Abstract
A One Health lens is increasingly significant to address the intertwined challenges in planetary health concerned with the health of humans, nonhuman animals, plants, and ecosystems. A One Health approach can benefit the public health systems in Africa that are overburdened by noncommunicable, infectious, and environmental diseases. Notably, the COVID-19 pandemic revealed the previously overlooked two-fold importance of pharmacogenetics (PGx), for individually tailored treatment of noncommunicable diseases and environmental pathogens. For example, dyslipidemia, a common cardiometabolic risk factor, has been identified as an independent COVID-19 severity risk factor. Observational data suggest that patients with COVID-19 infection receiving lipid-lowering therapy may have better outcomes. However, among African patients, the response to these drugs varies from patient to patient, pointing to the possible contribution of genetic variation in important pharmacogenes. The PGx of lipid-lowering therapies may underlie differences in treatment responses observed among dyslipidemia patients as well as patients comorbid with COVID-19 and dyslipidemia. Genetic variations in APOE, ABCB1, CETP, CYP2C9, CYP3A4, CYP3A5, HMGCR, LDLR, NPC1L1, and SLCO1B1 genes affect the pharmacogenomics of statins, and they have individually been linked to differential responses to dyslipidemia and COVID-19 treatment. African populations are underrepresented in PGx research. This leads to poor accounting of additional diverse genetic variants that could be important in understanding interindividual and between-population variations in therapeutic responses to dyslipidemia and COVID-19. This expert review examines and synthesizes the salient and priority PGx variations, as seen through a One Health lens in Africa, to improve and inform personalized medicine in both dyslipidemia and COVID-19.
Introduction
Dyslipidemia is a metabolic imbalance that predisposes individuals to atherosclerosis and cardiovascular disease (CVD). Dyslipidemia is progressively declining in developed countries but is on the increase in middle- and low-income countries. Emergence of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) and the COVID-19 pandemic in 2020 caused a planetary health disaster marked by significant morbidity and mortality riding on existing morbidities that severely impacted global socio-political and economical dynamics (Chen et al., 2020).
As of April 2, 2023, Africa accounted for 1,25% (n = 9,518,476) of SARS-CoV-2 infections, and 2,54% (n = 175,334) of the total deaths (Fig. 1A). These figures were significantly lower than the initially predicted COVID-19 mortality rate (Soko et al., 2022). This discrepancy has been attributed to several factors, including Africa's younger population. This also helps explain why Africa has the highest global rate of noncommunicable disease mortality before the age of 70 (Fig. 1B).
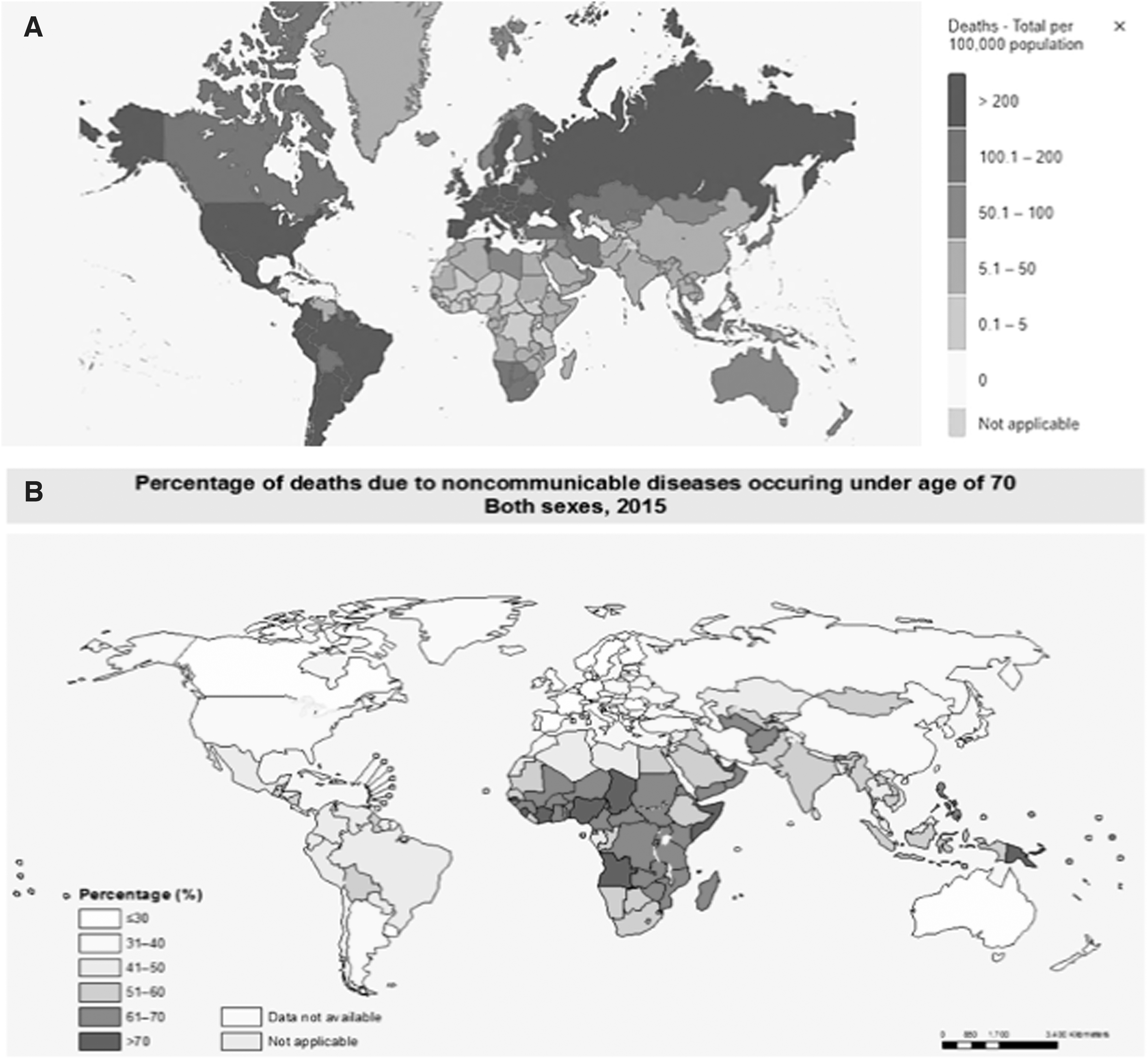
Noncommunicable diseases and their cardiometabolic risk factors are prevalent in Africa, due to rapidly changing socio-economic conditions. Comorbidity of noncommunicable disease with environmental pathogens calls for attention to the emerging literature on One Health and the interactions between the human host and the biological, social, and political ecosystems in which a person is embedded (Yetiskin, 2022).
The health of humans, nonhuman animals, plants, and planetary ecosystems is co-produced and interdependent as underlined by a One Health lens (Banerjee et al., 2023; Nissa et al., 2022). The need for integration of the health of humans and the health of the climate is also noteworthy in the contemporary context in the 21st century (Kalogirou et al., 2021).
The pooled prevalence of dyslipidemia across Africa is 52.8% (Obsa et al., 2022). Hence, many COVID-19 cases reported across Africa are likely to also have dyslipidemia as an underlying co-morbidity. Dyslipidemia was observed as an independent risk factor for severe COVID-19 infection (Wilk-Sledziewska et al., 2022), which accelerated mortality. A meta-analysis reported that patients with dyslipidemia and infected with SARS-Cov-2 had a 2.13-fold higher risk of mortality (Liu et al., 2020).
