Abstract
Epithelial–mesenchymal transition (EMT) has been thought to occur during early embryogenesis, and also the differentiation process of human embryonic stem (hES) cells. Spontaneous differentiation is sometimes observed at the peripheral of the hES cell colonies in conventional culture conditions, indicating that EMT occurs in hES cell culture. However, the triggering mechanism of EMT is not yet fully understood. The balance between self-renewal and differentiation of human pluripotent stem (hPS) cells is controlled by various signal pathways, including the fibroblast growth factor (FGF)-2. However, FGF-2 has a complex role for self-renewal of hES cells. FGF-2 activates phosphatidylinositol-3 kinase/AKT, mitogen-activated protein kinase/extracellular signal-regulated kinase-1/2 kinase, and also protein kinase C (PKC). Here, we showed that a PKC rapidly induced an early growth response protein-1 (EGR-1) in hES cells, which was followed by upregulation of EMT-related genes. Before the induction of EMT-related genes, EGR-1 was translocated into the nucleus, and then bound directly to the promoter region of SNAIL, which is a master regulator of EMT. SNAIL expression was attenuated by knockdown of EGR-1, but upregulated by ectopic expression of EGR-1. EGR-1 as the downstream signal of PKC might play a key role in EMT initiation during early differentiation of hES cells. This study would lead to a more robust understanding of the mechanisms underlying the balance between self-renewal and initiation of differentiation in hPS cells.
Introduction
T
In early embryogenesis, the EMT plays crucial roles in the differentiation of multiple tissues and organs and the formation of a body plan [31]. During the EMT process, E-cadherin is downregulated. Several transcription factors have been implicated in this repression, including SNAIL, SLUG, and TWIST1 [32 –37]. Actually, in hPS cell culture, spontaneous differentiation is sometimes observed with the expression of SNAIL and VIMENTIN at the peripheral of the hPS cell colonies in conventional culture conditions, indicating that EMT occurs in hPS cell culture [38 –41].
In this study, we investigated molecules involved in the cascade of PKC-induced EMT in hES cells using the minimal growth factor-defined culture medium hESF9, which includes FGF-2 as the sole growth factor to remove the effects of other growth factors on EMT [17]. The results showed that the early growth response protein-1 (EGR-1; also known as NGFI-A, KROX-24, ZIF268, or TIS8) was induced by a PKC activator, PMA, and bound to the regulatory region of SNAIL, resulting in the upregulation of SNAIL and EMT-related genes. PMA-induced SNAIL expression was attenuated by knockdown of EGR-1, whereas ectopic expression of EGR-1 induced EMT-related genes expression. These results indicated that a downstream effector of PKC signaling, EGR-1, contributed to the induction of EMT in hES cell differentiation. This study would lead to a more robust understanding of the mechanisms underlying the balance between self-renewal and initiation of differentiation in hPS cells.
Materials and Methods
Cell culture
The hES cell line, H9 [19,42] (WA09, WISC Bank; WiCell Research Institute), was routinely maintained as previously described [19]. For the experiment, the cells were seeded on a six-well plate (BD Falcon) coated with bovine fibronectin (FN; Sigma; 2 μg/cm2) in the hESF9 medium [17] consisting of the ESF basal medium (CSTI) [43] without 4-(2-hydroxyethyl)-1-piperazineethanesulfonic acid supplemented with
Immunocytochemistry
Immunocytochemistry was performed as previously described [4]. The image analysis was performed by IN Cell Analyzer 2000 and IN Cell Developer Toolbox software (GE Healthcare). The primary and secondary antibodies used are listed in Supplementary Table S1 (Supplementary Data are available online at
Real-time quantitative reverse transcription–polymerase chain reaction
Real-time quantitative reverse transcription–polymerase chain reaction (qRT-PCR) and real-time quantitative PCR (qPCR) were performed based on the SYBR Green gene expression technology in a 7300 Real Time PCR System (Applied Biosystems), according to the manufacturer's instructions. Specific primers used are listed in Supplementary Table S2.
DNA microarray
DNA microarray analysis was performed using the whole human genome DNA microarray 4x44K kit (ver.2.0) and a microarray scanner G2565BA (Agilent) according to the manufacturer's instructions (Agilent). The signal intensity data produced for each of the spots were analyzed using feature extraction (Agilent) and GeneSpring GX software (Agilent).
