Abstract
Abstract
Although the rate of breast cancer differs between women in Asian and Western countries, molecular genetics/genomics basis of this epidemiological observation remains elusive. Moreover, the intake of phytoestrogens is associated with a lower incidence of breast cancer. Genistein and daidzein are the primary soy isoflavones with a chemical structure similar to estrogens. Conceivably, the actions of phytoestrogens on gene expression signatures might mediate their postulated effects on breast cancer pathogenesis. The present study evaluated the transcriptional responsiveness of breast cancer cells to soy phytoestrogens using a whole-genome microarray-based approach. Human breast cancer cell lines and a fibrocystic breast cell line were treated with genistein or daidzein. We identified 278 and 334 differentially expressed genes after genistein or daidzein treatment, respectively, in estrogen-positive (MCF-7) and estrogen-negative (MDA-MB-231, MCF-10a) cells. Hierarchical clustering of this finding revealed a significant modulation, respectively, of 246 or 169 genes after genistein or daidzein exposures. Importantly, the molecular pathways for the differentially expressed genes included those that relate to cell communication, biodegradation of xenobiotics, lipid metabolism, signal transduction, and cell growth/death. These molecular observations collectively contribute to a growing knowledgebase on the putative mechanism(s) of action of phytoestrogens in breast cancer pathogenesis and chemoprevention.
Introduction
Isoflavones belong to the phytoestrogen family, which are a class of plant-derived compounds that are structurally similar to estrogens (Sirtori et al., 2005), and act as estrogen-like compounds having various biological effects (Trock et al., 2006). Genistein and daidzein are the primary isoflavones of soy that can bind and activate both estrogen receptors (α and β) that subsequently induce the transcription of estrogen-responsive target genes in a dose-dependent manner (Kuiper et al., 1998; Maggiolini et al., 2001; Satih et al., 2008). These isoflavones therefore compete with the physiological estrogens. Indeed, from a clinical standpoint, phytoestrogens from soy have unpredictable effects on breast cancer. Hence, despite the previously observed epidemiological differences in breast cancer prevalence among populations with variable dietary habits for soy product consumption, molecular genetics/genomics mechanisms of phytoestrogens in breast cancer pathogenesis remained elusive thus far. On the other hand, increasing technical feasibility to survey the global gene expression responses to phytoestrogen exposure might offer newer mechanistic insights in relation to their (patho)physiological effects.
The present study aimed to survey the genes that are differentially modulated after exposure to soy phytoestrogens. To this end, we examined the whole-genome expression using pangenomic microarrays in two different human breast cancer cell lines (MCF-7 and MDA-MB-231) and a human fibrocystic breast cell line (MCF-10a), after treatment with either genistein or daidzein for a period of 72 h. Subsequent analysis was performed according to the estrogen receptor (ER) status or the malignancy status of cell lines. That is, to assess the estrogen-like effects of phytoestrogens on the ER, we employed two types of cell lines: ER+ cell lines (MCF-7) and ER− cell lines (MCF-10a and MDA-MB-231). Moreover, these cell lines were also studied with microarrays according to the malignancy status: tumor cells (MCF-7 and MDA-MB-231) and fibrocystic cells (MCF-10a).