The mechanisms underlying the link between dyslipidemia and COVID-19 mortality are multifactorial, with the SARS-CoV-2 relying on membrane cholesterol to enter cells (Zapatero-Belinchón et al., 2021), and increased oxidative stress resulting in the oxidization of low-density lipoprotein (OxLDL), which contributes to the initiation and maintenance of vascular inflammatory processes that are a hallmark of COVID-19 infection (Trpkovic et al., 2015). Optimal control of cholesterol may, thus, potentially reduce the severity of CVDs associated with COVID-19 by protecting endothelial cells (Vuorio et al., 2021b).
Effective treatment of dyslipidemia is likely to reduce morbidity and mortality from CVDs. However, findings indicate that many patients are not achieving recommended lipid treatment targets (Blom et al., 2020b; Mulchandani et al., 2020). According to the CARDIO-TRACK survey from South Africa, only 33% of very high-risk patients reached their LDL-C goal of <1.8 mM on maximum-tolerated statin dose (Blom et al., 2020b), comparable to 34% among Egyptian patients (Taha et al., 2021).
HMG-CoA reductase inhibitors or statins are the first line of therapy for hypercholesterolemia, due to their affordability, accessibility, efficacy, and safety (Blom et al., 2020b; Elmekawy et al., 2021; Marcus et al., 2022). As a result, they are the most frequently prescribed lipid-lowering drugs (Mach et al., 2019). In African countries, most non-statin lipid-lowering medications remain unregistered or unaffordable (Marcus et al., 2022).
Conflicting observations seem to associate statin use with less severe illness and lower mortality from COVID-19 infection (Ayeh et al., 2021; Mbunge, 2020; Peymani et al., 2021). Thus, pharmacogenomics knowledge on dyslipidemia medications may advance precision outcomes of CVDs and COVID-19.
Pharmacogenetics (PGx) research has the potential to improve treatment outcomes through tailoring medications, including dosage and selection of drugs pre-emptively (Alfonsi et al., 2016; Dandara et al., 2014). Taking the One Health lenses, chemicals that are sprayed to agricultural products, as well as drugs given to animals, affect the way humans respond to medication due to variable exposures. For example, prescription guidelines for statin treatment formulated based on common mutations in SLCO1B1, ABCG2, and CYP2C9 pharmacogenetic status (Cooper-DeHoff et al., 2022) are likely to miss in some instances in African populations due to their premises being largely evidence from studies among non-African populations and variable ecosystem exposures that affect their effectiveness.
The two polymorphisms, ABCG2 c.421C>A (rs2231142) and SLCO1B1 c.521T>C (rs4149056), are rare in African populations (Aklillu et al., 2016; Soko et al., 2018), but important in Asian and European populations. In addition, as reported (Kuhnert et al., 2020), some pesticides such as chlorpyrifos-methyl, and tebuconazole, used in agriculture, significantly increased bovine ABCG2 (bABCG2) efflux activity compared with unexposed, further illustrating the importance of the One Health approach as this effect is also likely to happen in humans. Thus, the pharmacogenomics of statins is intrinsically linked to that of related chemicals used in agriculture that can affect the same enzyme systems.
The few PGx studies on lipid-lowering therapies conducted among African countries report on the effects of genetic variation in statin responses due to other mutations in genes such as ABCB1, ABCC2, CYP3A4, CYP3A5, CYP2C9, and SLCO1B1 (Muyambo et al., 2022; Radouani et al., 2020). The variability in dyslipidemia treatment-response observed among African populations cannot be accounted for by currently known genetic variants, indicating the need for more genetic characterization of African populations to discover contributory African-specific variants, as well as other unique environmental factors.
Indeed, a pharmacokinetics study conducted among Zimbabweans that involved whole exome sequencing discovered associations between rosuvastatin exposure and genetic variation in genes not previously prioritized for statin responses, such as ABCC2, SLC10A1, ABCB11, AHR, HNF4A, RXRA, and FOXA3 (Soko et al., 2018), findings that need further confirmation.
The regular discovery of “African-specific” variants shows that African populations are understudied, which has given “birth” to the “African Pharmacogenetic Consortium,” which seeks to promote PGx research and advocacy to unravel the extent of pharmacogenomic variation among African populations (Dandara et al., 2019).
As alluded to earlier, a One Health lens is increasingly significant to address the intertwined challenges in planetary health concerning the health of humans, nonhuman animals, plants, and ecosystems. A One Health approach can benefit the public health systems in Africa that are overburdened by noncommunicable, infectious, and environmental diseases.
The aim of the present expert review is to describe the drug profiles of three leading lipid-lowering therapies: (a) statins, (b) cholesterol absorption inhibitors, and (c) PCSK9-inhibitors; and their associated pharmacogenomics attributes. In addition, by highlighting the ways in which PGx can inform not only the treatment of dyslipidemia but also the conditions where dyslipidemia and COVID-19 are comorbid, the review contributes to a nascent body of literature on One Health and PGx.
PGx of Commonly Used Lipid-Lowering Therapies in African Populations
A literature search was conducted using the following databases: PubMed, Google Scholar, and PharmGkb. The keywords were used individually or in combination in the searches that included: “COVID-19 AND dyslipidemia,” “Pharmacogenetics AND COVID-19,” “Pharmacogenetics and One Health,” “simvastatin AND Africa,” “alirocumab AND Africa,” “Ezetimibe and Africa,” “ABCB1 and Africa.” The literature search in the earlier databases covered the timeline including and until June 15, 2023.
HMG-CoA Reductase Inhibitors
Statin use in Africa is ever-changing as new molecules and information on existing ones become available (Table 1) (Blais et al., 2021). Statins are used in the primary and secondary prevention of CVDs by lowering blood LDL-cholesterol (Marcus et al., 2022). Data from clinical trials show that for every 39 mg/dL (1.0 mM) decrease in LDL-C, there is about 22% reduction in major CVD events (Baigent et al., 2010).
List of FDA-Approved Lipid-Lowering Therapies Administered in African Countries and Their Key Pharmacogenes
HDL-C, high-density lipoprotein cholesterol; LDL-C, low-density lipoprotein cholesterol; TG, triglycerides; TC, total cholesterol; N/A, not applicable.
Statins are, therefore, essential for both the management of CVDs risk and dyslipidemia (Mach et al., 2019). Statins act by reversible, competitive inhibition of 3-hydroxyl-3-methylglutaryl-coenzyme A (HMG-CoA) reductase, the enzyme that catalyzes the conversion of HMG-CoA, the rate-determining step in the cholesterol biosynthetic pathway mevalonate (Maxwell et al., 2017).
Decreased intracellular cholesterol leads to the upregulation of LDL receptors and increased hepatocellular uptake of LDL. As more LDL is taken up into the hepatocytes, the concentration of plasma LDL-C falls. This effect is essential for CVD risk management, because LDL-C contributes to the development and progression of atherosclerotic CVD (Mach et al., 2019).
Statins have pleiotropic effects in addition to lowering lipids, which include improving endothelial function, anti-inflammation, anti-thrombotic, and anti-oxidative stress, functions that differentially impact different CVDs and COVID-19 outcomes (Vuorio et al., 2021a). For example, it has been shown that rosuvastatin use is associated with reduced hospitalization among COVID-19 patients (Israel et al., 2021).
