight alterations. Briefly, hiPSCs at 80% confluence were lifted using Versene and split at 3000 cells/well in a Nunclon™ Sphera™ 96-Well U-Shaped-Bottom Microplate (#174925, Thermo Scientific™) using StemFlex™ medium supplemented with 10 μM of Rho-associated protein kinases inhibitor (Y-27632, #10–2301, Focus Biomolecules). The cells were grown as embryoid bodies (EBs) for 7d, doing an adaptation to Neural Induction Medium (NIM) containing Dulbecco's Modified Eagle Medium/Nutrient Mixture F-12 (DMEM/F-12, #10565018, Gibco™), 1% N-2 supplement (#17502048, Gibco™), 1X Non-Essential Amino Acids Solution (NEAA, #11140050, Gibco™), 1X GlutaMAX™ Supplement (#35050038, Thermo Scientific™), and 2 mg/ml Heparin (#H3149, Sigma-Aldrich®). On d6, 1.5 nM of Bone Morphogenetic Protein-4 (#120-05, PeproTech®) was added to fresh NIM, and on d7 EBs were transferred to GFR Matrigel-coated 6-well plates to allow optic cup formation. The medium was replaced by half fresh NIM on d9, 12, and 15. On d16, the medium was replaced by RDM containing DMEM:F12 (3:1), 2% B-27™ minus vitamin A (#12587010, Gibco™), 1X NEAA, 1X GlutaMAX™, and 1X AB/AM and changed every 2 days. On d30, optic vesicles were manually dissected using a surgical scalpel (SM65A, Swann-Morton® Ltd) under the microscope EVOS™ XL Core (ThermoFisher Scientific). After dissection, organoids were cultured in suspension in 3D retinal differentiation medium (3D-RDM) containing DMEM:F12 (3:1), 2% B-27 minus vitamin A, 1X NEAA, 1X GlutaMAX™, 1X AB/AM, 5% Fetal Bovine Serum (FBS, #A5256801, Gibco™), 1:1000 Chemically Defined Lipid Concentrate (#11905031, Gibco™), and 100 μM Taurine (#T8691, Sigma-Aldrich®) and supplemented with 1 μM of all-
Western blotting
hiPSC or hiPSC-RPE was lysed on ice with Cell Lysis Buffer (#9803, Cell Signaling Technology®) containing Halt™ Protease and Phosphatase Inhibitor Cocktail (#78444, Thermo Scientific™) diluted to 1X. Pierce™ BCA Protein Assay Kit (#23225, Thermo Scientific™) was used to determine protein concentrations and 8% polyacrylamide gels were loaded with 30 µg of protein lysate, previously denaturated at 95°C for 5 min. Gels were then transferred onto 0.2 µm nitrocellulose membranes using Bio-rad Trans-Blot Turbo Transfer System. The following primary antibodies were used at the indicated dilutions: REP1 (#AB0123, SICGEN, 1:500), REP2 (#AB0132, SICGEN, 1:500), Oct3/4 (#sc-5279, Santa Cruz Biotechnology, 1:500), Nanog (#sc-134218, Santa Cruz Biotechnology, 1:1000), Sox-2 (#sc-365823, Santa Cruz Biotechnology, 1:1000), and peroxidase-conjugated β-Actin (#A3854, Sigma-Aldrich®, 1:25000). Membranes were blocked with 5% milk powder in Tris-Buffered Saline with Tween (TBS-T) before primary antibody incubation overnight, at 4°C with shaking. After three washes with TBS-T, HRP conjugated secondary antibodies [Donkey anti-Goat (#A16005, Invitrogen™, 1:5000) or Goat anti-Mouse (#554002, BD Pharmingen™, 1:5000)] was incubated for 1 h at room temperature (RT). Protein expression was detected by chemiluminescence using Amersham™ ECL™ Prime kit (#RPN2232, Cytiva) in a Bio-rad ChemiDoc Imaging System.
Immunocytochemistry
hiPSCs grown on laminin-coated coverslips were fixed for 15 min with 4% paraformaldehyde (PFA, #043368.9M, Thermo Scientific Chemicals), permeabilized with 0.2% Triton X-100 (TX-100, #T9284-100, Sigma-Aldrich®) in Phosphate-buffered saline (PBS) for 10 min, followed by 1 h blocking with 10% donkey serum (DS, #S30-100ML, Sigma-Aldrich®) in PBS at RT. Stem cells were then incubated for 2 h with primary antibodies OCT3/4 (#sc-5279, Santa Cruz Biotechnology, 1:100), SOX-2 (#sc-365823, Santa Cruz, 1:100), or Nanog (#sc-293121, Santa Cruz Biotechnology, 1:50) at 4°C overnight in blocking solution.
hiPSC-RPE was fixed either with methanol for 15 min on ice (for Claudin-19 and ZO-1 staining) or 4% PFA at RT (for phalloidin staining). Then, cells were permeabilized and blocked with perblock solution (0.05% Triton X-100 and 10% DS in PBS) for 1 h at RT. Cells were incubated with primary antibodies Claudin-19 (#365967, Santa Cruz Biotechnology, 1:100), ZO-1 (#40–2300, Invitrogen™, 1:100), or Alexa Fluor™ 568 Phalloidin F-actin probe (#A12380, Invitrogen™, 1:100) for 2 h at RT in perblock solution.