Chromatin immunoprecipitation assay
Chromatin immunoprecipitation (ChIP) assay was performed using the ChIP-IT Express kit (Active Motif) according to the manufacturer's instructions. Chromatin was precipitated with EGR-1 antibodies (Cell Signaling Technology) or H3K9ac antibodies (MAB Institute). The immunoprecipitated DNA samples were analyzed by qPCR. The SNAIL promoter was amplified with the primer pairs listed in Supplementary Table S2.
Construction of EGR-1 expression vector
The EGR-1 expression vector was constructed as follows. The EGR1-2A-eGFP fragment coding EGR-1 (NM_001964.2), a self-cleaving 2A peptide [44], and the enhanced green fluorescent protein (eGFP) were synthesized by the GeneArt gene synthesis service (Life Technologies). The synthesized fragment was inserted into the XhoI and NotI sites of the episomal pEBmulti-Hyg plasmid (Wako Pure Chemical Industries) to create the pEGR1-2A-eGFP plasmid. To generate the peGFP control vector, the pEGR1-2A-eGFP plasmid was digested with XhoI and SalI restriction enzymes, and the EGR1 fragment was removed from the pEGR1-2A-eGFP plasmid. The plasmids were verified by sequencing.
Transfection
Before transfection, the hES cells were incubated with ROCK inhibitor Y-27632 (10 μM) for 1 h and dissociated into single cells by a cell dissociation solution, TrypLE™ Select (Life Technologies). Pellets of 1×106 cells were mixed with 2 μg of the EGR-1 expression plasmid (pEB-EGR1-2A-eGFP) or control plasmid (peGFP) in 100 μL of Neon R buffer solution (Life Technologies). The cell suspension was transferred to a cuvette and electroporated using a Neon Transfection system (Life Technologies) with program (voltage 1,050, width 30, and pulses 2) according to the manufacturer's protocol. The electroporated cells were plated onto a six-well plate coated with FN in the hESF9 medium supplemented with ROCK inhibitor (10 μM). The eGFP-positive cells were selected on hygromycin B (200 μg/mL) for 6 days in the hESF9 medium.
Transfections with siRNA
Transfections with siRNA targeting human EGR-1 (SMARTpool ON-TARGETplus, L-006526-00) or nontargeting control siRNA (ON-TARGETplus Non-targeting Pool, D-001810-10) were performed using Dharmafect1 (Dharmacon) as previously described [4]. Total RNAs or proteins were extracted for analysis 72 h after the fast transfection.
Western blot analyses
Western blot analyses were performed as described previously [4]. The protein was separated by 12.5% sodium dodeyl sulfate–polyacrylamide gel electrophoresis and transferred to polyvinylidene fluoride membranes (Millipore). The membranes were reacted with primary antibodies, peroxidase-conjugated secondary antibodies, and ECL Plus reagent (GE Healthcare). Protein bands were visualized using the LAS-4000 imager (Fujifilm). The primary antibodies used are listed in the Supporting Information Table S2.
Cell imaging analysis
The images of eGFP-positive cells in culture were captured in a cell imaging system, BioStation CT (Nikon Instruments, Inc.) at 37°C 10% CO2. The images were analyzed by a software CL-Quant (Nikon Instruments, Inc.).
Results
EMT induction of hES cells by PMA
To confirm whether a PKC activator, PMA, induces EMT in hES cells, PMA was added into the culture of H9 hES cells grown in the defined culture conditions. The colony morphology of H9 cells was changed within 24 h after PMA addition. Compared with the control (Fig. 1A[a]), the packed colonies of undifferentiated H9 hES cells came loose and flatter cells scattered out (Fig. 1A[b]). PMA addition together with the PKC inhibitor GFX did not change the colony morphology of H9 hES cells (Fig. 1A[c]). The scattered cells were positive for SSEA-1 and VIMENTIN, and negative for NANOG or E-cadherin, and vice versa for the cells treated with PMA together with GFX (Fig. 1A[d–i]). These results indicated that the activation of PKC induced the EMT process.
