Abstract
Extinction rates are rising, and current conservation technologies may not be adequate for reducing species losses. Future conservation efforts may be aided by the generation of induced pluripotent stem cells (iPSCs) from highly endangered species. Generation of a set of iPSCs from multiple members of a species can capture some of the dwindling genetic diversity of a disappearing species. We generated iPSCs from fibroblasts cryopreserved in the Frozen Zoo®: nine genetically diverse individuals of the functionally extinct northern white rhinoceros (Ceratotherium simum cottoni) and two from the closely related southern white rhinoceros (Ceratotherium simum simum). We used a nonintegrating Sendai virus reprogramming method and developed analyses to confirm the cells' pluripotency and differentiation potential. This work is the first step of a long-term interdisciplinary plan to apply assisted reproduction techniques to the conservation of this highly endangered species. Advances in iPSC differentiation may enable generation of gametes in vitro from deceased and nonreproductive individuals that could be used to repopulate the species.
Introduction
The earth is currently experiencing global wide extinction levels that are higher than any previously experienced, termed “the sixth extinction” [1], which has ultimately been caused by human activities. Attempts to save species through habitat conservation, captive breeding, and education programs have helped some species such as the giant panda (Ailuropoda melanoleuca) [2] and the California condor (Gymnogyps californianus) [3]. However, for species with small population sizes and low genetic diversity this may not be enough to maintain a stable population. Technological advances in stem cell research may allow us to augment these more traditional conservation efforts through the generation of in vitro gametes from cryopreserved cell lines.
All five rhinoceros species and their subspecies are listed on the International Union for Conservation of Species (IUCN) Red List [4] (Supplementary Table S1), with poaching and habitat loss the most destructive to these species. Between 2013 and 2017, more than 1,000 rhinoceros were poached each year in South Africa alone for the illegal trade in their horn [5,6]; poaching has diminished recently, but still continues [7]. Rhinoceros horn is seen as a status symbol and erroneously believed to have medicinal value in several cultures, which has continued to drive the illegal trade [8].
The northern white rhinoceros (Ceratotherium simum cottoni: NWR) and southern white rhinoceros (Ceratotherium simum: SWR) are considered to be either separate species or subspecies [9,10], and each was hunted to near extinction in the last century. Both forms survived severe bottlenecks during the 20th century; SWR dwindled to fewer than 100 in the early part of the century [11] and only about 15 NWR remained by the 1980s [12]. The SWR population recovered, growing to ∼18,000 individuals, and is now the most abundant form of rhinoceros, although still considered near threatened by the IUCN due to continued poaching. Unfortunately, the NWR was unable to rebound being exposed to civil war, extreme poaching, and habitat loss and fragmentation, which prevented the species from recovering [13].
The NWR is now functionally extinct, with only two nonreproductive females remaining, a mother-daughter pair, living under armed guard on a preserve in Kenya, Ol Pejeta. Even if these animals could reproduce, they are unlikely to sustain a viable population. However, as technology advances, we may have other options. In 2015, we met with colleagues worldwide to outline a plan to save the NWR from extinction through the use of cellular and assisted reproduction techniques [14]. The key to applying these new techniques to endangered species is to have viable cellular material available.
The Frozen Zoo® [15,16] (San Diego Zoo Global), a repository of biomaterials established in 1975, contains fibroblast cell lines from 12 NWR individuals, eight of which are unrelated, and of which five never reproduced. These cell lines were established opportunistically from skin biopsies, cryopreserved from 1979 to 2016, and capture a high level of genetic diversity from this species [17]. Supplementary Fig. S1 shows the pedigree of the available cell lines. With the development of methods to convert fibroblasts into induced pluripotent stem cells (iPSCs) [18], these cryopreserved cells have the potential for genetic rescue applications.
iPSCs can divide indefinitely in culture and are capable of differentiation into any cell type in the body, including gametes [19 –21]. Thus, using iPSCs, it may be possible to eventually apply assisted reproduction methods to generate new, genetically unique, individuals of an extinct species. Rhinoceros belong to the mammalian Order Perissodactyla, odd-toed ungulates, which include horses, rhinoceros, and tapirs. There has been limited effort in generating iPSCs for this order, and work has mostly focused on therapeutic uses for veterinary medical applications in horses, Equus caballus [22,23].
In 2011, we reported the first generation of iPSCs from an endangered species, using integrating lentiviral vectors carrying the human reprogramming factors [POU5F1 (also known as OCT4), SOX2, MYC, and KLF4] for reprogramming NWRs and a nonhuman primate, the drill [24]. Since transgene integration can disrupt recipient genomes and integrated reprogramming factors can be reactivated, this approach is not suitable for making genomically unaltered iPSC-derived gametes for use in assisted reproduction technology.
In this study, we report the generation of transgene-free iPSCs from nine NWR and two SWR individuals, using improved reproducible methods for rhinoceros cells. Using these methods, we generated iPSCs from nine NWR individuals and two SWRs using nonintegrating Sendai vectors. We optimized reprogramming methods, developed species-specific culture conditions, and identified markers of pluripotency and differentiation for rhinoceros cells.
Materials and Methods
Fibroblasts
NWRs and SWR fibroblast cell lines were obtained from the San Diego Zoo Institute for Conservation Research Frozen Zoo biobank and expanded. No IACUC approval was required for the transfer of previously established cell lines. Cells were cultured at 37°C with 5% CO2 in either Dulbecco's modified Eagle's medium (DMEM) 1X+GlutaMAX™ (Gibco; supplemented with 10% fetal bovine serum, and 1% MEM nonessential amino acids; NEAA, Gibco) or a 1:1 mix of the supplemented DMEM and Fibroblast Growth Medium (FGM™-2; Lonza). The additional proprietary growth factors and supplements (including 0.5% recombinant human fibroblast growth factor-b (rhFGF-b) and 0.05% recombinant human insulin) in the FGM-2 formulation have been shown to improve the growth of rhinoceros fibroblast cells [25,26] and have been key to the growth and biobanking of many rhinoceros species in the Frozen Zoo. We refer to this medium as 50:50 throughout.
iPSC culture
iPSC medium
Basic iPSC medium contained DMEM:F12 (1:1; Gibco) with 20% KnockOut™ Serum Replacement (KOSR; Gibco), 1% NEAA, and 1% GlutaMAX. We supplemented daily with 12 ng/mL basic fibroblast growth factor (bFGF): (Biopioneer) and 0.1 nM β-mercaptoethanol (Gibco).
KB medium (rhinoceros iPSCs)
During the optimization, we adapted the basic iPSC medium formulation to incorporate the rhinoceros-favored FGM-2 medium. We used the same final concentrations of all components, substituting the DMEM:F12 with equal parts DMEM 1X+GlutaMAX and FGM.
Initial reprogramming conditions
Angalifu
In our first reprogramming experiments, fibroblasts from “Angalifu” (laboratory no. KB 9947), passage 10, were reprogrammed using the CytoTune™ 2.0 Kit (Thermo Fisher Scientific). Cells were thawed 2 days before the start of reprogramming, plated on 0.1% gelatin in supplemented DMEM at an approximate cell density of 13,000 cells/cm2 (∼70% confluent on day 0), and transduced with the recommended multiplicity of infection (MOI) for each vector for human cell reprogramming (ie, KOS-5, MYC-5, and KLF4-3).
The medium was changed according to the protocol. In brief, fibroblast medium (supplemented DMEM) was changed within 24 h postinfection and every other day until replating. The cells were replated 5 days after transduction using 0.05% Trypsin-ethylenediaminetetraacetic acid (EDTA) (Gibco), 2 days earlier than recommended due to overgrowth of the cells. Cells were plated at a density of 180 cells/cm2 onto irradiated (mitotically inactivated) CF-1 mouse embryonic fibroblast (MEF) feeder layers or 5,300 cells/cm2 onto Geltrex™ (Gibco). All cultures were changed to iPSC medium 24 h later and subsequently fed every day.