After washing cells with PBS, hiPSC or hiPSC-RPE was incubated for 1 h with secondary antibodies anti-Mouse or anti-Rabbit (#A-21202 and #A-21206, respectively, Invitrogen™, 1:500), followed by 5 min with DAPI (#D9542-5MG, Sigma-Aldrich®, 1:10 000). Coverslips were mounted with VECTASHIELD® Antifade Mounting Medium (#H-1000-10, Vector laboratories) and images were acquired with LSM980 (Zeiss) confocal microscope.
Brightfield images of hiPSC colonies, EBs, and ROs at different developmental stages (from d3 to d140 with PR development) were acquired using an EVOS™ XL Core digital microscope.
Phagocytosis assay
Porcine POS were isolated as described previously, 17 covalently labeled with Alexa Fluor 488 (A488), and stored at −80°C. Labeled POS were defrosted and washed with PBS by three centrifugation rounds of 10 min, at 5000 rpm, at 4°C. After each centrifugation, POS-A488 were resuspended in 1 mL sterile PBS, and kept with limited light exposure for a maximum of one week. Before RPE cell feeding, POS were sonicated three times for 5 min. hiPSC-RPE were incubated overnight with a 10% POS-A488 preparation diluted in X-VIVO, in a 5% CO2 incubator at 37°C. The next day, cells were fixed and stained with phalloidin, as described in the previous section.
Transepithelial electrical resistance
hiPSC-RPE cultures from RBi CHM+ and RBi CHM− lines were plated onto laminin-coated transwell inserts (#3460, Corning®) at a density of ∼1x105 cells cm−2. X-VIVO medium was changed twice per week before measurements for 35d. TEER was measured using the EVOM™ epithelial volt-ohmmeter (#EVM-MT-03-01, World precision instruments). Data were represented using a 4-parameter logistic regression with 95% confidence intervals (CI).
Transmission electron microscopy (TEM)
All reagents and materials were purchased from Electron Microscopy Sciences, unless otherwise stated. Cells on glass coverslips were fixed in 2% PFA, and 2% glutaraldehyde in 0.1M phosphate buffer (PB) at pH 7.4 overnight at 4°C. After washing with PB, specimens were postfixed with 1% osmium tetroxide and 1.5% potassium ferrocyanide in distilled water for 1 h on ice, and then incubated with 1% tannic acid for 30 min at RT. Specimens were subsequently dehydrated with a series of increasing ethanol concentrations (50%, 70%, 90%, and 2 × 100%) before infiltrating and embedding in Epon resin (EMbed 812). After polymerizing at 65°C overnight, resin blocks were sectioned at 60–70 nm using a Reichart Ultracut S ultramicrotome (Leica) and a diamond knife (Diatome), and sections collected on copper mesh grids. Sections on grids were poststained with uranyl acetate and Reynold’s lead citrate and imaged using a Hitachi H‐7650 TEM equipped with an AMT XR41 M digital camera.
Quantitative PCR analysis
mRNA from hiPSC, hiPSC-RPE, or EBs (pool of 5) was extracted using RNeasy Mini Kit (#74104, Quiagen), according to manufacturer’s instructions. Next, 1 μg of purified mRNA was reverse transcribed into cDNA using SuperScript™ II Reverse Transcriptase (#18064022, Invitrogen™). Reverse transcriptase quantitative polymerase chain reaction (RT-qPCR) was carried out in a QuantStudio™ 5 system (Applied Biosystems™) using PowerUp™ SYBR™ Green Master Mix (#A25742, Applied Biosystems™), following manufacturer’s instructions. The primer pairs (forward/reverse) used are described in Table 1 and relative gene expression results were analyzed using 2-ΔΔCt method,
18
normalized to the housekeeping gene
Human Primer Sequences Used in RT-qPCR
RT-qPCR, Reverse transcriptase quantitative polymerase chain reaction
Karyotyping
hiPSC colonies at 60–80% confluency were incubated with 0.1 µg/mL KaryoMAX™ Colcemid™ (#15212012, Gibco™) for 1:30–2 h before collection using TrypLE. The suspension was centrifuged, and the pellet was treated with prewarmed 75 mM KCl hypotonic solution for 20 min at 37°C. Then, cells were fixed using a methanol-acetic acid solution at a 3:1 ratio followed by three 15 min centrifugations at 1250 rpm until the solution became translucid with a white pellet. G-banding at 400–550 band resolution was used to assess hiPSC genome integrity, with at least 20 metaphase spreads examined (Genomed, Lisbon, Portugal).
