Abstract
Prostaglandin E2 (PGE2) has recently gained attention in the field of regenerative medicine because of the beneficial effects of this molecule on stem cell proliferation and migration. Furthermore, PGE2 has the ability to mitigate immune rejection and fibrosis. In the colon and kidney, PGE2 induces YAP1, a transcription factor critical for cardiac regeneration. Establishing a similar connection in stem cells that can be transplanted in the heart could lead to the development of more effective therapeutics. In this report, we identify the effects of PGE2 on neonatal Islet-1+ stem cells. These stem cells synthesize PGE2, which functions by stimulating the transcription of the extracellular matrix protein Agrin. Agrin upregulates YAP1. Consequently, both YAP1 and Agrin are induced by PGE2 treatment. Our study shows that PGE2 upregulated the expression of both YAP1 and Agrin in Islet-1+ stem cells through the EP4 receptor and stimulated proliferation using the same mechanisms. PGE2 administration further elevated the expression of stemness markers and the matrix metalloproteinase MMP9, a key regulator of remodeling in the extracellular matrix post-injury. The expression of PGE2 in neonatal Islet-1+ cells is a factor which contributes to improving the functional efficacy of these cells for cardiac repair.
Introduction
Understanding the underlying mechanisms that make certain stem cells more efficacious is a critical area of research as it could lead to improvements in the selection, isolation, and enhancement of stem cells for therapeutic use. One challenge present in nearly all stem cell transplants is rejection. Rejection not only eliminates the transplanted cells, but the resulting immune response can also damage the surrounding tissue, exasperating the original injury. Immunosuppressants administered over the long term can have a substantial negative impact on patient health. Therefore, it is prudent to identify stem cell therapeutic options, which require minimal to no immunosuppressant use.
Allogeneic mesenchymal stem cells are initially tolerated but are rejected as they begin to differentiate. 1 Administration of prostaglandin E2 (PGE2) prevents rejection of these cells. PGE2 is an extremely potent, but transient, immunomodulator. PGE2 regulates activation and exerts anti-inflammatory effects on several types of leukocytes, including monocytes, Natural Killer cells, T cells, and macrophages. 1 –4 Furthermore, PGE2 induces angiogenesis, enhances proliferation, activates native stem cells, and mitigates fibrosis. 1,5 –8 PGE2 has a vast array of functions that vary depending on the microenvironment it operates within and which of its four G-coupled receptors (EP1–EP4) it binds to. Many of the pro-regenerative PGE2-associated functions are specifically tied to activation of PGE2 receptor 4 (EP4). 9 –11 These aspects make PGE2 and EP4 extremely attractive targets in cardiac regeneration as the adult heart has limited capacity for self-repair and is prone to fibrosis postinfarct.
In the colon and kidney, regeneration has been achieved through PGE2-mediated upregulation of the transcription factor YAP1 (yes-associated protein 1). 12,13 As YAP1 can facilitate functional recovery in infarcted hearts, identification of an equivalent pathway in a cardiac model would be of great interest. 14,15 In the heart, YAP1 is activated by the extracellular matrix (ECM) protein Agrin. 16,17 Agrin is expressed in neonatal cardiac tissue and has the ability to improve cardiac function, reduce infarct size, and mitigate fibrosis postmyocardial infarction through its interaction with YAP1. 16,18 Stem cells that continuously express PGE2, YAP1, and Agrin therefore have the potential to facilitate sustained cardiac repair.
Islet-1+ (Insulin gene enhancer protein ISL-1) stem cells express PGE2, YAP1, and Agrin and are enriched in cardiovascular tissue during the neonatal period. 16,19,20 The use of these cells for cardiac regeneration is of particular interest since Islet-1 is a transcription factor required for cardiac development during early embryogenesis and Islet-1+ stem cells differentiate into all cardiac cell types, including cardiomyocytes, smooth muscle, endothelial cells, and pacemaker cells as shown by lineage tracing and differentiation in vitro. 21 –24 In addition, Islet-1+ cells facilitate cardiac repair in small and large animal models. 24 –26 Given the challenges associated with acquisition and ethical concerns when using human embryonic stem cells (hESCs) and hESC-derived cardiovascular stem cells, other sources of Islet-1+ stem cell populations are of great interest. Neonatally derived Islet-1+ stem cells can be rapidly isolated and cloned as pure populations from cardiovascular tissue, and they facilitate cardiac repair in the sheep model of myocardial infarction. 27,28
The unique and well-established capacity of neonates to repair cardiac tissue after injury has intrigued investigators who are interested in improving outcomes in regenerative therapy. The mechanistic basis for this brief period of enhanced repair has not been fully elucidated. Neonatal Islet-1+ cardiovascular stem cells express Agrin and YAP1, but their numbers decline shortly after birth. 16,19,20 Our laboratory is interested in the identification of the unique features associated with neonatally-derived Islet-1+ stem cells. In this report, we have identified a novel signaling pathway in Islet-1+ stem cells, in which PGE2 elevates both YAP1 and Agrin through the EP4 receptor. Neonatal Islet-1+ stem cells secrete prostaglandin synthases and PGE2 receptors. PGE2 stimulation of neonatal Islet-1+ stem cells activates several transcription factors associated with stemness. The interaction of PGE2 in neonatal Islet-1+ stem cells with EP4 is responsible for elevated levels of YAP1 and Agrin, as well as the improved proliferation and enhanced stemness identified in these cells during the pro-regenerative window that occurs during the first few days after birth.