Seven colonies, across both conditions, were picked using a needle between days 14 and 16 post-transduction and plated onto MEFs. Plates became severely overgrown after day 16, resulting in high cell death; therefore, we did not maintain the plates for the full 4 weeks recommended by the protocol. We successfully obtained one clone from this reprogramming attempt, which originated from a Geltrex coated plate. The remaining colonies were either partially reprogrammed or outcompeted by the carried over fibroblasts.
Feeder-free culture conditions
This rhinoceros iPSC clone was also successfully adapted to feeder free conditions using mTeSR™1 (Stem Cell Technologies) medium and the Geltrex substrate. Previous attempts with less nutrient rich and more defined media such as mTeSR-E8™ (Stem Cell Technologies) failed to support growth of the rhinoceros cells. Cells grown in feeder-free conditions were passaged using StemPro® Accutase® (Gibco).
Sendai virus
The Sendai virus appeared to have little to no cytotoxicity for NWR fibroblast cells, which we needed to account for during the initial seeding, as severe overgrowth of the cells frequently occurred. Clearance of the Sendai virus from the cells was tested using the published primer sequences provided in the kit instructions. Positive controls were collected during the replating on day 5 and compared to the iPSCs collected at passage 6, 11, 16, and 21.
Optimized reprogramming conditions
Additional reprogramming focused on optimizing the protocol and was successful in generating iPSCs from seven additional NWR fibroblast cell lines, as well as two SWRs. These individuals' fibroblasts were plated on 0.1% gelatin in 50:50 medium at a range of approximate cell densities from 4,800 to 10,600 cells/cm2 instead of the initial protocol recommendation of 21,000–32,000 cells/cm2. This lower cell density is similar to published densities used for retroviral reprogramming in horse [22,27] and the previous retroviral reprogramming in endangered species [24], as well as the updated protocol from Thermo Fisher Scientific.
Wells were chosen at ∼50% confluency, the lower end of the initially recommended range of 50%–80% confluency. Cell density was too low to reliably count, so cell number was estimated based upon 36 h of growth (∼1.5 doublings) and infected with the recommended MOI of KOS-5, MYC-5, and KLF4-3. Additional wells were transduced with 1.5 times (7.5 - 7.5 - 4.5) and 2 times (10 - 10 - 6) the recommended MOI. See Table 1 for successful combinations of cell density and MOI for each rhinoceros.
Northern White Rhinoceros and Southern White Rhinoceros Fibroblast Cell Lines Used in These Experiments and the Resulting Induced Pluripotent Stem Cell Clones
Those individuals with estimated dates of birth (est) were wild caught. SD-WAP refers to the San Diego Zoo Safari Park, Escondido, CA. Zoo Dvůr Králové is in the Czech Republic (Štefánikova 1029, 544 01 Dvůr Králové nad Labem, Czechia). Chromosome complement was provided with the cell line from the Frozen Zoo® and is used for comparison to the derived induced pluripotent stem cell lines. Italicized clones were derived from alternate MOIs and/or initial plating density, also indicated by italics.
MOI, multiplicity of infection; NA, not applicable; NWR, northern white rhinoceros; SB, studbook; SD-WAP, San Diego Zoo Safari Park; SWR, southern white rhinoceros.
The medium was changed according to the protocol. In brief, fibroblast medium (50:50) was changed within 24 h post-transduction and every other day until replating. The cells were replated using TrypLE™ Select (Gibco) 1–2 days earlier than recommended. Cells for the feeder-free condition were replated on day 5 post-transduction at a density of 4,300 cells/cm2, and cells on MEFs were replated on day 6 post-transduction at a density of 360 cells/cm2. All MEF cultures were changed to KB Medium, and feeder-free cultures were changed to mTeSR1 (Stem Cell Technologies) 24 h later and subsequently fed every day.
Multiple clones were picked between days 16 and 23 post-transduction, with the most successful lines picked between day 16 and 19. Overall, clones picked from feeder layers were more successful, resulting in more clones and fewer losses to differentiation. All clones were expanded until at least passage 10 and then cryopreserved in 10% dimethyl sulfoxide (Sigma) and 90% KOSR using a CoolCell® LX control rate freezing container (BioCision) at −80°C for a minimum of 4 h before being transferred to nitrogen vapor holding.
The difference in growth rate of the initial fibroblast cells between the initial and optimized reprogramming efforts is most likely caused by the lack of 50:50 medium used during the initial reprogramming.
Spontaneous embryoid body-induced differentiation
Embryoid bodies [28] (EBs) were made by either passaging feeder-dependent colonies with a StemPro EZPassage™ (Thermo Fisher) tool or passaging feeder-free colonies with Accutase. One to three wells of a six-well plate, depending on density, were passaged into one well of an ultralow attachment six-well plate. These cells were fed with 50:50 medium every other day for 7 days and then transferred to a plate coated with 0.1% gelatin for seven more days to continue the differentiation. The cells were subsequently fixed for immunocytochemistry [ICC] or trypsinized, rinsed, and frozen at −80°C in Dulbecco's phosphate buffered saline (DPBS) for RNA extraction and analysis.
Immunocytochemistry
Cells were washed in DPBS, fixed in 4% paraformaldehyde for 10 min at room temperature, and blocked in 5% bovine serum albumin (BSA) and 0.3% Triton X-100 for 30 min at room temperature. Primary antibodies were applied according to the dilutions listed in Supplementary Table S2 and incubated at 4°C overnight.
Cells were then rinsed thrice in DPBS, and the secondary antibody was applied at a 1:1,000 dilution and incubated for 1 h at room temperature. Cells were washed three additional times with DPBS. Pluripotency and germ layer markers were counterstained with 4′,6-diamidino-2-phenylindole (Invitrogen P36962, ProLong™ Diamond antifade mountant). Images were acquired on an IN Cell Analyzer 6000 (GE Healthcare) or a Keyence BZ-X800E (Keyence Corporation of America).
Flow cytometry
EBs were collected by rinsing with DPBS and treated with 0.05% trypsin at 37°C for 5 min. After digestion, cells were collected and centrifuged at 0.3 relative centrifugal force for 5 min. The pellet was resuspended in flow buffer containing 1% bovine serum in DPBS and incubated for 15 min on ice with 1% fetal bovine serum in DPBS containing the following antibodies at a 1:100 dilution: PE anti-human TNAP Antibody (BioLegend 327305) and FITC anti-human CD38 Antibody (BioLegend 303503). Cells were then washed thrice with the flow buffer, and the resulting single cell suspension was passed through a 40 μm filter and analyzed using a NovoCyte cytometer. Seven-amino-actinomycin D Viability Staining Solution (Thermo Fisher Scientific 00-6993-50) was used as a marker of viability.
Karyotyping
Fibroblasts were harvested from one T25 flask for karyotyping after 0–30 min in 0.02 μg/mL colcemid (KaryoMAX®; Gibco), depending on the culture, by selectively trypsinizing (0.05% Trypsin-EDTA; Gibco) to remove 1/3–1/2 of the cells. Cells were pelleted and resuspended in 5 mL of 0.0675 M potassium chloride and incubated at 37°C for 25 min. Cells were fixed in five parts methanol to one part acetic acid (5:1 fix) and washed thrice by centrifugation. Fixed cell suspension was dropped onto prepared slides in a humidified chamber (42%) at 27°C. Spreads were stained with Giemsa for 7 min and then imaged and karyotyped using the CytoVision System (Leica Biosystems). All fibroblast lines were karyotyped after initial banking and again after expansion for reprogramming, providing a standard for comparing the iPSCs.
iPSCs were harvested from one well of a 6-well plate after 30–60 min in 0.02 μg/mL colcemid using Accutase for feeder-free conditions or 0.05% trypsin-EDTA for feeder conditions and treated the same as the fibroblasts. Any contamination from the feeder layer was easily identifiable as mouse chromosomes instead of NWR and eliminated from the analysis.
