Abstract
Cardiosphere derived cells (CDC) are present in the human heart and include heterogeneous cell populations of cardiac progenitor cells, multipotent progenitors that play critical roles in the physiological and pathological turnover of heart tissue. Little is known about the molecular pathways that control the differentiation of CDC. In this study, we examined the role of Notch 1/J kappa-recombining binding protein (RBPJ) signaling, a critical cell-fate decision pathway, in CDC differentiation. We isolated CDC from mouse cardiospheres and analyzed the differentiation of transduced cells expressing the Notch1 intracellular domain (N1-ICD), the active form of Notch1, using a terminal differentiation marker polymerase chain reaction (PCR) array. We found that Notch1 primarily supported the differentiation of CDC into smooth muscle cells (SMC), as demonstrated by the upreguation of key SMC proteins, including smooth muscle myosin heavy chain (Myh11) and SM22α (Tagln), in N1-ICD expressing CDC. Conversely, genetic ablation of RBPJ in CDC diminished the expression of SMC differentiation markers, confirming that SMC differentiation CDC is dependent on RBPJ. Finally, in vivo experiments demonstrate enhanced numbers of smooth muscle actin-expressing implanted cells after an injection of N1-ICD-expressing CDC into ischemic myocardium (44±8/high power field (hpf) vs. 11±4/high power field (hpf), n=7 sections, P<0.05). Taken together, these results provide strong evidence that Notch1 promotes SMC differentiation of CDC through an RBPJ-dependent signaling pathway in vitro, which may have important implications for progenitor cell-mediated angiogenesis.
Introduction
R
The Notch1 signaling pathway is evolutionarily conserved from invertebrates to vertebrates and is involved in cell-fate decisions during development [17 –20]. Notch1 signaling is activated via juxtacrine binding of an adjacent cell's Delta-like or Jagged ligands with the Notch receptor. On activation, the Notch1 intracellular domain (N1-ICD) is released from the membrane receptor via enzymatic cleavage. The N1-ICD can function as a transcriptional activator, translocating into the nucleus and forming a DNA-binding complex with J kappa-recombining binding protein (RBPJ) to control the downstream effectors of Notch1 signaling, such as Hes1. Notch1 signaling is important in regulating CPC differentiation depending on the cell type. For example, activated Notch1 can promote the early specification of CPC to a highly proliferative myocyte phenotype via Nkx2.5 upregulation [21]. However, Notch signaling also regulates the SMC differentiation of epicardium-derived cells [20], which play a crucial role in the formation of the coronary vasculature.
CDC are comprised of progenitors from different sources and, therefore, constitute a heterogeneous cell population. To date, little is known about the role of Notch1 signaling in CDC differentiation. In this study, we tested the hypothesis that Notch1 activation can promote cardiac lineage cell differentiation of CDC using novel terminal differentiation polymerase chain reaction (PCR) array technology. We found that Notch1 strongly promotes SMC differentiation of CDC in vitro, and that an injection of N1-ICD-expressing CDC leads to enhanced numbers of implanted cells differentiating into SMC in a mouse myocardial infarction model.
Materials and Methods
CDC culture
Mouse CDC were isolated as previously described [22] using a protocol approved by the Institutional Animal Care and Use Committee of the University of Cincinnati. Briefly, adult mouse hearts were minced to 1 mm3 in size and plated on laminin-coated dishes for 2 weeks. The round, phase-bright migrating cells were harvested and filtered with 40 μm cell strainers to avoid heart tissue contamination. Cells were cultured in poly-
Flow cytometry
Flow cytometric analyses were performed as previously described [22]. Briefly, cultured cells were blocked with 5% bovine serum albumin, and stained with a panel of antibodies including Sca-1, c-kit, and CD90 (BD Biosciences, San Jose, CA), or isotype controls (Vector Laboratories, Burlingame, CA), respectively. The cells were subjected to flow cytometry using a BD LSRII flow cytometer and BD FACSDivaTM software.
Plasmid construct and generation of recombinant adenoviruses
The adeno-backbone plasmid (pAdtrack-CMV-N1-ICD) was produced by amplifying 2.4-kb of human N1-ICD cDNA from EF.hICN1.Ubc.GFP(Addgene 17626) via PCR with primers containing Not1 and Xho1 restriction sites on the ends (Forward PCR primer: AAGGAAAAAA GCGGCCGC GCCACC ATGCGGCGGCAGCATGGCCA. Reverse PCR primer: CTAG CTCGAG TTACTTGAACGCCTCCGGGA.). The N1-ICD fragment was subcloned into the Not1-Xho1 site of the pAdtrack-CMV vector (Addgene plasmid 16405). Recombinant adenovirus Adtrack-CMV-N1-ICD and Adtrack-CMV were generated by using the AdEasy-1 expression system, as previously described [25]. Briefly, parental empty pAdtrack-CMV bearing GFP, and pAdtrack-CMV-N1-ICD vectors bearing N1-ICD and GFP, were first linearized with PmeI, and then, introduced into BJ5183-AD-1 electroporation competent cells (Agilant Technologies, La Jolla, CA) by electroporation. Recombinant adenoviral plasmids were recovered from E. coli and introduced into AD-293 cells by Fugene HD (Roche Applied Science, Indianapolis, IN). The recombinant virus was propagated in AD-293 cells. The viruses were purified and concentrated using the Adeno MINI Purification ViraKit (VIRAPUR, LLC, San Diego, CA). Cells seeded on dishes were infected with adenovirus at a multiplicity of infection of 500.
