Abstract
Background:
The coronavirus disease 2019 (COVID-19) pandemic, first observed in December 2019 caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), has influenced every individual on the planet. The virus has influenced our lifestyle, education, economy, and the environment. Though the vaccines against COVID-19 have protected against the disease, new strains of the SARS-CoV-2 virus (e.g., Omicron BA.2.12.1, BA.4, and BA.5) have lowered the efficiency of the parent vaccines. There is still no effective therapy for the treatment of the disease. Understanding the protein structure of the virus may lead to the development of effective therapies for the disease. We recently mapped the structural proteins and non-structural proteins of SARS-CoV-2. The accessory proteins (open reading frames, Orfs) of SARS-CoV-2 modulate the host environment to favor virus replication. This paper reports mapping the accessory proteins (Orfs) of SARS-CoV-2.
Method:
Using bioinformatics, we mapped the accessory proteins (Orfs) of SARS-CoV-2.
Result:
Computational modeling predicted that the accessory proteins Orf3a, Orf7a, and Orf7b are transmembrane proteins.
Conclusion:
Bioinformatics tools were used to map the structure of the accessory proteins of SARS-CoV-2. The accessory proteins (Orfs) Orf3a, Orf7a, and Orf7b are transmembrane proteins.
Introduction
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) is the RNA virus responsible for the coronavirus disease 2019 (COVID-19). Millions of people have been infected with COVID-19 since December 2019. As of October 2022, the pandemic has resulted in more than 630 million infections and 6.5 million deaths (https://coronavirus.jhu.edu/map.html). The disease has impacted every country of our planet. The virus has infected humans, companion animals and wild animals and has led to huge economic loss.1,2
Though there are vaccines to protect against the disease, even after 3 years, there are no effective therapies for the treatment of the disease.1 Current vaccines are based on the spike protein (S). The parent COVID-19 vaccines are not effective against mutant strains of the virus (e.g., Omicron BA.2.12.1, BA.4, and BA.5); hence, there is a need to develop new and potent vaccines to protect against multiple strains of SARS-CoV-2.
It is still not understood the functions of all the proteins of SARS-CoV-2. The structural proteins of SARS-CoV-2 include membrane glycoprotein (M), envelope protein (E), nucleocapsid protein (N), and the spike protein (S). In the host cell, the Orf1ab is cleaved to form 16 nonstructural proteins (nsps) that is involved in viral replication and inhibition of innate immunity.2 We demonstrated recently that nsp3, nsp4, and nsp6 are transmembrane proteins.2
There are several accessory genes, including the Orf3a, Orf6, Orf7a, Orf7b, Orf8, and Orf10, that are interspaced among the structural proteins of SARS-CoV-2. The accessory proteins are involved in viral replication and inhibition of host immune defenses.3 Understanding the structure of the accessory proteins may be useful to design new therapies and vaccines. The development of new strains of SARS-CoV-2 (variants of concern, VOC) warrants the development of new and effective vaccines. This paper reports mapping the accessory proteins of SARS-CoV-2 that may aid in designing novel therapies, antiviral, and vaccines to protect against COVID-19.
Materials and Methods
SARS-CoV-2 accessory protein structure
The accessory protein sequences of the SARS-CoV-2 were downloaded from the NCBI (https://www.ncbi.nlm.nih.gov/protein/) protein database. The accessory proteins of SARS-CoV-2 include ORF3a (accession No. QNH88661), ORF6 (accession No. QNH88664), ORF7a (accession No. QWW27599), ORF7b (accession No. BDB04007), ORF8 (accession No. QNH88667), and ORF10 (accession No. QNH88669).
Protein modeling
To determine the snake diagram model of the SARS-CoV-2 accessory proteins, we used Protter (http://wlab.ethz.ch/protter). For the three-dimensional homology modeling of the SARS-CoV-2 accessory proteins, we employed the iterative threading assembly refinement (I-TASSER) (https://zhanglab.ccmb.med.umich.edu/I-TASSER/) with default settings. In addition, we also used different protein modeling software including Phyre 2 (http://www.sbg.bio.ic.ac.uk/∼phyre2/html/page.cgi?id=index) and SWISS-MODEL (https://swissmodel.expasy.org/) for confirmation of the predicted protein structures. The protein sequences of SARS-CoV-2 were entered in FASTA format. We used PyMol (Schrodinger, USA) for protein visualization.