Materials and Methods
RNAseq analysis of human Islet-1+ stem cell clones and extracellular vesicles
Total RNA was generated from neonatal Islet-1+ stem cell clones, which were isolated from discarded cardiovascular tissue derived from 8- to 30-day-old patients, under Institutional Review Board (IRB) approval. 28,29 In short, cardiac tissue was digested with collagenase, passed through a 40 μm cell strainer, and cells were cloned by limiting dilution. Islet-1+ stem cells were characterized by flow cytometry based on their expression of PDGFR (Platelet-derived growth factor receptor), KDR (Kinase insert domain receptor), CD90 (Cluster of Differentiation 90), cKIT (tyrosine-protein kinase KIT), Islet-1, SSEA-4 (stage-specific embryonic antigen-4), CD34 (Hematopoietic Progenitor Cell Antigen CD34), and CLASS I HLA. 28,30 Their stemness profile was further characterized via real-time quantitative polymerase chain reaction (RT-qPCR) based on their expression of early stage markers, including SALL4, SOX2, EPCAM, TBX5, GATA4, and TFAP2C. 30 The extracellular vesicles presented in this study were isolated using an ExoQuick-TC® kit (System Biosciences, Palo Alto, CA) as previously described. 29 Ribo-Zero rRNA removal kit (Illumina, San Diego, CA) and an Ion Total RNA-Seq Kit v2 (ThermoFisher Scientific, Waltham, MA) were used to prepare cDNA libraries. Samples were purified using nucleic acid binding beads (Ambion, Austin, TX). Fragmented RNA was purified and hybridized to ion adapters before ligase addition, then treated with reverse transcriptase. cDNA libraries were generated and then amplified by PCR using Platinum PCR SuperMix High Fidelity and Ion Xpress Barcode primers. The size and distribution of the cDNA libraries were determined using Agilent 2100 Bioanalyzer after purification with nucleic acid binding beads and buffers (Ambion).
An Agilent dsDNA High Sensitivity kit (Agilent, Santa Clara, CA) was used to test the quality of final libraries. Samples were enriched using the Ion PI™ Hi-Q™ OT2 solutions 200 Kit and were run on an Ion OneTouch ES instrument. Sequencing was performed on an Ion Proton sequencer using an Ion torrent RNAseq run. Raw sequence files were then aligned to the human genome (Hg19) reference sequence by the alignment plugin tool in Ion Torrent Suite software. Quality of the resulting Binary Alignment Map files was assessed using the Strand NGS program, and low-quality reads were excluded based on alignment score. The remaining aligned reads were normalized and quantified using the DESeq algorithm in the Strand NGS software.
Regulation of the PGE2 signaling pathway in vitro
Islet-1+ stem cell clones were cultured at 37°C with 5% carbon dioxide and 95% oxygen using stem cell growth media generated with 66% M199 Life Technologies (Carlsbad, CA), 22% EGM2 Media (LONZA, Basel, Switzerland), 10% GenClone Fetal Bovine Serum from Genesee Scientific (El Cajon, CA), 1% PEN/STREP (100X) (Life Technologies, Carlsbad, CA), and 1% MEM NEAA (100X) (Life Technologies, Carlsbad). PGE2 (Cat No. #14010 Cayman Chemical, Ann Arbor, MI) was reconstituted with dimethyl sulfoxide (DMSO) (CORNING, Corning, NY), and the optimal concentration was determined in preliminary experiments. An equivalent amount of DMSO was added to control treatment groups. A measure of 10 μM of a competitive EP4 antagonist L161,982, Cat No. 10011565 (Cayman Chemical, Ann Arbor, MI) and 3 μM Verteporfin, a YAP1 inhibitor, Cat No. 5305 (Tocris, Bristol, England), were added to the cells 2 h before addition of 1 μM PGE2. A measure of 1 μM of the EP4 agonist L-902,688, Cat No. 10007712 (Cayman Chemical) and 1 μM PGE2 were added simultaneously.
RNA and cDNA preparation
Cells were lysed in QIAzol, and RNA was prepared using the miRNeasy Mini Kit (QIAGEN, Valencia, CA) following the manufacturer’s protocols. RNA quality and quantity were assessed using a NanoDrop 2000 spectrophotometer (ThermoFisher Scientific, Rochester, NY) and gel electrophoresis. cDNA was prepared using SuperScript III (Life Technologies, Carlsbad, CA).
Quantitative RT-PCR
RT-qPCR was performed using iTaq™ Universal SYBR® Green Supermix (Bio-Rad, Hercules, CA) and was run using a Bio-Rad CFX96 Touch Real-Time PCR Detection System (Bio-Rad). The run conditions were 94°C for 10 min and 45 cycles of 94°C for 15 s, 58°C for 60 s, and 72°C for 30 s. RT-qPCR products were verified using gel electrophoresis to confirm correct gene amplification. Primers were designed using the National Center for Biotechnology Information Primer-BLAST system and were manufactured by Integrated DNA Technologies (Coralville, IA). Primer sequences are listed in Supplementary Table S1.