Quantitative reverse transcription-polymerase chain reaction and PCR arrays
Total RNA was extracted from adenovirus infected CDC GFP and CDC N1-ICD using the RNeasy Plus Micro kit (QIAGEN, Valencia, CA) following the manufacturer's instructions. Approximately 1ug of total RNA was used for cDNA synthesis using the BluePrint 1st Strand cDNA Synthesis Kit (Takara Bio, Inc., Madison, WI) following the manufacturer's instructions. The cDNA synthesized was used to perform quantitative PCR using Mouse Terminal Differentiation Markers RT2 Profiler PCR Array (catalog no. PAMM-048A) on an Mx3000P Real-Time PCR System (Agilent Technologies, Santa Clara, CA) using the SensiMix SYBR kit (Bioline, Tauton, MA) according to the manufacturer's instructions. The data were analyzed using the RT2 profiler PCR Array Data Analyzer template provided by Sabioscience. Gene expression was compared between CDC GFP and CDC N1-ICD , and genes that showed more than 2.5-fold difference in expression were identified. We verified results using primers listed in Table 1. For each sample, the gene of interest was normalized to glyceraldehyde 3-phosphate dehydrogenase (GAPDH) before the calculation of relative fold up- or down-regulation in the transcription levels compared with CDC GFP .
Western blotting
Cell lysate supernatants were prepared, resolved on a 7%–12% sodium dodecyl sulfate bis-tris gel, and transferred to an Immobilon® FL PVDF membrane (Millipore, Billerica, MA). For Odyssey technology, the membrane was blocked with Odyssey blocking buffer (LICOR Biosciences, Lincoln, NE) and probed with rabbit anti-Transgelin (1:500, 22kda, SC-50446; Santa Cruz Biotechnology, Santa Cruz, CA), rabbit anti-MYH11 (H-44) (1:500, 200kda, SC-98705; Santa Cruz Biotechnology), mouse anti-GAPDH (1:2,000, MAB374; Millipore), and then, IRDye 680 goat anti-rabbit IgG or IRDye 800 Goat anti-mouse IgG at 1:5,000 (LICOR Biosciences). Probed blots were scanned using an Odyssey infrared imager.
Immunofluorescence staining
CDC cells infected with Adeno-empty vector or Adeno-N1-ICD were plated on 8-well chamber slides (BD Biosciences) and fixed with 4% paraformaldehyde. After blocking nonspecific binding with 10% goat serum in phosphate-buffered saline, the cells were incubated with rabbit anti-Transgelin (1:50; Santa Cruz Biotechnology), rabbit anti-MYH11 (H-44) (1:50; Santa Cruz Biotechnology), rabbit anti-smooth muscle actin (SMA, 1:100; NeoMarkers, Fremont, CA), and rabbit anti-activated Notch1 antibodies (1:100; Abcam) at 4°C overnight, washed 3×in phosphate buffered saline with Tween 20, and incubated with Alexa 555-conjugated anti-rabbit secondary antibodies (Invitrogen). Slides were mounted using VECTASHIELD HardSet Mounting Medium with DAPI (Vector Laboratories). The staining was analyzed by a Zeiss 710 Laser Scanning Microscope (Carl Zeiss, Thornwood, NY).
Cell labeling, myocardial infarction, and intramyocardial cell delivery
CDC were genetically engineered to express Adeno-N1-ICD and Adeno-GFP. Male C57/BL6 mice (2-month-old) were anesthetized with ketamine/xylazine (100 mg/kg/10 mg/kg, i.p.) and mechanically ventilated. Myocardial infarction was induced via ligation of the left anterior descending coronary artery 2 mm from the tip of the normally positioned left atrium as previously described (n=6) [22]. Ten minutes after ligation, a 20 μL solution containing 5×105 cells in DMEM was injected intramyocardially 1 mm above the ligation site. Animals were handled according to approved protocols and animal welfare regulations of the Institutional Animal Care and Use Committee of the University of Cincinnati.
In vivo differentiation of Adeno-N1-ICD modified CDC
Two weeks after surgically induced myocardial infarction and intramyocardial injection of CDC, mouse hearts were harvested, embedded in optimal cutting temperature compound, snap frozen, cut into 5-μm sections, and immunostained with anti–cardiac troponin I (1:100, SC-15368; Santa Cruz Biotechnology, Inc.), anti–von Willebrand factor (1:100, F3520; Sigma), or anti– SMA (1:100, RB-9010-P; NeoMarkers) antibodies. Primary antibodies were detected via secondary staining with goat anti-rabbit Alexa Fluor 555–conjugated. Nuclei were counterstained with both DAPI (Vector Laboratories) and Draq5. The staining was analyzed by a Zeiss 710 Laser Scanning Microscope (Carl Zeiss).
Statistical analyses
Values are presented as mean±standard deviation, as indicated in the legend. Comparison was evaluated by Student's t-test between 2 groups and 1-way analysis of variance among multiple groups. In all analyses, P<0.05 was considered statistically significant.
Results
In vitro studies
Isolation and phenotypic characterization of CDC
Cardiospheres were grown from enzymatically digested adult hearts, and the CDC were isolated using a hematopoietic lineage-depletion cocktail (Fig. 1A). Surface marker expression was profiled by flow cytometry. About 80% of the sorted cells were positive for Sca-1 expression, 62% cells were positive for CD90, and 4% were positive for c-kit (Fig. 1B). These data indicate that CDC represent a subpopulation of cells that are positive for Sca-1and CD90.