The mechanisms by which statins may potentially influence the outcome of COVID-19 patients are not fully understood. However, it is postulated that inhibiting HMG-CoA reductase reduces the synthesis of isoprenoids, which are essential for the prenylation of signaling molecules that regulate pro-inflammatory pathways. This effectively reduces the expression of various pro-inflammatory cytokines such as IL-6, IL-8, and MCP-1 causing anti-inflammatory effects (Lefer, 2002). In addition, statins have also been demonstrated to enhance endothelial function through the production of nitric oxide, a potent vasodilator that prevents platelet aggregation, also accounting for statins' anti-thrombotic effects (Violi et al., 2013).
Statins have generally shown favorable outcomes in several viral infections, including Avian influenza, H1N1, supporting the hypothesis that they may have similar anti-viral effects on the causative agent SARS-CoV-2, the causative agent for COVID-19 (Minz et al., 2020).
The multiple, and at times conflicting effects of statins on various other comorbidities is linked to potential drug-drug interactions (DDIs), which became apparent during the COVID-19 pandemic when various drugs where trialed. DDIs have been linked to impaired statin bioavailability, through inhibition or induction (Hirota et al., 2020). For example, remdesivir, a COVID-19 FDA-approved drug, presented a high potential for DDIs because it is a substrate of CYP enzymes, and transporters such as OATP1B1 and P-glycoprotein, which participate in the disposition of statins (Ambrus et al., 2021; Deb et al., 2021).
Similarly, Paxlovid (contains nirmatrelvir and ritonavir), another FDA-approved drug for COVID-19, has the potential for DDIs, especially due to its ritonavir component that inhibits CYP3A enzymes, leading to increased plasma concentrations and a longer half-life for drugs that are substrates of CYP3A4. Hence, the FDA has precautionary warnings on the co-administration of CYP3A4 inhibitors (i.e., ritonavir) and statins due to the possibility of increased risk of statin-associated muscle symptoms (SAMS) (Law and Rudnicka, 2006).
What is not covered in this analysis is the interaction of statins with chemicals used in agriculture and poultry, as both can be substrates of pharmacogenes or through inhibition and induction, which has the effect of “blurring” phenotypes expected from purely genetic variation.
SAMS are the most common adverse effects of statins, and they are associated with most statin-intolerance or discontinuation cases (Muñoz-Blanco et al., 2022). SAMS differ in clinical manifestations and severity spanning from muscle stiffness, tenderness, or cramps with normal CPK levels (myalgia) to muscle weakness with or without elevated CPK levels (myopathy) to the most severe form of SAMS-rhabdomyolysis, which is characterized by muscle necrosis, elevated CPK levels, and rarely death (Ward et al., 2019).
However, rhabdomyolysis has been reported in a few COVID-19 patients (Anklesaria et al., 2020; Byler et al., 2021). The frequency of SAMS ranges from 1–5% in clinical trials to 7–29% in clinical practice (Vrablik et al., 2014). The disparities in these rates are likely due to varying ascertainment and the nocebo effect, as shown by the SAMSON trial, a crossover trial on side effect patterns in placebo and no treatment groups (Muñoz-Blanco et al., 2022).
Significant inter-individual variability exists among SAMS patients, which has been linked to underlying host genetics, among other factors (Cooper-DeHoff et al., 2022; Soko et al., 2018). The key pharmacogenes associated with the pharmacokinetics and pharmacodynamics of statins include ABCB1, ABCG2, APOE, CETP, CYP3A4, CYP3A5, CYP2C9, HMGCR, LDLR, PCSK9, and SLCO1B1 (Table 1).
3-Hydroxyl-3-methylglutaryl-coenzyme A reductase
The gene coding for 3-Hydroxyl-3-methylglutaryl-coenzyme A reductase, HMGCR, is very important with respect to the pharmacodynamics of statins. The gene is located on chromosome 5q13.3–14 (Sundaresan et al., 1989), and 23 exons. Alternative splicing of exon 13 in the HMGCR gene has been identified as the main molecular mechanism affecting statin response (Medina et al., 2008).
HMGCR g.27506T>G (rs17238540, referred to as SNP29) and HMGCR g.331648A>T (rs17244841, referred to as SNP12), are the most notable single nucleotide polymorphisms (SNPs) that have been associated with differential statin response (Chasman et al., 2004; Medina et al., 2011). Carriers of the rs17238540G variant present with sub-optimal responses to statins, among patients treated with pravastatin (Chasman et al., 2004). The rs17238540T variant has been associated with increased LDL-C and T/C reduction on simvastatin and pravastatin. The G/T heterozygote carriers have been reported to present with decreased lipid responses for both simvastatin and pravastatin.
Further, the rs17244841A/A genotype has been associated with increased LDL-C reduction when pravastatin is used (Chasman et al., 2004). SNP29 and SNP12, along with HMGCR g.23092A>G (rs3846662, referred to as SNP 20144) form haplotype H7 (Chasman et al., 2004) (Table 2), which is associated with poor or reduced response to simvastatin and pravastatin. SNP 20144 by itself has not been shown to be significantly associated with statin response but regulates alternative splicing of HMGCR exon 13 giving rise to the complete omission of exon 13 (Chasman et al., 2004).
Effects of Haplotypes for APOE, HMGCR, and SLCO1B1 in Statin Response
The prevalence of the rs3846662G variant allele is estimated at 6% among Africans according to the 1000 Genome Project (Martin et al., 2023). However, a study among Tunisians reported a 25% frequency for rs3846662G variant showing the extent of genetic diversity among African populations (Table 3) (Jmel et al., 2018). The HMGCR rs3846662G variant is found in complete linkage disequilibrium (D’ = 1.0; r2 = 100%) with another HMGCR SNP, rs12916C>T (Chien et al., 2010).
Distribution of Genetic Variants That Affect Lipid Lipid-Lowering Therapies
SNP, single nucleotide polymorphism.
Chien et al. (2010) also reported that rs12916C>T polymorphism reduced LDL-C lowering effects among its C/C carriers on various statins in Chinese hypercholesterolemic patients, a finding supported by observations among Indian patients on atorvastatin (Poduri et al., 2010).
A study by Medina et al. (2008) reported that the response to simvastatin-based therapy was more affected by HMGCR polymorphisms in African Americans than what was observed among their European counterparts. However, a study from Egypt recently found no significant association between rs12916T and atorvastatin response in type 2 diabetic dyslipidemia patients (Abd El-Kader et al., 2019). This study reported a 71% frequency of the variant allele of rs12916T.
An inverse correlation with severe COVID-19 and HMGCR rs12916 has been reported (Schooling et al., 2020). Another HMGCR haplotype H2 is characterized by the combination of common alleles across 11 tag SNPs (Medina et al., 2008), and it is found at a frequency of 32% among African Americans and has not yet been reported among Africans. Combined carriage of H2 and H7 is associated with decreased LDL-C lowering efficacy on statins compared with either H2 or H7 carriers alone.
Finally, HMGCR expression and HMGCR-mediated LDL cholesterol levels have been favorably linked with the incidence of COVID-19 hospitalization, indicating that HMGCR inhibition may have a protective impact against COVID-19 hospitalization (Huang et al., 2021).