Cardiac differentiation
NWR iPSCs were differentiated into cardiomyocytes using a human kit (PSC Cardiomyocyte Differentiation Kit:Gibco), using the supplied protocol. In brief, feeder-free iPSCs were plated at a density of 6,400–7,900 cells/cm2 on Geltrex in mTeSR1. The use of Rho kinase (ROCK) inhibitors was not necessary for cell survival during plating. Successful wells achieved ∼20% confluency on the first day of differentiation, slightly less than the minimum of the range of 30%–90%. Medium was changed every other day, and cells began to contract ∼9 days after the start of differentiation.
RNA extraction
Cell pellets were obtained for RNA from feeder-dependent conditions using dispase (0.2 μg/mL; Gibco), allowing the colonies to settle, and rinsing twice in DPBS to remove as many MEFs as possible or from feeder-free conditions using Accutase. RNA was then extracted from these pellets using the mirVana™ miRNA Kit (Ambion; Invitrogen) according to the supplied protocol.
Primer design
Genes of interest (GOI) were identified from whole genome RNAseq data from NWR fibroblasts, iPSCs, and EBs (unpublished data) that were aligned to cow and dog genomes for annotation. The cow and dog were chosen based on the quality of their genome assembly and annotation, which is lacking in rhinoceros at this time. We then aligned them to the SWR genome (
Quantitative reverse transcriptase–polymerase chain reaction
RNA was DNase treated using ezDNase™ enzyme (Thermo Fisher) following the published protocol. cDNA was immediately synthesized using the iScript™ Select cDNA Synthesis Kit (Bio-Rad) with random primers following the published protocol and stored in TE buffer. Pluripotency markers were analyzed at a concentration of 0.3 ng cDNA, while germ layer and cardiac markers were analyzed at 3 ng. All polymerase chain reactions (PCRs) were carried out using the SsoAdvanced™ Universal SYBR® Green Supermix Kit (Bio-Rad) according to the published protocol with an annealing temperature of 60°C for 15 s and primers specifically designed for rhinoceros (Supplementary Table S3). Primer concentration was checked for all primers and optimized for ACTB to be 250 nM forward and 300 nM reverse, while all the others are used at 200 nM. To eliminate any nonspecific signal from primer-dimer formation, an additional 80°C read step for 5 s was added after annealing. By reading the signal at 80°C instead of 60°C, all smaller amplicons would have been denatured and would not contribute to expression levels. The temperature was chosen based on the melt curves of the GOI and was 1.5°C below the lowest observed GOI melt temperature. Primer specificity and efficiency were checked with a standard curve of mixed cell types and melt curve analysis. Each sample was run in triplicate, and the expression levels were normalized to ACTB (actin beta), which is an accepted housekeeping gene in horse [23,31] and human [32] iPSC analyses. Samples that did not amplify strongly enough to produce a melt curve were subsequently removed from the analysis.
Loci in which genomic amplification was observed in the no reverse transcriptase (NRT) reaction with a Cq value <5 cycles from the sample were corrected using the following formula [33]:
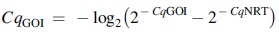
Results
Generation of white rhinoceros iPSCs
We generated iPSC lines from 11 rhinoceros individuals (nine NWRs and two SWRs) across four reprogramming experiments by modifying a Sendai virus reprogramming protocol designed for human cells (CytoTune 2.0, Thermo Fisher Scientific). We increased the efficiency of the human protocol by optimizing the growth medium, the viral titer, and the cell plating density. See Table 1 for the fibroblast cell line information and successful reprogramming conditions and Supplementary Table S4 for a summary of the clones obtained from each of the four reprogramming experiments and additional experimental information.
Because the rhinoceros cells were resistant to the viral toxicity that affects human cells, using the human protocol resulted in severe cell overgrowth during reprogramming. We reduced the plating density and increased the viral concentration to 1.5–2 times the recommended starting MOI. See Supplementary Table S2 for the details for each experiment. Under the modified conditions, we were able to obtain iPSC clones from all but one of the fibroblast lines. The only fibroblast line that was not successfully reprogrammed was one established in 1979 (Lucy KB 3731).
iPSC clones arose in populations cultured on mitotically inactivated MEF feeder layers or Geltrex (Gibco), but only the feeder layer cultures produced stable pluripotent cell lines. Feeder layer-free clones either spontaneously differentiated or were only partially reprogrammed (Supplementary Fig. S2). However, once the clones stabilized, it was possible to adapt them to feeder-free growth conditions on Geltrex in mTeSR1 medium (Stem Cell Technologies). The rhinoceros iPSC colonies had morphology that is distinctive and consistent with our previous observations [24]. The iPSC colonies were flat, like human PSC colonies, but were more irregular in shape (Fig. 1).
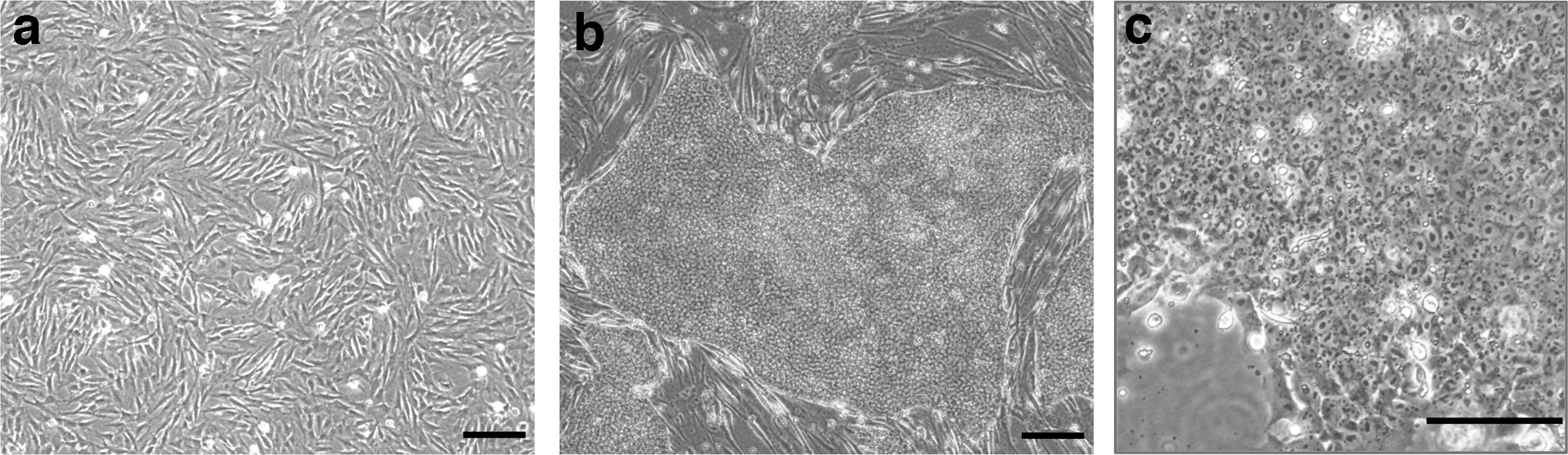
Cell morphology of rhinoceros cells
We explored KLF4 MOI optimization and the addition of small molecules (sodium butyrate, a histone deacetylase inhibitor [34], and SB431542 [35], a transforming growth factor beta inhibitor), but did not see consistent results. SWR iPSC line 21409-c550 (Wallis) was reprogrammed with the use of SB431542 from day 8 of reprogramming until 1 day after the clone was picked. Reducing the MOI of KLF4 to 1.5 or 0.75 was also successful for NWR clones 17626-c5101 and 17626-c5103 (Fatu) and SWR 21409-c550 (Wallis).