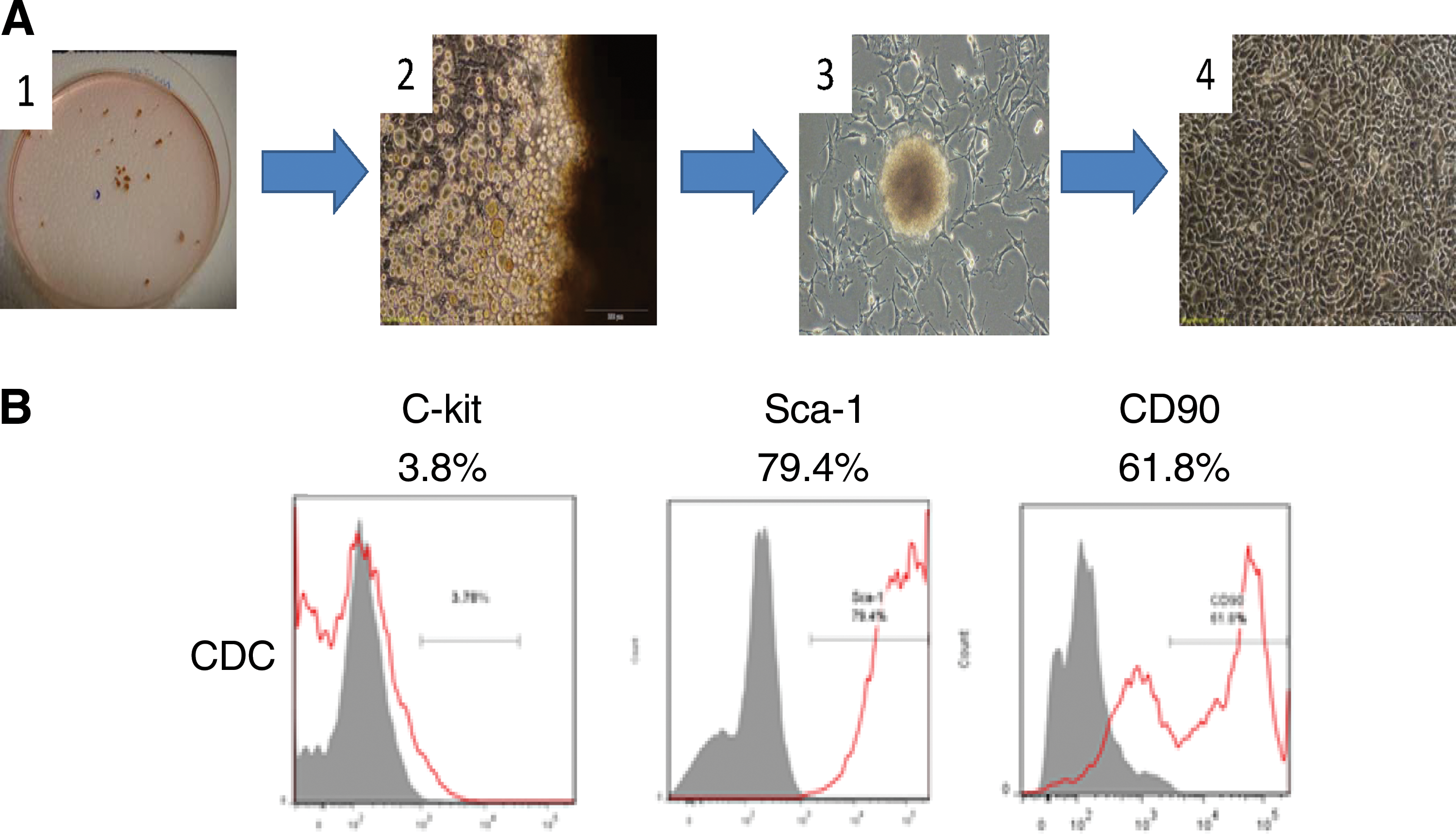
Isolation, expansion, and phenotypic characterization of cardiosphere derived cells (CDCs).
CDC differentiation induced by overexpression of constitutively active N1-ICD
To assess the differentiation of Notch1-activated CDC, we analyzed the expression of a number of terminal differentiation marker genes that cover 13 organs, using the PCR array. From the genes analyzed, the expression of many cardiovascular markers, including SMC (Tagln and Myh11), endothelium markers (Pecam1 and Nrp1), cardiomyocyte markers (Nppa and Myh7), and chemokine receptors for stem cell homing (CXCR4), were significantly upregulated in N1-ICD overexpressing CDC (>2.5-fold, Fig. 2). Among all the upregulated genes, Tagln and Myh11, the SMC markers, were the most strongly upregulated, indicating that SMC differentiation is the main outcome of Notch1-activated CDC. Interestingly, Notch1 activation resulted in the downregulation of genes in 5 other systems, including markers for osteoblasts, cholangiocytes, beta cells, oligodendrocytes, and podocytes. We performed PCR using primer sets distinct from the commercial array, which confirm the significantly increased Tagln and Myh11 mRNA expression in Notch1-activated CDCs (Fig. 3A).