Apolipoprotein E
The Apolipoprotein E (APOE) gene is found on chromosome 19q13.32 in a cluster with apolipoprotein C1 (APOC1) and apolipoprotein C2 (APOC2). It consists of 4 exons and encodes protein with 299 amino acids (Hoek et al., 2008). APOE is synthesized in the liver where it modulates lipid metabolism by acting as a ligand for low-density lipoprotein receptor (LDLR) facilitating the clearance of very low-density lipoprotein (VLDL) and chylomicron remnants (Nieminen et al., 2008). The APOE gene is highly polymorphic.
Its most common SNPs are APOE c.526 C > T (rs7412) and APOE c. 388T>C (rs429358) that result in three isoforms, namely, E2, E3, and E4 (Sacks, 2015), which are defined by the three haplotypes: e2, e3, and e4 (Table 2) (Utermann et al., 1977). The haplotypes are represented by the following genotype combinations: rs7412T-rs429358T (e2), rs7412C-rs429358T (e3), and rs7412C-rs429358C (e4) (Fullerton et al., 2000).
Homozygosity for the e2 isoform of the APOE gene is associated with familial dysbetalipoproteinemia, a type of dyslipidemia characterized by elevated plasma cholesterol and triglycerides due to impaired clearance of remnant lipoproteins (Sacks, 2015). The e4 isoform is associated with increased risk of atherosclerosis (Li et al., 2016; Phillips, 2014) and COVID-19 (Ostendorf et al., 2022), whereas e3 is considered the wild type or normal form.
All APOE isoforms are associated with varying responses to statin therapy due to their LDLR binding affinity (E4>E3>>E2) (Anoop et al., 2010) with APOE ɛ4 as major genetic risk factor for CVD among the three isoforms. There are multiple potential explanations for elevated cholesterol levels associated with carrying E4, which include reduced LDLR expression due to accelerated hepatic uptake of apoeE4-enriched VLDL.
Patient carriers of E2 present with a better response to statins compared with E4 carriers (Ordovas et al., 1995; Zhang et al., 2019; Zintzaras et al., 2009). However, the findings are inconsistent (Lagos et al., 2015). Candidate gene studies have associated E2 and E3 with increased response to atorvastatin (Ostendorf et al., 2022), pravastatin (Mega et al., 2009), rosuvastatin (Kononov et al., 2021), and simvastatin (Wanmasae et al., 2017) compared with E4 among Asian and European populations.
The Scandinavian Simvastatin Survival Study (4S trial) found that E4 carriers benefited the most from statin treatment in terms of reduced CVD mortality risk (Gerdes et al., 2000). Contrastingly, there was no correlation between APOE genotypes and statin responses in the Regression Growth Evaluation Statin Study (REGRESS), which included 815 men with CHD (Maitland-Van Der Zee et al., 2006). The distribution of the haplotypes among Africans are, on average, e2 (12%), e3 (71%), and e4 (18–22%) (Atadzhanov et al., 2014; Ndadza et al., 2021).
Cholesterol ester transferase protein
The Cholesterol Ester Transferase Protein (CETP) gene consists of 17 exons encompassing 25 kbp on chromosome 16q13 (Jap et al., 2002). It produces the CETP protein that transfers cholesterol esters from high-density lipoprotein (HDL) to apolipoprotein B-containing particles in exchange for triglycerides, lowering HDL cholesterol (HDL-C).
Several CETP genetic variants have been associated with dyslipidemia, especially the atherogenic dyslipidemia lipoprotein phenotype characterized by reduced HDL-C and increased triglyceride levels (Srisawasdi et al., 2021). Further, reduced HDL-C levels (Li et al., 2021) and a high triglycerides (TG)/HDL-C ratio have been associated with worse survival in COVID-19 patients (Peng et al., 2022). Several CETP polymorphisms, including rs3764261C>A, rs4783961G>A, rs5882G>A, and rs708272G>A, have been linked to variations in the response to statin therapy (Anagnostopoulou et al., 2007; Bercovich et al., 2006; Leusink et al., 2016).
The rs708272 (commonly known as TaqI B polymorphism) is an intron 1 alteration that results in alleles B1 and B2 where allele B1 is characterized by high CETP and low HDL, and allele B2 results in low CETP concentration and high HDL (Kuivenhoven et al., 1998).
A study among Egyptians showed that the rs708272A variant influences HDL-C levels and is associated with premature coronary artery disease (Abd El-Aziz et al., 2014). The study reported a significant association between the B2B2 genotype and increased HDL-C levels and reduced total cholesterol (TC), TG, and LDL-C, a finding supported by observations among black South African women (Ellman et al., 2015). Frequencies for the genotypes were for B1/B1 27% (black women) and 31% (white women) and for B2/B2 13% (black women) and 14% (white women), respectively (Ellman et al., 2015).
Lower TC, LDL-C and higher HDL-C concentrations were associated with the LPL X allele and lower LDL-C concentrations were associated with the CETP B2 allele among black African women. The CETP rs3764261A variant has also been associated with significantly increased HDL among Algerians (Meroufel et al., 2015). The rs3764261A variant allele is found in 20–65% of Africans, which is comparable to the frequencies found in Europeans and Asians (Li et al., 2008).
Eleven more CETP variations have been significantly linked to HDL-C levels among Ghanaians, with the strongest links seen on CETP rs17231520 and rs34065661 (Agongo et al., 2020). The rs17231520 polymorphism showed an additive effect, with serum HDL-C levels increasing in response to the variant A allele. CETP polymorphisms have been associated with protective effects against atherosclerosis through their effects on HDL-C. Hence, CETP inhibitors were evaluated as potential cardiovascular therapies although none has yet made it to the market (Armitage et al., 2019).
Solute-like carrier organic anion transporter 1B1
The SLCO1B1 gene is located on chromosome 12p12.1, spanning 108.59kb in size and has 14 coding and 1 non-coding exon (Stojaković and Igić, 2013). It encodes 691 amino acids that form 12 transmembrane helices, including the hepatic organic anion transporter 1B1 (OATP1B1) (Niemi et al., 2011). Many endogenous substances and xenobiotics, including statins, are absorbed by liver cells through the action of this transporter (Ho et al., 2006).
Statins are actively taken up by hepatocytes through OATP1B1, OATP1B3, OATP2B1, and members of the solute carrier family 10 playing minor roles (Ho et al., 2006; Pasanen et al., 2006). The expression or function of the OATP1B1 protein is reduced by mutations in the SLCO1B1 gene limiting the hepatic absorption of substrates such as statins (Niemi et al., 2011). In turn, this impairs hepatic clearance of drugs, including statins, increasing the likelihood of side effects that are due to increased systemic exposure.
The two SNPs that have received the greatest attention are SLCO1B1 c.521T>C (rs4149056) and SLCO1B1 c.388A>G (rs2306283), both of which have been repeatedly linked to the effectiveness and safety of statins (Chasman et al., 2004; Ramsey et al., 2014). The two variants rs4149056C and rs2306283G form important haplotypes (Table 2) such as *1B:(rs2306283G- rs4149056T, -p. Asn130Asp), *5:(rs2306283A-rs4149056C p. Val17Ala), and *15:(rs2306283G-c. rs4149056C, p. Asn130Asp, pVal174Ala) that have been associated with varied statin response.
The most significant haplotype is SLCO1B1*5, which has been linked to decreased transport function in vitro and increased systemic statin exposure in vivo, increasing the risk of statin myopathy (Chasman et al., 2012; Hirota et al., 2020; Vrablik et al., 2014). Further, the combination of genotypes of these variants gives rise to six phenotypes, namely increased function, normal function, decreased function, possible decreased function, poor function, and intermediate function.