Individuals Angalifu (KB 9947) and Fatu (KB 17626) have been reprogrammed at least twice, with Angalifu's fibroblasts producing three additional, as yet uncharacterized, clones during the fourth reprogramming experiment. We have also been successful in cryopreserving the remaining transduced fibroblasts during the replating step, thawing at a later time and obtaining additional clones [36] (Saut 9939-c5101 and three additional clones for Nasima 8174, not yet characterized at the time of submission).
The rhinoceros iPSCs were maintained in feeder and feeder-free conditions in the presence of bFGF, similar to human iPSCs. However, unlike human PSCs [37], the rhinoceros iPSCs could be passaged as single cells without the addition of Rho-associated coiled-coil protein kinase (ROCK) inhibitors. We were able to maintain a rhinoceros iPSC clone in feeder-free conditions for 53 passages and still successfully differentiate it into cardiac lineage cells, indicating that long-term culture did not affect the differentiation potential of this iPSC line.
While Sendai vector does not integrate into the genome, it can be retained in cells through multiple passages. We found that varying the MOI led to more variable retention times (Supplementary Fig. S3b). The exogenous Sendai virus cassette sequences were undetectable in the cells of Angalifu (9947-c501) by passage 11 when the recommended MOI for human cells of 5:5:3 was used (Supplementary Fig. S3a). But two clones obtained from the lower KLF4 MOI of 1.5 (Fatu-c5101 and c5103) still showed expression of the KOS and c-MYC cassettes at passage 20 and passage 14, respectively, while the clone obtained with the 0.75 KLF4 MOI (Wallis-c550) showed complete clearance by passage 12.
Of note, the PCR primers used for detecting viral sequences showed a high amount of nonspecific amplification in nontransduced rhinoceros fibroblasts, including a nonspecific peak at 500 bp in the KOS cassette, multiple bands in the KLF4 cassette, and a faint band at 500 bp in the MYC cassette (Supplementary Fig. S3a–c). Following a protocol supplied for the CytoTune 2.0 Kit, we shifted several clones that retained viral sequence (Fatu-c5101, Fatu-c5103, and Suni-c508) to 38.5°C for 5 days, which resulted in complete clearance of the reprogramming cassettes (Supplementary Fig. S3c).
Many of our samples continued to indicate the presence of the Sendai backbone (SeV) by RT-PCR without expression of the reprogramming cassettes. To confirm that the virus was no longer active in these cells we performed ICC to detect the Sendai virus, which indicated that only cells that were positive for a reprogramming cassette (KOS, KLF4, or MYC) were positive for the virus. Samples that were positive for Sendai backbone only by RT-PCR were negative for protein expression from the virus (Supplementary Fig. S4).
To determine whether the reprogramming affected the chromosome complement of the cell lines, 20–40 metaphase spreads for each clone were counted and a random male and female iPSC line was karyotyped (Supplementary Table S5 and Supplementary Fig. S5).
Inherent variation within chromosome morphology of the p-arm is common in both white rhinoceros subspecies [25], and length is influenced by the time of colcemid exposure. Due to the high chromosome number (82) and small chromosome size, karyotyping does not definitively reveal any but the most gross chromosomal changes within the cell lines. Previous work shows examples of fibroblast karyotypes and a discussion of the Robertsonian translocation that is present in three of the NWR fibroblast lines [25]. After reference genomes are established for these species, we will use genome resequencing to assess the stability of the lines at much higher resolution.
Pluripotency of rhinoceros iPSCs
Unlike iPSC lines from other species, predominantly domestic species [38], the rhinoceros iPSCs did not require continued expression of exogenous reprogramming factors to maintain pluripotency. ICC of Angalifu 9947-c501 or Najin 8173-c505 feeder-free iPSCs confirmed positive expression of canonical markers of pluripotency [OCT4 (also known as POU5F1), SOX2, NANOG, and LIN28a], which have been previously shown as reliable markers in rhinoceros [24,39] or identified in this study (Fig. 2: see Supplementary Table S2 for all antibodies tested and Supplementary Fig. S6 for secondary antibody controls). The rhinoceros cells also bound the lectin, UEA1, that labels PSCs from multiple species [40]. In addition, we found that the pluripotency-associated human cell surface markers SSEA3 and SSEA4 were detectable on rhinoceros iPSCs.
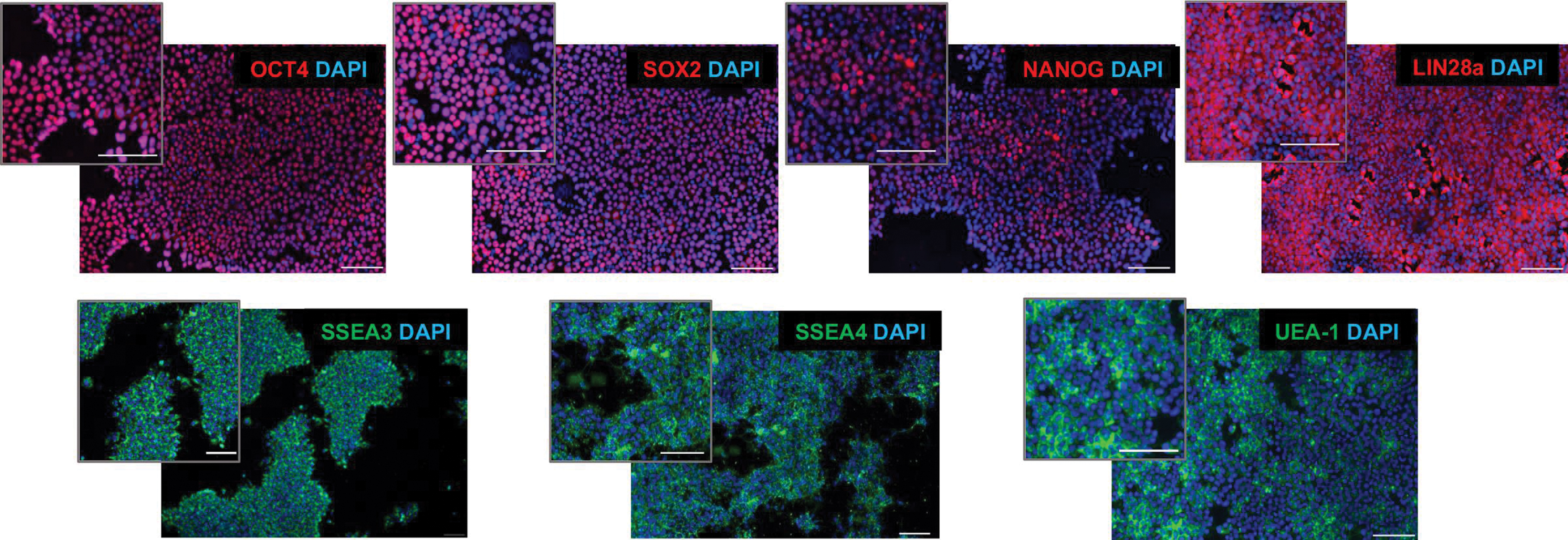
ICC for pluripotency-associated markers in NWR iPSCs. Nuclear or cytoplasmic markers are shown in the top row, and cell surface markers are indicated in the second row. All nuclei are counterstained with DAPI. All images were taken of feeder-free cell lines. Scale bars = 100 μm. DAPI, 4′,6-diamidino-2-phenylindole; ICC, immunocytochemistry. Color images are available online.