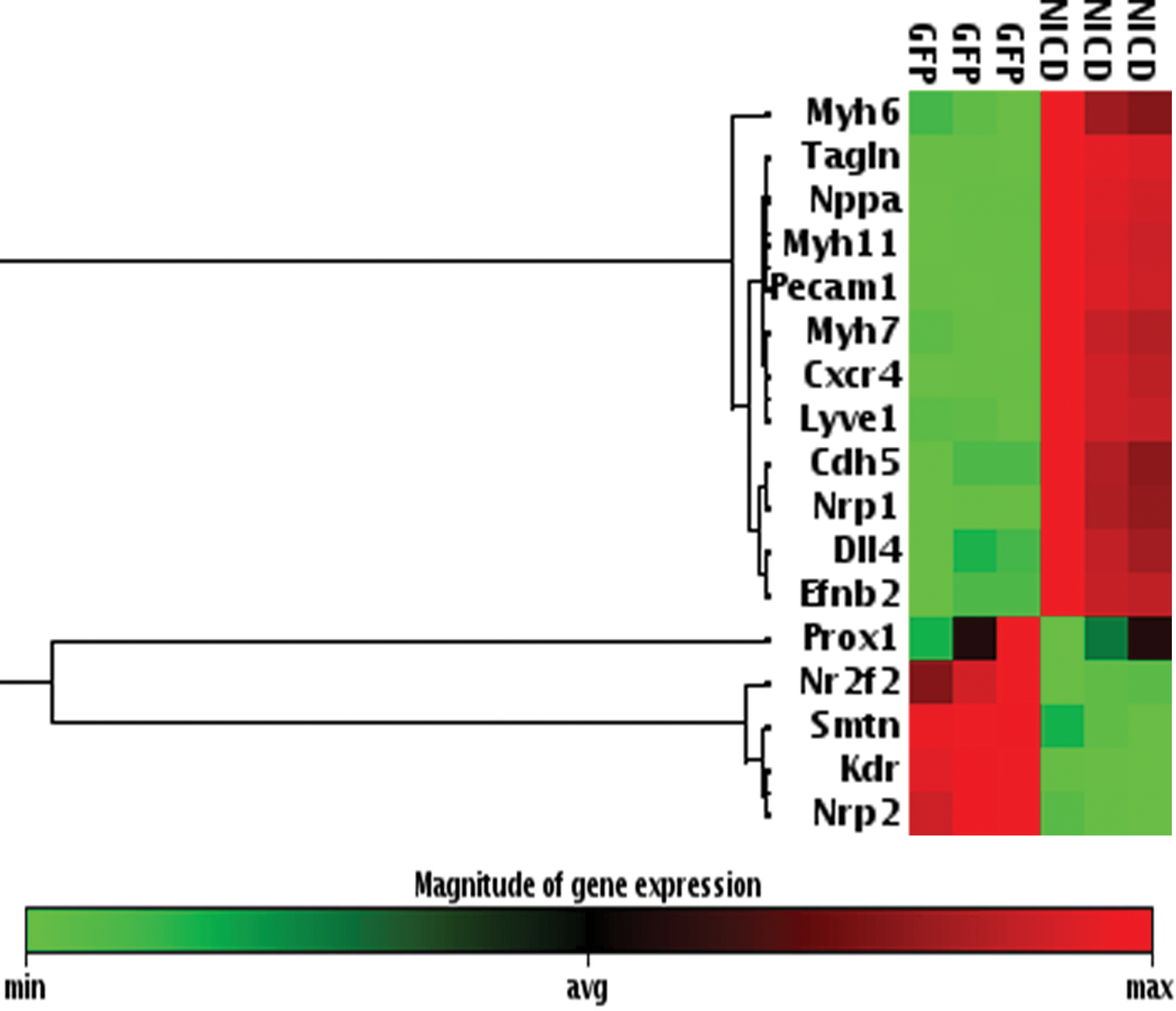
Heat map of cardiovascular genes identified as significantly altered in Notch1-activated CDC, using reverse transcription (RT)-polymerase chain reaction (PCR)–based mouse Terminal Differentiation Marker system. Color images available online at
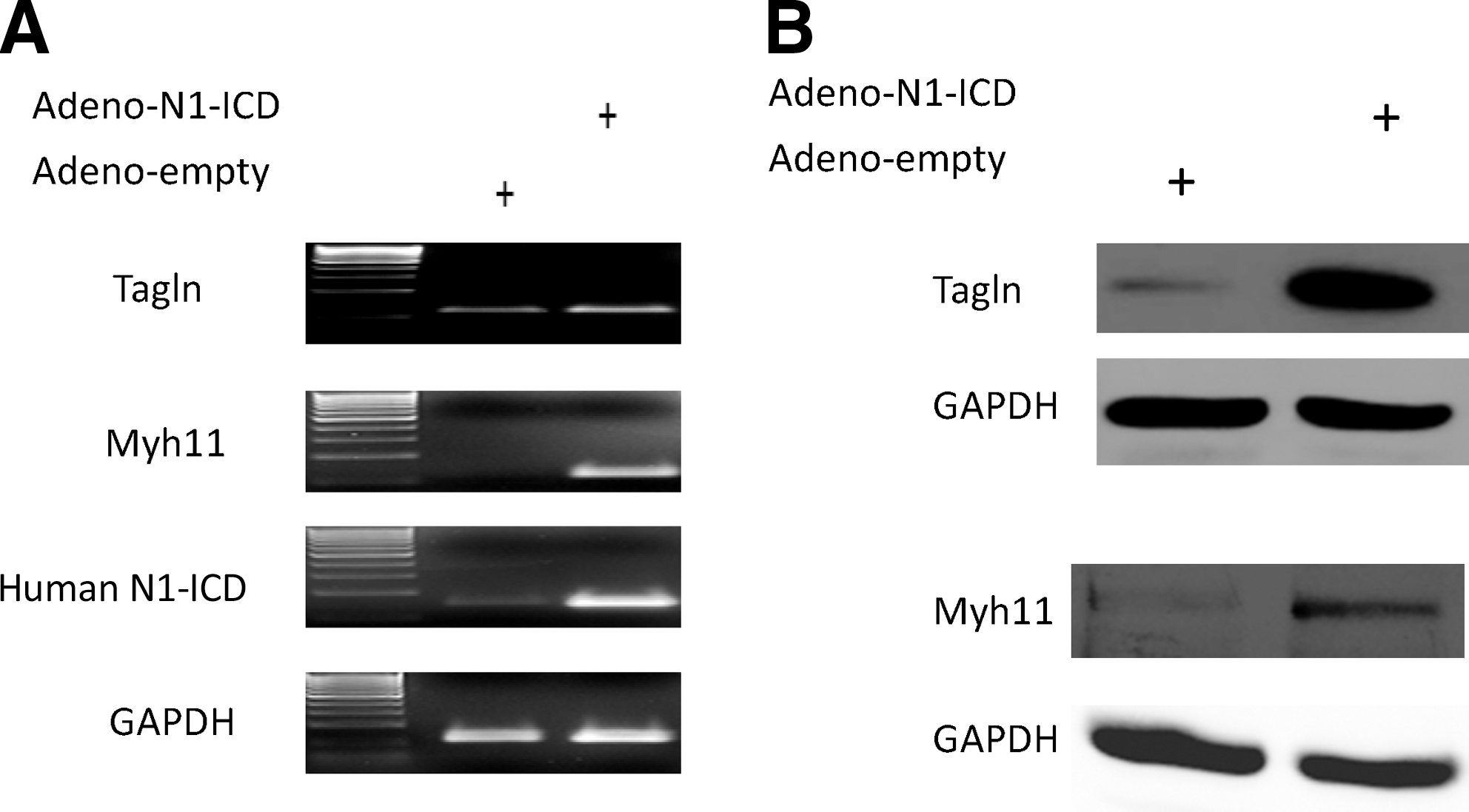
The constitutively active form of Notch1 intracellular domain (N1-ICD) indiscriminately up-regulates both Tagln and Myh11 mRNA and protein expression in CDC.
Both Tagln and Myh11 protein expression is induced by overexpression of constitutively active N1-ICD
Due to posttranscriptional regulation, mRNA expression does not always lead to corresponding increases in protein accumulation [26], Thus, we evaluated whether Tagln and Myh11 mRNA up-regulation is accompanied by an expression