Methods
Cell culture, phytoestrogen treatments, and RNA extraction
MCF-7 and MDA-MB-231 breast tumor cell lines were established from a pleural effusion of patients with invasive breast carcinoma (Cailleau et al., 1974; Soule et al., 1973). The MCF-10a cell line came from breast tissue of a patient with fibrocystic breast disease (Soule et al., 1990). The three human cell lines were obtained from the American Type Culture Collection (ATCC, Rockville, MD, USA) (Table 1). MCF-7, an ER+ cell line, was grown in RPMI 1640 media and supplemented with 2 mM L-glutamine (Invitrogen, Carlsbad, CA, USA), 20 μg/mL gentamycin (Panpharma, Fougères, France), 10% fetal bovine serum (FBS) (Invitrogen), 0.04 IU/mL insulin (Novo Nordisk, Princeton, NJ, USA) in a humidified atmosphere at 37°C containing 5% CO2. MCF-10a, an ER− cell line, was maintained in DMEM-F12 (Invitrogen) containing 10% horse serum (Invitrogen), 2 mM L-glutamine, 20 μg/mL gentamycin (Panpharma), 20 ng/mL epidermal growth factor (Sigma, St. Louis, MO, USA), 100 ng/mL cholera toxin (Sigma), 0,25 IU/mL insulin (Novo Nordisk), and 0.5 μg/mL hydrocortisone (Sigma) at 37°C with 5% CO2. Growth medium for MDA-MB-231, an ER− cell line, was Leibovitch L-15 media with 15% FBS, 20 μg/mL gentamycin (Panpharma), 2 mM L-glutamine in a 37°C humidified atmosphere without CO2. Isoflavone supplementations were carried out for 72 h with 18.5 μM genistein or 78.5 μM daidzein dissolved in DMSO (Sigma). Notably, these selected concentrations were previously determined to correspond to the 50% inhibition of the proliferation (IC50) (Vissac-Sabatier et al., 2003). As controls, cell lines were also reseeded in medium without isoflavone treatments. After 72-h treatments, cells were washed three times with phosphate-buffered saline (PBS) and lysed in 5 mL of RNA-PLUS (MP Biomedicals, Solon, OH, USA) according to the manufacturer's protocol. Recovered RNA was quantified using a Nanodrop 6000 spectrophotometer (Nanodrop Technology®, Cambridge, UK), and RNA integrity was assessed using a 2100 Bioanalyser (Agilent, Palo Alto, CA, USA). RNA samples with an RNA integrity number (RIN) greater than 9 were used for further analysis. We used an equimolar-pooled mixture of each test RNA sample for the reference RNA for two-color microarray technology.
cDNA synthesis, labeling, and hybridization
Pangenomic microarrays (G4112A) were provided by Agilent®. Hybridization was performed on 44K Human Whole-Genome 60-mer oligo-chips (G4112A, Agilent Technologies, Santa Clara, CA, USA) in a rotary oven (4,000 rpm, 60°C, 16 h). One microgram of total RNA was amplified and labeled using Low RNA Input Fluorescent Linear Amplification Kit PLUS two-color (5188-5340, Agilent®), in accordance with the manufacturer's instructions. Each amplified cDNA was fluorescently labeled with Cy5-CTP for treated samples and Cy3-CTP for control samples. Both Cy3- and Cy5-labeled probes were mixed and hybridized to pangenomic arrays in a competitive reaction. Slides were washed using a wash buffer kit (5188-5327, Agilent®) and a stabilization solution (5185-5979, Agilent®). These experiments were performed in triplicate. The fluorescent intensities were scanned with a 418 GMS scanner (Affymetrix®) and quantitative values for the signals were calculated using GenePix Pro 6 (Axon®) software. Expression ratios between treated and untreated cells were obtained for each spot in a GenePix file.
Analysis of gene expression
For each gene, the ratio of Cy3 to Cy5 signal intensities (Cy3/Cy5) was calculated and log2-converted using LimmaGUI 1.7.0 software through the R 2.2.0 software (Wettenhall and Smyth, 2004). Then, a background correction was performed by an Edwards's correction. Normalizations were performed within and between arrays by a global lowess method (Edwards, 2003), based on the local estimation of intensities and a regression calculation weighted toward similar spots (Fig. 1). The gene probes used as negative controls were removed from the data set using a differential expression cutoff of twofold and hierarchical clusters were performed with TIGR MeV version 4.0.0 software using the complete-linkage Pearson correlation clustering method (Fig. 2). Clusters were color coded: red for upregulation and green for downregulation compared to the control group. Expression was scheduled with a volcano-plot (Fig. 3), which demonstrated the amount of overexpressed and underexpressed genes. Significant genes were clustered by the t-test using the method by Tusher and colleagues (2001) with a K-nearest neighbors imputer of 10 neighbors. (Chalabi et al., 2006, 2007). Therefore, significantly expressed genes were classified according to implicated biological pathways by using EASE version 2.0 software (Hosack et al., 2003) and Gene Ontology (GO) database using Bonferroni correction and False Discovery Rate controls (Hosack et al., 2003).