Recent CPIC guidelines have recommendations for all statins based on the phenotypes described earlier, whereas previously the recommendations were only based on the poor function phenotype and simvastatin (Cooper-DeHoff et al., 2022; Ramsey et al., 2014). The inheritance and impact of the rs4149056C variant seems ethnic-specific. For instance, this variant has been found to increase the exposure of atorvastatin and rosuvastatin in Asian populations compared with Europeans (Birmingham et al., 2015; Lee et al., 2005).
A study among Zimbabweans found that the variant rs4149056C plays a minor or no role in rosuvastatin disposition in black Zimbabweans due to its low frequency among Africans (0%) compared with Asians (0.15) and Europeans (0.20) (Soko et al., 2018). In addition, the study showed greater inter-individual variability that was not explained by any of the known common variants, suggesting the existence of novel African-specific variants in this gene or other genes.
In a South African study, the prevalence of rs4149056C variant was significantly higher in black African participants with hypercholesterolemia (53.3%) compared with Europeans (15.3%) (De Beer et al., 2021). Further, there were no differences in the frequencies of rs4363657C and rs2306283G among Africans when comparing healthy controls to hypercholesterolemic participants.
SLCO1B1 g.89595T>C (rs4363657) is another polymorphism that has been implicated in statin-associated toxicity. It is a non-coding variant and is in almost complete linkage disequilibrium with both rs4149056 (r2 = 0.95) and rs2306283 (r2 = 89) (Nagy et al., 2015). The Study of the Effectiveness of Additional Reductions in Cholesterol and Homocysteine (SEARCH) using a genome wide association (GWAS) approach reported significant association with statin intolerance (Link et al., 2008).
This variant was again identified in the JUPITER trial along with another intronic SNP, rs12317268 A>G, and both were reported to reduce LDL-C lowering response (Chasman et al., 2012). There is not much variation in the distribution of the rs4363657C and rs12317268G variants between the world populations (Table 3).
OATP1B1 has been shown to be inhibited by ritonavir (Larson et al., 2014). The polymorphism rs4149056 has been associated with decreased transport activity in vivo, variations of lopinavir plasma concentrations, and 37% lower clearance. Reduced clearance is associated with decreased hepatocyte absorption, which results in higher lopinavir plasma concentrations among C allele homozygotes (Hartkoorn et al., 2010). According to Gong and Kim (2013), majority of SLCO1B1 polymorphisms (rs56101265, rs56061388, rs72559745, rs72559746, rs55901008, rs59502379, and rs56199088) may be associated with variable response to remdesivir. All these variants are associated with reduced transporter function (Gong and Kim, 2013). Therefore, these PGx associations should be considered in future COVD-19 PGx trials.
Cytochrome P450 family 3A4/5 (CYP3A4 and CYP3A5)
CYP3A4 and CYP3A5 are homologous genes found on chromosome 7q21–22, both containing 13 exons, spanning 27 kbp and 32 kbp, respectively (Zeigler-Johnson et al., 2004). They code for enzymes responsible for phase 1 metabolism of lipophilic statins and over 75% of other therapeutic drugs (Canestaro et al., 2014; Rendic and Carlo, 1997).
CYP3A4 is the predominant gene accounting for about 30% of hepatic CYP expression (Daly, 2006), and the distribution and frequencies of its genetic variants vary qualitatively and quantitatively across world populations (Muyambo et al., 2022; Swart et al., 2012). The significant relevance of CYP3A4 in the COVID-19 treatment is that it is downregulated by cytokines (IL-6) during the inflammatory response via the JAK/STAT pathway (Jover et al., 2002).
The most studied SNP of CYP3A4 is g.-392A>G (rs2740574, CYP3A4*1B) that affects CYP3A4 expression. The reduced function allele has been associated with increased risk of statin-induced myopathy and rhabdomyolysis in patients treated with atorvastatin (Wilke et al., 2005). According to PGx surveys conducted in African countries, the prevalence of the rs2740574G variant ranges from 19% to 74% in Central Africa, Eastern Africa, Southern Africa, and Western Africa to as high as 76–89% in Northern Africa (Fernández-Santander et al., 2016; Gaibar et al., 2018; Kassogue et al., 2014).
The second important SNP of CYP3A4 is c.-392 A>G (rs2242480, CYP3A4*1G), which occurs predominantly among Africans (85–94%) (Kassogue et al., 2014; Swart et al., 2012) compared with Europeans (8%) (Hirota et al., 2020). Studies have shown that the presence of rs2242480G variants is associated with decreased response to atorvastatin among Chinese but not simvastatin (He et al., 2014). A third SNP, c.522–191G>A (rs35599367, CYP3A4*22), has also been linked to lipid-lowering effects on simvastatin.
Patients with A variant allele have increased reduction in TC or LDL-C when compared with G/G carriers (Elens et al., 2011). However, it is very rare among Africans (0%) and was observed in 3% of Mixed Ancestry South Africans in a recent study (Muyambo et al., 2022), implying that the CYP3A4 rs35599367A variant is unlikely to play a major role in the PGx of statins in African countries. The drugs that have been approved in the treatment of COVID-19 are metabolized by cytochrome P450 enzymes.
CYP3A5 is an important gene, and its most studied SNP is CYP3A5c. 6986A>G (rs776746; CYP3A5*3), a splice acceptor variant that causes a premature termination codon and thus a non-functional protein. In a study aimed at investigating the influence of genetic variants on risk and COVID-19 outcome, CYP3A5*3 was not significantly associated with COVID-19 (Salem Hareedy et al., 2021).
CYP3A5*3/*3 carriers have been reported to present with increased response to simvastatin and atorvastatin (Kim et al., 2007; Willrich et al., 2008), as well as decreased simvastatin clearance (Kim et al., 2007). When treated with atorvastatin or rosuvastatin, carriers have presented with increased risk of myalgia and muscle damage (Ramakumari et al., 2018; Wilke et al., 2005) due to increased exposure.
The frequency of the rs776746G variant allele is more prevalent among Europeans (95%) than Africans (15%) (Muyambo et al., 2022), and the effects of this allele may be less severe in Africans. The A allele, on the other hand, has been linked to a higher simvastatin clearance rate (Tsamandouras et al., 2014); this could imply that African patients are more likely to experience reduced lipid-lowering effects from statins.
CYP3A5 c.624 G > A (rs10264272, CYP3A5*6) and CYP3A5 c.1035dup (rs41303343, CYP3A5*7) are two other important polymorphisms in this gene that have not been associated with statin disposition (Saiz-Rodríguez et al., 2020). CYP3A5*6 is an exonic mutation that gives rise to an aberrantly spliced mRNA lacking exon 7, and CYP3A5*7 is a frame-shift mutation that results in a null allele (Adehin et al., 2017). In Africa, allele frequencies for CYP3A5*6 and CYP3A5*7 range between 5–25% and 0–21%, respectively (Bains et al., 2013).
Moreover, variation in CYP3A5 expression has been shown to affect tacrolimus dosing among black South Africans, requiring higher dosage than recommended (Muller et al., 2020). Variations in CYP3A4 and CYP3A5 polymorphisms should be investigated in relation to statin dosing among Africans, as these genes encode enzymes responsible for the majority of statin metabolism, particularly atorvastatin and simvastatin, which are still the most prescribed statins in the region.