of the cognate proteins. Accordingly, cells infected with the Adeno-N1-ICD vector or Adeno- empty vector were subjected to Western blot with anti-Tagln, anti-Myh11, and anti-N1-ICD antibodies. Western blot showed that Tagln and Myh11 levels were upregulated by N1-ICD overexpression (Fig. 3B), demonstrating that N1-ICD can induce SMC differentiation of CDC. In addition, as depicted in Fig. 4A–F, immunofluorescent staining revealed that Tagln and My11 expression was clearly detected only in the cytoplasm of Notch1-activated CDC.
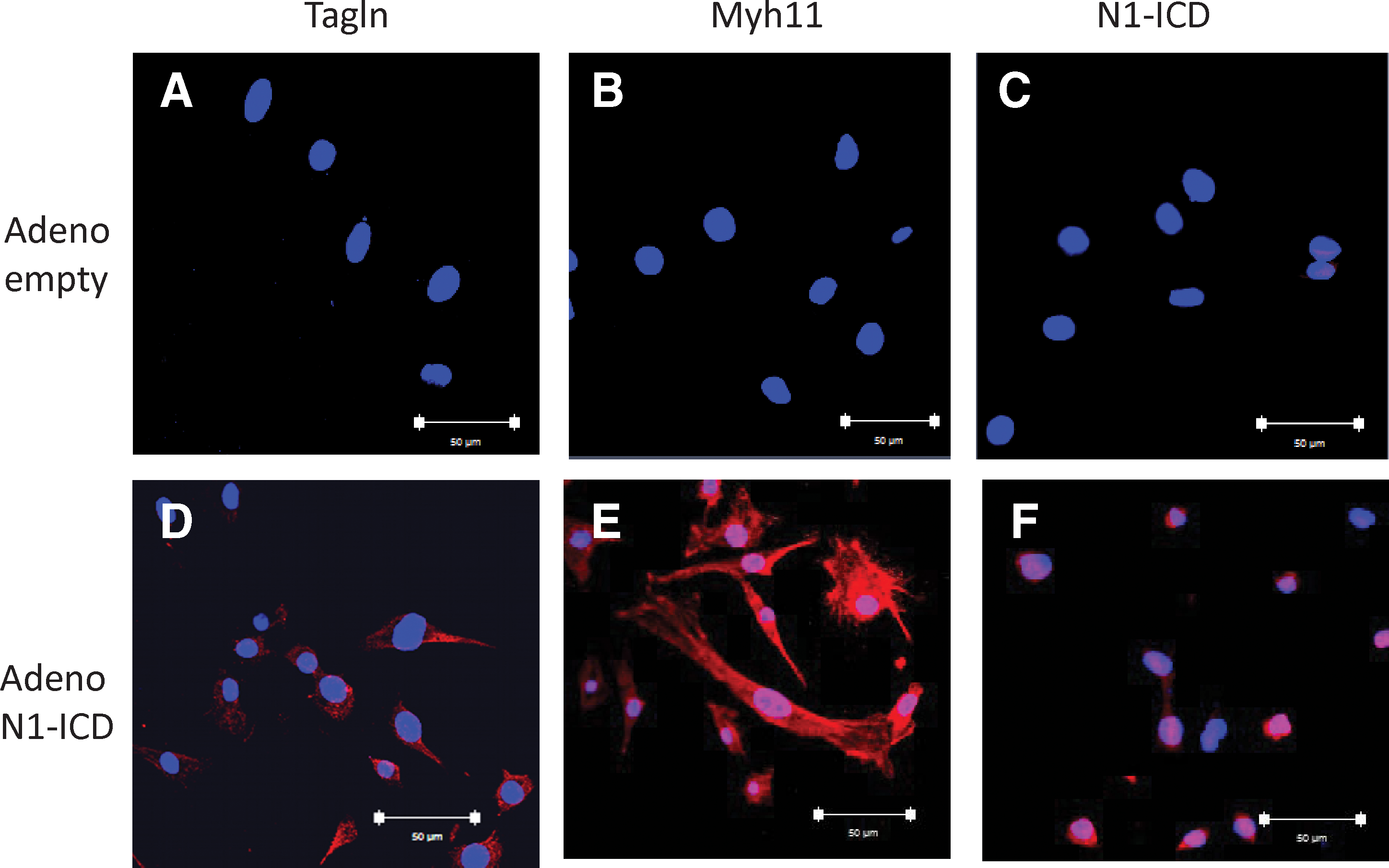
Tagln and Myh11 protein expression in CDCs overexpressing N1-ICD. Cells were infected with adeno-N1-ICD or adeno-empty vector, and immunofluorescence assay was performed using specific anti-Tagln, anti-Myh11, or anti-N1-ICD antibody.
