Abstract
Background:
Although previous observational studies suggest a potential association between gut microbiota (GM) and knee osteoarthritis (KOA), the causal relationships remain unclear, particularly concerning the role of blood metabolites (BMs) as potential mediators. Elucidating these interactions is crucial for understanding the mechanisms underlying KOA progression and may inform the development of novel therapeutic strategies.
Objective:
This study aimed to determine the causal relationship between GM and KOA and to quantify the potential mediating role of BMs.
Methods:
Instrumental variables (IVs) for GM and BMs were retrieved from the MiBioGen consortium and metabolomics genome-wide association studies (GWAS) databases. KOA-associated single-nucleotide polymorphisms were sourced from the FinnGen consortium. Inverse-variance weighted approach was utilized as the main analytical method for Mendelian randomization (MR) analysis, complemented by MR-Egger, simple mode, weighted mode, and weighted median methods. The causal relationships between GM, BMs, and KOA were sequentially analyzed by multivariate MR. False discovery rate correction was applied to account for multiple comparisons in the MR results. Sensitivity analyses and reverse MR analysis were also conducted to verify the reliability of the findings. Finally, a two-step approach was employed to determine the proportion of BMs mediating the effects of GM on KOA.
Results:
MR analysis identified seven gut microbial species that are causally associated with KOA. Additionally, MR analysis of 1091 BMs and 309 metabolite ratios revealed 13 metabolites that influence the risk of KOA. Through two-step analysis, three BMs were identified as mediators of the effects of two GMs on KOA. Among them, 6-hydroxyindole sulfate exhibited the highest mediation percentage (10.26%), followed by N-formylanthranilic acid (6.55%). Sensitivity and reverse causality analyses further supported the robustness of these findings.
Conclusion:
This research identified specific GMs and BMs that have a causal association with KOA. These findings provide critical insights into how GM may influence KOA risk by modulating specific metabolites, which could be valuable for the targeted treatment and prevention of KOA.
Introduction
Knee osteoarthritis (KOA) is a primary contributor to pain and disability among the elderly worldwide, significantly contributing to socioeconomic burdens. 1,2 The prevalence of KOA is rising, primarily due to longer life expectancy and increasing obesity rates. 3 Despite its high prevalence, many individuals with KOA do not receive early and effective treatment. Since KOA is currently incurable, treatment strategies mainly focus on symptom relief, which underscores the need for early diagnosis. However, early detection of KOA is challenging because of the weak correlation between pain and structural damage. Recent genome-wide association studies (GWAS) have uncovered various genetic loci related to KOA, providing new insights into its genetic etiology. 4,5
The human gastrointestinal tract hosts trillions of symbiotic bacteria, which play a crucial role in maintaining health. 6 Advances in high-throughput sequencing technologies have suggested the possible role of the gut microbiota (GM) in skeletal health. Gut microorganisms are responsible for various physiological functions, including metabolism, immune response, infection resistance, and neurotransmitter production. 7 An imbalance in gut bacteria can lead to conditions such as diabetes, obesity, and metabolic disorders. 8 Growing evidence also suggests that gut microbes play a role in bone metabolism, although the exact mechanisms are still unclear. As our understanding of GM deepens, it may become possible to detect early KOA pathogenesis through analysis of microbial populations. 9
During the past decade, studies on the human microbiome have demonstrated that our intestinal symbionts play a crucial role in maintaining overall physiological balance by regulating immunological responses within the gastrointestinal tract and beyond. 10 Disruptions in intestinal flora may contribute to the development of KOA by influencing metabolic, hormonal, and immune balance. 11 Several reports have documented a close association between GM and KOA. For instance, Lorenzo and coworkers showed significant alterations in the GM and oral microbiota of individuals with KOA and rheumatoid arthritis. 12 Additionally, a study has found that polymorphisms in the vitamin D receptor gene are related to symmetrical hand KOA in the Finnish population. 13
As functional intermediates under environmental exposure, blood metabolites (BMs) are not only byproducts of the body’s pathophysiological processes but also serve as signaling molecules that regulate cell proliferation, differentiation, and apoptosis. 14 They reflect an organism’s physiological state in response to genetic or environmental changes and can predict or influence disease onset and progression. Recent studies have used machine learning models to predict KOA risk, incorporating specific BM levels as predictive factors. 15,16 An observational study indicated that moxibustion may regulate KOA-related cartilage synthesis and degradation through amino acid metabolic pathways, including proline, tryptophan, glutamate, and histidine, exerting anti-inflammatory, analgesic, and cartilage-protective effects. 17 In addition, exploratory studies on KOA-related pain symptoms have revealed significant differences in serum metabolic profiles between KOA patients and healthy individuals. Notably, serum levels of 8,9-epoxyeicosatrienoic acid and 14,15-dihydroxyeicosatrienoic acid were positively correlated with pain severity. 18 Although multiple confounding factors such as age, behavior, and weight contribute to KOA progression, these findings establish a significant association between BMs and KOA development. Dysregulated blood metabolomics may represent a key underlying mechanism in KOA pathogenesis, highlighting the need to identify relevant metabolites to facilitate more personalized therapeutic interventions for KOA patients.