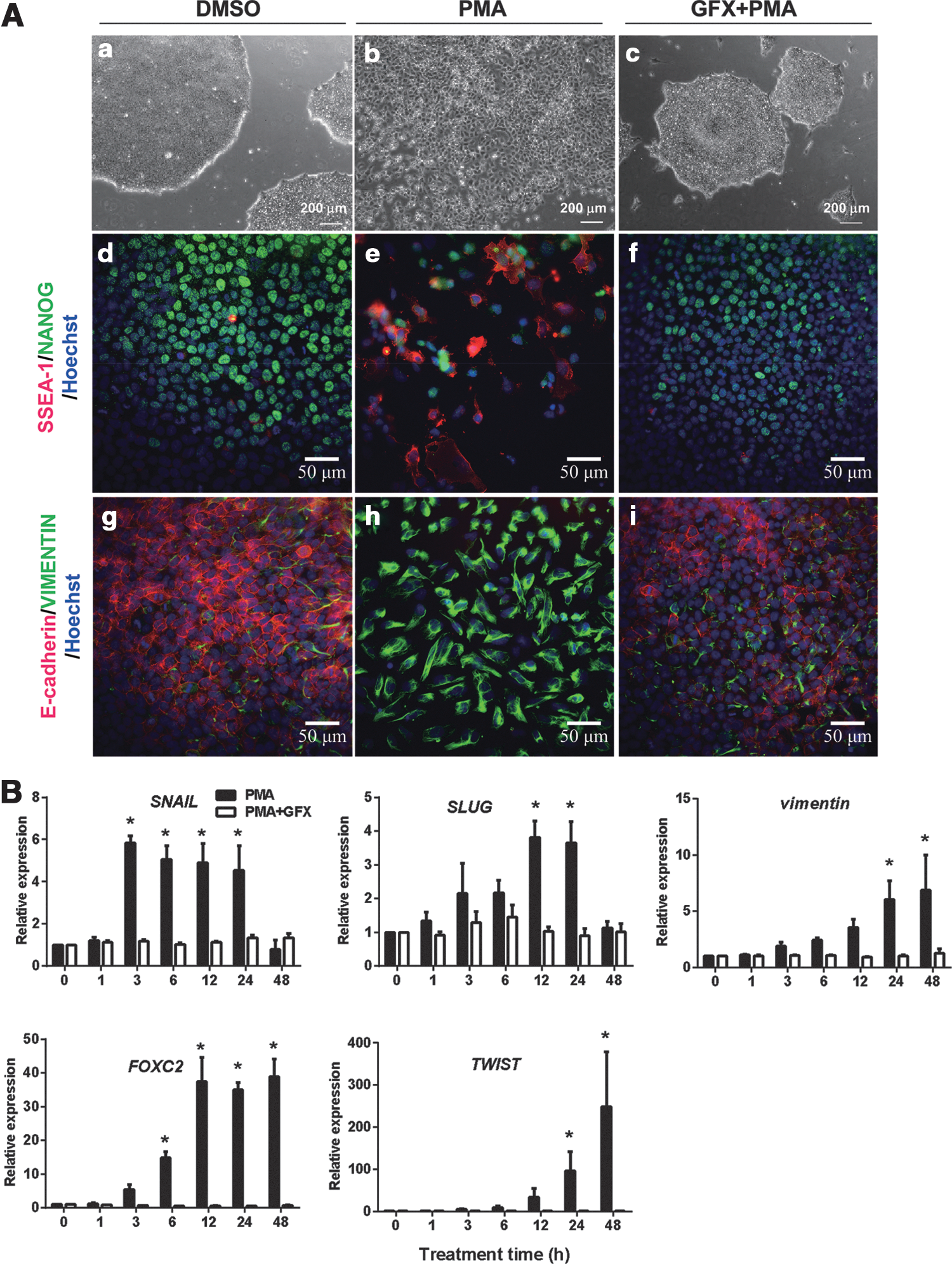
Phorbol 12-myristate 13-acetate (PMA)-induced epithelial–mesenchymal transition (EMT) process in human embryonic stem (hES) cells.
To determine whether PMA promotes the expression of EMT-related genes or lineage-specific cell differentiation-associated genes, H9 hES cells were cultured in the presence of PMA with or without GFX for 48 h, and then for 4 more days after changing to a fresh medium without PMA or GFX (Supplementary Fig. S1A). qRT-PCR analysis showed that the expression of SNAIL rapidly increased 3 h after PMA treatment (Fig. 1B). After the SNAIL expression, the expression of SLUG, VIMENTIN, FOXC2, and TWIST significantly increased in a time-dependent manner in the cells treated with PMA. After the gene expression, the expression of an ExEn marker gene SOX7 and endoderm marker genes SOX17, EOMES, GATA4, and GATA6 was significantly increased in the cells (Supplementary Fig. S1B, C). On the contrary, the expression of a primitive streak marker gene GSC (Supplementary Fig. S1C) or ectoderm genes MSI1, SOX9, MAP2, Nestin, and PAX3 was not induced by PMA (Supplementary Fig. S1D). PMA addition together with GFX did not change the expression profile of EMT-related genes or cell differentiation marker genes. These results confirmed that the activation of PKC induced EMT-related genes and led to ExEn differentiation.