We also analyzed all iPSC lines using rhinoceros sequence-specific quantitative reverse transcriptase PCR (qRT-PCR) primers for genes associated with pluripotency (Fig. 3: see Supplementary Table S3 for a list of primers used in these experiments). As has been noted with other species [41], we saw variation in pluripotency-associated markers in NWR compared to mouse or human. NANOG, which is intermittently expressed in mouse and human PSCs [38 –40], was highly expressed in all the iPSC lines analyzed. As observed in our earlier work [24], KLF4 was expressed at low level by rhinoceros iPSCs. PRDM14 and SALL4 showed consistently high expression and may be fulfilling the role of KLF in the rhinoceros pluripotency network (for a review of KLF4 in pluripotency see Bialkowska 2017 [42]). LIN28a was expressed at high levels and is also highly expressed in PSCs from the related perissodactyl, domestic horse [43]. However, TERT and ZFP42 (also known as REX1) are expressed in the horse [43], but were undetectable in the rhinoceros iPSCs.
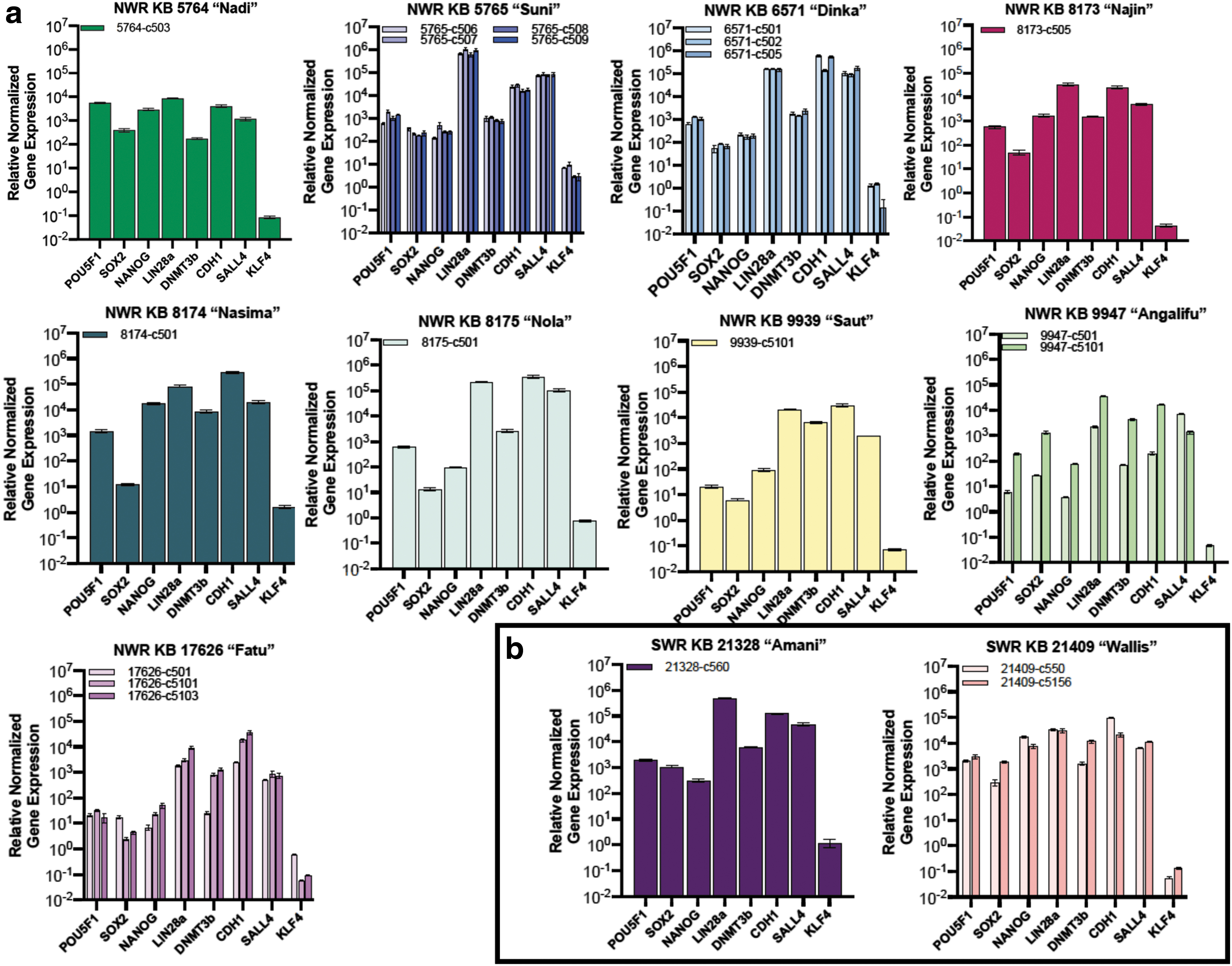
qRT-PCR analyses with rhino-specific primers for all iPSC clones confirmed that CDH1, DNMT3b, LIN28a, NANOG, POU5F1, SOX2, and SALL4 are expressed by rhinoceros iPSCs. Gene expression was normalized to ACTB and controlled to the source fibroblasts.
Differentiation of rhinoceros iPSCs
The ability of pluripotent cells to differentiate into cells of all three germ layers is a functional indicator of pluripotency. The rhinoceros iPSCs spontaneously differentiated in EBs (Fig. 4) and expressed markers for each of the three germ layers, identified by ICC (Fig. 5) and qRT-PCR (Fig. 6).
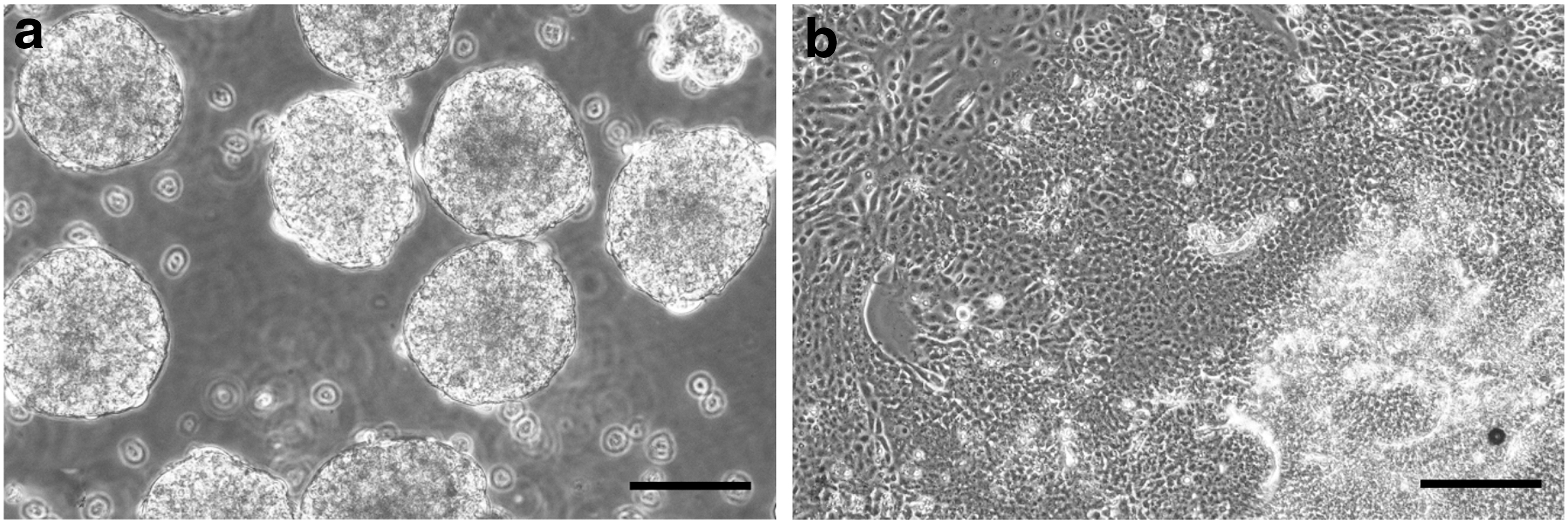
NWR embryoid bodies.