A recent large-scale association study highlighted the enrichment of microbiome loci in metabolic, nutritional, and environmental pathways, identifying causal roles for GM in conditions such as rheumatoid arthritis and ulcerative colitis. 19 This presents an important opportunity to evaluate the causal associations between specific GM and OA. Although a two-sample Mendelian randomization (MR) study evaluated the overall relationships between GM and KOA, the findings were inconclusive. 20 The complexity and diversity of microbiome composition suggest that it may be inappropriate to consider the gut microbiome as a single entity. Interestingly, research suggests that GM may influence KOA by altering BM levels. 21 Considering the potential links between GM, BMs, and KOA, we examined the causal relationships among these factors to identify potential targets for risk stratification and to clarify the underlying mechanism of KOA for further study.
Methods
Study design
Figure 1 presents the research design diagram. We performed a large two-step, two-sample MR study to examine the association between the genetically determined GM, BMs, and KOA. Initially, we performed MR analysis to examine the following causal associations: (i) GM and KOA and (ii) BMs and KOA. Additionally, to further examine whether GM can impact the onset of KOA through metabolic pathways, we carried out a second-step MR analysis. This analysis focused on the GM species and BMs identified as causally linked to KOA. The MR study design requires the fulfillment of three key assumptions: (i) the instrumental variables (IVs) must be significantly related to GM and BMs (association assumption); (ii) the IVs must be independent of any confounders between GM, BMs, and KOA (independence assumption); and (iii) the IVs must affect KOA only through their effects on GM and BMs, without any direct pathways (exclusivity assumption). This research adhered to the Strengthening the Reporting of Observational Studies in Epidemiology guidelines.
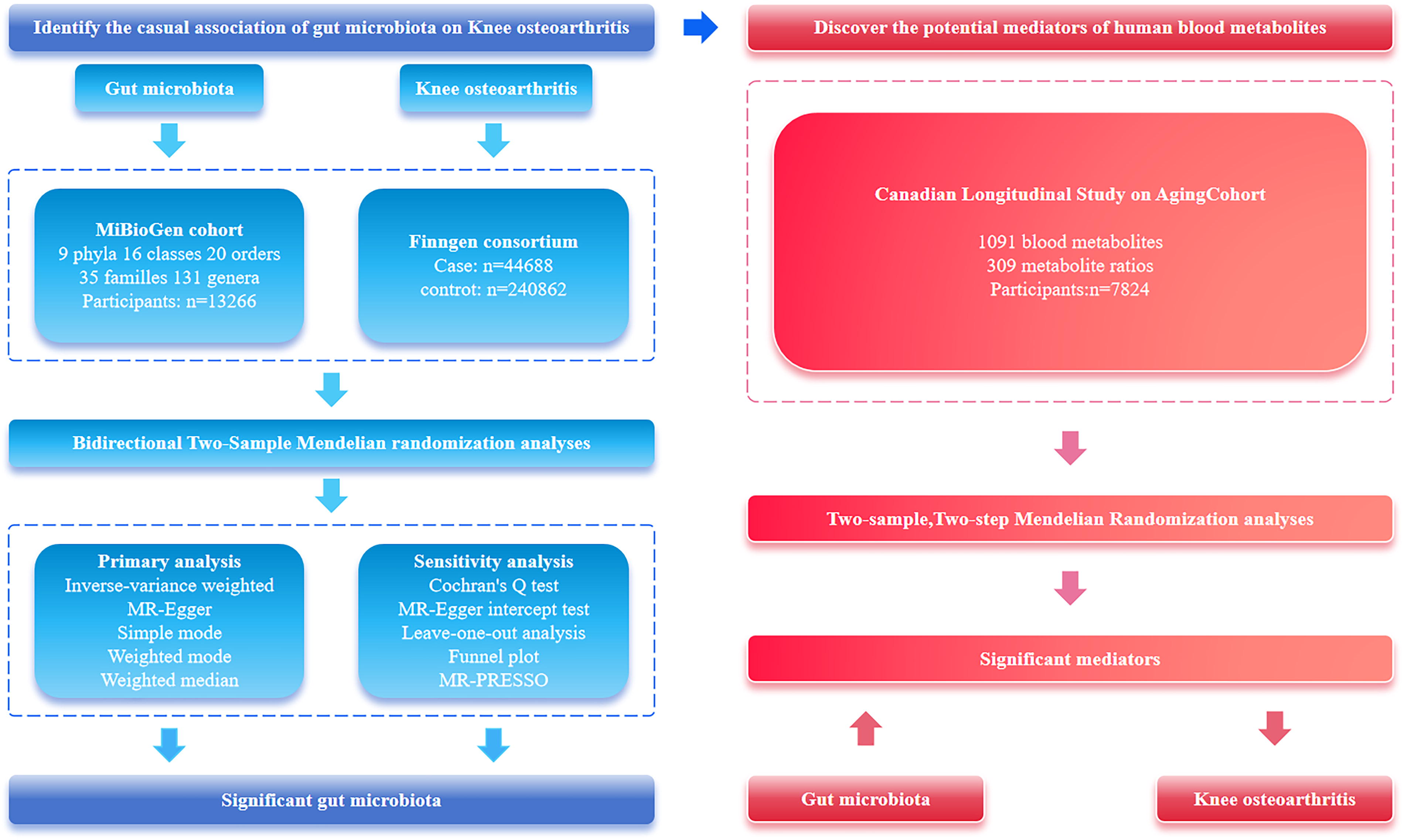
Study design of the two-stage Mendelian randomization analysis.