The expression of EGR-1
To determine PKC downstream molecules involved in the EMT initiation, differences in gene expression profiles among the cells treated with DMSO (control), PMA, or PMA together with GFX were analyzed by using DNA microarrays (Fig. 2A, B). The cells were treated with PMA for 1 h, in which time period, the SNAIL expression was before increase as shown in Fig. 1B. We previously reported that GFX, which is a selective inhibitor of PKC-α, β, γ, δ, and ζ negated PMA-induced differentiation of hPS cells, whereas Gö6976, which is a selective inhibitor of PKC-α, β, and γ isoforms could not counteract the effect of PMA [4]. Therefore, among genes influenced by PMA (fold change ≥5.0), genes of which expression were different between the cells treated PMA or together with GFX (fold change ≥5.0) were considered to be involved in EMT (Fig. 2C). Accordingly, the analysis nominated 26 activated genes and 2 repressed genes as the candidates for PKC downstream genes (Supplementary Table S3). Among them, we found that a zinc finger transcription factor EGR gene family members, EGR-1, EGR-2, EGR-3, and EGR-4 were increased in the PMA-treated cells. The expression levels of EGR-1 were significantly higher in the EGR gene family members (Fig. 2D). Therefore, we focused on EGR-1, which is reported to be involved in the EMT process in epithelial cells [45,46].
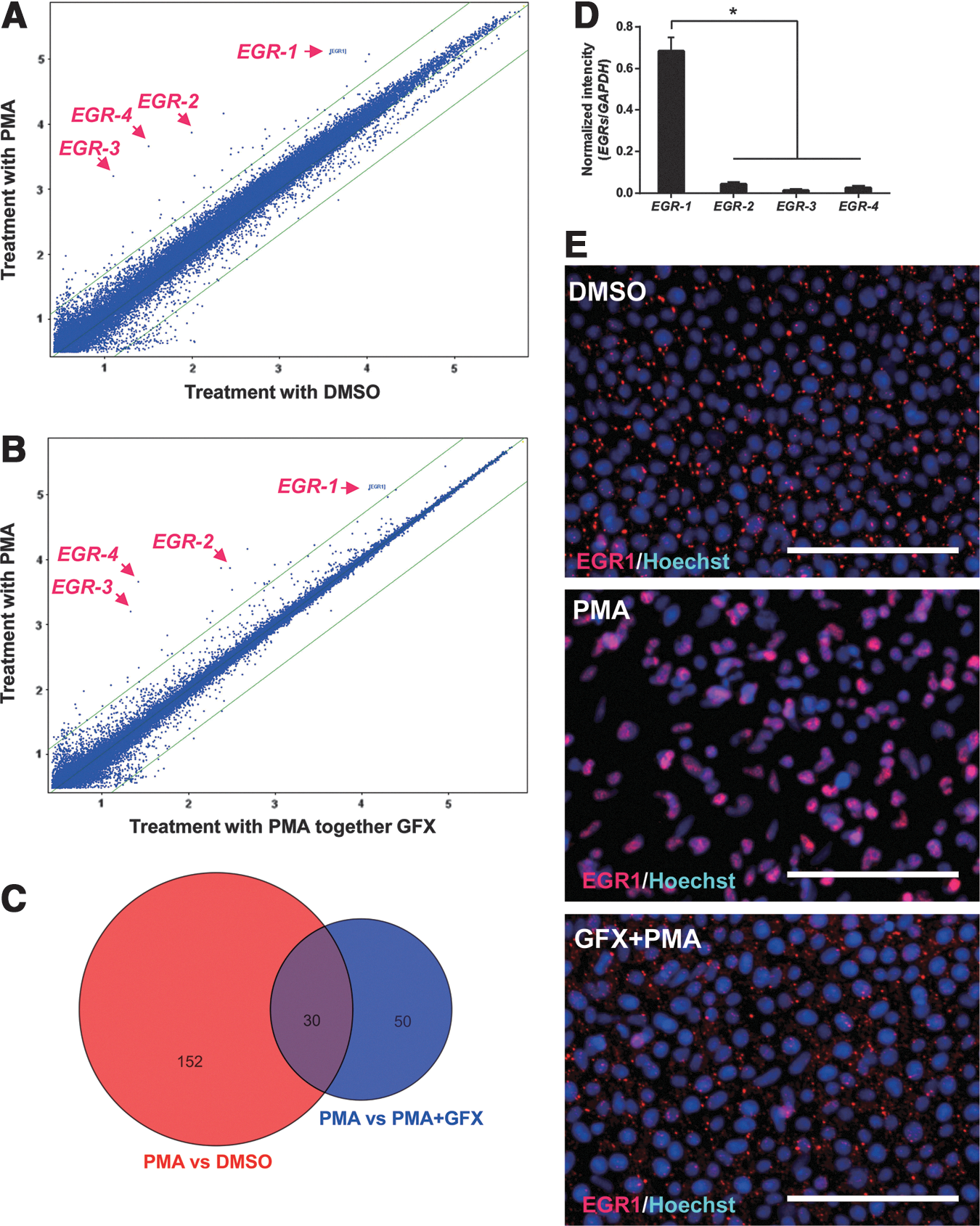
Early growth response protein-1 (EGR-1) increases in the hES cells treated with PMA.