Statistical analysis
Data were analyzed using GraphPad Prism 9 software and is presented as mean ± SEM from at least
Results
Generation and characterization of CHM and control isogenic hiPSCs
In this work, we used two strategies to develop isogenic hiPSC lines to model CHM disease. In Strategy 1, we used a healthy donor cell line RBi001-A (henceforth named RBi CHM+) to produce a CHM mutant line targeting
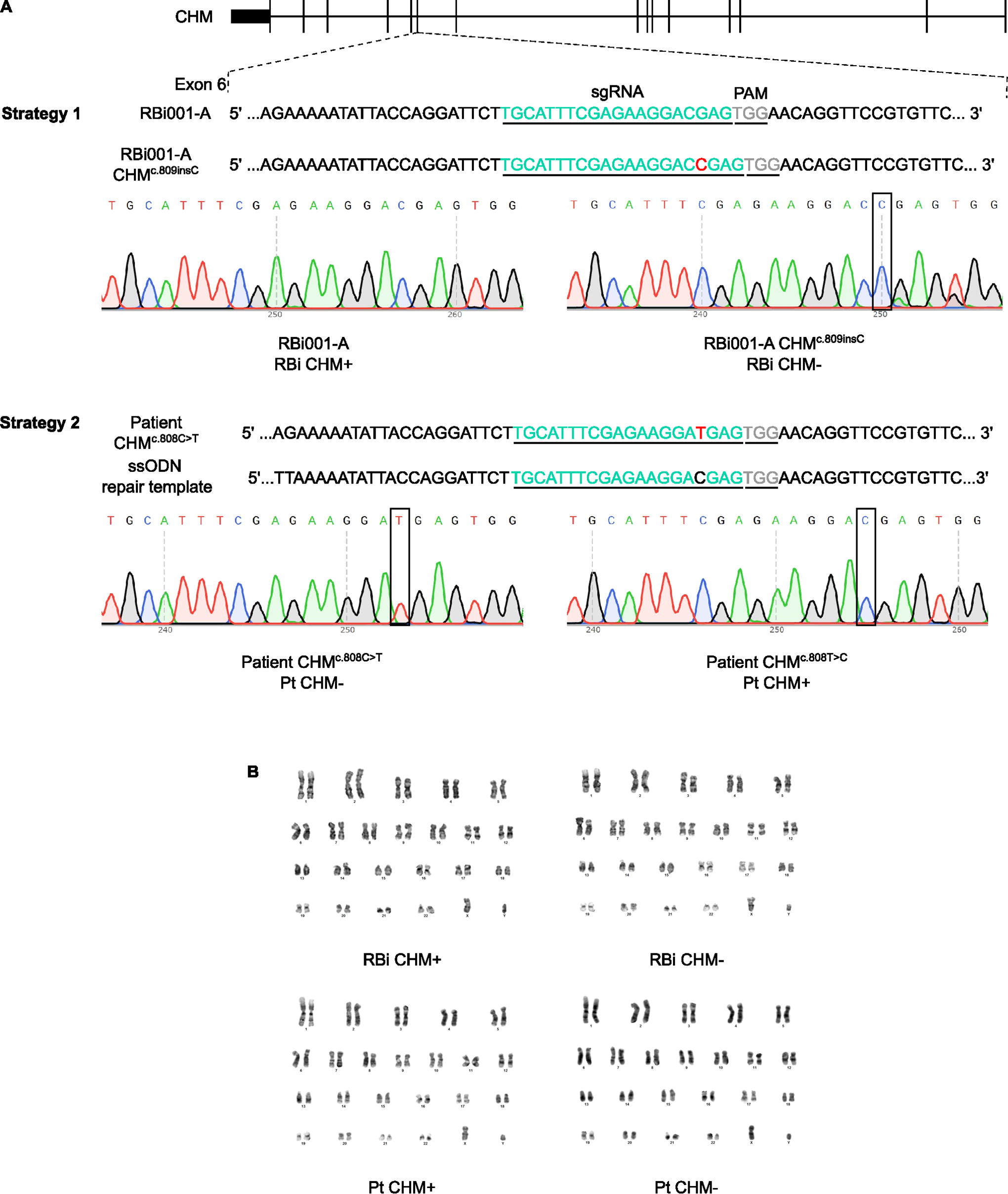
CRISPR/Cas9 strategies to generate isogenic hiPSC lines to model CHM disease.
