Abstract

M
EpCAM++CD49flow/− (luminal) cells were isolated by fluorescent-activated cell sorting (FACS) from normal virgin adult female C57Bl/6J mouse mammary glands (Fig. 1A), transduced at 40% efficiency with a library of DNA-barcoded green fluorescent protein-encoding lentiviruses, and 1.3 × 105 cells injected with Matrigel into a single cleared fat pad in each of two mice (Fig. 1B) as previously described for mouse basal cell transplants [4]. The purity of the transplanted cells determined by reanalyzing an aliquot of the sorted cells showed >98% still displayed a luminal phenotype, with a <0.2% chance of being contaminating basal (EpCAMlowCD49f+) cells that could subsequently contribute to clone formation.
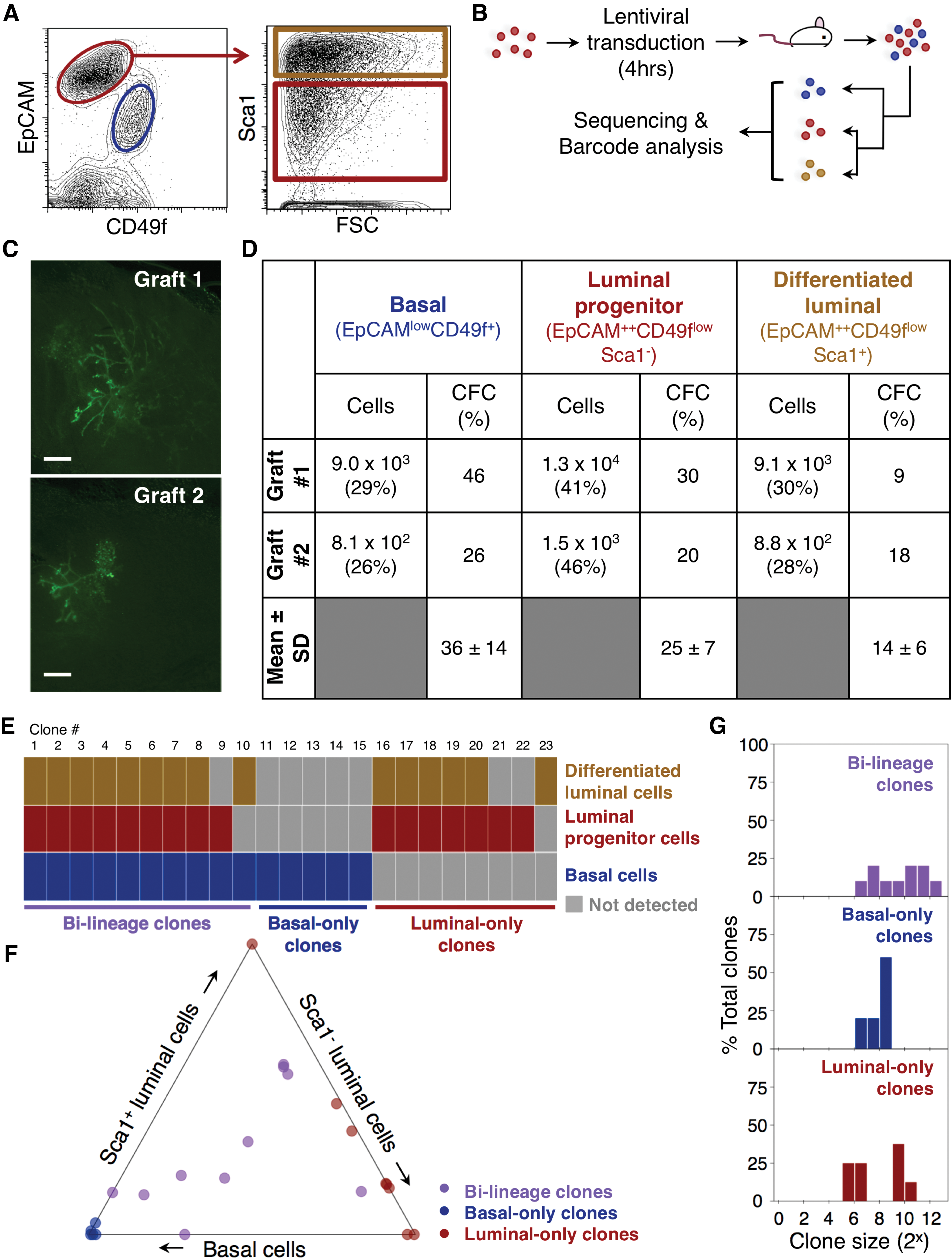
After 8 weeks, whole-mount microscopy revealed complete branched GFP+ structures had been regenerated in both mice (Fig. 1C). Isolation of the EpCAMlowCD49f+ basal cells, EpCAM++CD49flow/−Sca1− luminal progenitor cells, and EpCAM++CD49flow/−Sca1+ terminally differentiated luminal cells from the enzymatically dissociated fat pads (Fig. 1A) revealed that these three subsets were present in similar proportions in both fat pads (>2-fold more Sca1− and Sca1+ luminal cells than basal cells, with the Sca1− luminal progenitor fraction more prominent; Fig. 1D), despite a 10-fold difference in the total number of mammary cells recovered from each fat pad (3.1 × 104 vs. 3.2 × 103). A normal adult mammary gland would have a higher proportion of differentiated luminal cells compared with luminal progenitor cells [2], suggesting these regenerated glands may be less mature.
The colony-forming cell (CFC) content in each harvested subset was also largely restricted to the basal and luminal progenitor fractions (frequencies of ∼36% and ∼25%, respectively), compared with the EpCAM++CD49flow/−Sca1+ cells (∼14%; Fig. 1D). Thus, despite large differences in the overall size of the glands generated from the transplanted cells, their progeny distributions were similar to one another, and also to those found in the normal adult mouse mammary gland [3].
DNA sequencing and barcode analysis [4] detected 14 and 9 clones in the 2 regenerated structures (Fig. 1E). FACS isolation of the different phenotypes in each structure before DNA extraction from them showed that 10 clones (7 and 3 clones in the 2 regenerated structures) were bilineage and had a mean size of ∼2 × 103 cells each. Five clones contained only basal cells and eight only luminal cells (Fig. 1F) with smaller mean sizes of ∼300 and ∼500 cells per clone, respectively; Fig. 1G).
These findings illustrate the power of cellular barcoding to analyze the clonal outputs of cells with limited growth potential—here applied to rare (∼10−4) transplantable cells isolated from the luminal compartment of the normal adult mouse mammary gland.
Footnotes
Acknowledgments
The authors thank G. Edin, M. Hale, and D. Wilkinson for excellent technical support; D. Ko and W. Xu (Flow Cytometry Facility, Terry Fox Laboratory) for assistance with flow cytometry; and T. MacDonald for assistance with rodent husbandry. This work was supported by grants from the Canadian Cancer Society (grant no. 702851) to C.J.E., a CIHR Vanier Canada Graduate Scholarship to L.V.N., a Canadian Institute of Health Research Banting and Best Graduate Fellowship to M.M., and a University of British Columbia Graduate Fellowship to S.B.
Author Disclosure Statement
The authors have no competing interests to disclose.