To confirm the expression of EGR-1 in the cells treated with PMA, immunocytochemistry analysis was performed using the anti-EGR-1 antibody (Fig. 2E). The result indicated EGR-1 protein expression in the cells 1 h after PMA treatment and accumulation of EGR-1 in the nucleus. EGR-1 protein expression was at low levels in the cells treated with PMA together with GFX. From these results, it was confirmed that EGR-1 expression was induced by PMA in hES cells.
Relation between EGR-1 and SNAIL
To investigate the induced expression level of EGR-1 in the cells treated with PMA, qRT-PCR analysis was performed. The results showed that the EGR-1 expression peaked at 1 h after PMA treatment (Fig. 3A) and the induction of SNAIL expression peaked at 3 h after PMA treatment (Fig. 3B).
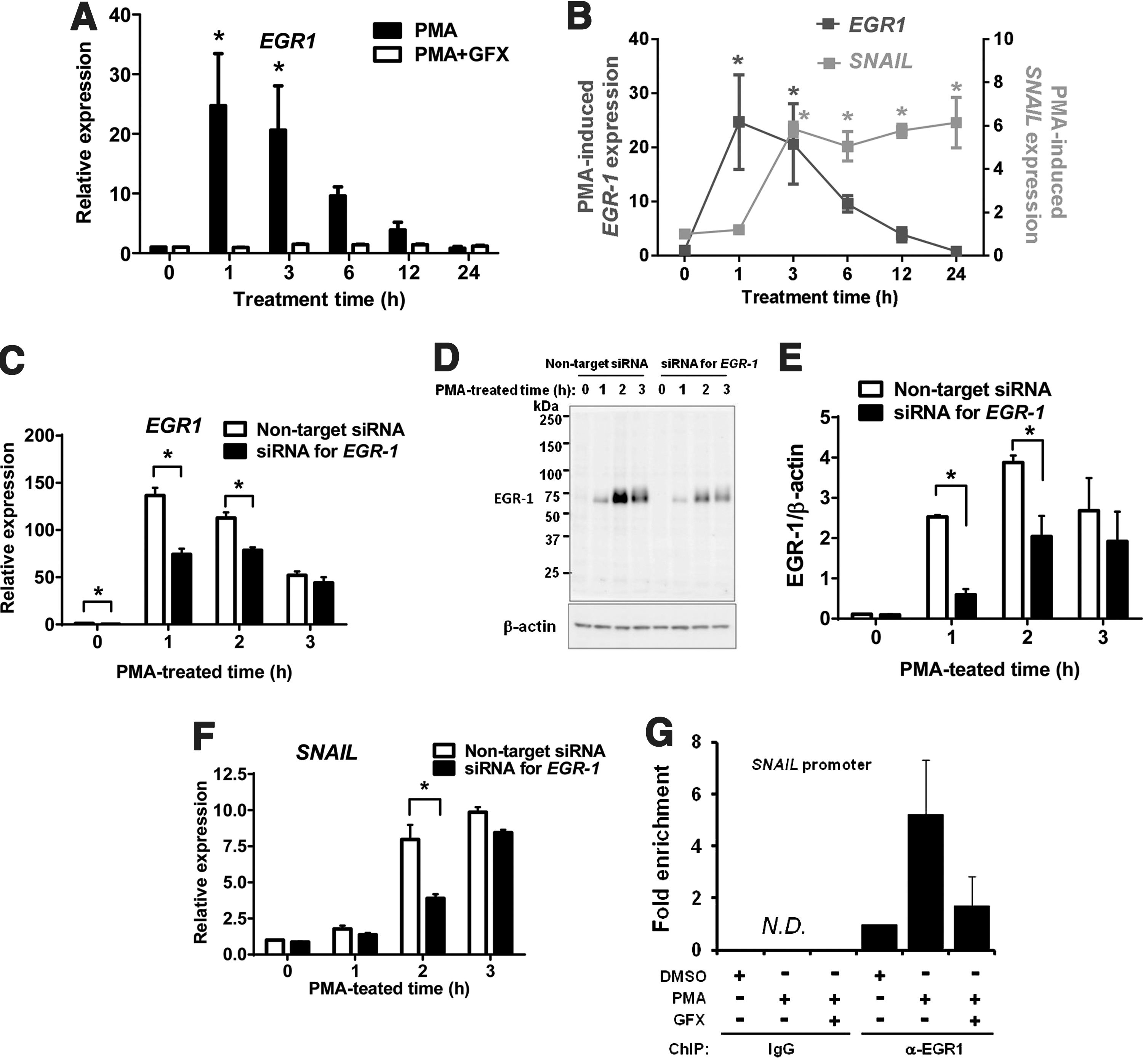
PMA-induced EGR-1 expression in hES cells.