Tagln and Myh11 expression in CDC is RBPJ dependent
To test the hypothesis that RBPJ is required for SMC differentiation induced by overexpression of N1-ICD, we employed the MEF cell line OT11, which carries homozygous mutations at the RBPJ loci, and its parental wild-type cell line, OT13. The RBP-J−/−(OT11) and WT (OT13) fibroblast cell lines are reliable model systems that study RBP-J dependent signaling in cell differentiation. Overexpression of N1-ICD by adenoviral-mediated gene transfer induced both Tagln and Myh11 (27-fold) gene expression in OT13 cells (Fig. 5A, B). In contrast, overexpression of N1-ICD failed to induce either Tagln or Myh11 gene expression in OT11 cells (Fig. 5A, B). Hes1, the known target of Notch, was used as positive control.
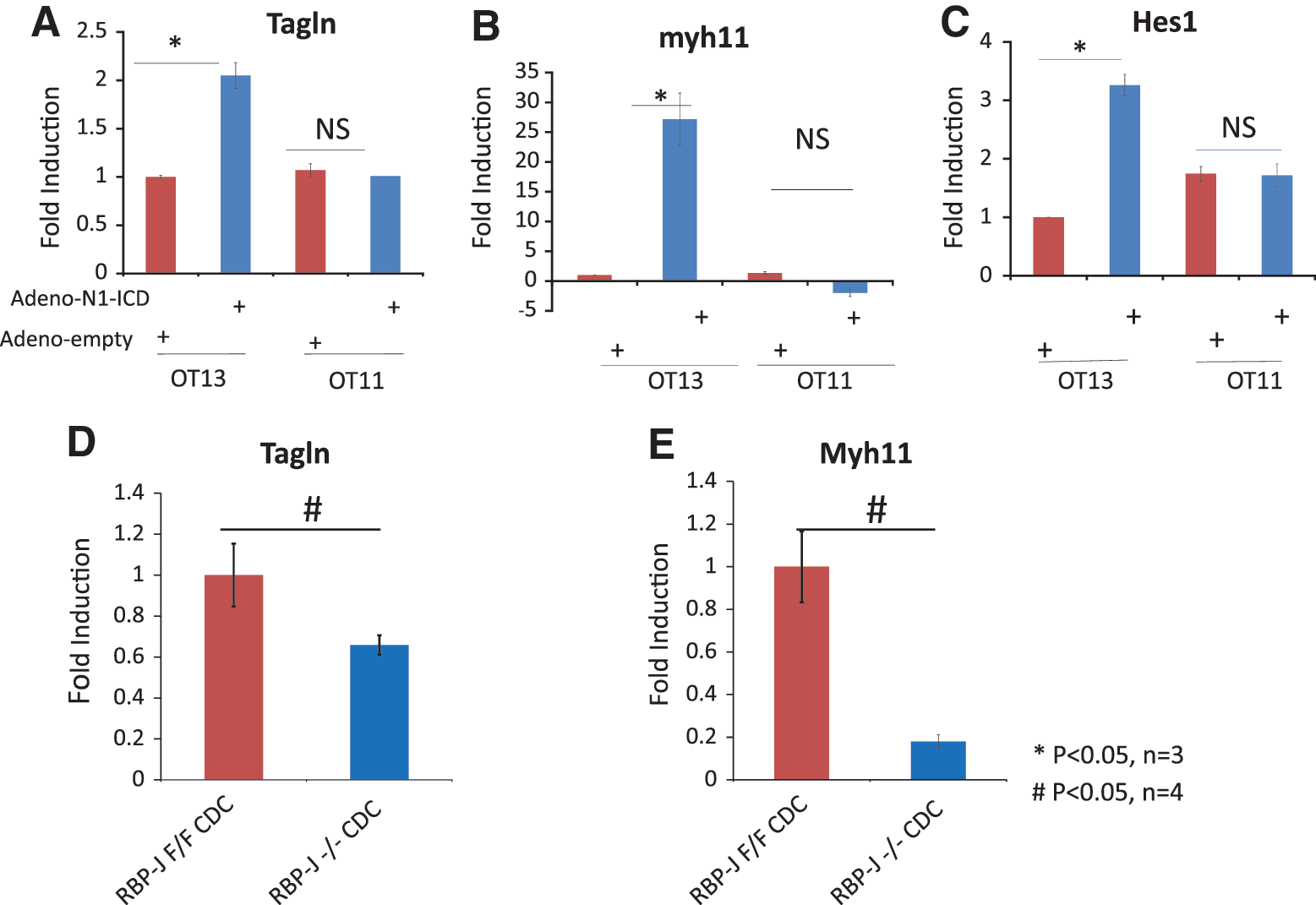
J kappa-recombining binding protein (RBPJ) mediates the increased Tagln and Myh11 expression induced by Notch1 activation.