Cytochrome P450 family 2C9 (CYP2C9)
The CYP2C9 gene is a component of the cluster CYP2C8-CYP2C9-CYP2C19-CYP2C18 found in a 500 kb region on chromosome 10q24 and produces an enzyme important in the oxidation of statins, particularly fluvastatin (Van Booven et al., 2010). CYP2C9 is the second most abundant cytochrome P450 family enzyme, after CYP3A4, and it is expressed in the liver. Genetic variation in CYP2C9 alters the metabolic pathways of its substrates, affecting their bioavailability (Hirvensalo et al., 2019).
CYP2C9 c.430C>T (rs1799853, CYP2C9*2) and CYP2C9 c.1075A>C (rs1057910, CYP2C9*3) are the most studied, and they are associated with reduced CYP2C9 function that has been associated with increased systemic fluvastatin (Hirvensalo et al., 2019; Zhou et al., 2012) and pitavastatin exposure (Zhou et al., 2013). CYP2C9 reduced-function variants have been linked to increased risk of adverse events in patients treated with fluvastatin following kidney transplantation (Miroševic Skvrce et al., 2013).
The minor allele frequencies of all three SNPs differ across all populations and range from 0% to 9% in African populations (Nthontho et al., 2022). CPIC guidelines are available for fluvastatin and CYP2C9 (Cooper-DeHoff et al., 2022). CYP2C9 genetic polymorphisms are more relevant in warfarin PGx, where several African-specific variants (e.g., CYP2C9 *5, *6, *8 and *11) have been recently incorporated into the African warfarin PGx algorithm (Ndadza et al., 2021).
Warfarin, an anticoagulant, is one of the medications used to treat blood clotting issues linked to COVID-19, but additional research is needed to determine how CYP2C9 genetic variation affects COVID-19 treatment.
ATP binding cassette B1
The ABCB1 gene is located on chromosome Chr7q21.12 and codes for 32 exons, spanning 209.6 kb (Dean et al., 2001). It forms part of the 49 ATP-binding cassette (ABC) gene family members that encode channel and transporter proteins, including p-glycoprotein, an ATP-independent drug efflux pump for xenobiotic compounds (Ambudkar et al., 2003). It is one of the seven ATP-driven membrane transporters expressed at all tissue barriers (Ambudkar et al., 2003). ABCB1 is a vital efflux drug transporter that has been associated with several DDIs.
This enzyme is thought to act as an enhancer for hydrochloroquine and azithromycin and is responsible for the synergistic impact of the combination of both medications in the treatment of COVID-19 (Scherrmann, 2020). ABCB1 common variants include ABCB1 c.1236T>C (rs1128503), c.2677T>G (rs2032582), and c.3435C>T (rs1045642), which have been shown to affect the exposure to statins (Dagli-Hernandez et al., 2021; Fukunaga et al., 2016).
The rs1045642T variant has been significantly associated with increased total cholesterol reduction on simvastatin treatment, and increased risk of myalgia for patients on atorvastatin (Keskitalo et al., 2008). The rs1045642T variant is found in 40% of South Africans of Mixed Ancestry and in 57% of East/Central Africans while occurring in about 10% among other African populations (Muyambo et al., 2022), thus showing the heterogeneity of African populations.
The rs1128503C variant has been significantly associated with an increased reduction of total cholesterol and a reduced risk of myalgia in patients treated with simvastatin (Becker et al., 2009(Table 3). The ABCB1 rs2032582T is associated with risk of atorvastatin-induced liver injury among Japanese patients (Fukunaga et al., 2016).
The effects of these variants on statin disposition have not been characterized in African patients. Nevertheless, ABCB1 polymorphisms have been implicated in the safety and efficacy of various antiretroviral drugs such as lopinavir and ritonavir (Biswas, 2021), efavirenz (Chala et al., 2021; Ngaimisi et al., 2013; Sinxadi et al., 2015), and nevirapine (Langaee et al., 2021) and in COVID-19 patients for some of these studies.
ATP binding cassette G2
The ABCG2 gene is localized on chromosome Chr4q22.1 comprising 21 exons, spanning over 66 kbp (Robey et al., 2009) and encodes the breast cancer resistance protein, which is an ABC efflux transporter (Mao and Unadkat, 2015). This transporter plays an essential role in drug response and has a broad substrate specificity to efflux xenobiotics, including statins (Soko et al., 2016).
ABCG2 has been shown to affect rosuvastatin bioavailability since it is mostly involved in its excretion from the liver back to the lumen. For statins, CPIC guidelines currently only have recommendations for ABCG2 and rosuvastatin (Cooper-DeHoff et al., 2022) as no significant associations have been reported for atorvastatin, simvastatin, pitavastatin, and pravastatin (Bai et al., 2019; Bakar et al., 2018; Lee et al., 2019). The A allele of its most studied polymorphism, ABCG2 c.421C>A (rs2231142) has been associated with reduced protein expression and increased plasma levels of rosuvastatin (Cooper-DeHoff et al., 2022).
It has also been associated with an increased risk of SAMS and liver damage in patients treated with rosuvastatin. Frequencies for rs2231142A variant range between 10–15% among Europeans, 25–35% among Asians, and 0–8% among Africans (Cooper-DeHoff et al., 2022; Mao and Unadkat, 2015; Niemi et al., 2011). Further, a detailed study from our group (Soko et al., 2018) reported no significant differences in the frequencies of the minor rs2231142A variant across nine ethnic African groups, with an average prevalence of 2%.
It is, thus, reasonable to postulate that ABCG2 rs2231142 plays a minor role in the pharmacogenomics of rosuvastatin in African populations. Both lopinavir and ritonavir have been shown to strongly inhibit the ABCG2 transporter (Kumar and Trivedi, 2021), which would possibly impact the effectiveness of patients on both statins and Paxlovid, the approved COVID-19 drug.
Cholesterol Absorption Inhibitors
Cholesterol absorption inhibitors such as ezetimibe are a second-line therapy for dyslipidemia (Table 1), often used after patients fail to reach target lipid levels on the maximum tolerated dose of statin monotherapy (Mach et al., 2019). Ezetimibe is a more recently licensed lipid-lowering medication, and its availability has been reported in Egypt (Reda et al., 2020) and South Africa (Blom et al., 2020a) on the African continent.
Ezetimibe works by inhibiting intestinal cholesterol uptake by inhibiting the Niemann-Pick C1-like 1 (NPC1L1) transporter, thereby reducing cholesterol delivery to the liver and upregulating LDLR expression, ultimately resulting in lower LDL-C levels overall. Ezetimibe also reduces TC, ApoB, and TG but increases HDL-C in patients with hypercholesterolemia and has been shown to have beneficial effects on CVD outcomes.
However, ezetimibe remains underutilized due to its costs and practitioner complacency (Blom et al., 2020a). Clinical trials have shown that ezetimibe monotherapy reduces LDL-C by 15–22% with relatively high interindividual variation, but concomitant use of ezetimibe with statins reduces LDL-C levels by an additional 21–27% (Auton et al., 2015).
Ezetimibe was first approved by the FDA in 2002 and since then combination agents made up of ezetimibe+atorvastatin/simvastatin/rosuvastatin have also been approved (Sharma and Kaur, 2003). In Africa, ezetimibe first appeared with the approval of Ezetrol in 2005 in South Africa (Fivaz and Noble, 2006).