From these data above, it was predicted that the PMA-induced EGR-1 might directly regulate the induction of SNAIL. To determine the relationship between EGR-1 and SNAIL, the cells were transfected with siRNA targeting EGR-1. When the cells were treated with PMA for 3 h, the expression of PMA-induced EGR-1 mRNA (Fig. 3C) and its protein levels (Fig. 3D, E) were significantly reduced at 1–2 h PMA treatment time, and then SNAIL expression (Fig. 3F) was significantly reduced at 2 h PMA treatment time, in the cells transfected with siRNA of EGR-1 compared with those in the cells transfected with nontarget siRNA. Then, we searched the ChIP-on-chip data for EGR-1, provided by the FANTOM 4 database (
EMT induction by ectopic EGR-1
To confirm the function of EGR-1 as an EMT inducer in hES cells, an episomal vector carrying a CAG promoter-driven EGR1-2A-eGFP was transiently transfected into H9 cells. The expression of ectopic EGR-1 in H9 hES cells 24 h after the transfection was confirmed by western blotting analysis using the specific anti-EGR-1 antibody (Fig. 4A). The expression of SNAIL was significantly increased in the cells expressing ectopic EGR-1 with or without GF109203X (GFX) (Fig. 4B). Furthermore, the cells expressing ectopic EGR-1-2A-eGFP were selected on hygromycin B for 6 days, colonies became loose and scattered flatter cells. Control cell colonies maintained undifferentiated morphology (Fig. 4C). The result by qRT-PCR showed that the expression of EMT-related genes SNAIL, SLUG, and FOXC2 were significantly upregulated in the cells expressing ectopic EGR-1, but the expression of E-cadherin was significantly downregulated (Fig. 4D). These results indicated that the expression of EGR-1 induced EMT in hES cells.
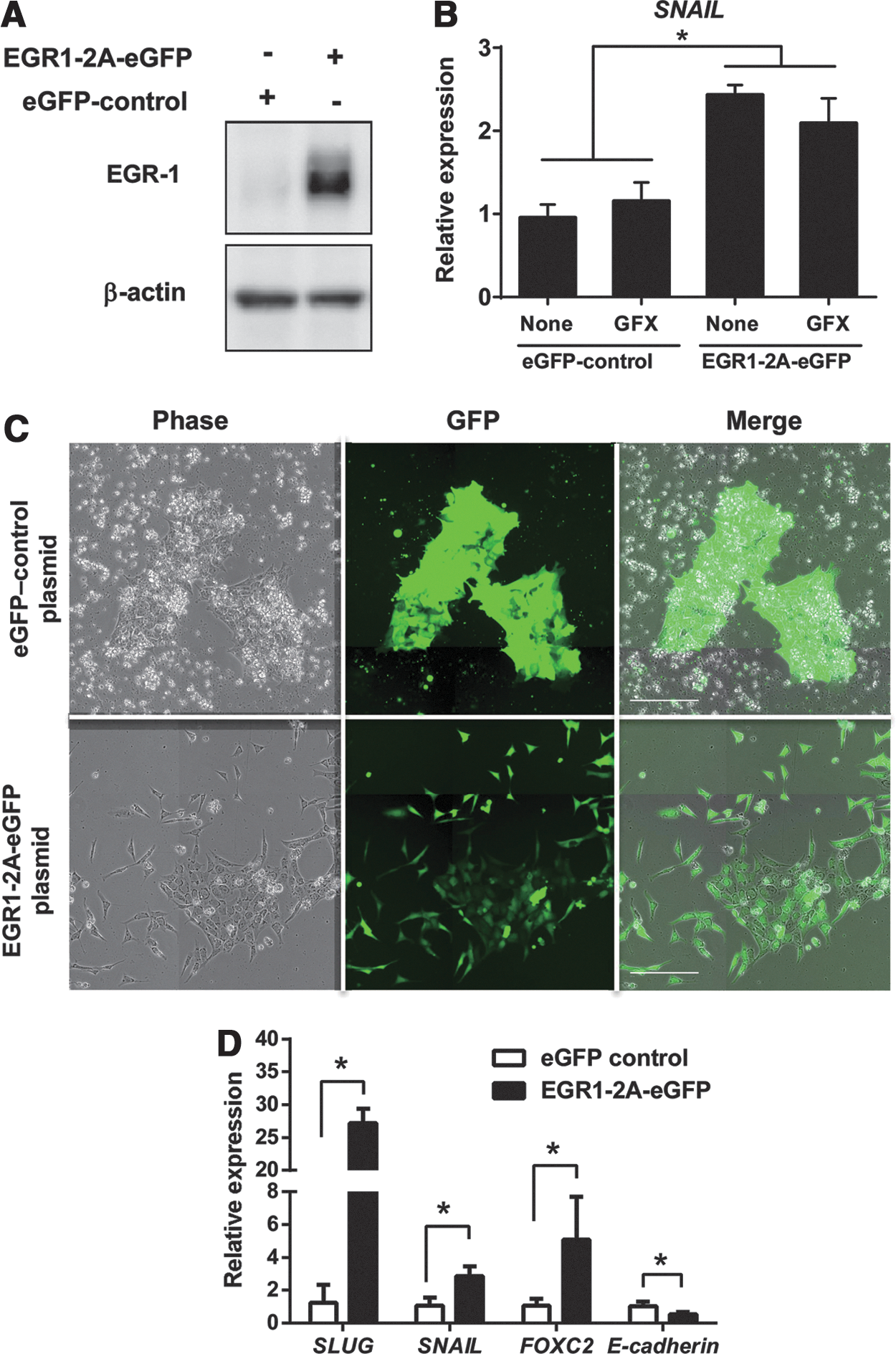
EMT-related gene expression by ectopic EGR-1.
Discussion