Next, we analyzed the morphology and expression of pluripotency markers in the generated lines. All hiPSC lines exhibited typical pluripotent stem cell morphology (Fig. 2A), and expression of the pluripotency markers Oct-3/4, Nanog, and Sox-2 (Fig. 2B, D). Similarly, we observed that the mRNA expression of stemness markers (
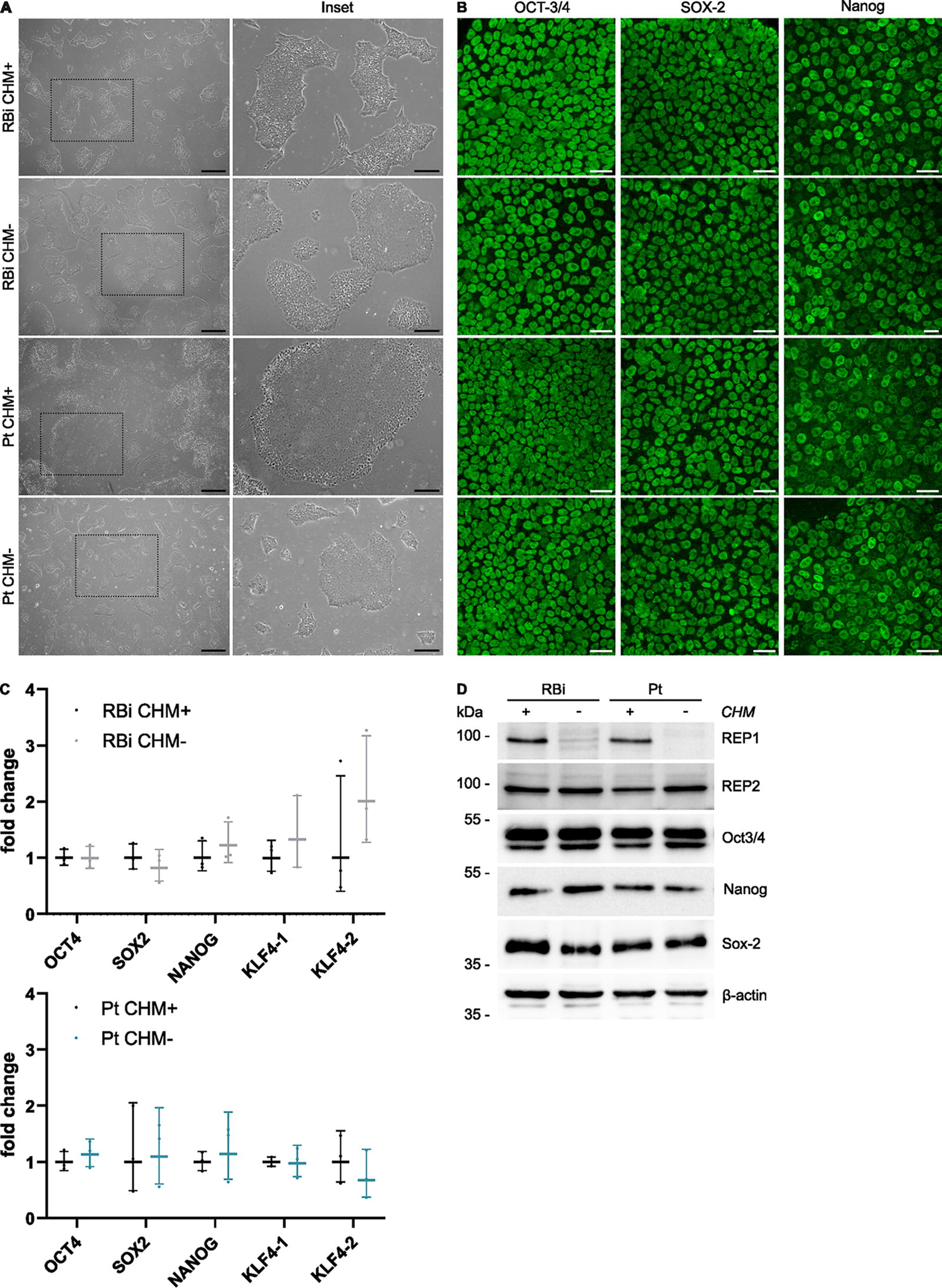
Characterization of the engineered CHM hiPSC lines.
REP1 deficiency in humans leads to a retinal exclusive phenotype, due to the activity of its isoform REP2 in remaining tissues, which is encoded by the CHM-like (
hiPSC CHM isogenic lines recapitulate key RPE features
CHM pathophysiology is complex, affecting three interdependent retinal layers (choroid, photoreceptors [PRs], and RPE), with the RPE playing a critical role in disease progression. 6 RPE cells provide crucial support to the PRs by providing nutrients and performing POS phagocytosis, among other functions. 9,10 Considering the importance of the RPE for CHM progression, we evaluated the capacity of our newly developed cell lines to acquire an RPE phenotype.