Data sources
GWAS data were acquired from the MiBioGen consortium (https://mibiogen.gcc.rug.nl). The MiBioGen project aims to explore the causal relationships between human genetics and GM compositions and represents the most extensive multiethnic GWAS in this field to date. 19 The consortium analyzed genotyping data from 13,266 subjects of European descent, alongside 16S rRNA gene sequencing profiles of their GM, identifying 122,110 genetic loci. Following the exclusion of unidentifiable taxa, GWAS data for 211 GM species were acquired, covering 131 genera, 35 families, 16 classes, 20 orders, and 9 phyla.
GWAS data on BMs were retrieved from the Metabolomics-GWAS-Server (https://metabolomics.helmholtz-muenchen.de/gwas/), including data on 1091 BMs and 309 metabolite ratios. Among these metabolites, 107 remain unidentified due to unknown chemical properties, while the others have been chemically categorized into 8 major groups of metabolic pathways: energy, cofactors and vitamins, amino acids, nucleotides, carbohydrates, peptides, lipids, and xenobiotic metabolism. This dataset is the largest species-level genomics study of human BMs to date, involving 7824 participants of European descent. 22
The summary data for KOA were provided by the FinnGen consortium under the dataset identifier “finngen_R9_M13_ARTHROSIS_KNEE.” This dataset included 285,500 samples (44,688 cases and 240,862 controls), with a total of 20,167,485 single-nucleotide polymorphisms (SNPs) genotyped. Further details are available on the FinnGen project website (https://www.finngen.fi/en/). Additional information on each GWAS data source can be found in Supplementary Table S1.
Selection of genetic IVs
The fundamental aspect of MR studies is the use of SNPs as IVs. To ensure the selection of optimal IVs, several rigorous quality control steps were implemented. First, SNPs were chosen based on a strong relationship with the exposure (p < 1 × 10−5) and no linkage disequilibrium (r 2 < 0.001 and clump window >10,000 kb). 23,24 Second, F-statistics were calculated using the formula: F = R 2 × (n − 1 − k)/([1 − R 2] × k), where R 2 represents the proportion of variance explained by the instrument, N is the sample size, and K is the number of SNPs included in the instrument. An F-statistic greater than 10 indicates sufficient statistical power. 25 Third, SNPs with a minor-allele frequency of ≤0.01 were excluded. 26 Additionally, SNPs were harmonized between exposure and outcome datasets, removing those with palindromic sequences and allelic inconsistencies (e.g., C/T vs. C/A). The finalized set of SNPs was then used for MR analysis.
Statistical analysis
To determine the causal relationships between GM, BMs, and KOA, a two-sample MR analysis was conducted. This research utilized five distinct MR approaches to evaluate the causal effects of GM and BMs on KOA, each approach operating under different assumptions for horizontal pleiotropy. The inverse-variance weighted (IVW) approach, known for its robustness and widespread use in MR studies, was employed to minimize bias due to heterogeneity. 27 Therefore, IVW-based estimates were used to identify GMs and BMs that have a causal influence on KOA.
In addition, complementary analyses were performed using the weighted median, weighted mode, simple mode, and MR-Egger methods. MR-Egger regression was specifically applied to adjust the IVW method for pleiotropy and correct for pleiotropic bias. 28 The weighted median approach provided robust MR estimates when <50% of the IVs exhibited pleiotropic effects. 29 While the simple mode method is less powerful than IVW, it is robust against pleiotropy. The weighted mode method, however, can be sensitive to challenges related to bandwidth selection for mode estimation.
To assess the mediating effect of BMs, we first calculated the total effect (β_all) of GM on KOA using the IVW method. The mediating effect (β1β2) was then derived as the product of the beta coefficient for GM on BMs (β1) and the beta coefficient for GM on KOA (β2). The mediation ratio was expressed as β1β2/β_all, quantifying the extent to which BMs mediate the causal pathway between GM and KOA.
Statistical analysis and data visualization were conducted using the R software (version 4.0.2), TwoSampleMR (version 0.5.6), 30 MR Pleiotropy RESidual Sum and Outlier (MR-PRESSO), 31 and forestploter packages.
Sensitivity analysis
Sensitivity analyses were performed to verify the reliability of the results and to address any possible breaches of the MR assumptions. First, horizontal pleiotropy was examined using the Egger intercept and the MR-PRESSO approaches, which are based on SNP-level data and global heterogeneity estimates. 32 To evaluate the heterogeneity of the MR analysis, the Cochran’s Q test was used, with p < 0.05 meaning significant heterogeneity. 33 Next, leave-one-out (LOO) analysis was carried out to examine the stability of the results. This involved removing each SNP one by one and re-executing the MR analysis to determine if the overall findings were unduly affected by any single SNP. 34 A funnel plot was also created to detect and visualize potential publication bias.