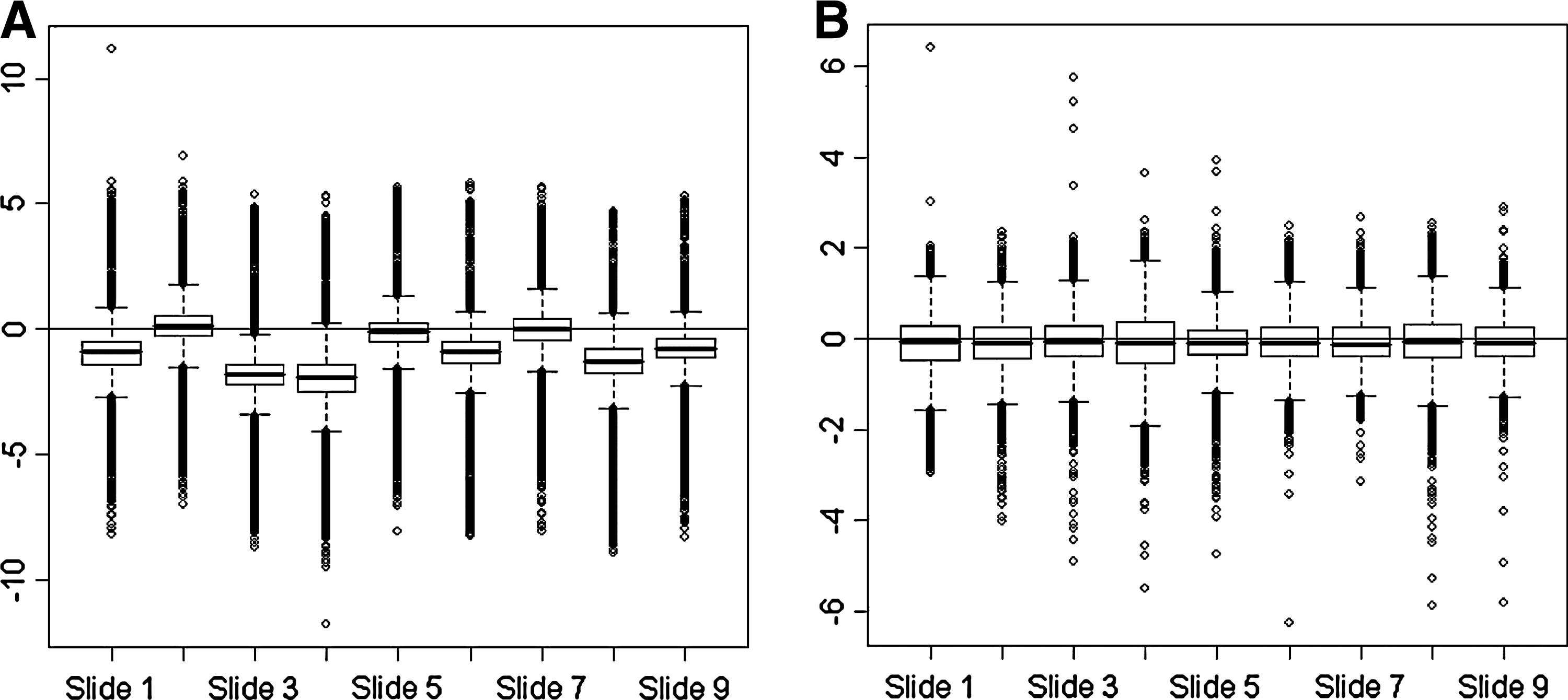
M Box Plot for all slides (
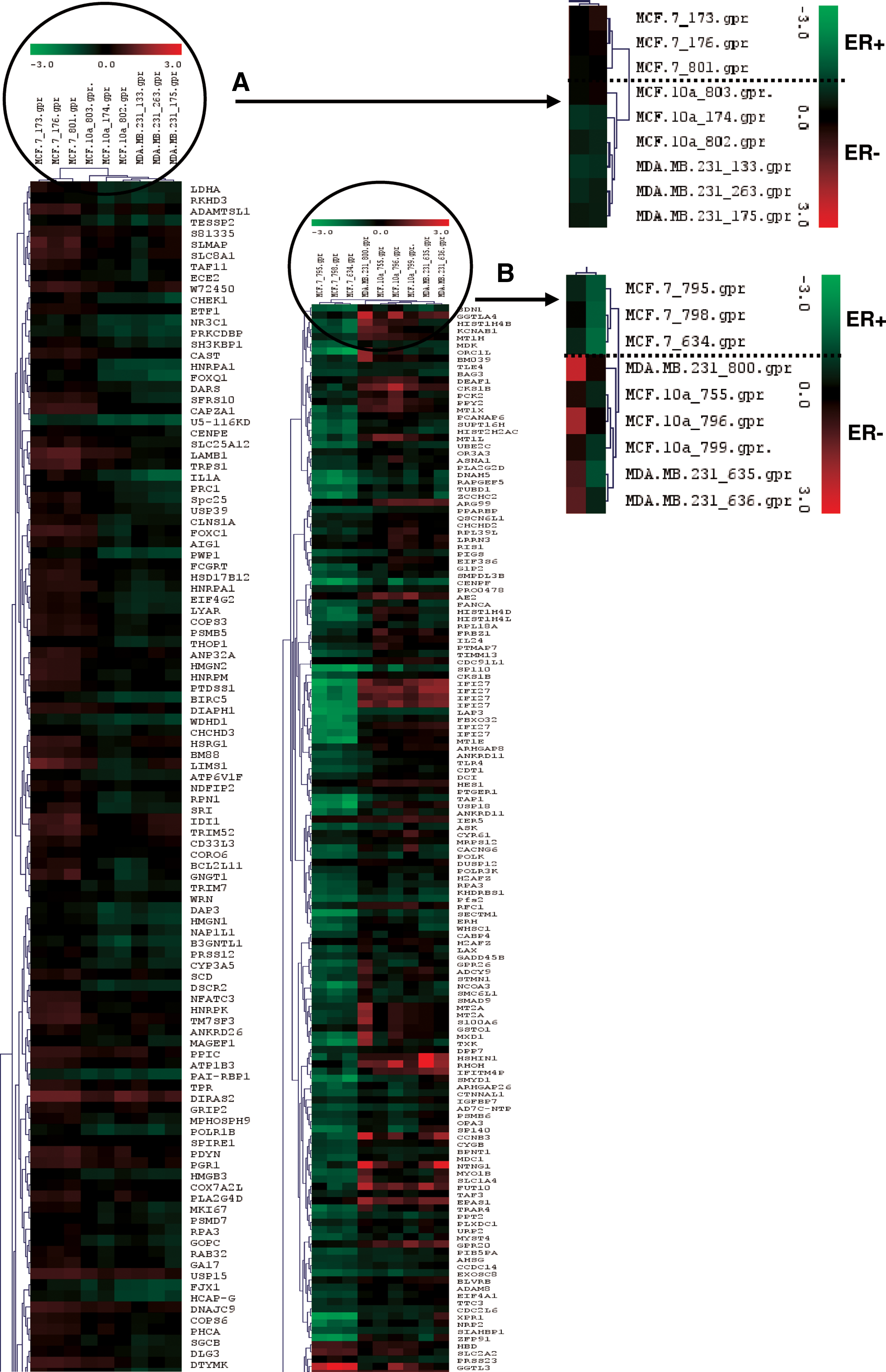
Hierarchical clustering of the gene expression profiles of (
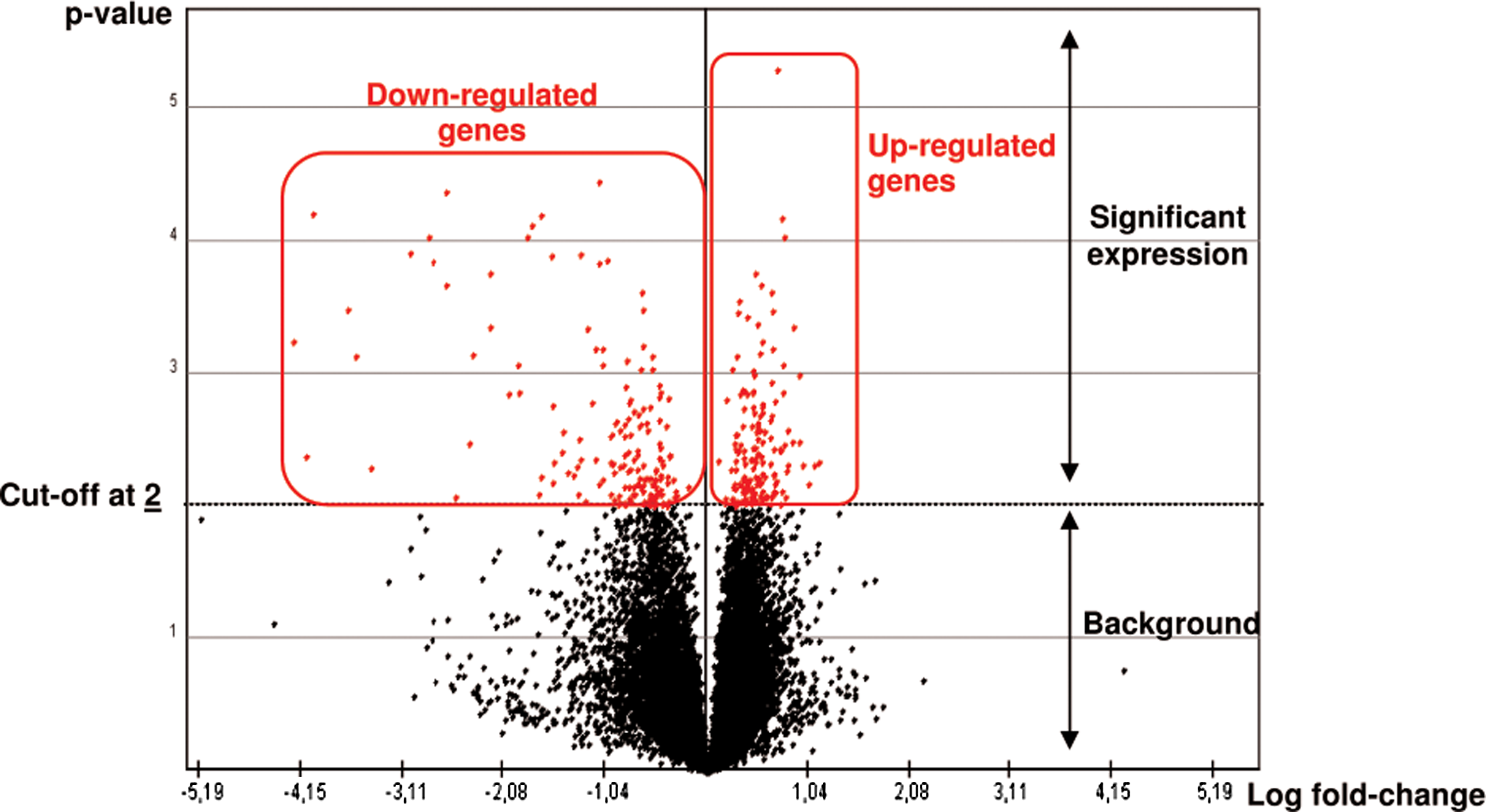
Volcano-plot of gene expression profile using a differential expression cutoff of twofold. Example of genistein exposure according to the ER status discrimination between the MCF-7, MDA-MB-231 breast cancer cell lines, and MCF-10a fibrocystic breast cell line, performed with TIGR MeV version 4.0.0 using a t-test method. Genes represented over the cutoff line were considered modulated as overexpressed on the right and underexpressed on the left.
Results
Hierarchical clusters and differentially expressed genes