We were able to successfully differentiate the four hiPSC lines into RPE, which exhibited the typical polygonal morphology and polarization, as shown by the expression of the tight junction markers Zonula Occludens-1 (ZO-1) and Claudin-19 at the interface between adjacent cells (Fig. 3A). Polarization between apical and basolateral membranes was confirmed by measuring the TEER of one of the hiPSC-RPE pairs, with RBi CHM+ reaching a predicted value of 285.2 Ω cm−2 (231.9, 384.7) and RBi CHM− reaching 192.0 Ω cm−2 (159.4, 231.6), considering a 95% CI (Fig. 3C).
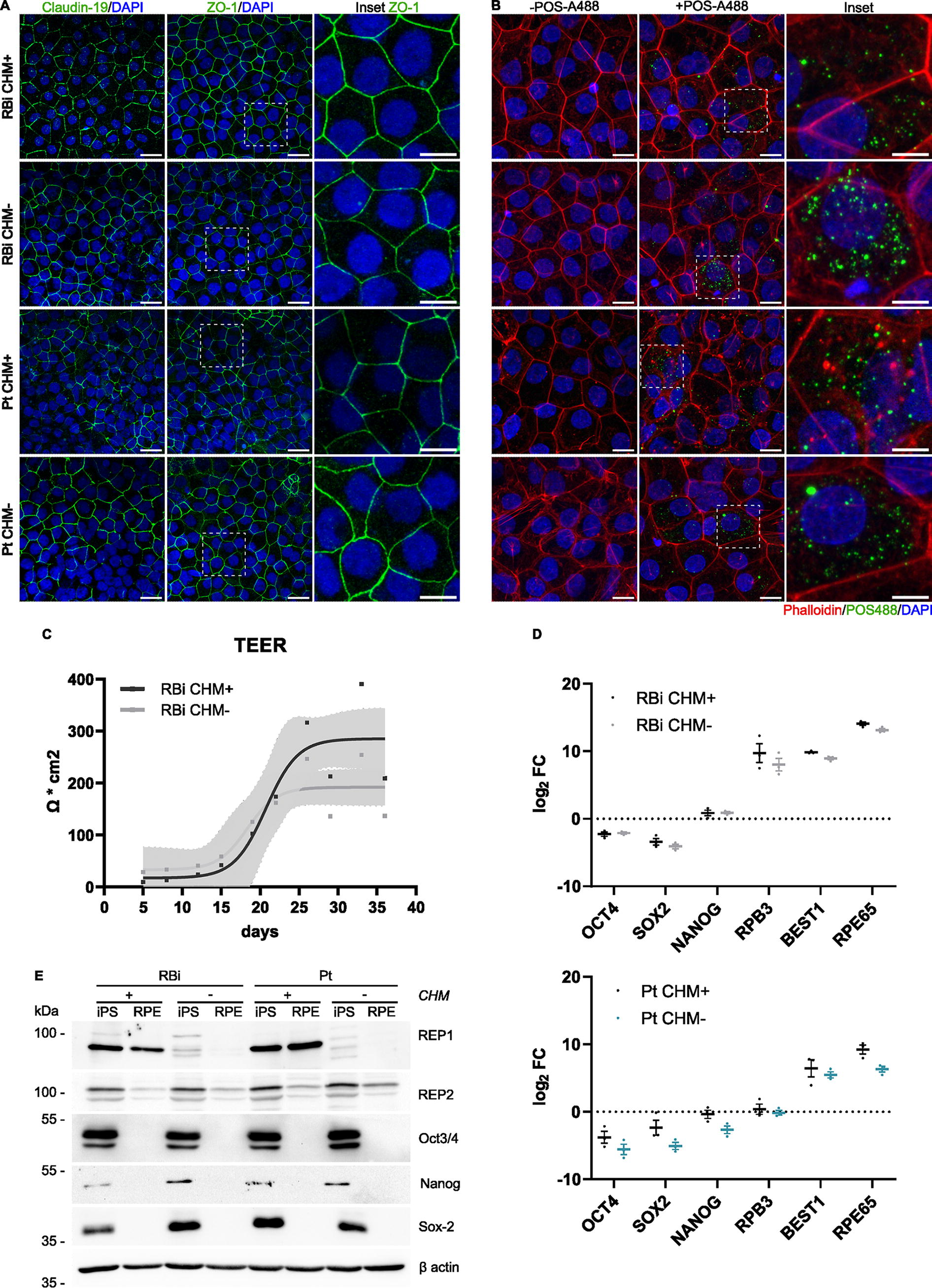
hiPSC CHM lines can be successfully differentiated into RPE.