Protein purification and analysis
Islet-1+ stem cells were treated for 4 h with PGE2 exclusively or PGE2 + L-161,982 in addition to a matched DMSO control. The L-161,982 was preincubated for 2 h before PGE2 addition. At 4 h after PGE2 treatment, plates were aspirated, washed with cold phosphate-buffered saline, and incubated on ice with cold trypsin (ThermoFisher Scientific, Waltham, MA) until cells were detached. The cells were resuspended in a protein lysis buffer solution, which consisted of RIPA buffer, 0.5M of ethylenediaminetetraacetic acid, protease inhibitor cocktail, sodium orthovanadate, and sodium fluoride. This solution was agitated for 1 h at 4°C. The samples were centrifuged at 14,000 g and aliquoted. Protein concentration was quantified using the Micro BCA Protein Assay Kit (ThermoFisher Scientific, Waltham, MA) and measured using spectrophotometer. The gel-free western blotting system, Simple Wes (ProteinSimple, San Jose, CA), was used to quantify specific protein expression following the associated protocol provided by the manufacturer. Antibodies and their dilutions are shown in Supplementary Table S2 (Cell Signaling Technology, Danvers, MA).
Proliferation assay
Cells were seeded at 5,000 cells/well in a 96-well plate with 100 µL of media. The five treatment groups included a media and DMSO control (CORNING), 1 µM PGE2 Cat No. 14010 (Cayman Chemical), 1 µM L-902,688 (Cayman Chemical), 10 µM L161,982 + 1 µM PGE2, and 3 µM of Verteporfin + 1 µM PGE2 (Tocris). Verteporfin and L-902,688 were added 2 h before PGE2. At 4 h after PGE2 was added, plates were treated with 10 µL water-soluble tetrazolium salt (WST) reagent provided in Biovision’s Quick Cell Proliferation Colorimetric Assay Kit Plus (Milpitas, CA). The plate was incubated for an additional 4 h and then briefly placed on a shaker (Benchmark Scientific, Sayreville, NJ). Absorbance measurements were generated using a BIO-TEK uQuant microplate spectrophotometer (BioTek, Winooski, VT).
Statistics
RT-qPCR data, ProteinSimple data, and spectrophotometer readings were analyzed using Excel, R, and PRISM software and are reported here as the mean ± standard error. Gene expression was calculated using the 2−ΔΔCT method. Normality was determined using a Shapiro–Wilk test. Differences between control and treated datasets were determined using unpaired t-tests to determine statistical significance. To compare non-normal datasets, Mann–Whitney U test was performed to analyze statistical significance. P values <0.05 were considered statistically significant. Beta-actin and GAPDH were used as housekeeping controls to normalize the expression of genes and proteins of interest.