To account for multiple comparisons and hypothesis testing, we applied the false discovery rate (FDR) correction using the Benjamini–Hochberg procedure. A p value <0.05 was considered suggestive of causality, whereas an FDR-adjusted p value <0.05 provided confirmatory evidence of association. This approach minimizes false positives while preserving statistical power to detect meaningful associations. 35
Results
Genetic IV selection results
After filtering for strong correlations (p < 1 × 10−5) and ensuring independence (r 2 < 0.001), significant and independent SNPs were included in the analysis. This process resulted in the selection of 2033 SNPs as IVs for 211 GM species and 34,930 SNPs as IVs for 1400 BMs. Each SNP related to GM and BMs had an F-statistic >10, suggesting a low risk of weak instrument bias influencing the results.
Association between GM and KOA
The MR analysis identified substantial causal associations between KOA and seven intestinal species included in the study, with genetic assessments indicating either positive or negative associations (p < 0.05). At the genus level, IVW estimations demonstrated that higher genetically predicted levels of Anaerostipes (Odds ratio (OR): 1.09, 95% CI: 1.01–1.18, p = 0.026; FDR = 0.046), Coprococcus3 (OR: 1.16, 95% CI: 1.00–1.35, p = 0.039; FDR = 0.048), FamilyXIIIAD3011 group (OR: 1.08, 95% CI: 1.00–1.16, p = 0.046; FDR = 0.048), Ruminococcaceae UCG002 (OR: 1.06, 95% CI: 1.00–1.13, p = 0.034; FDR = 0.047), and Ruminococcaceae UCG005 (OR: 1.11, 95% CI: 1.02–1.21, p = 0.011; FDR = 0.036) were related to higher KOA risk. Conversely, an increased genetic predisposition toward Ruminococcaceae UCG003 (OR: 0.89, 95% CI: 0.80–0.99, p = 0.032; FDR = 0.048) was associated with lower KOA risk. At the family level, the genetically predicted higher abundance of Peptococcaceae (OR: 0.89, 95% CI: 0.82–0.96, p = 0.002; FDR = 0.012) was markedly related to lower KOA risk. The robustness of the MR effect estimates was further confirmed by complementary methods. Figure 2 displays scatter plots of the causal estimates, while Figure 3 presents a forest plot depicting the causal relationships between GM and KOA as determined by the MR analysis.
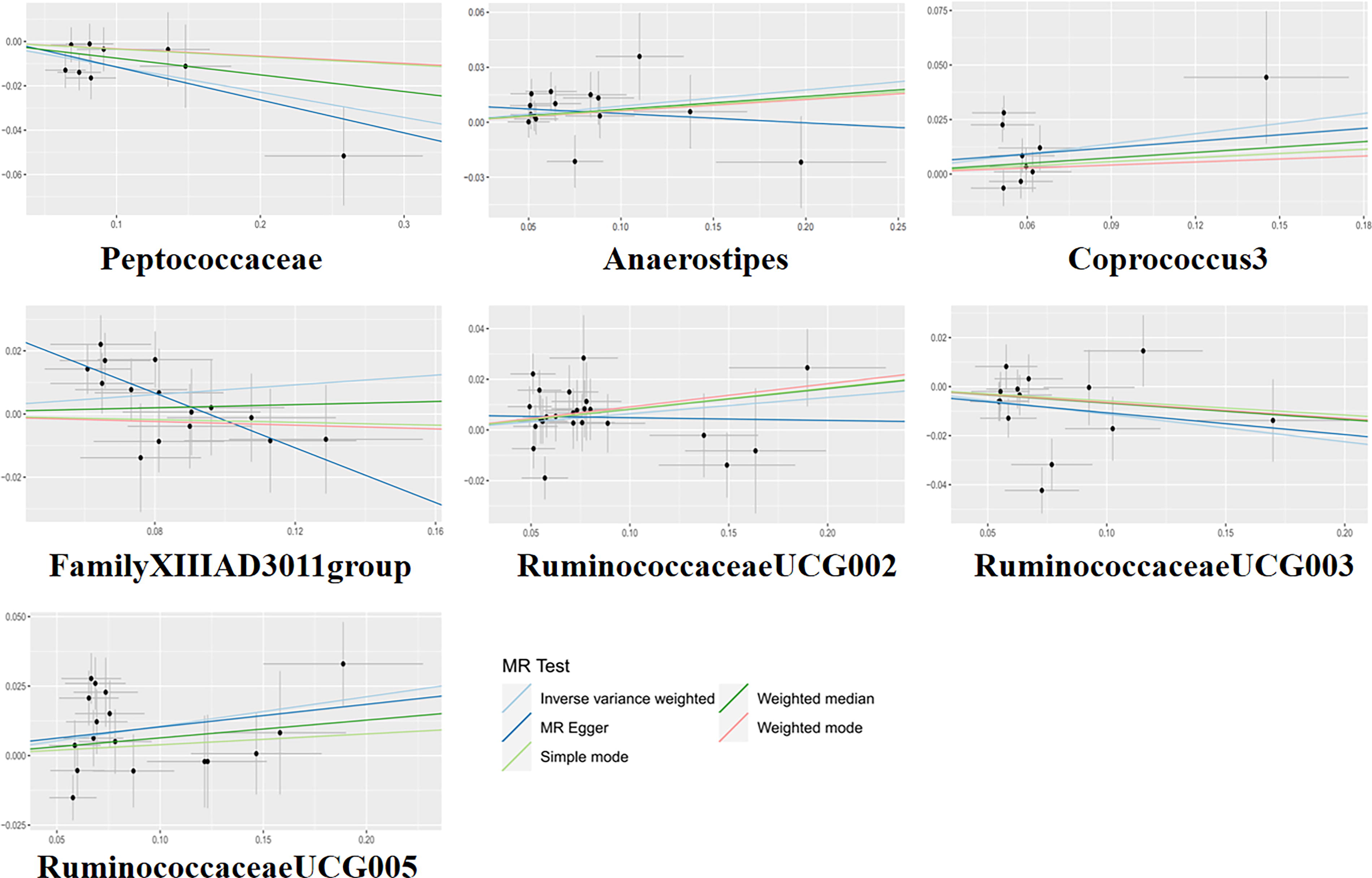
The scatter plots for the causal association between GM and KOA. GM, gut microbiota; KOA, knee osteoarthritis.