We confirmed that the expression of pluripotent markers in hiPSC-RPE is greatly reduced (Fig. 3D, E), compared to RPE-specific genes (
Another hallmark of RPE cells is the presence of melanin pigment, and using transmission electron microscopy (TEM), we could observe vesicles of all (eu)melanogenesis stages, including stage I to stage IV melanosomes, in all the cell lines studied (Fig. 4A). Interestingly, when subculturing our hiPSC-RPE, we observed that cells lacking
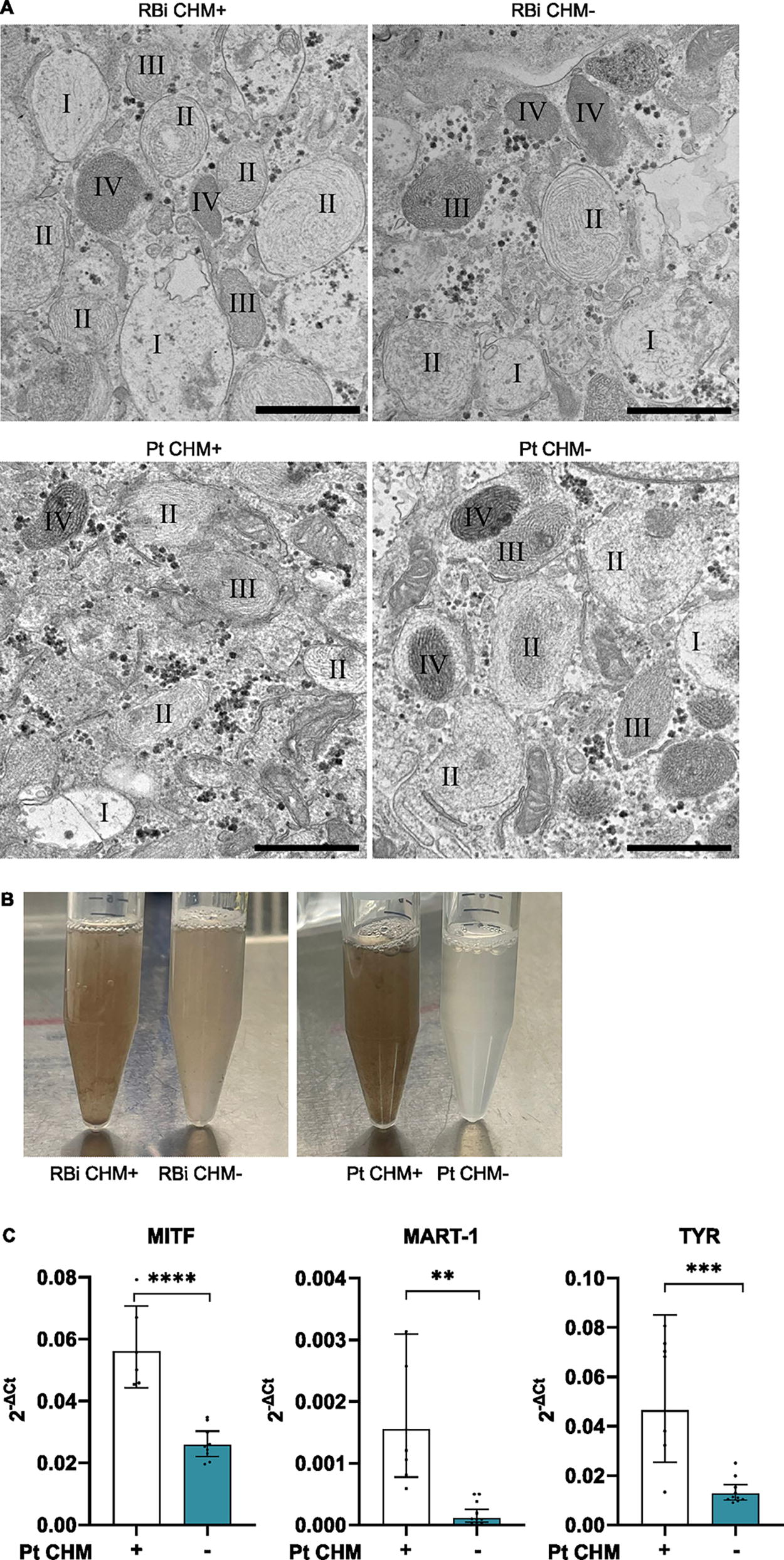
Pigmentation appears to be differentially expressed between healthy and CHM lines.