IRB approval
The IRB of Loma Linda University approved a protocol (5110115) for the use and acquisition of discarded cardiac tissue acquired as a byproduct of cardiovascular surgery with a waiver of informed consent. No identifiable patient information was collected. Clonal cell populations were generated from this tissue for further study as previously reported. 28
Results
Subpopulations of neonatal Islet-1+ stem cell clones preferentially express PGE2 signaling components
The transcriptome of Islet-1+ clones isolated from cardiac tissue collected from neonates varying in age from 8 days to 1 month postpartum was examined using RNAseq analysis. The 8-day clone, which was isolated during the pro-regenerative window, expressed elevated levels of PGE2 signaling components in addition to elevated levels of transcripts associated with proliferation, stemness, and ECM reorganization, respectively (Fig. 1A–D). The extracellular vesicle content of a neonatal Islet-1+ stem cell clone, isolated within the pro-regenerative window, further demonstrated enriched PGE2-related transcript content (Fig. 1E).
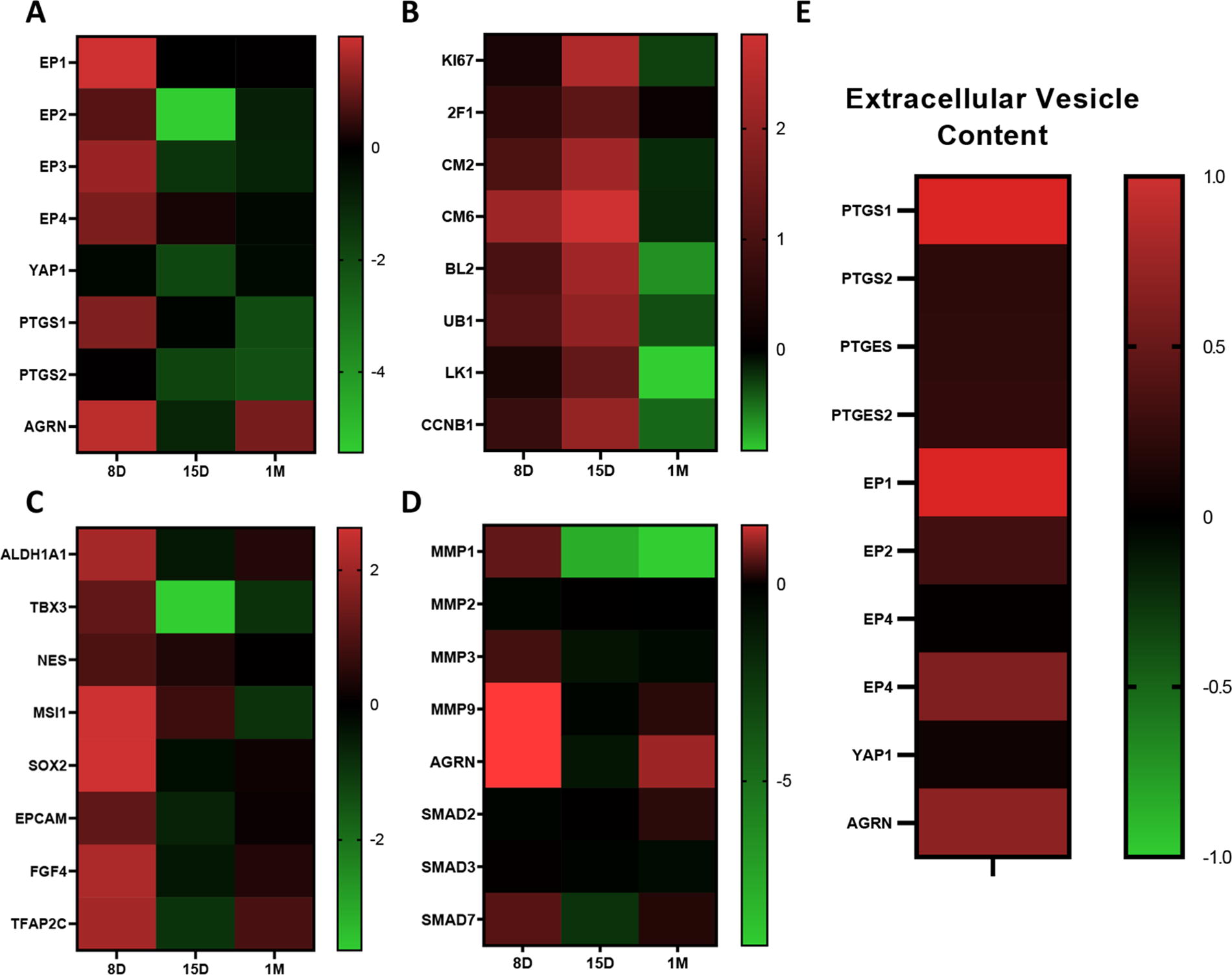
Select neonatal Islet-1+ stem cell clones preferentially express PGE2 signaling components. RNAseq analysis demonstrated that the Islet-1+ stem cell clone derived from the 8-day-old neonate expressed the highest levels of PGE2-signaling components and Agrin as shown in the heatmap
PGE2 induces stemness markers and MMP9
Neonatal Islet-1+ stem cells were treated with 1 μM PGE2 for 4 h. RT-qPCR analysis demonstrated that PGE2 stimulation upregulated the expression of several well-established stemness markers, including NES, SOX2, EPCAM, MIXL1, ETV2, and OCT4 (Fig. 2A). Transcripts encoding MMP9 and MMP2, two key metalloproteases with pivotal roles in cardiac remodeling, were elevated in response to PGE2. 31 –33 MMP9, in particular, plays a key role in both initiating and resolving inflammation, which is contingent on the cell type and stage of injury. 31,32 Our results identified a significant 4-fold induction of MMP9 but not MMP2 post PGE2 treatment, suggesting that PGE2 treatment influences ECM reorganization (Fig. 2B).
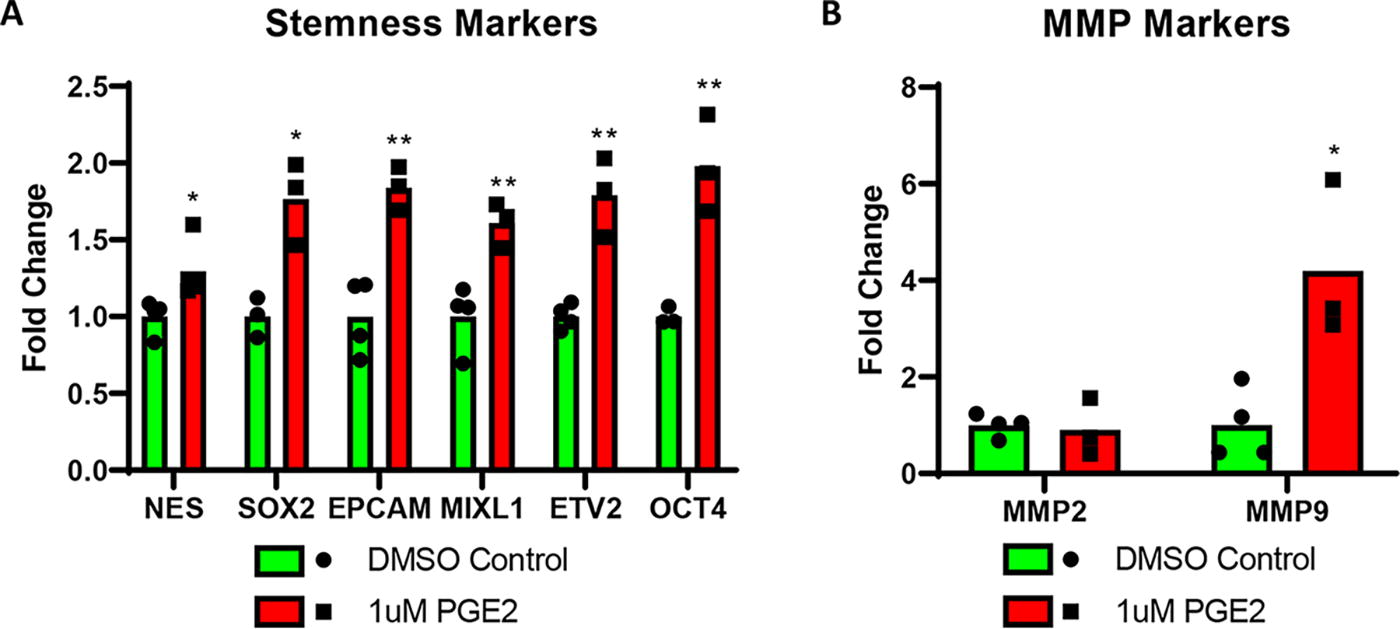
Prostaglandin E2 induces stemness markers and MMP9. A 4 h treatment with 1 μM PGE2 demonstrated a significant induction of numerous stemness markers