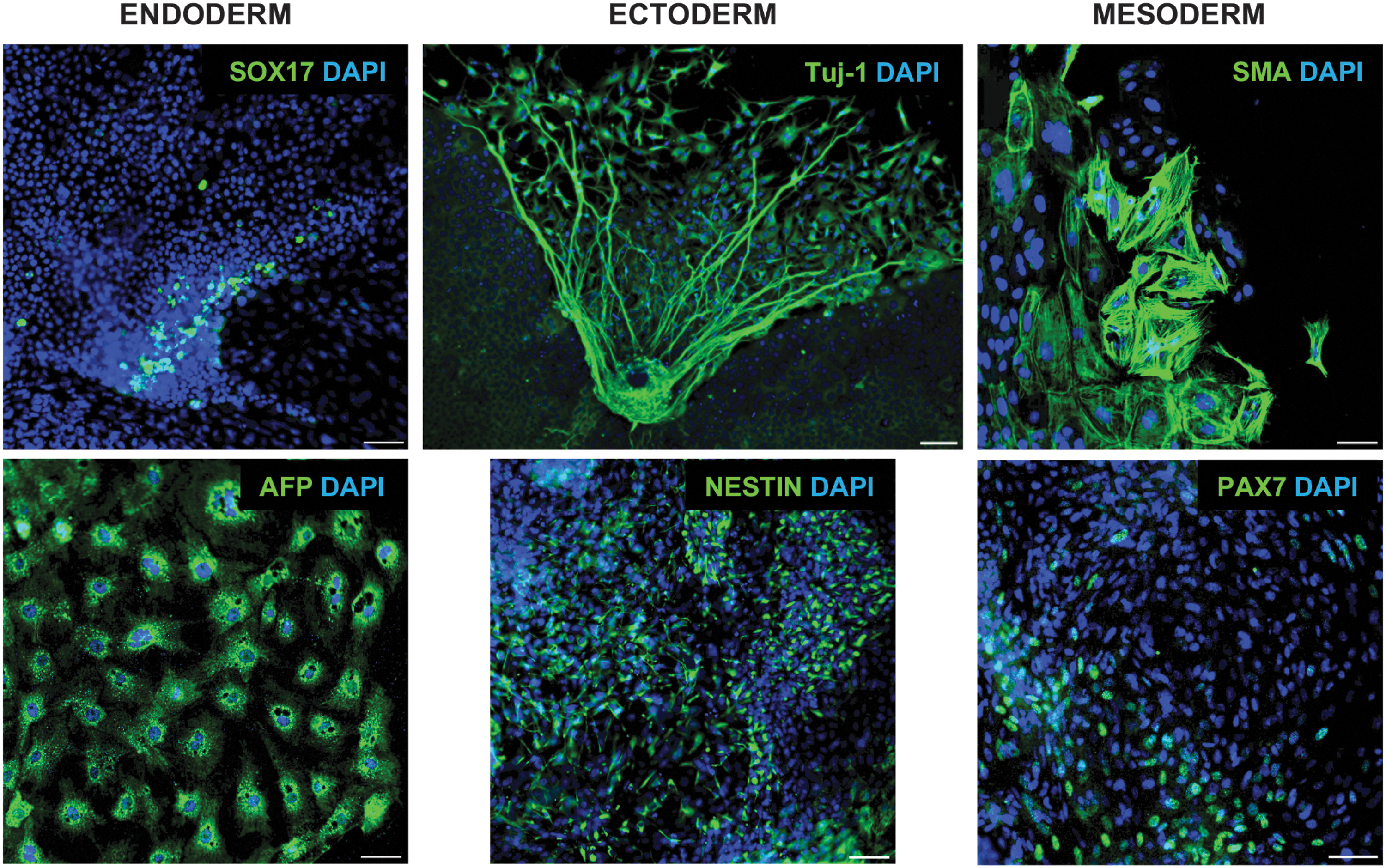
Three-germ-layer differentiation potential was confirmed using known human ICC markers for endoderm, ectoderm, and mesoderm in cells from NWR EBs that had differentiated for 2 weeks. All nuclei are counterstained with DAPI. Scale bar = 100 μm. Color images are available online.
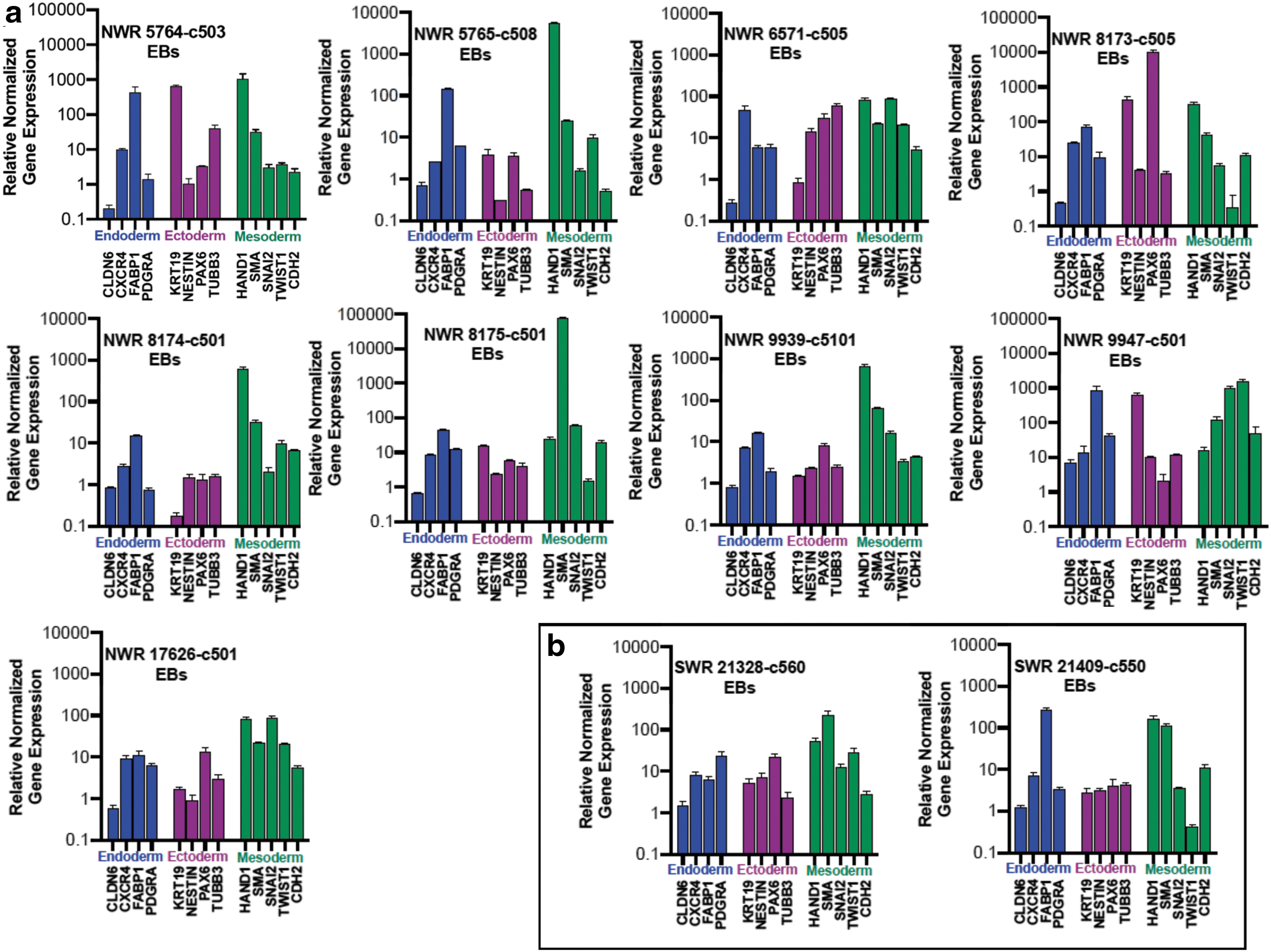
EB-induced differentiation was confirmed with qRT-PCR for rhinoceros-specific primers for multiple markers for each germ layer.