We used multiple bioinformatics software for the prediction of transmembrane regions of membrane proteins. Transmembrane helix prediction (TMHMM) (transmembrane hidden Markov model) was used for the prediction of transmembrane domains (www.cbs.dtu.dk/services/TMHMM/). Phobius was used for the prediction of transmembrane topology and signal peptides from the amino acid sequence of proteins (https://phobius.sbc.su.se/).4
Results
The evolution of VOC among the SARS-CoV-2 virus warrants the development of novel therapies and vaccines to protect against new strains of the virus. As yet there are no therapies for the treatment of COVID-19. Social distancing, wearing mask, and vaccination are the only strategies to protect against COVID-19.1,2,5 There is a need to develop new therapies and vaccines to protect against the disease.
Using bioinformatics tools, we mapped the structure of structural proteins and nsps of SARS-CoV-2. We demonstrated that the membrane (M) protein of the virus could function as a glucose transporter.1 We also demonstrated that nsp3, nsp4, and nsp6 are transmembrane proteins.2 In this paper, using bioinformatic tools, we mapped the structure of the accessory proteins (Orf3a, Orf6, Orf7a, Orf7b, Orf8, and Orf10) of SARS-CoV-2.
In our previous paper, we mapped the structure of nsps.2 We used Orf3a as a reference protein. The Orf3a is a transmembrane protein with three transmembrane domains and a long and short luminal domain jutting into the ER lumen (Fig.1). The bioinformatic tools Protter, TMHMM, and Phobius demonstrated that Orf3a has three transmembrane domains. The data confirmed that the largest accessory protein Orf3a is a transmembrane protein.
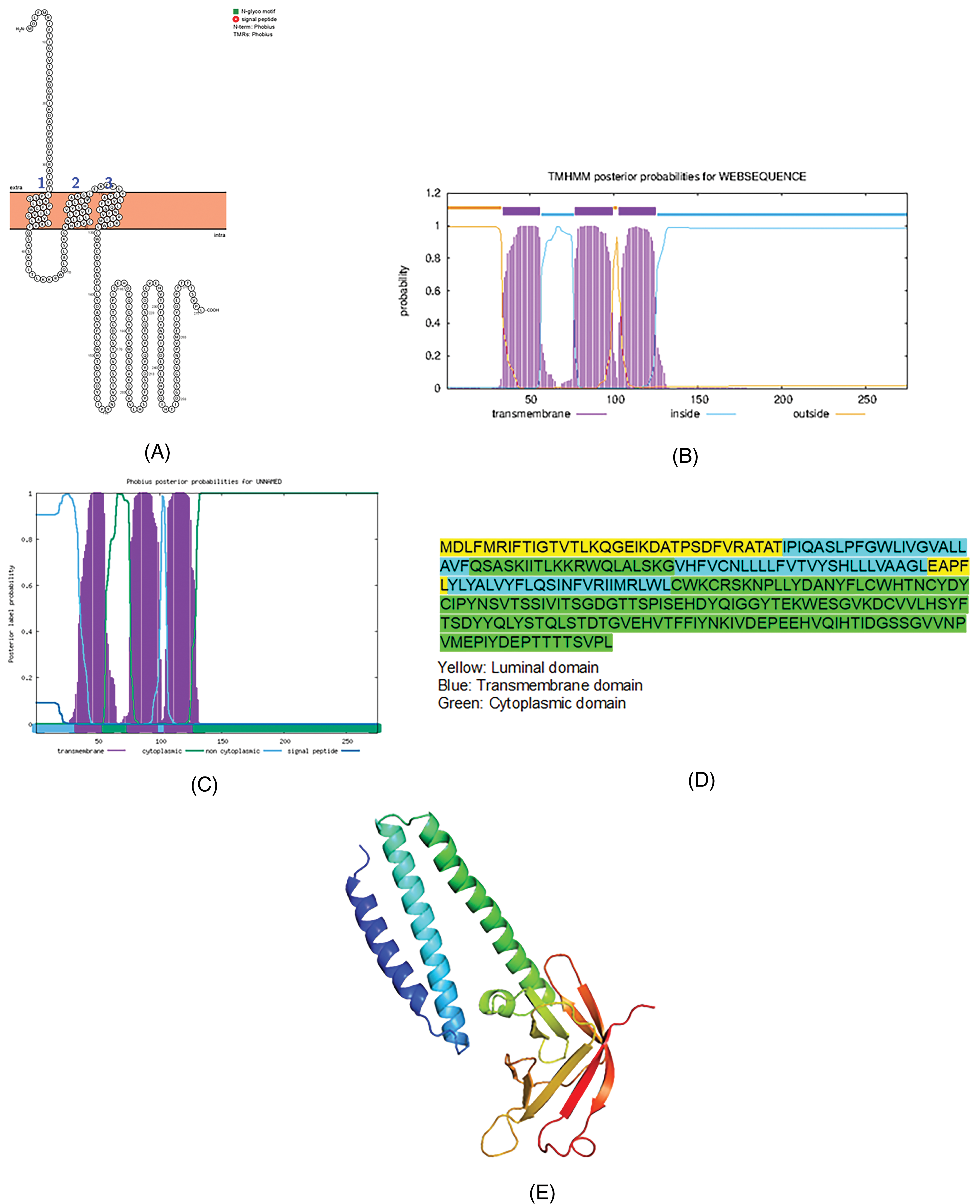
The topology of the accessory protein Orf3a of SARS-CoV-2. (A) The topology of Orf3a of SARS-CoV-2 determined using Protter. (B) TMHMM of Orf3a. (C) The topology of Orf3a as determined by Phobius database. (D) The domains (cytoplasmic, transmembrane, and luminal) of Orf3a of SARS-CoV-2. (E) The predicted Orf3a protein structure of SARS-CoV-2 (ribbon diagram) determined using the software Phyre2 and visualized by PyMol. The Orf3a figure was used as a model in our previous paper2 for reference purposes.
Of the protein modeling software used, Phyre2 and I-TASSER provided better protein models. In this manuscript, we use the data retrieved from Phyre-2. The bioinformatic tools predicted that the accessory protein Orf6 is a cytosolic protein and not a transmembrane protein (Fig.2).
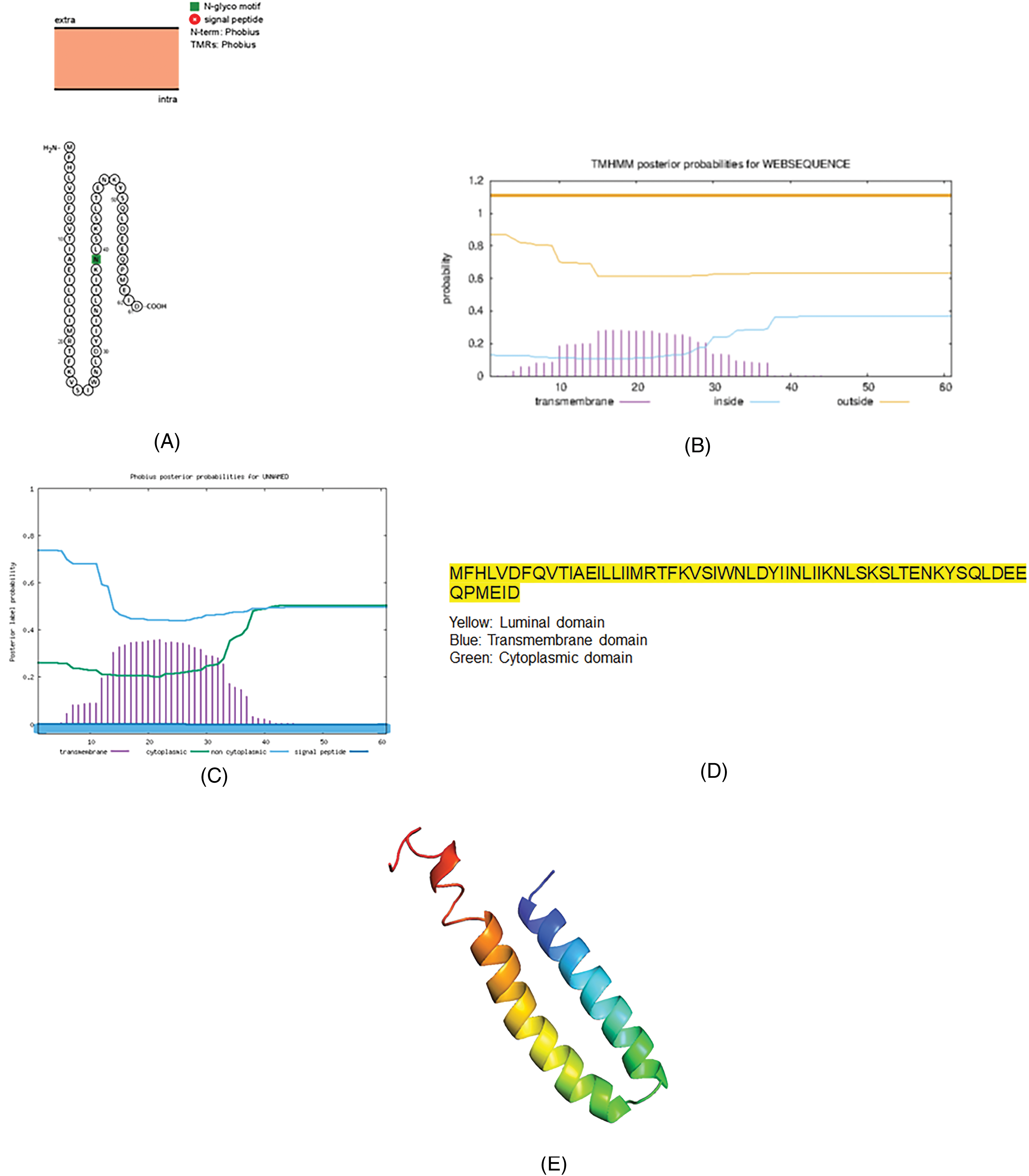
The topology of the accessory protein Orf6 of SARS-CoV-2. (A) The topology of Orf6 of SARS-CoV-2 determined using Protter. (B) TMHMM of Orf6. (C) The topology of Orf6 as determined by the Phobius database. (D) The domains (cytoplasmic, transmembrane, and luminal) of Orf6 of SARS-CoV-2. (E) The predicted Orf6 protein structure of SARS-CoV-2 (ribbon diagram) determined using the software Phyre2 and visualized by PyMol.