PGE2 induces YAP1 and Agrin in neonatal Islet-1+ stem cells
To determine whether PGE2 activates YAP1 in our model, neonatal Islet-1+ stem cells were treated with 1 μM PGE2 over a 1-week period. RT-qPCR revealed that YAP1, Agrin, and EP4 were all elevated, peaking at ∼72 h and subsequently declining (Fig. 3A–C). At 144 h, all three transcripts of interest had significantly decreased compared with their peak. A media change at 72 h was necessary to sustain the 144 h time point. The media was replaced with either control media + DMSO or media with the same initial concentration of PGE2. When the additional PGE2 was applied, the cells were unable to further induce or reactivate YAP1, suggesting that a tightly regulated pathway exists that cannot be persistently upregulated (Supplementary Fig. S1). Additional studies were conducted at shorter time points to determine the initial sequence of activation following PGE2 treatment.
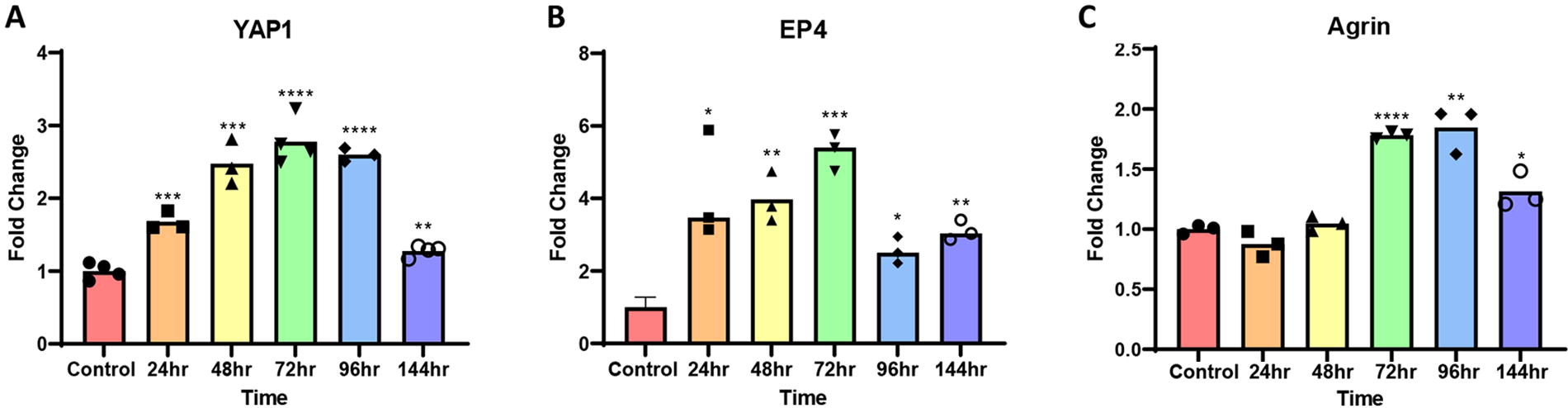
Prostaglandin E2 transiently induces YAP1, EP4, and Agrin. Treatment of neonatal Islet-1+ stem cells with 1 μM PGE2 induced transcripts encoding YAP1
PGE2 induces YAP1 and Agrin in neonatal Islet-1+ stem cells through the EP4 receptor
At 4 h after PGE2 activation, YAP1 transcript and protein levels were induced along with transcript levels of CTGF, a marker of intranuclear YAP1 activity (Fig. 4A–D). To elucidate the role of EP4 as the PGE2 receptor responsible for the downstream induction of YAP1 and Agrin, and to assess whether activated YAP1 plays an active or coincidental role in modulating these components, both EP4 and YAP1 were blocked with antagonists. A measure of 10 μM of L-161,982, a competitive EP4 antagonist, and 3 μM Verteporfin, a YAP1 inhibitor that functions by sequestering YAP1 in the cytoplasm, were cocultured with the cells 2 h before PGE2 activation. 34 Both the EP4 and YAP1 inhibition blocked YAP1 induction despite PGE2 treatment (Fig. 4A, C, D). CTGF was also downregulated by the competitive EP4 antagonist (Fig. 4B). At 8 h, EP4, prostaglandin synthase 1 (PTGS1), prostaglandin synthase 2 (PTGS2), and Agrin were all significantly induced by PGE2. This effect was blocked by the addition of both the EP4 and YAP1 antagonist. In addition, YAP1, EP4, and Agrin were all significantly downregulated compared with their respective DMSO controls, suggesting that YAP1 and EP4 inhibition blocks an intrinsic cyclical pathway in these cells.