In addition, we demonstrate that two NWR iPSC clones readily generated beating cardiomyocytes using a human differentiation protocol. These findings suggest that the cardiac differentiation pathway is conserved between humans and rhinoceros. See Fig. 7 for cardiac confirmation through ICC and qRT-PCR and Supplementary Video S1.
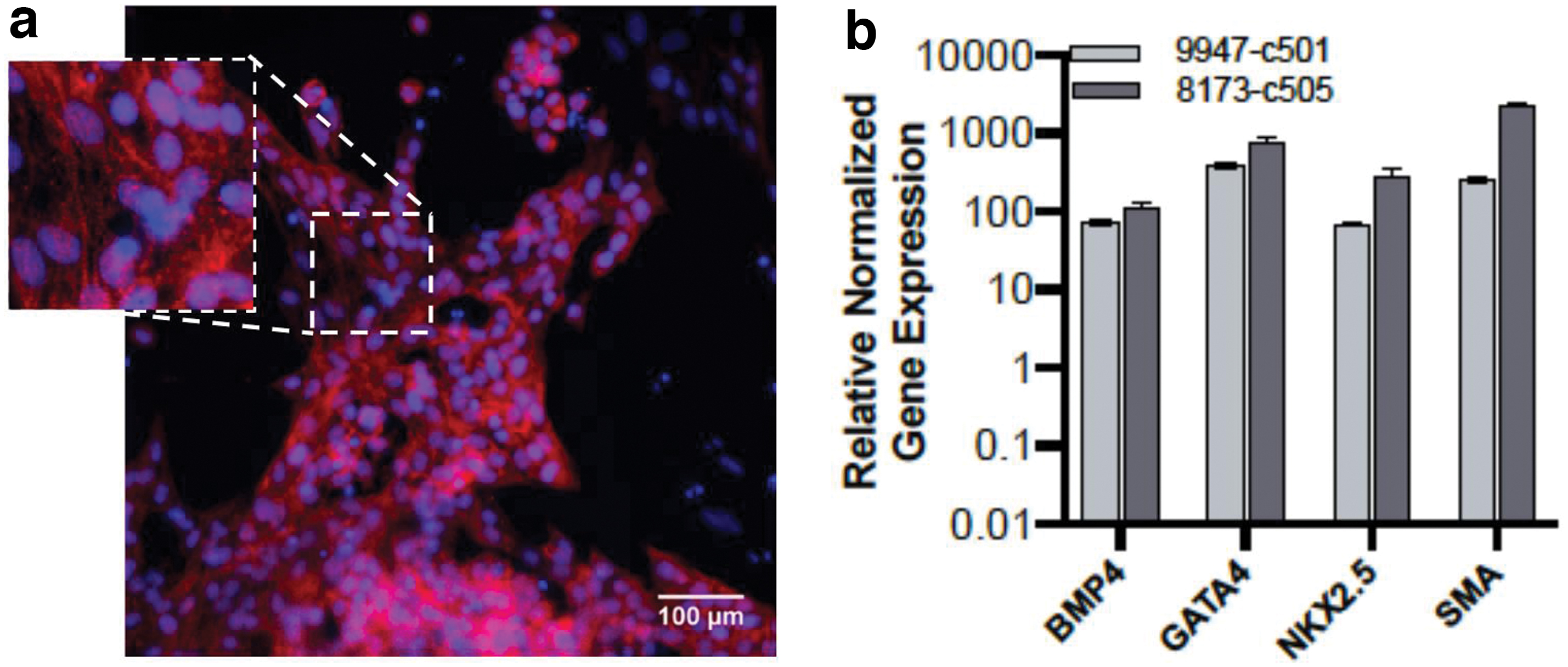
Differentiation of NWR iPSCs into cardiomyocytes.
Since our preferred approach for use of the rhinoceros iPSCs is genetic rescue through the generation of gametes from NWR iPSCs, we wanted to determine whether they can differentiate into primordial germ cells (PGCs), the precursors of mature oocytes and sperm. We analyzed the spontaneously differentiated EBs derived from male and female NWR and SWR iPSCs for expression of early markers of the PGC specification network. The rhinoceros cells showed expression of several genes associated with early PGC competency and development: BMP4 (bone morphogenic protein 4), PRDM1 (PR/SET domain 1, also known as BLIMP1), PRDM14 (PR/SET domain 14, also known as PFM11), and TFAP2C (transcription factor AP-2 gamma) (Fig. 8).
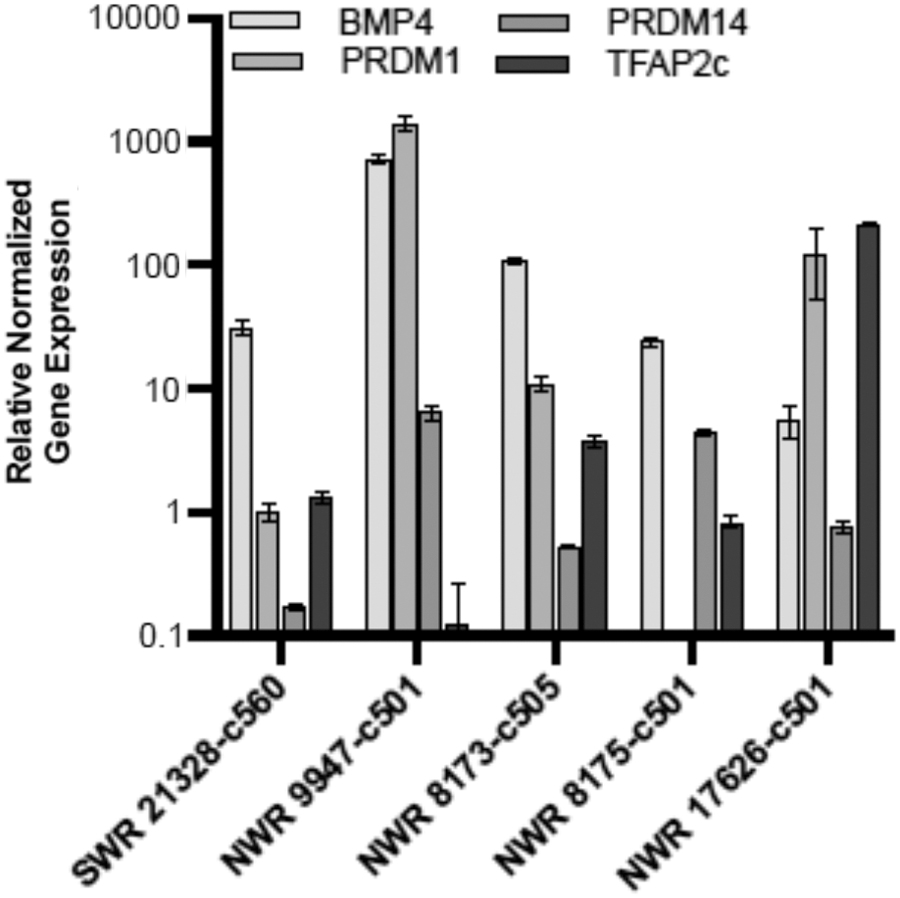
qRT-PCR expression of markers associated with PGC development in EB-derived populations from one SWR and four NWRs. Gene expression was normalized to ACTB and controlled to the source cell type, iPSCs. Error bars indicate the standard deviation of the triplicate Cq values. PGC, primordial germ cell.
We also detected colocalized PDPN (podoplanin) and SOX17 (SRY-box 17) by ICC in a small number of cells within our EB cultures (Fig. 9). PDPN is reported to be a conserved marker of PGCs in cynomolgus monkey and humans [44], while SOX17 is an early pathway catalyst in human PGC development [45].