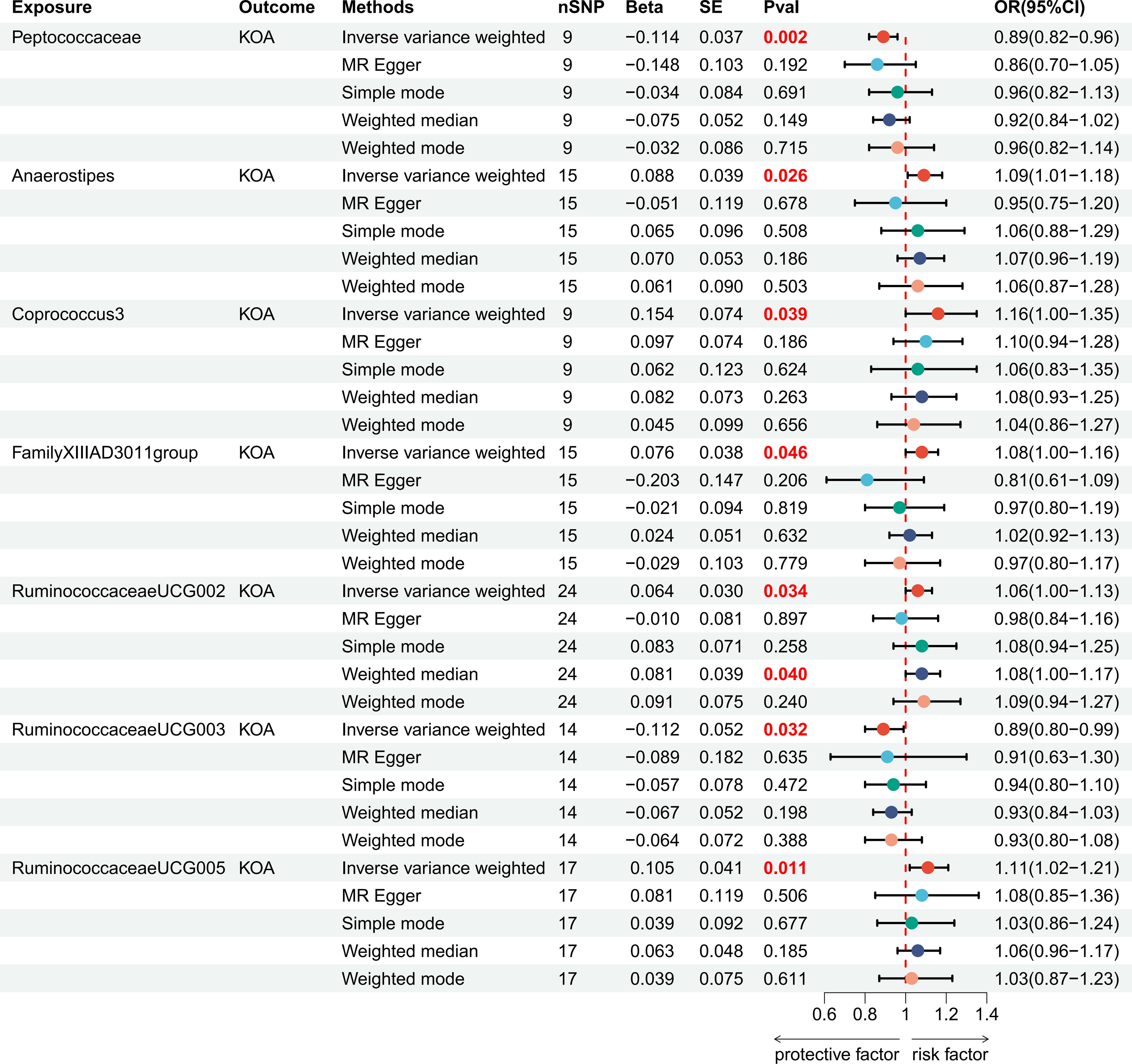
Forest plot of the associations between genetically determined GM with the risks of KOA.