Embryoid bodies and retinal organoids obtained from hiPSC CHM isogenic lines
To explore the potential of the hiPSC CHM lines to recapitulate the self-organization of early embryogenesis, the four hiPSC lines were differentiated into embryoid bodies (EBs) and retinal organoids (ROs). During EB formation, no morphological change was detected between EBs derived from hiPSCs lacking
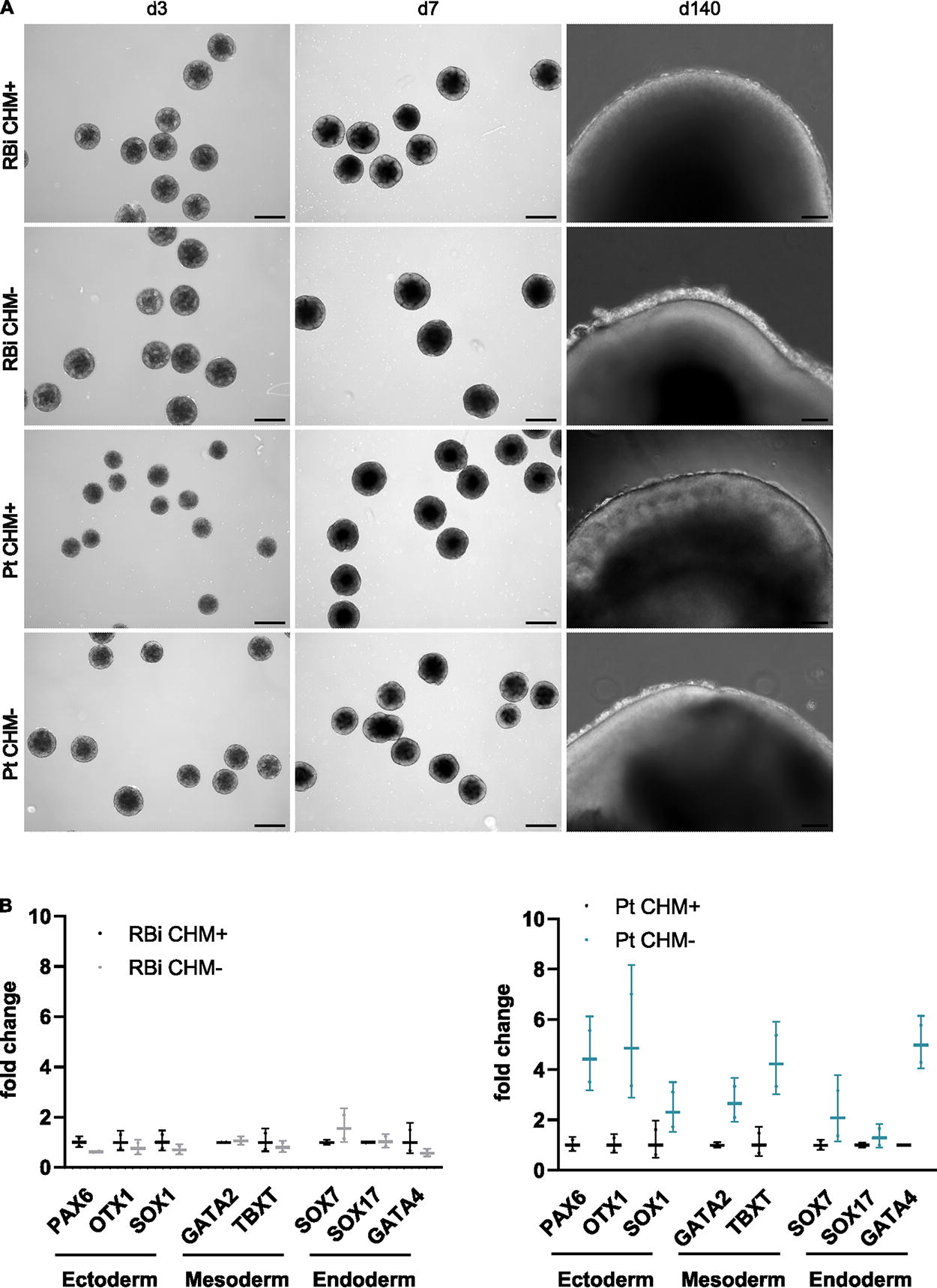
hiPSC CHM lines can effectively form EBs and differentiate into ROs.
Discussion
The complex pathophysiology of the CHM degenerative process involves the interaction between the RPE, choroid, and PRs, and depends heavily on RPE cells. 6 The aim of this work was to generate new cellular models to study the RPE in the context of CHM pathology, taking advantage of hiPSCs and CRISPR/Cas9 technology to develop isogenic models of the disease. In the long term, we believe that these models constitute a powerful platform to delve into the mechanisms of RPE degeneration in CHM, as well as other diseases of the RPE, which are still unclear.
The RPE is thought to be one of the primary sites of degeneration in CHM, being situated between the PRs and choroid. 6 Therefore, to accurately model CHM cellular phenotypes, we derived our newly developed hiPSCs into RPE and show that these cells reproduce key RPE features. These include the expression of tight junction markers ZO-1 and Claudin-19, demonstrating the typical RPE polygonal morphology, along with polarization, as shown by the TEER increase from 20d of differentiation onwards, observed for the RBi CHM pair. Moreover, all hiPSC-RPE express RPE-specific genes, as well as melanosomes in all stages of melanosome maturation, and have the capacity to phagocytose POS.