To test the hypothesis that endogenous Notch signaling is required for SMC differentiation of CDC (in the absence of overexpression of N1-ICD), we employed RBPJ (−/−) CDC, which carry a homozygous deletion of the RBPJ loci. RBPJ (fl/fl) CDC were used as a control. RBPJ−/−CDC displayed significantly reduced Tagln and Myh11 expression compared with RBPJ fl/fl control (Fig. 5D, E), confirming that differentiation of CDC into SMC depends on endogenous Notch signaling.
In vivo studies
Differentiation of Sca-1+ cells in injured myocardium
Our previous studies demonstrated the multipotent capacity of Sca-1+ CDC, which can differentiate into cardiomyocytes, endothelial cells, and SMC in vivo [22]. To investigate the effects of overexpression of N1-ICD in CDC in vivo, we injected N1-ICD-CDC, or GFP-CDC as a control, intramyocardial in syngenic mice after inducing myocardial infarction. Donor cells were detected in the myocardium by laser confocal microscopy 2 weeks after cell injection. As compared with GFP-CDC control, N1-ICD-CDC yielded more cells that stained positively for SMA (44±8/high power field (hpf) in N1-ICD-CDC vs. 11±4/hpf in GFP-CDC, n=7 sections, P<0.05, Fig. 6A, B). Notably, N1-ICD modified CDC maintained their multipotency in vivo, also differentiating into cardiomyocytes (Fig. 6C, Supplementary Fig. S1; Supplementary Data are available online at
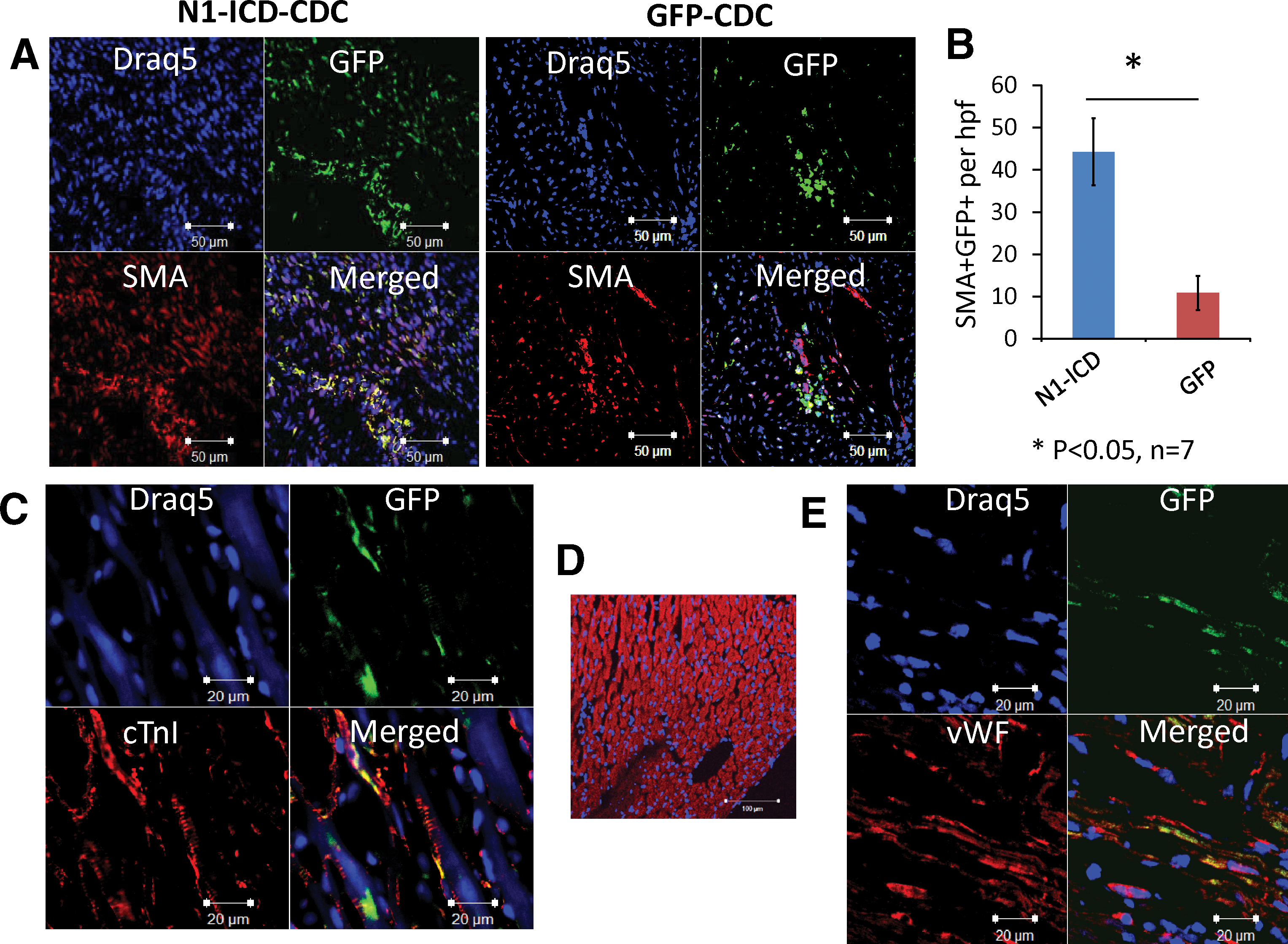
In vivo differentiation of Adeno-N1-ICD modified CDC. One month after surgical MI and cell injection, hearts were harvested and sectioned, then sections were immunostained for the lineage-specific proteins (red) cardiac troponin I (cTnI), von Willebrand factor (vWF), or smooth muscle actin (SMA); nuclei were counterstained with Draq5 (blue).