Orf7a is predicted to be a transmembrane protein with a signal peptide at the N-terminus. Orf7a has a single transmembrane domain, a long N-terminal domain (NTD), and a small C-terminal domain (Fig.3).

The topology of the accessory protein Orf7a of SARS-CoV-2. (A) The topology of Orf7a of SARS-CoV-2 determined using Protter. (B) TMHMM of Orf7a. (C) The topology of Orf7a as determined by the Phobius database. (D) The domains (cytoplasmic, transmembrane, and luminal) of Orf7a of SARS-CoV-2. (E) The predicted Orf7a protein structure of SARS-CoV-2 (ribbon diagram) determined using the software Phyre2 and visualized by PyMol.
The accessory protein, Orf7b, is the smallest transmembrane protein, as predicted by bioinformatics (Fig.4). All the bioinformatics software has predicted Orf7b as a transmembrane protein.

The topology of the accessory protein Orf7b of SARS-CoV-2. (A) The topology of Orf7b of SARS-CoV-2 determined using Protter. (B) TMHMM of Orf7b. (C) The topology of Orf7b as determined by the Phobius database (D) The domains (cytoplasmic, transmembrane, and luminal) of Orf7b of SARS-CoV-2. (E) The predicted Orf7b protein structure of SARS-CoV-2 (ribbon diagram) determined using the software Phyre2 and visualized by PyMol.
Orf8 is not a transmembrane protein as predicted by different bioinformatics software. The protein has a signal peptide at the NTD (Fig.5).