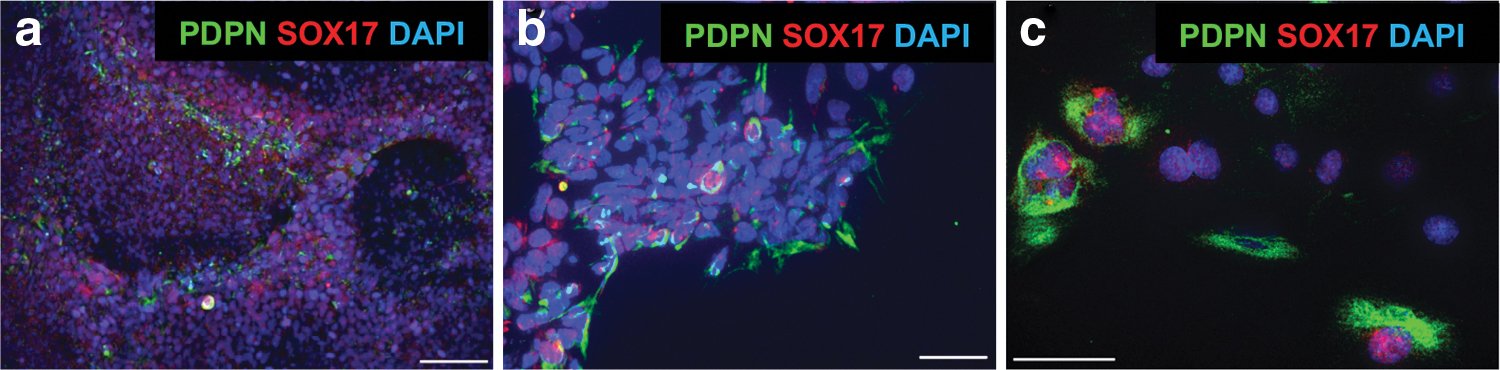
Immunocytochemistry of spontaneous differentiation of cells with PGC-associated markers from NWR EBs, detected with human (SOX17) and conserved human/cynomolgus monkey (PDPN) markers.
Discussion
Many species are on the brink of extinction due to the activities of humans. Notable is the mammalian Family Rhinocerotidae, of which all five species are endangered. The NWRs are of particular concern because with only two individuals remaining in 2020, this unique form of rhinoceros is functionally extinct. It is too late for the NWR to be saved by conventional conservation efforts, and the only hope for the survival of this species is introduction of new, unconventional experimental strategies. One of these strategies is to take advantage of the virtually unlimited differentiation capacity of PSCs using them to generate gametes that could be incorporated into assisted reproduction technologies.
We modified existing methods to generate iPSCs from NWR fibroblasts stored in the Frozen Zoo biorepository at the San Diego Zoo Institute for Conservation Research. Among the thousands of cryopreserved specimens in the Frozen Zoo are skin fibroblasts cultured from 12 different NWR individuals. In 2011 we reported the first generation of iPSCs from any endangered species, including iPSCs from one NWR, Fatu; those iPSCs were generated using the available methods at the time, integrating lentiviral vectors carrying the human reprogramming factors [24].
Fatu's iPSCs are not suitable for reproductive use because integration of transgenes disrupts the genome and reactivation of integrated reprogramming factors has been shown to lead to tumors in mice. When nonintegrating RNA virus vectors became available several years ago, we began to adapt the methods developed for human cells for use in NWR fibroblasts.
There were several challenges in adapting the human protocols for generating and characterizing iPSCs to the rhinoceros, including adjustment of cell density and identification of antibodies that cross-react with the rhinoceros homologs of pluripotency and lineage-associated proteins. Development of primers for qRT-PCR was hampered by not having a high quality, annotated NWR reference genome.
Our preliminary results for four NWR lines were reported as a preprint in 2017 [46]. Now we report optimized methods for generation and characterization of iPSCs from fibroblasts of nine NWRs and two SWRs that had been stored in liquid nitrogen for as long as 40 years. The 9 iPSC lines capture a genetic diversity greater than the two living animals and banked gametes. This is the first step to genetic rescue of a species through the use of reprogramming technology.
We are aware that mutations arising from cell culture in the iPSCs could compromise their value as germ cells. It is unlikely that the reprogramming itself would have introduced deleterious mutations [47], but since PSCs have unlimited proliferative ability, harmful mutations and copy number variants can arise over time in culture [48 –50]. A comprehensive genomic reference sequence of the NWR and SWR is in progress and will be used to compare with whole genome sequencing of the iPSC lines.
The long-term goal of our multidisciplinary project is to generate gametes from NWR iPSCs that can be used in assisted reproductive technologies that are currently being implemented for rhinoceros by other members of our multidisciplinary team [51,52]. There has been steady progress in development of gametes from iPSCs of other species. Human PGCs, the precursors of sperm and eggs, can spontaneously develop during EB differentiation [53], and our preliminary analyses suggest that our iPSCs produce a small number of cells with characteristics of PGCs. Recent research has reported the generation of functional gametes from iPSCs in mouse [20], and key PGC developmental pathways have been identified in human [21,53,54] and cynomolgus monkey [44]. Recently, transcription factor-driven oocyte development from PSCs has been reported [55].
The reprogramming and characterization strategy reported here should serve as a template for iPSC generation of additional endangered species. We see the potential for use in other rhinoceros species, as well as in other groups such as endangered cats and black-footed ferrets, and for research in veterinary medicine. By generating iPSCs, we have created a renewable resource for research and the potential recovery of endangered species. Furthermore, the information we and colleagues are obtaining about all aspects of rhinoceros reproduction [56,57] will aid conservation of other species.
Footnotes
Acknowledgments
The authors thank all members of the Frozen Zoo team (especially Arlene Kumamoto, Marlys Houck, Julie Fronczek, and Suellen Charter) for biobanking the fibroblast cells, karyotyping, and for cell culture recommendations and Tate Tunstall and Cynthia Steiner for bioinformatics support. The authors thank members of the Loring Lab (including Ronald Coleman, Roy Williams, Candace Lynch, Ha Tran, Andres Bratt-Leal, Eveline Lee, Ai Zhang, and Jason Stein) and Björn Brändl and Franz-Josef Müller for advice and helpful discussions. The authors are grateful to the San Diego Zoo Global and San Diego Zoo Wildlife Conservancy donors, including Anne and Christopher Lewis, and the Robert Kleberg and John and Beverly Stauffer Foundations for funding and to Uma Lakshmipathy from Thermo Fisher Scientific for generously providing supplies. The authors thank Genea BioCells (La Jolla, CA) and Keyence Corporation of America for the use of their imaging systems. The work would not be possible without the prescience and vision of Dr. Kurt Benirschke, who founded the Frozen Zoo in 1975.
Data Availability Statement
The data supporting the qRT-PCR findings are available from the corresponding author upon reasonable request. All other data are accessible from the article or supplementary materials.
Author Disclosure Statement
M.L.K., S.M.F., T.D.N., I.V.A., S.E.P., and O.A.R. have no disclosures. J.F.L. is a founder and has stock options in Aspen Neuroscience. C.G.P. has stock options in Aspen Neuroscience.
Funding Information
Project was supported by San Diego Zoo Global and donations to San Diego Zoo Global Wildlife Conservancy in support of the Northern White Rhino Initiative. We gratefully acknowledge funding from the Seaver Institute, the Robert and Helen Kleberg Foundation, and the Alice C. Tyler Perpetual Trust. We thank the California Institute for Regenerative Medicine (CIRM) for providing laboratory training for TDN and CGP (CL1-00502).
Supplementary Material
Supplementary Figure S1
Supplementary Figure S2
Supplementary Figure S3
Supplementary Figure S4
Supplementary Figure S5
Supplementary Figure S6
Supplementary Table S1
Supplementary Table S2
Supplementary Table S3
Supplementary Table S4
Supplementary Table S5
Supplementary Video S1
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