REP1 participates in the prenylation of Rab GTPases, a post-translational modification catalyzed by Rab geranylgeranyltransferase type II (also called GGTase-II) and required for proper cellular localization, membrane association, and protein–protein interactions of Rabs.
20
Rab proteins are involved in intracellular vesicle trafficking, making REP1 essential for this process, particularly for melanosome trafficking.
21
We observed that hiPSC-RPE lacking REP1 have less pigment than their control counterparts to the naked eye, and therefore we explored the possibility of a pigmentation deficiency in these cells. Indeed, some genes of the melanogenesis pathway are decreased in the Pt CHM− line, compared to the isogenic control Pt CHM+. Although no difference was observed in melanin formation by electron microscopy, and all stages of melanosome vesicles were present, the decreased expression of melanogenesis genes is in line with findings in zebrafish CHM mutant models (
Following synthesis, Rab proteins are escorted either by REP1 or its paralog REP2, presenting them to GGTase-II for prenylation. Subsequently, Rabs are inserted in their target membranes and activated, where they can perform their functions in membrane trafficking, vesicle formation, movement, and fusion. While REP2 also supports Rab prenylation, it does so with a different specificity than REP1. Rab proteins are of particular importance in the retina since they are involved in POS phagocytosis and trafficking of melanosomes to apical microvilli, contributing to RPE homeostasis. Hence, mutations that affect REP1 lead to a prenylation deficit, disrupting these pathways, a characteristic of CHM pathogenesis. 20
Interestingly, in our hiPSC-RPE models, we observed that REP2 protein expression decreased after RPE differentiation when compared to their undifferentiated counterparts. In agreement, Fry and colleagues, by using fibroblasts from CHM patients, mention that a higher protein loading was necessary for REP2 detection. 20 This observation hints at a possible lower REP2 expression when compared to REP1, which could be more noticeable in fully differentiated RPE cells than in fibroblasts. Altogether, these data suggest that the inability of REP2 to compensate for the loss of REP1 in CHM could be due to a combination of lower expression and Rab GTPase substrate specificity, resulting in a primary eye phenotype. 7,20
Finally, we show that these new hiPSC CHM isogenic models can differentiate into EBs, suggesting they are able to replicate the self-organization of early embryogenesis. The EBs formed express typical markers from the endodermal, mesodermal, and ectodermal lineages. Differentiation was further directed into ROs, which are becoming potent models for investigating retinal development and illnesses, as well as toxicology, and novel treatments for retinal diseases. 23 These 3D models are extremely valuable because they are also derived from hiPSCs and can be obtained from individuals suffering from specific disorders. ROs have the advantage of forming a 3D structure, mimicking all neuroretina cell types, including Müller cells and astrocytes, similar to the human retina. 23 Importantly, we observe that the organoids formed using our model cells develop PRs, known to be both photosensitive and phototransductive, enabling diverse types of studies such as advanced electrophysiology. 23
In conclusion, we developed pairs of isogenic hiPSC CHM lines representing new models, which can now be used to further explore the intricate mechanisms of retinal degeneration of this blinding disease.
Footnotes
Acknowledgments
The authors are grateful to David Gamm Lab (Wisconsin University) for the gift of a CHM male patient-derived hiPSC line CRFi001-A, and the Molecular Mechanisms of Disease Lab, the Ocular Low-Cost Gene Therapy Lab, and the Degeneration and Aging Lab for their advice and support. The authors thank the Cell Culture facility at NOVA Medical School (NMS), the scientific and technical assistance of T. Pereira from the NMS Microscopy facility. They thank the Electron Microscopy Facility at Instituto Gulbenkian de Ciência. They also acknowledge the local slaughterhouse (CASO — Centro de Abate de Suínos do Oeste) that generously provided porcine eyes.
Author Disclosure Statement
The authors state that the research was conducted without any commercial or financial relationship that could be regarded as a potential conflict of interests.
Funding Information
The authors declare financial support was received for the research, authorship, and/or publication of this article. Research was supported by Fundação para a Ciência e Tecnologia (FCT)—Portugal (including iNOVA4Health projects UIDB/04462/2020, UIDP/04462/2020, and associated laboratory LS4FUTURE with reference LA/P/0087/2020). A.F.F. is funded by FCT PhD studentship (2022.12254.BD). M.L.S. is funded by the individual grant CEECIND/01536/2018. A.S.F. postdoctoral contract is funded by “La Caixa Foundation” (NASCENT HR22-00569). P.A. is supported by grant EXPL/MED-OUT/0599/2021, funded by national funds from FCT/MCTES, and individual grant CEECIND/03862/2020. L.L. is a recipient of a Choroideremia Research Foundation postdoctoral fellowship, Throssell-Hillier Research Award. Research at the Center for the Unknown is supported by Fundação Champalimaud.