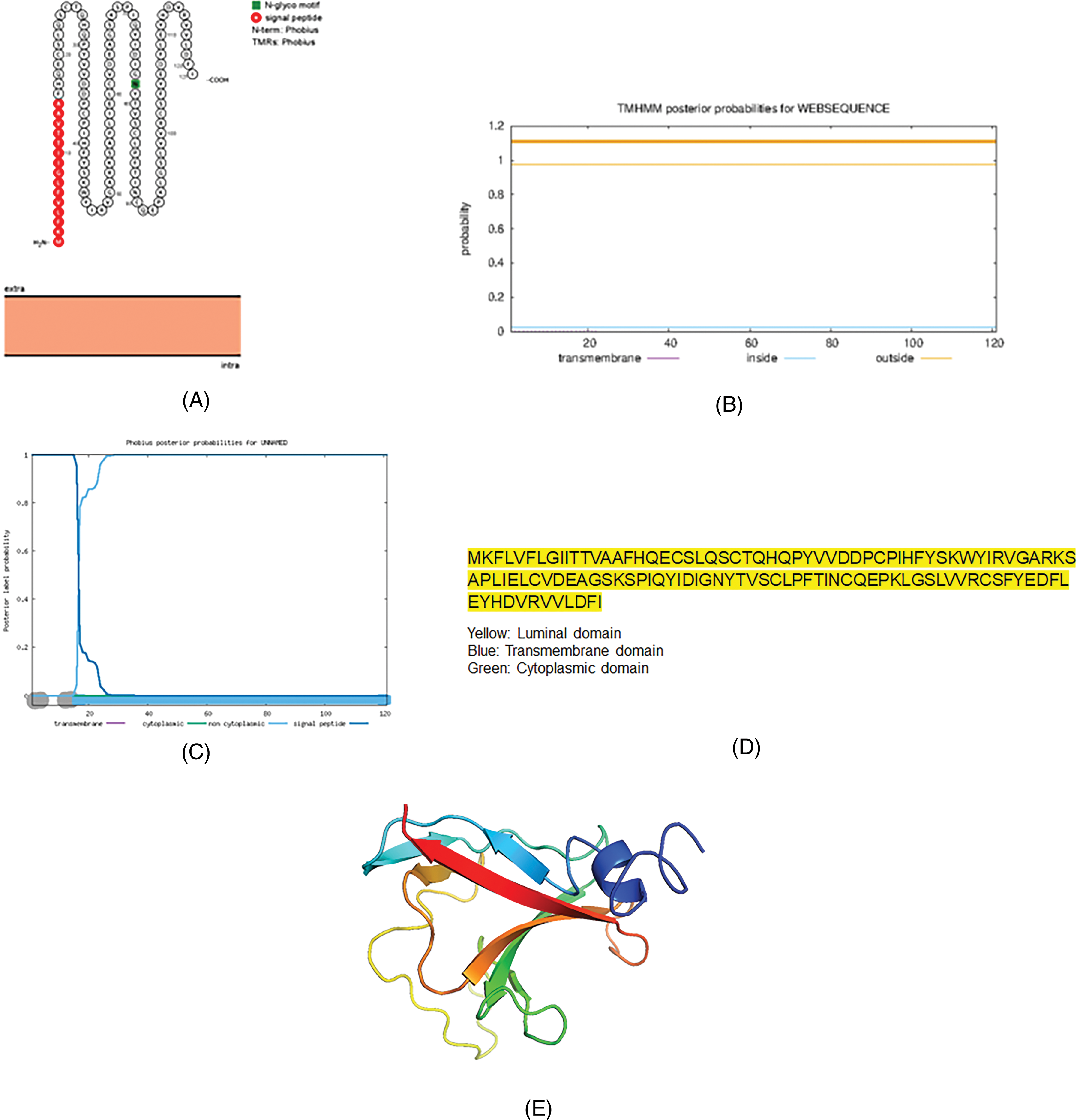
The topology of the accessory protein Orf8 of SARS-CoV-2. (A) The topology of Orf8 of SARS-CoV-2 determined using Protter. (B) TMHMM of Orf8. (C) The topology of Orf8 as determined by Phobius database. (D) The domains (cytoplasmic, transmembrane, and luminal) of Orf8 of SARS-CoV-2. (E) The predicted Orf8 protein structure of SARS-CoV-2 (ribbon diagram) determined using the software Phyre2 and visualized by PyMol.
The final accessory protein, Orf10, is also not a transmembrane protein. It is the smallest accessory protein (Fig.6).