Importantly, reverse MR analysis did not identify any correlations between KOA and GM, suggesting that our MR results may not be affected by reverse causation. Cochran’s Q test indicated no evidence of heterogeneity, while both MR-Egger intercept and MR-PRESSO global tests revealed no indication of directional pleiotropy bias. Moreover, LOO analysis did not detect any SNP outliers, further validating the consistency and durability of the MR analysis. The funnel plots for all GM indicators were symmetrical, indicating a low likelihood of publication bias in this study. Comprehensive data from the reverse MR and sensitivity analyses can be found in Supplementary Tables S2 and S3 and Supplementary Figures S1 and Figure S2.
Association between BMs and KOA
Of the 1400 BMs analyzed, 13 were identified with suggestive associations with KOA risk. This includes 11 metabolites with known chemical identities and 2 with unknown identities. Based on IVW estimates, increased abundances of 4-guanidinobutanoate (OR: 1.04, 95% CI: 1.00–1.08, p = 0.022; FDR = 0.046), N-acetylglycine (OR: 1.07, 95% CI: 1.03–1.11, p = 0.001; FDR = 0.003), 4-hydroxy-2-oxoglutaric acid (OR: 1.05, 95% CI: 1.01–1.10, p = 0.008; FDR = 0.034), N-formylanthranilic acid (OR: 1.04, 95% CI: 1.01–1.07, p = 0.006; FDR = 0.030), 2-methylserine (OR: 1.02, 95% CI: 1.00–1.04, p = 0.001; FDR = 0.012), sphingadienine (OR: 1.04, 95% CI: 1.00–1.08, p = 0.038; FDR = 0.048), sucrose (OR: 1.03, 95% CI: 1.00–1.07, p = 0.023; FDR = 0.046), and glycine (OR: 1.05, 95% CI: 1.01–1.08, p = 0.001; FDR = 0.012) were found to be associated with higher KOA risk. On the contrary, higher levels of 6-hydroxyindole sulfate (OR: 0.93, 95% CI: 0.88–0.99, p = 0.025; FDR = 0.046), 4-hydroxyphenylpyruvate (OR: 0.96, 95% CI: 0.92–0.99, p = 0.048; FDR = 0.049), xylose (OR: 0.95, 95% CI: 0.91–0.99, p = 0.041; FDR = 0.048), X-12707 (OR: 0.95, 95% CI: 0.92–0.99, p = 0.017; FDR = 0.044), and X-25109 (OR: 0.95, 95% CI: 0.91–0.99, p = 0.036; FDR = 0.048) were protective against KOA. Furthermore, reverse MR analysis indicated no causal relationships between KOA and these BMs.
Complementary methods and sensitivity analyses demonstrated an absence of horizontal pleiotropy or outliers, indicating the stability and robustness of this MR study. These 13 metabolites warrant further investigation. The forest plots summarizing these results are presented in Figure 4, while the outcomes of the complementary methods, reverse MR, and sensitivity tests are demonstrated in Supplementary Tables S4, S5, and S6.
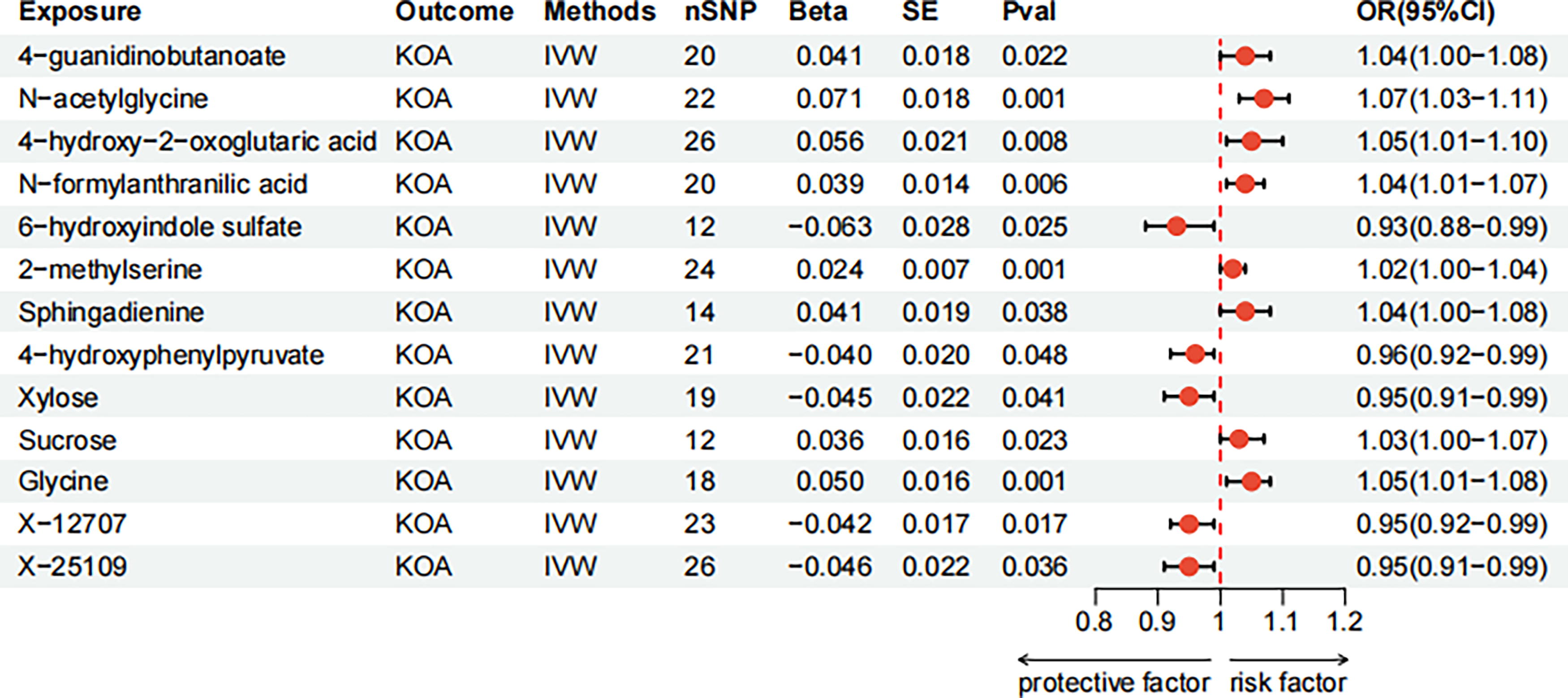
Forest plot of the associations between genetically determined blood metabolites with the risks of KOA.