Discussion
The results presented here demonstrate that Notch1 activation promotes differentiation of CDC into SMC in vitro in an RBPJ-dependent fashion. An injection of N1-ICD-expressing CDC into the ischemic myocardium yields enhanced numbers of implanted cells differentiating into SMC in vivo. To the best of our knowledge, this is the first study examining Notch-mediated CDC differentiation.
CDC constitute a heterogeneous population of progenitors with potential for heart regeneration. To identify the profile of CDC differentiation induced by constitutive Notch-1 activation, we performed reverse transcription (RT)-PCR array analysis of terminal differentiation markers spanning 13 organs and tissues. We observed significant increases in SMC specific mRNA and protein expression in N1-ICD-CDC. Recently, Notch1 has been reported to modulate cell differentiation in the proepicardium and adjacent pericardial mesoderm, and to be required for coronary vessel formation and myocardial growth during embryonic heart development [27]. Since recent reports indicate that epicardial tissue is a major source of CPC, in this regard, our data support a role for Notch1 in endogenous cardiac progenitor commitment to SMC differentiation in coronary vessel formation. Pathophysiologically, a similar mechanism could be operative during vessel regeneration in ischemic heart disease. Recent studies have shown that activated CPC can regenerate conductive, small, and intermediate-sized arteries and capillary structures and improve cardiac function in chronically injured rat hearts long after infarction and scar formation [6,7].
Data are conflicting regarding SMC differentiation of Notch1 activated cells. Several in vitro studies indicated that Notch1 signaling inhibits SMC differentiation of C3H10T1/2 fibroblasts [28] and human aortic SMC [29]. On the other hand, recent reports show that Notch1 activation promotes SMC differentiation of human aortic endothelial cells and primary human foreskin fibroblasts via RBPJ directly binding to the alpha SMA promoter [17,30]. Likewise, we observed that SMC differentiation is RBPJ dependent. In RBPJ knockout CDC, the basal expression of SMC markers was significantly lower compared with the RBPJ floxed control CDC. Therefore, the role of Notch1 in SMC differentiation may be cell specific. Our study specifically addresses the role of Notch1 in differentiation of CDC, a promising cardiac progenitor source. We uncovered Myh11 and Tagln as targets of Notch1 signaling via forced expression of the constitutively active N1-ICD in CDC.
Our in vivo experiments suggest that N1-ICD modified CDC yield considerably more implanted cells that express SMA, a marker of mature, differentiated SMC, suggesting a strong correlation between Notch activation and differentiation of implanted CDC into SMC. On the other hand, it is evident that not all implanted N1-ICD-CDC express SMA, suggesting that N1-ICD could regulate differentiation of CDC into other lineages in vivo and/or modulate cell survival, implantation, proliferation, or interactions with the micro-environment. In addition, our experiments utilized adenoviral-mediated gene transfer, which produces transient expression of the transgene for a period of about 2 weeks. Sustained, as compared with transient, Notch activation could have profoundly different effects on the fate of implanted CDC. Further studies will be required to resolve these issues.
For the long-term cell culture over 78 days, we did not check the karyotype of the CDC, but we checked the possibility of cancer and/or cancer stem cells using CDC within 10 passages, and we did not observe obvious tumor growth at 4 months after an intramuscular injection in NOD/SCID mice, while we observed obvious tumor formation in mice injected with the same amount of mouse-induced pluripotent stem cells at 3 weeks after cell injection (Supplementary Fig. S2); thus, it seems that the possibility of cancer stem cells or cell line in CDC within 10 passage is low.
From our understanding of stem cell biology, Notch-1 regulation of SMC differentiation of CDC has potentially far-reaching implications. After myocardial infarction, the vasculature in the infarct zone is impaired [31 –33], which is directly related to survival [31]. Therefore, reestablishing blood flow to the damaged area is a critical step in restoring oxygen and nutrient supply to the cardiac matrix. Our data suggest that Notch1 activation promotes the SMC commitment of CDC in vitro. Further studies examining the long-term effect of Notch1/RBPJ activation in CDC differentiation and postinfarction angiogenesis are warranted.
Footnotes
Acknowledgment
This work was supported by the American Heart Association Beginning Grant-in-Aid 0765094Y (to Y.T.); NIH grant HL086555 (to Y.T.), and NIH grants HL076684 and HL62984 (to N.L.W.).
Author Disclosure Statement
There is no conflict of interest in connection with this article.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