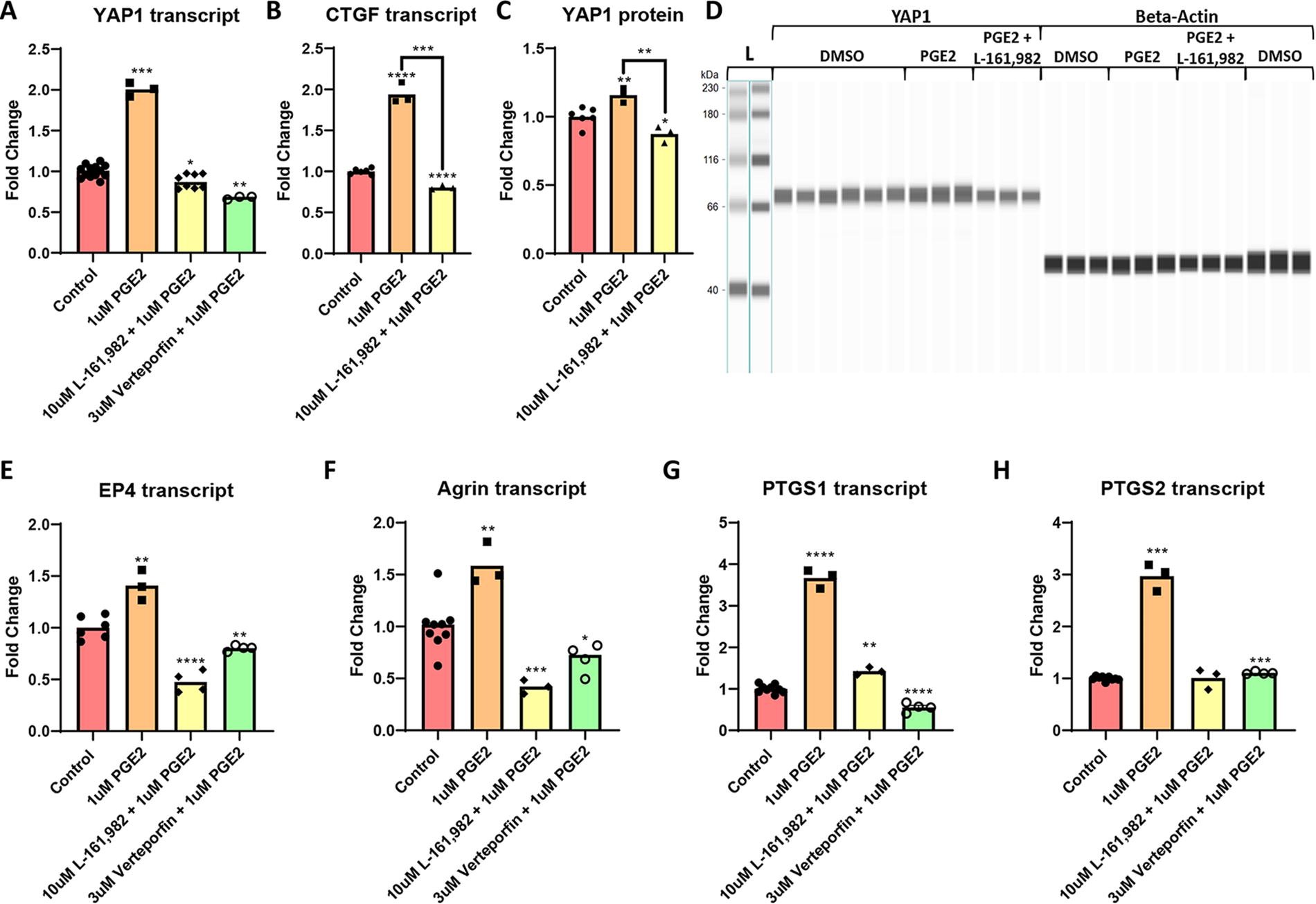
Prostaglandin E2 induces YAP1 and Agrin in neonatal Islet-1+ stem cells through EP4. At 4 h after PGE2 treatment, RT-qPCR analysis revealed that YAP1 and CTGF transcript expression was elevated. Administration of the EP4 antagonist, 10 μM L-161,982, blocked this response. The YAP1 inhibitor, Verteporfin, prevented PGE2-mediated YAP1 transcript induction
PGE2-mediated activation of EP4 and YAP1 induces proliferation
Our proposed model (Fig. 5A) visualizes a mechanism whereby PGE2 induces YAP1 through the EP4 receptor. Subsequent YAP1 activation leads to an increase in expression of Agrin, PTGS1/2, EP4, and YAP1. To expand on the functional impact of these changes, a proliferation assay was conducted. At 8 h post PGE2 treatment, proliferation was induced. The EP4 antagonist L-161,982 and Verteporfin, a YAP1 antagonist, blocked this proliferative response. Addition of an EP4 agonist, L-902,688, induced proliferation (Fig. 5B). These findings demonstrate that PGE2 induces proliferation through the EP4 receptor and subsequent YAP1 activation in our model. Both YAP1 and EP4 inhibition reduce proliferation compared with the DMSO control, indicating that this pathway is a naturally active component of the neonatal Islet-1+ stem cell proliferative pathway.