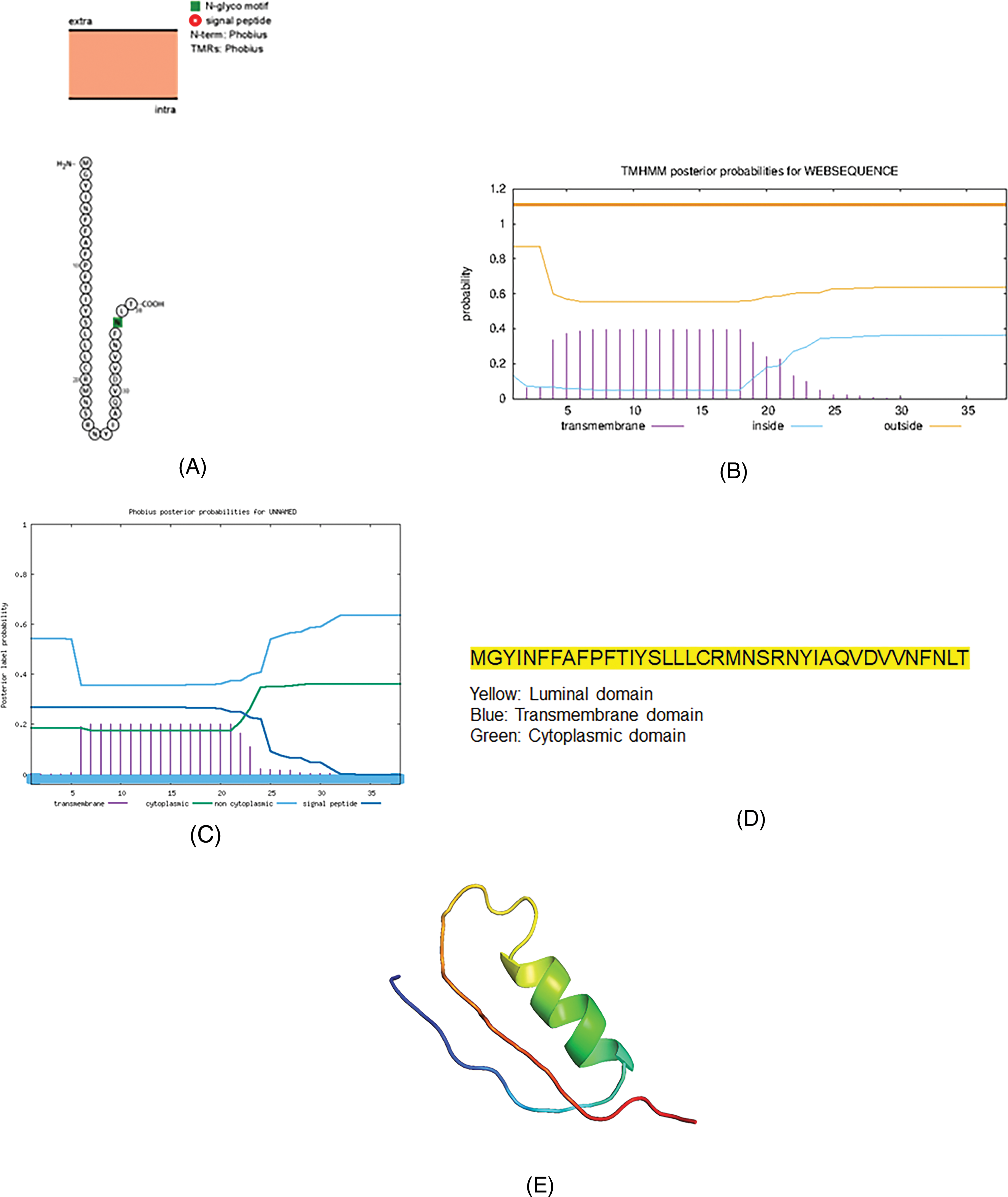
The topology of the accessory protein Orf10 of SARS-CoV-2. (A) The topology of Orf10 of SARS-CoV-2 determined using Protter. (B) TMHMM of Orf10. (C) The topology of Orf10 as determined by Phobius database. (D) The domains (cytoplasmic, transmembrane, and luminal) of Orf10 of SARS-CoV-2. (E) The predicted Orf10 protein structure of SARS-CoV-2 (ribbon diagram) determined using the software Phyre2 and visualized by PyMol.
Our study demonstrates that the accessory proteins Orf3a, Orf7a, and Orf7b are transmembrane proteins.
Discussion
The COVID-19 disease is the first pandemic disease of the twenty-first century that has impacted every family of our planet. The disease is also observed in companion animals and wild animals.5 The pandemic has resulted in travel bans, change in lifestyle, ban on crowding, maintaining safe distance, and wearing mask while traveling outside home. The disease has negatively impacted the economy of all the countries.1,5 The development of vaccines against the disease has provided relief; however, arise of VOC of SARS-SoV-2 warrants the development of new therapies and vaccines to protect against the virus. It is still not understood why SARS-CoV-2 turned out to be a successful virus causing pandemic.