Mediated MR analysis
Building on the GM and BMs previously identified as having significant causal relationships with KOA, we conducted further MR analysis. In this analysis, GMs were treated as exposures and BMs as outcomes to establish their associations. The MR analysis indicated no obvious relationship between five of the GM species (Peptococcaceae, Anaerostipes, FamilyXIIIAD3011 group, Ruminococcaceae UCG002, and Ruminococcaceae UCG005) and the BMs. Notably, we identified three causal relationships between two GMs and three BMs. Specifically, the IVW approach demonstrated a causal association between Coprococcus 3 and 2-methylserine (OR: 0.80, 95% CI: 0.64–0.99, p = 0.041; FDR = 0.048). Moreover, Ruminococcaceae UCG003 exhibited a significant causal relationship with N-formylanthranilic acid (OR: 0.82, 95% CI: 0.70–0.96, p = 0.017; FDR = 0.044) and 6-hydroxyindole sulfate (OR: 0.83, 95% CI: 0.69–0.99, p = 0.045; FDR = 0.048). Complementary methods and sensitivity analyses confirmed the reliability of these MR findings, with an absence of unbalanced horizontal pleiotropy or heterogeneity. Comprehensive data of the MR and sensitivity testing are provided in Supplementary Tables S7 and S8.
Calculation of the potential mediation effect
In the mediation analysis, we excluded factors that were not causally influenced by GM or did not exhibit a causal effect on KOA. Ultimately, our findings identified 2-methylserine, N-formylanthranilic acid, and 6-hydroxyindole sulfate as significant mediators of the association between GM-related traits and KOA. To quantify the mediation effect, we computed the total effect (β_all) of GM on KOA, the effect of GM on BMs (β1), and the effect of BMs on KOA (β2). The indirect effect was determined as the product of β1 and β2 (β1β2), and the mediation proportion was calculated by dividing the indirect effect by the total effect. The results indicated that three BMs mediated the relationship between two GMs and KOA (Fig. 5). Among them, 6-hydroxyindole sulfate exhibited the highest mediation effect, accounting for 10.26% of the total effect in the inhibition of KOA progression by Ruminococcaceae UCG003. This was followed by N-formylanthranilic acid (6.55%) and 2-methylserine (3.42%).
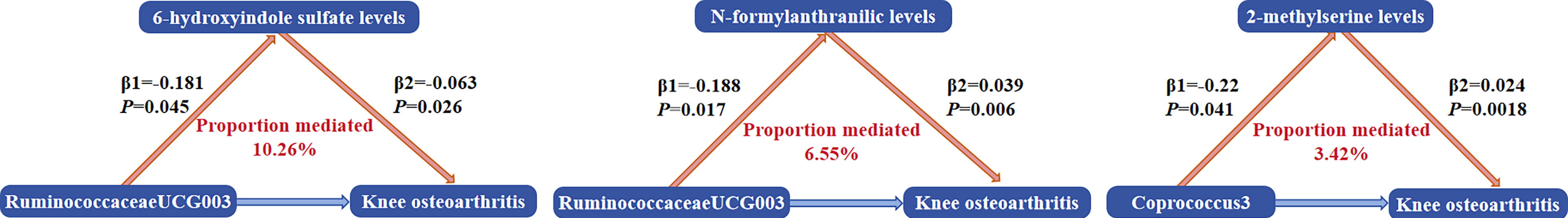
The proportions of each significant blood metabolite mediating from the corresponding gut microbiota to KOA.
Discussion
Numerous studies have revealed a potential connection between GM and KOA risk. 36,37 The gut–bone axis, serving as a “link” between bone metabolism and the gastrointestinal tract, can improve bone metabolism by releasing estrogen-like substances, serotonin, and other small molecules through GM, as well as regulating calcium and phosphorus metabolism. 38 These findings raise the possibility that the microbiome may influence the onset and progression of KOA. Nonetheless, the precise role of GM in the pathophysiology of KOA is still being explored.