In this study, we showed that PMA rapidly induced the expression of EGR-1 and accumulation of EGR-1 into the nucleus, which resulted in the expression of SNAIL in hES cells. PMA-induced SNAIL expression was attenuated by knockdown of EGR-1, whereas ectopic EGR-1 expression induced EMT-related genes expression, resulting in the induction of the EMT process in hES cells. Furthermore, EGR-1 may function in hES-specific lineage cell differentiation. The ChIP-on-ChIP database of EGR-1, provided by the FANTOM 4 database (
The previous study reported that the SNAIL protein is phosphorylated by GSK-3β, and then the phosphorylated SNAIL protein binds to the promoter region of E-cadherin and downregulates the transcript of E-cadherin, leading to the EMT process [48]. These findings implied that the function of SNAIL protein induced by EGR-1 might be regulated by GSK-3β in the nucleus [48]. Although GSK-3β is generally considered to be a cytoplasmic protein, previous reports suggest that GSK-3β also functions in the nucleus [49 –51]. It means that Wnt signaling promotes EMT in hPS cells. We have previously reported that phosphorylation of GSK-3β is induced by FGF-2-activated PKC and that this process might be related to hPS cell differentiation [4]. This is consistent with previous studies that canonical Wnt signaling does not appear to promote stem cell maintenance [52,53]. However, this conclusion contradicts the findings of previous studies, which demonstrated that canonical Wnt signaling supports self-renewal of stem cells [6,38]. A recent study has shown the dual function of Wnt signaling in hES cells, suggesting that the pathways of self-renewal or differentiation are dependent on the presence of hES cell supporting factors [7,30,54,55]. From these studies, GSK-3 has emerged as an important regulator of undifferentiated state in hPS cells.