Ezetimibe is mostly well tolerated with minor adverse effects such as headaches, fatigue, runny nose, and rarely rhabdomyolysis when used in combination with high statin doses. For instance, it has been reported that the combination medication of ezetimibe, atorvastatin, remdesivir, and tacrolimus was associated with toxic myopathy and liver damage in a patient with COVID-19 (Sabljić and Bašić-Jukić, 2022).
However, another study identified ezetimibe along with rosuvastatin to be associated with significantly reduced odds for COVID-19 hospitalization (Israel et al., 2021). Rosuvastatin was observed to raise 25-OH vitamin D levels, which could be mediated through the Niemann-Pick C1 like 1 (NPC1L1) membrane transporter implicated in vitamin D absorption (Israel et al., 2021). In other research, vitamin D insufficiency has been linked to an increased risk of COVID-19 (Israel et al., 2021).
The pharmacokinetics of ezetimibe are not yet well understood but in vitro studies have shown that the enterohepatic circulation and excretion of ezetimibe is dependent on ABCC3, ABCC2, and ABCG2; ezetimibe glucuronide is believed to be a substrate of SLCO1B1 and SLCO2B1 (Oswald et al., 2008). Ezetimibe is converted to ezetimibe glucuronide by UGT enzymes: UGT1A1, UGT1A3, and UGT2B15 (Oswald et al., 2008). Thus, polymorphisms in any of the earlier cited genes that affect the function of the gene product are likely to affect the way patients respond to ezetimibe.
Ezetimibe is not metabolized by cytochrome P450 enzymes and thus has fewer potential DDIs (Oswald et al., 2008). Due to the ezetimibe/rosuvastatin and ezetimibe/rosuvastatin combination drugs, the Swiss Agency for Therapeutic Products (Swissmedic) has come up with drug-gene PGx annotation for ABCG2 and SLCO1B1 (Gencer and Mach, 2016).
The labels recommend reducing dosages of these combination drugs for SLCO1B1 c.521T>C and ABCG2 c.421C>A carriers because these polymorphisms may lead to increased exposure to the statins. The few PGx studies available on ezetimibe are focused on the NPC1L1 gene and its potential role in lowering lipids (Hegele et al., 2005; Liao et al., 2021). Further, at the time of this literature review, there were no PGx studies on ezetimibe from an African country.
Niemann-pick C1 like 1
The NPC1L1 gene is localized to chromosome 7p13 and contains 20 exons, spanning ∼29 kb (Davies et al., 2000). NPC1L1 encodes the Niemann-Pick C1 like 1 protein, a key transmembrane protein expressed in the gastrointestinal tract epithelial cells and on hepatocytes. NPC1L1 is a key regulator of cholesterol absorption and the target molecule of ezetimibe binding in vivo. Several variants have been associated with interindividual variation in response to ezetimibe treatment (Hegele et al., 2005). Specifically, SNPs on the NPC1L1 gene have been shown to have a 15% greater LDL-C lowering response to ezetimibe compared with non-carriers (Simon et al., 2005).
Of note, NPC1L1 g.1679 C > G (rs2072183) has been associated with decreased response to statins in European patients with CVDs and hypercholesterolemia (Smit et al., 2016). This effect has also been confirmed in Asians but in response to hyzetimibe combination with atorvastatin (Liao et al., 2022). Hyzetimibe is a second-in-class, orally administered cholesterol absorption inhibitor developed in China and approved for marketing by the National Medical Products Administration of China (NMPA) in June 2021 (Liao et al., 2022).
Clinical trials assessing hyzetimibe have been mostly conducted among Chinese (Chen et al., 2022; Liao et al., 2021; Ruan et al., 2014). The C/C genotype of g.1679 C > G (rs2072183) SNP was associated with increased reduction of LDL-C on hyzetimibe monotherapy while the G/C genotype was found to induce better LDL-C reduction on the atorvastatin/hyzetimibe combination (Tremblay et al., 2011). The rs2072183C>G SNP is in linkage disequilibrium with NPC1L1 -762T>C(rs2073548), which is known to affect transcriptional levels of NPC1L1, thus altering cholesterol absorption (Liao et al., 2022).
The second important SNP of this gene is NPC1L1 c.-133A>G (rs17655652), as its C/C genotype has been associated with an increased reduction in LDL-C in healthy males treated with pravastatin compared with the C/T and T/T genotype carriers (Lu et al., 2016). The NPC1L1 rs17655652G variant has a low frequency among Africans (3%), compared with both Europeans (29%) and Asians (23%) (Table 3). Therefore, these studies show that polymorphisms in the NPC1L1 gene could potentially result in different responses in different populations, and that more studies are needed among Africans.
Proprotein Convertase Subtilisin/Kexin Type 9 Inhibitors
Proprotein convertase subtilisin/Kexin type 9 (PCSK9) protein is expressed in the liver and is encoded by the PCSK9 gene on chromosome 1 (Seidah et al., 2003). PCSK9 binds extracellularly to LDLRs, preventing LDLRs recirculation to the cell surface, thus leading to their degradation. LDLR binds to LDL-particles, facilitating their absorption into the enterocytes from extracellular fluids, thus decreasing circulating LDL-particles (Cohen et al., 2006; Seidah et al., 2003).
Overexpression of PCSK9 leads to reduced LDLR on the cell surface and high LDL-C; hence, when PCSK9 is inhibited, it leads to increased LDLR present and lower LDL-C (Weinreich and Frishman, 2014). Regarding PCSK9's effects on viral infection, it has been demonstrated that PCSK9 promotes the degradation of LDLR-related protein (LRP1), where increased expression of LRP1 is a protective reaction to viral infection (Scicali et al., 2020).
Therefore, PCSK9 may boost viral infectivity by causing LRP1 degradation, and its inhibition may be helpful to reduce the infectivity of certain viruses, including COVID-19 (Scicali et al., 2020). PCSK9 inhibitors are the newest and supposedly safest lipid-lowering drugs approved by the FDA (Table 1) (Zimmerman, 2015).
There are currently two FDA-approved PCSK9 inhibitors, namely, alirocumab and evolocumab that are fully human monoclonal antibodies (Mach et al., 2019). In Africa, both PCSK9 inhibitors are available in South Africa (Klug et al., 2018), and their use in Egypt is still ambiguous but they have been strongly recommended by the Egyptian Association of Vascular Biology and Atherosclerosis (Bytyçi et al., 2022). They have been shown to significantly reduce LDL-C levels by up to 60% either as monotherapy or by 73% in combination with a maximally tolerated statin (Mach et al., 2019).
The COVID-19 pandemic caused a reduction in PCSK9-inhibitor prescription (Seijas-Amigo et al., 2022) and adherence (Caso et al., 2022). A third PCSK9 inhibiting drug called inclisiran has been recently approved by the FDA for the treatment of hypercholesterolemia (Lamba et al., 2012). Inclisiran is a small interfering double-stranded RNA (siRNA) that binds to RNA-induced silencing complex, thus silencing mRNA coding for PCSK9.
Results from phase 3 studies, ORION-9, −10, and −11, demonstrated the efficacy of this agent in lowering LDL-C (Chiu et al., 1995). Inclisiran is not yet available in any African country but is undergoing clinical trials in South Africa (Elserafy et al., 2022). However, PCSK9 inhibitors are suggested as a third-line therapy by treatment recommendations for patients who are at extremely high risk of severe cardiovascular events and who have not achieved target lipid levels on combination therapy of statin and ezetimibe due to their costs (Klug et al., 2018; Reda et al., 2020).