In this research, MR analysis was carried out to elucidate the causal associations among GM, BMs, and KOA. The use of MR analysis is particularly valuable as it minimizes the likelihood of confounding factors that often complicate observational studies. To the best of our knowledge, this is the first study to explore the potential role of metabolite traits in mediating the causal pathway between GM and KOA. Through a comprehensive, large-scale MR analysis, we identified N-formylanthranilic acid and 2-methylserine as mediators of the causal influence of Ruminococcaceae UCG003 and Coprococcus 3 on KOA, respectively. Additionally, 6-hydroxyindole sulfate was found to mediate the pathway from Ruminococcaceae UCG003 to KOA.
The association between Ruminococcaceae UCG003 and a reduced risk of KOA is consistent with previous research 39 suggesting that Ruminococcaceae family members are typically advantageous for gut health, possibly due to their role in producing short-chain fatty acids (SCFAs) such as butyrate, which exhibit anti-inflammatory effects. 40 In contrast, the association of Coprococcus3 with a high KOA risk diverges from some studies that have linked Coprococcus species to positive health outcomes, including increased SCFA production. This inconsistency emphasizes the necessity for additional research to elucidate the context-dependent effects of these bacteria on health.
This research contributes further genetic evidence in humans connecting specific metabolic disorders to the onset of KOA. For example, we identified elevated levels of glycine and N-acetylarginine as common risk factors for KOA. 41 Glycine may influence cartilage structures and functions, while N-acetylarginine is involved in nitric oxide pathways and inflammatory reactions. The protective effects observed for metabolites such as 6-hydroxyindole sulfate further suggest that metabolic regulation plays a complex role in KOA progression. These results highlight the intricate interactions between GM and host metabolism, indicating that gut bacteria might influence KOA by modulating metabolic pathways. This could involve the production or regulation of metabolites that impact inflammation or other physiological processes relevant to KOA. For instance, some metabolites identified in this study, such as glycine and N-acetylglycine, have been associated with inflammatory processes, 42 which are critical in the pathogenesis of KOA.
The study’s findings may have various clinical implications. First, they recommend that modifying GM composition could offer a novel therapeutic approach for preventing or managing KOA. For example, probiotics, prebiotics, or dietary interventions designed to enhance beneficial bacteria such as Ruminococcaceae UCG003 might help reduce KOA risk. Additionally, monitoring BM levels could facilitate the identification of individuals at higher risk for KOA, allowing earlier and potentially more effective interventions. The study also highlights the potential for personalized medicine in KOA treatment, as tailoring interventions to address specific microbial imbalances might be more effective than generalized treatments, given the variability in GM among individuals.
Nevertheless, this research has several limitations. It did not account for environmental factors such as diet and medication use, which can impact both GM and metabolite levels. Furthermore, the exact role of the identified GM species in KOA remains speculative. Although the associations found are statistically significant, experimental validation is necessary to confirm the causal role of these bacteria in KOA pathogenesis. Future research could include germ-free mouse models or clinical trials assessing the effects of microbiota-targeted interventions on KOA outcomes.
Conclusion
Our study is the first to comprehensively assess the causal relationships among GM, BMs, and KOA. These findings highlight a potential mechanism linking GM to KOA, wherein GM influences KOA progression through BMs. These results provide new insights into microbiome-based therapies for KOA and metabolite-targeted treatment strategies.
Footnotes
Patient and Public Involvement
Patients and the public were not involved in the design, conduct, reporting, and dissemination plans of this article.
Availability of Data and Materials
The datasets used and/or analyzed during the current study are available from the corresponding author upon reasonable request.
Ethics Approval and Consent to Participate
Data were collected from public databases. And there is no ethical approval necessary.
Authors’ Contributions
Z.W., C.Z., Y.Z., and H.X.: Conceptualized and designed the study. Z.W., L.Z., J.C., and Y.W.: Provided the package codes in R language and analyzed the data in the study. J.D., M.L., and L.Z.: Gave constructive suggestions when writing the article. All authors were involved in drafting the article or revising it critically for important intellectual content, and all authors approved the final version to be published.
Author Disclosure Statement
The authors declare no competing interests.
Funding Information
This work was supported by the Henan Province Chinese Medicine Scientific Research Special Project (No. 2024ZY3060, 2024ZY2096), the Central Plains Thousand Talents Program-Central Plains Famous Doctors (No. ZYQR201912120), the 2022 Central Plains Talent Plan (Talent Education Series)-Central Plains Youth Top Talent Project (No. Yu Talent Office [2022] No. 5), the Henan Provincial International Science and Technology Cooperation Project (No. 252102521021), the Henan Provincial Science and Technology Research and Development Program (Joint Fund Project) (No. 242301420104), Henan Province Key Research and Development and Promotion Special Project (Science and Technology Research) (No. 232102311203), Henan Province Traditional Chinese Medicine “Double First-Class” Scientific Research Project (No. HSRP-DFCTCM-2023-7-09) and Henan University of Chinese Medicine Graduate Research Innovation Ability Improvement Plan Project (2023KYCX071).
Supplementary Material
Supplementary Figure S1
Supplementary Figure S2
Supplementary Figure S3
Supplementary Figure S4
Supplementary Table S1
Supplementary Table S2
Supplementary Table S3
Supplementary Table S4
Supplementary Table S5
Supplementary Table S6
Supplementary Table S7
Supplementary Table S8
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