The bioinformatics software available online are important tools that aid in protein modeling and to a limited extend unravel the function. We used the tools to determine the protein structure of the structural proteins S, M, E, and N.1 We demonstrated that the M protein resembles a glucose transporter.1 Later, we modeled the nsps of SARS-CoV-2. We showed that nsp3, nsp4, and nsp6 are transmembrane proteins.2 In this paper, using bioinformatics tools, we model the accessory proteins (Orfs) of SARS-CoV-2. We demonstrate that the SARS-CoV-2 accessory proteins Orf3a, Orf7a, and Orf7b are transmembrane proteins.
The role of SARS-CoV-2 accessory proteins in viral pathogenesis and viral replication is not completely understood.3 The accessory proteins work as antagonists of interferon (IFN)-signaling during viral infection, viralpathogenesis, apoptotic inducers, or antiviral suppressors.3
The accessory protein ORF3a induces cell death through apoptosis, necrosis, and pyroptosis leading to tissue damage of the host.2,6 In addition, ORF3a could trigger cytokine storm to promote pro-inflammatory cytokines and chemokines.6,7 Our study demonstrated that Orf3a is a transmembrane protein. ORF3a is a viroporin that activates the NLRP3-inflammasome that contributes to virus release.8,9 The structure of ORF3a is similar to the M protein.6,8 The authors are of the opinion that Orf3a has a shared ancestral origin.
ORF6 proteins can block IFN signaling10–12 and can act as a virulence factor that modulates nucleocytoplasmic trafficking to accelerate viral replication, progressing to disease.12 Lee etal.10 reported that Orf6 is localized to the endoplasmic reticulum, autophagosome, and lysosomal membranes. Wong etal.13 were of the opinion that ORF6 is a peripheral membrane protein, as opposed to being a transmembrane protein. Our study also confirms that Orf6 is not a transmembrane protein.
ORF7a is an immunomodulating factor for immune cell binding and triggers dramatic inflammatory responses.14 ORF7a efficiently binds to CD14+ monocytes and contributes to the recruitment of monocytes to infected lungs during COVID-19. ORF7a may suppress the antigen-presenting ability of these monocytes.14 Our study demonstrates that Orf7a is a transmembrane protein.
The accessory protein ORF7b promotes the expression of inflammatory cytokines that may induce apoptosis.15 ORF7b interferes with important cellular processes that involve leucine-zipper formation and contributes to heart arrythmias, odor loss, impaired oxygen uptake, and intestinal dysfunction.16 Our study using bioinformatics tools demonstrates that Orf7b is a transmembrane protein.
Orf8 is an accessory protein that has been proposed to interfere with immune responses.17 Orf8 promotes the expression of pro-inflammatory factors thereby acting as a contributing factor to cytokine storm during COVID-19 infection. Orf8 is also not a transmembrane protein as determined by bioinformatics analysis.
The accessory protein, ORF10, suppresses the expression of type I IFN (IFN-I) genes and IFN-stimulated genes.18 ORF10 is known to impair cilia function, thereby leading to loss of smell and taste that are symptoms of COVID-19.19 Hassan etal.20 predicted that Orf10 is a noncytoplasmic protein. Our bioinformatics analysis also confirms that Orf10 is a noncytoplasmic protein.
A feature of the accessory proteins is the presence of the amino acid methionine (M) at the N-terminus of each protein. The M is also found in the N-terminus of all the structural proteins of SARS-CoV-2.1 Of the nsp, only NSP-1 has M at the N-terminus; majority of the nsps have alanine (A) or serine (S) at the N-terminus.
The Omicron variant of concern has 37 mutations in the spike protein, which is responsible for host cell entry. Most of these mutations are in two domains: the receptor-binding domain (RBD) and the NTD.21 The SARS-CoV-2 Omicron variant has higher affinity for human angiotensin-converting enzyme 2 (ACE2) than the Delta variant due to a significant number of mutations in the SARS-CoV-2 RBD. Based on docking studies, the Q493R, N501Y, S371L, S373P, S375F, Q498R, and T478K mutations contribute significantly to high binding affinity with human ACE2.22
Mutations in the accessory genes (ORFs) of SARS-CoV-2 contribute to pathogenesis in the host through interference with innate immune signaling.23 However, it is clearly not understood the mutations in the accessory proteins of Omicron variant of SARS-CoV-2.
Overall, our bioinformatic analyses demonstrate that the accessory proteins Orf3a, Orf7a, and Orf7b are transmembrane proteins. Our protein modeling could lead to the development of better therapies and vaccines.
Footnotes
Acknowledgments
The author acknowledges Abraham Thomas Foundation and Women’s Board of Lankenau for providing the resources for this work.