In addition, alirocumab was demonstrated to result in a considerable LDL-C lowering and better achievement of LDL-C target values in the ODYSSEY FH1 and FH2 trials, demonstrating the efficacy of PCSK9 inhibitors for individuals with primary hypercholesterolemia (Kastelein et al., 2015). Other lipid effects of PCSK9 inhibitors include reducing TG levels and Lp (a) and increasing HDL and ApoA-1. In the FOURIER trial, treating patients with atherosclerotic CVD, and LDL-C levels of 1.8 mM (70 mg/dL) with evolocumab did not reduce the risk of CV mortality but only reduced the risk of the primary CV endpoints by 15% (Ference et al., 2018).
While in the ODYSSEY Outcomes trial, alirocumab was associated with a significant reduction in all-cause mortality, this was an exploratory outcome and not supported by any significant impact on CV death (Szarek et al., 2019). Both trials confirmed the safety of both drugs, apart from reports of adverse reactions at the injection site, such as itching and flu-like symptoms.
Since both drugs are fully human antibodies, they also have low potential for developing auto-antibodies. PCSK9 inhibitors are injected subcutaneously once every 2 weeks or monthly depending on the prescribed dosage (Ference et al., 2018). They presently have no reported DDIs with orally administered drugs possibly because they are not molecules subject to hepatic metabolism.
This makes the pharmacokinetics of PCSK9-inhibitors interesting as they exhibit non-linear kinetics; hence, there are a few PGx studies globally. Lastly, even though PCSK9 inhibitors are highly effective medications that can reduce LDL-C and CVD events in addition to statin and/or ezetimibe treatment, their use in African nations with scarce health care resources is currently restricted by the costs of the treatment.
PCSK9 consists of 13 exons and produces two isoforms through alternative splicing (Varret et al., 2008). Gain-of-function mutations increase LDLR degradation, resulting in hypercholesterolemia (Sun et al., 2005). Loss-of-function mutations reduce LDLR degradation, resulting in hypocholesterolemia (Langsted et al., 2016). Individuals with a gain of function of PCSK9 are expected to have a greater response to PCSK9 inhibitors, because they have increased LDLR degradation. Its most reported SNP, PCSK9 c.137G>T (rs11591147) has been significantly associated with LDL-C response when treated with statins such as atorvastatin (Thompson et al., 2005), but this SNP has only been reported among Europeans (2%) and is rare among African ancestry (Table 3).
A second important SNP is c.1274A>G (rs28362261), which is a loss-of-function mutation that has also been associated with statin response among African Americans and not Americans of European descent (Feng et al., 2017). It is found in 1.4% of Nigerians (Cohen et al., 2006) and 3.7% of Zimbabweans (Hooper et al., 2007). Hooper et al. (2007) also discovered the loss-of-function C679X, which is more common in Africans with allele frequencies ranging from 1% to 3.7% and may have a greater effect on PCSK9 inhibitor disposition in Africans.
Therefore, loss-of-function mutations in PCSK9 seem to have significant effects on statin therapy in patients and may also affect PCSK9 inhibitor therapy in patients with dyslipidemia in Africa. Patients with COVID-19 and healthy controls have been shown not to significantly differ in their expression of the PCSK9 gene (Yang et al., 2022).
Low-density-lipoprotein receptor
The LDLR gene encodes the LDLR protein that mediates endocytosis of LDL particles (Südhof et al., 1985). It is found on chromosome 19p13.2 encompassing 18 exons and 860 amino acids, spanning 5.3 kb. LDLR is predominantly expressed in hepatic cells, where it generates six isoforms via alternative splicing (Zghair et al., 2021).
The use of PCSK9 inhibitors increases LDLR protein expression, which increases LDL-C clearance from the plasma and is particularly indicated in individuals with familial hypercholesterolemia (FH). Response to PCSK9 inhibitors in homozygous FH individuals is influenced by residual functionality of the mutated LDLR (Hindi et al., 2021). Patients with no LDLR function (receptor-negative mutations) do not respond to evolocumab (Stein et al., 2013).
Further, a few of the LDLR SNPs, rs5925, rs688, rs1433099, and rs2738466, have been linked to variable-response statin outcomes in Europeans (Lagos et al., 2015; Polisecki et al., 2008; Thompson et al., 2005). The frequencies of respective variants among African populations are 15%, 4%, 38% and 16% for rs5925, rs688, rs1433099, and rs2738466, respectively (Table 3). In vitro hydroxychloroquine (HCQ) treatment and SARS-CoV-2 infection datasets showed a considerable upregulation of the LDLR gene, indicating that both HCQ treatment and SARS-CoV-2 infection changes the transcription of genes involved in lipid metabolism (Corley et al., 2020; Usenko et al., 2022).
Conclusions and Outlook
This expert review examined and synthesized the priority PGx variations as seen through a One Health lens in Africa to improve and inform personalized medicine in both dyslipidemia and COVID-19. Dyslipidemia is increasing across Africa, further pushing high the burden of CVDs. The biggest worry is the variability observed among dyslipidemia patients on how they respond to the different treatment regimens.
The COVID-19 pandemic worsened the situation, as dyslipidemia was one of the risk factors increasing mortality among COVID-19 patients. Further, the treatments/drugs that were indicated for SARS-CoV-2 posed potential DDI problems. Thus, pharmacogenomics knowledge among African populations may help in the understanding of where potential DDIs are likely, aiding in pre-emptive steps that could lead to improved patient outcomes.
The PGx of lipid-lowering treatments have been extensively studied in American, Asian, and European populations; however, such studies are severely lacking among African populations. There are several reasons for the disparity of research among African populations, with the main reason being limited research in genomics among Africans due to both lack of funding and limited expertise in genomics (Wonkam et al., 2022).
However, African populations should not be ignored with respect to how genomic variation may contribute to better health. For instance, some polymorphisms such as SLCO1B1 c.521T>C, which are more frequent and therefore important in European and Asian populations, have been shown to have a reduced role in Africans. Instead, polymorphisms such as ABCB1 c.3435 C>T, CYP3A4 c. −392 A>G, and CYP3A5 c.3705C>T, which are more frequent in African populations, but have not yet been studied in terms of statin response in this population group, need to be evaluated.
Lastly, as African researchers and based in Africa, we call for the inclusion of African populations in clinical trials so that the diverse African genomic background is considered and accounted for at the earliest stages of drug development and clinical trials.
Authors' Contribution
Z.L.: conceptualized (lead), formally analyzed, wrote original draft, reviewed, and edited manuscript. N.D.S.: conceptualized (supporting), formally analyzed, reviewed, and edited manuscript. B.D.: conceptualized (supporting), formally analyzed, reviewed, and edited manuscript. S.M.: validated, reviewed manuscript. E.J.: validated, reviewed manuscript. B.R.: formally analyzed, reviewed, and edited manuscript. J.B.: validated, reviewed manuscript, funding acquisition. P.S.: validated, reviewed manuscript. M.T.D.: validated, reviewed manuscript. C.D.: conceptualized (lead), formally analyzed, reviewed, and edited the literature review. All authors made a significant intellectual contribution to the final version and have read and approved the manuscript for publication.
Footnotes
Author Disclosure Statement
The authors declare they have no conflicting financial interests.
Funding Information
The Pharmacogenomics and Drug Metabolism Research Group is funded as part of the UCT/South African Medical Research Council Extramural Unit.