GSK-3β has been reported to be controlled by various growth factors, including insulin, IGF, FGF-2, and WNT3A [4,22,29,56,57]. We reported that in hPS cells, FGF-2-induced GSK-3β phosphorylation is inhibited in the presence of the PKC inhibitor, resulting in the inhibition of the PKC/GSK-3β signaling pathway to support the maintenance of an undifferentiated state [4]. From the findings above, it is implied that FGF-2 induces the expression of EGR-1 through the PKC signaling pathway, EGR-1 induced the SNAIL protein, and then PKC/GSK-3β activates SNAIL, leading to the EMT process [48] (Fig. 5). However, if FGF-2 continues to amplify the EMT process, hPS cells should rapidly differentiate. Inhibitory signaling related to the dual function of GSK-3β might exist in hPS cell culture and it can balance the FGF-2-induced EMT process to maintain the undifferentiated state although the PKC inhibitor can artificially stabilize the balance between undifferentiated state and cell differentiation. Further investigation of mechanisms involved in the regulation of PKC/EGR1/SNAIL and PKC/GSK-3β activity should be performed in future.
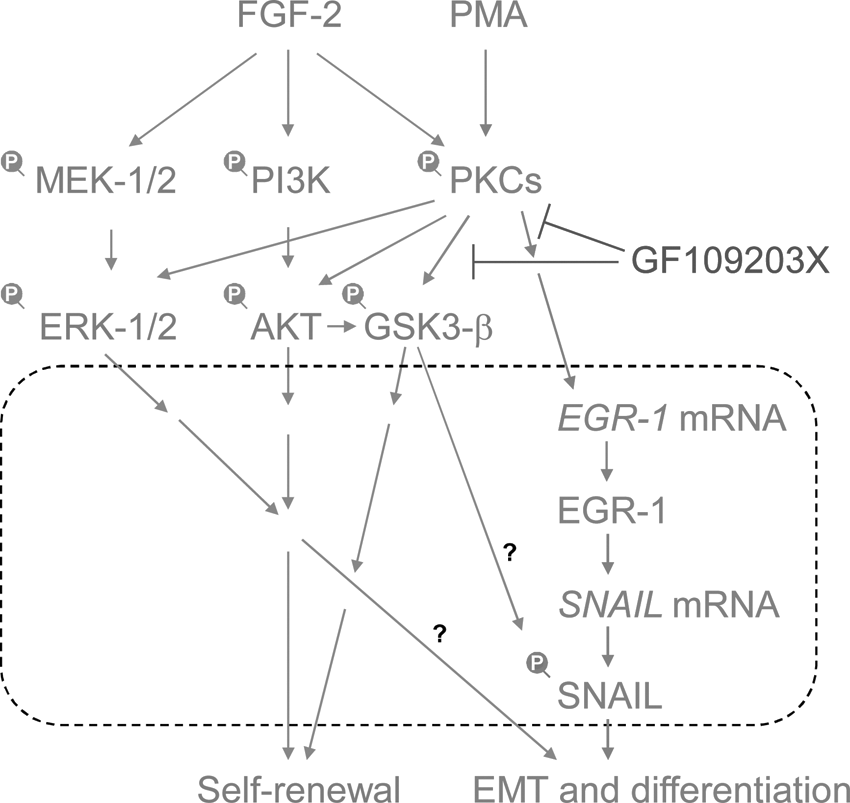
A model of EMT-triggering pathways using protein kinase C (PKC) in fibroblast growth factor (FGF-2) signaling. PMA induces PKCs, and then induces EGR-1. EGR-1 directly induces the expression of SNAIL mRNA, leading to the EMT process [48]. GF109203X, a selective inhibitor of PKCs, negated the PKC-induced EGR-1/SNAIL expression, EMT process, and differentiation of human pluripotent stem cells. PKC-EGR-1-SNAIL pathways promote EMT and cell differentiation. Dot line box indicates the process in the nucleus.
This study clarified that PKC activation induced the EMT process through the EGR-1/SNAIL pathway suggesting that EGR-1 plays a role in the differentiation of hES cells. This finding would lead to a better understanding of the precise mechanism regulating the balance between undifferentiated state and cell differentiation in hPS cells.
Footnotes
Acknowledgments
This study was supported by grants-in-aid from the Ministry of Health, Labor and Welfare of Japan to M.K.F., the Ministry of Education, Culture, Sports, Science and Technology of Japan to M.K.F. and M.K., and the New Energy and Industrial Technology Development Organization (NEDO) of Japan to M.K.F. We thank Hiroko Matsumura, Ayaka Fujiki, Naoko Ueda, Yujung Liu, Daiki Tateyama, and Minako Okamura for excellent technical support, and Takayuki Fukuda and Kana Yanagihara for technical support and discussion.
Author Disclosure Statement
All the authors have read and approved the article, and hereby declare that none of them has any competing interest.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