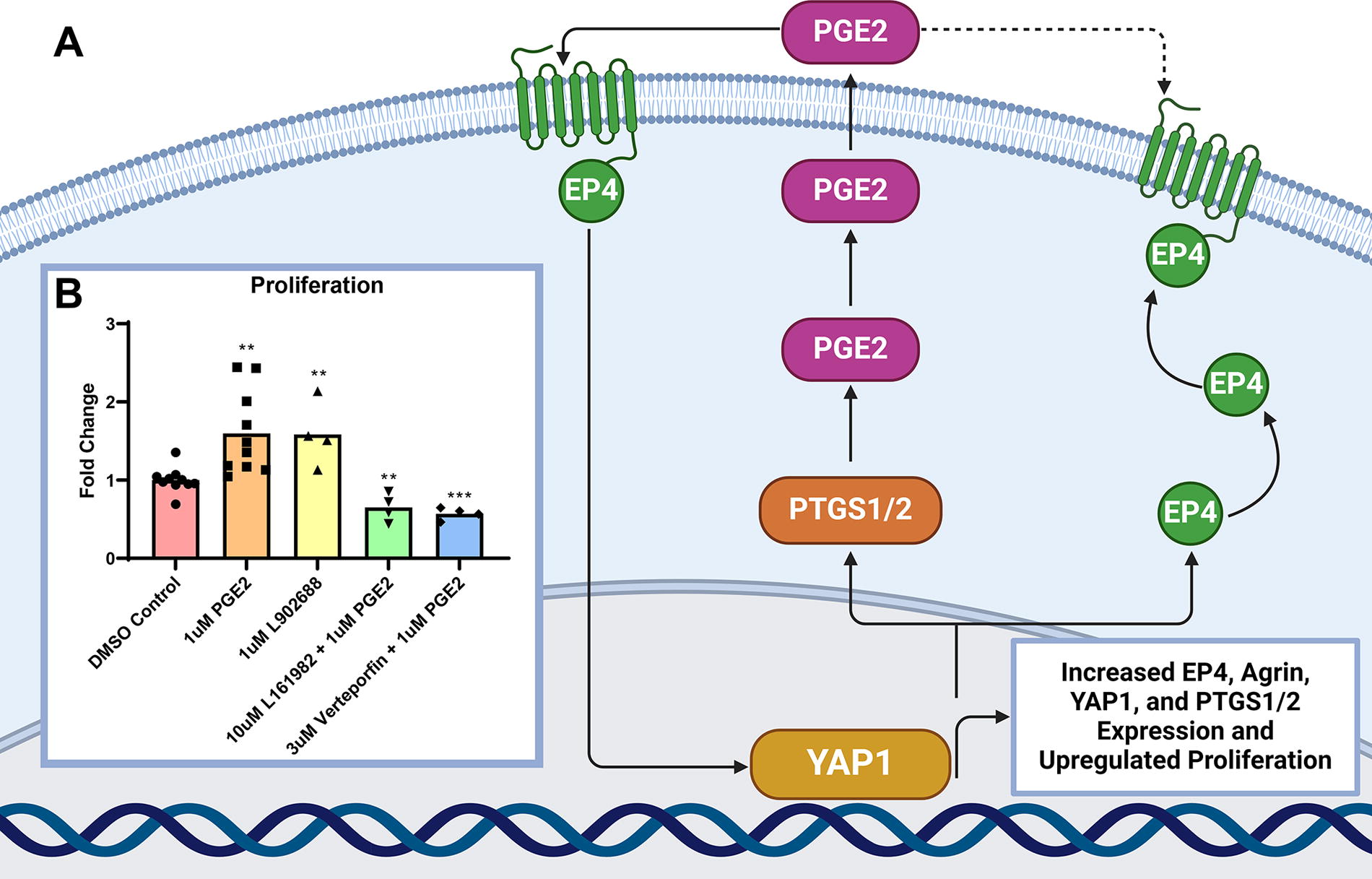
Prostaglandin E2 induces proliferation through EP4 and YAP1. A pathway visualizing the cyclical pathway in which PGE2 induces YAP1 and Agrin through EP4 and YAP1 was generated using Biorender.com
Discussion
In this report, we have described a novel pathway in which PGE2 induces YAP1 and Agrin through EP4 in neonatal Islet-1+ stem cells. This is the first report to our knowledge to describe any causal relationship between PGE2 and Agrin and the first to connect PGE2 and YAP1 in a cardiac system. Cells that can utilize this pathway may be more beneficial therapeutically given the numerous documented benefits of YAP1, Agrin, and PGE2 in wound repair and regeneration. PGE2 induces cardiomyocyte proliferation, migration, and angiogenesis; activates endogenous stem cells; inhibits fibrosis; and has extremely potent immunomodulatory capabilities that can inhibit rejection of implanted allogeneic stem cells. 1,5 –7,9,10 PGE2 prevents the generation of alloantibodies and modulates pro-regenerative M2 macrophage polarization and recruitment via the EP4 receptor. 1,35 –38 EP4 agonists reduce infarct size postmyocardial infarction by regulating cardiac fibroblasts. 9,11 PGE2 and EP4 agonists have been shown to protect the heart from ischemia/reperfusion injury via EP4. 39 Our model demonstrates that the pathway activated by EP4 results in a feedback loop whereby YAP1 and Agrin expression are induced, and prostaglandin synthase expression is elevated. Activation of this pathway stimulates the production of proteins expressed during the pro-regenerative neonatal window and may induce protective and regenerative effects in the heart.