Using a differential expression cutoff of twofold (TIGR MeV), differentially expressed genes were identified between ER+ cells (MCF-7) and ER− cells (MDA-MB-231 and MCF-10a). Hierarchical clustering showed 278 and 334 differentially expressed genes after genistein or daidzein 72-h treatments (Fig. 2).
Differential gene expressions were also examined based on the tumor (MCF-7 and MDA-MB-231) and nontumor (MCF-10a) status. Hierarchical clustering revealed a significant modulation of 246 genes after genistein 72-h treatment and a significant modulation of 169 genes after daidzein exposure.
Molecular pathways modulated
The different gene lists determined to be significantly modulated were subjected to EASE version 2.0. software. EASE, developed by Hosack et al. (2003), is a software application that facilitates the biological interpretation of gene lists derived from the results of microarray analyses. EASE provides statistical methods for discovering enriched biological themes within gene lists, generates gene annotation tables, and enables automated linking to online analysis tools. Subsequently, the genes were related to molecular pathways via KEGG database (Table 2). Two types of contrasts, ER+ versus ER−, and tumor versus nontumor status were used to compare the effects of genistein and daidzein on genomic pathways. After genistein exposure, 24 total pathways were dysregulated, 17 pathways common to both ER− status and tumor status. After daidzein exposure, 22 total pathways were dysregulated, 13 common to both ER− status and tumor status. The major functional categories for these gene lists included various molecular pathways such as cell growth and death, biodegradation of xenobiotics, as well as cell communication, lipid metabolism, and signal transduction.