YAP1 and Agrin are key regulators of cardiac repair that are expressed at elevated levels in neonatal cardiac tissue during the pro-regenerative window. 16,20 While YAP1 alone can induce cardiac regeneration in vivo, this is contingent upon YAP1 being able to translocate into the nucleus where it can act as a transcription factor. 14 –17 Only dephosphorylated YAP1 can accomplish this while phosphorylated YAP1 is sequestered in the cytoplasm by dystrophin-glycoprotein complex (DGC). 16,17 In neonates, Agrin binds to DGC in cardiomyocytes, releasing YAP1, and activating dedifferentiation and proliferation in mature cardiomyocytes. 16,17 These events stimulate nondividing cardiomyocytes to dedifferentiate into the proliferative neonatal phenotype. 16,17 In our study, PGE2 induces YAP1, as well as OCT4, SOX2, and CTGF, which have been established as downstream markers of active YAP1. 40 –42 Consequently, PGE2-mediated activation leads to increased cell proliferation and elevated levels of early stemness markers. In mammary stem cells, the PGE2/EP4 axis maintains the stem cell population through activation of a similar pathway. 43 In this model, EP4 antagonism converts basal mammary stem cells from a mesenchymal/stem cell state to a nonbasal state, which has diminished functional capacity and reduced expression of stem cell markers. 43 In our model, this pathway maintains stemness via YAP1 release through Agrin. In neonates, Agrin not only promotes de-differentiation but also functions to mitigate fibrosis, reduce infarct size, and inhibit adverse remodeling. 16 –18 Stemness has been shown to improve engraftment and cardiomyocyte differentiation. 44 We may therefore infer that Islet-1+ stem cells treated with PGE2, which exhibit an enhanced stemness profile, are potentially capable of demonstrating enhanced functional repair.
Studies defining the ways in which PGE2 influences stem cells have demonstrated a predominantly positive relationship. PGE2 has been shown to enhance hematopoietic stem cell homing, survival, and proliferation. 45 Furthermore, dimethyl PGE2 has been shown to enhance hematopoietic stem cell transplantation with a high degree of safety in a preclinical nonhuman primate study. 46 PGE2 regulates hematopoietic stem cells directly through EP4, a finding analogous with our results. 47 In addition, PGE2 has been shown to be required to maintain the self-renewal potential of mesenchymal stem cells. 48 In cardiac models, previous studies have demonstrated that PGE2 can influence native cardiac stem cells and enhance cardiomyocyte replenishment. 5,8 Therefore, implanting PGE2-producing stem cells could enhance repair. Dhingra et al. compared stem cell transplantation with and without supplemental PGE2 and found that allogeneic mesenchymal stem cells administered without supplemental PGE2 were rejected as they differentiated but transplantation of a PGE2 hydrogel in this model alleviated the allogeneic immune response, prevented rejection, and restored cardiac function 5 weeks post-transplant. 1
The hESC-derived, SSEA-1+ Islet-1+ progenitors used by Menasché’s group involve the application of stem cells at a similar stage of differentiation as those reported in our current study, using cells with comparable markers. The major differences lie in acquisition methods and the clonal nature of the cells that were isolated from neonates in our study. Studies done using purified, hESC-derived Islet-1+ stem cells reduce scar tissue post-transplantation in a nonhuman primate model and demonstrated significant improvement in cardiac function in early clinical trials using a purified population with no evidence of tumor formation. 26,49 Purification is necessary since unpurified, cardiac-committed cells developed teratomas in nonhuman primates. 26 The purified clonal neonatal Islet-1+ cells examined in vivo by our research group have similarly demonstrated no evidence of teratoma formation two months post-transplant. 27
Our study provides new insight into one of the mechanisms by which neonatal Islet-1+ stem cells, isolated during the pro-regenerative window, upregulate transcripts associated with stemness, pluripotency, and cardiac repair. We report the novel finding that these Islet-1+ stem cells produce prostaglandin that functions to activate both YAP1 and Agrin through the EP4 receptor. This pathway is active by default in our Islet-1+ stem cells and is naturally self-regulated, although it can be enhanced and inhibited. Considering that persistently active YAP1 has been linked to carcinogenesis, the fact that this pathway cannot be continuously upregulated suggests that our Islet-1+ cells may be a safe and effective option for stem cell transplant. 12 Our findings suggest that screening for PGE2 expression in stem cells selected for cardiac repair could lead to the development of more effective therapeutics.
Footnotes
Acknowledgment
The authors thank Victor Camberos for optimizing the PGE2 concentrations used in this study.
Data Availability Statement
The presented transcriptomic data are available in Supplementary Table S3. Additional data will be made available upon reasonable request.
Author Disclosure Statement
The authors declare that they have no conflicts of interest to report.
Funding Information
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the
Supplementary Material
Supplementary Figure S1
Supplementary Table S1
Supplementary Table S2
Supplementary Table S3