These data were obtained using EASE version 2.0software and related to molecular pathways via KEGG database. The differentially expressed genes through ER discrimination are shown in black, and those through tumor discrimination in gray.
After genistein treatment, the apoptosis related genes ASK and GADD45B were downregulated, whereas CYCS and IkB were upregulated. The differentially expressed genes related to xenobiotic biodegradation were CYP4B1 (downregulated with genistein) CYP2A6, CYP1B1 (upregulated with daidzein), and CYP3A5 (downregulated with daidzein). The genes involved in cell communication were RAC2 and VASP (downregulated with genistein). One gene involved in lipid metabolism was AKR1C3 (downregulated with genistein). Finally, one gene related with signal transduction pathway was CDKN3 (downregulated with genistein and daidzein) (Table 3).
Selected genes with fold changes through DNA microarrays assays were repeated three times after the treatment of MCF-7, MDA-MB-231 breast cancer cell lines, and MCF-10a fibrocystic breast cell line with genistein (black) and daidzein (gray). Results were first analysed by TIGR MeV version 4.0.0 using a differential expression cutoff of twofold. Results are statistically significant (ap < 0.01; bp < 0.05) by the Student's t-test.
Discussion
Breast cancer has become the most common malignancy among Western women (Rice and Whitehead, 2006), whereas its incidence in Asian countries such as in Japan is only about one-third of that in Western populations. This marked difference has often been attributed to a much higher dietary intake of soy phytoestrogens. However, despite the large number of epidemiologic studies conducted, the attendant mechanisms warrant further molecular insights (Martinez et al., 2006; McCarty, 2006; Messina et al., 2006).
Breast cancer is a steroid hormone responsive tumor type that is particularly estrogen-responsive. Investigations have been carried out on the gene expression profiles in response to phytoestrogens and their putative involvement in breast cancer pathogenesis. Ise et al. (2005) examined a panel of phytoestrogens and their effects on expression of a total of 172 estrogen responsive genes, using a customized DNA microarray on the MCF-7 cell line. Those expression profiles were then compared with that for 17β-estradiol (E2) using correlation coefficients. In another study, Dip et al. (2008) investigated the gene expression induced by soy products such zas deconjugated phytoestrogens using a whole genome microarray approach on T47D and MCF-7 cell lines. These two comparative studies concluded that there was a partial overlap between the gene expression patterns elicited by E2 and phytoestrogens for the first one, and a stereotyped expression fingerprint in breast cancer cells containing ERα for the last one. Our study consisted of a global gene expression profiling by DNA microarray analysis using specifically genistein (18.5 μM) and daidzein (78.5 μM) on two breast cancer (MCF-7 and MDA-MB-231) cell lines and a fibrocystic (MCF-10a) cell line. The breast cell lines used herein exhibit different ER status. Hence, we compared the observed gene expression changes in relation to the ER status and malignancy status. In the first case, genistein treatment modified expression of 278 genes, and 246 genes in the second. Daidzein exposure modified the expression, respectively, of 334 and 169 genes with ER and tumor status discriminations. The different genes that were significantly modulated were subjected to the EASE version 2.0 software and related to the molecular pathways via the KEGG database. This highlighted important functional categories such as cell growth and apoptosis, biodegradation of xenobiotic and cell communication, as well as lipid metabolism and signal transduction.
We observed modulation of the apoptosis-related genes: ASK, GADD45β, IκB, and CYCS. ASK and GADD45β were downregulated after genistein treatment, whereas CYCS was upregulated, suggesting a pro-apoptotic effect also observed with IκB after the daidzein exposure. Divya and Pillai (2006) have shown that curcumin blocked IκB phosphorylation and degradation, leading to abrogation of NFκB activation. Our data suggested that daidzein could act in the same way as an anti-inflammatory and pro-apoptotic nutrient agent.
In a recent study, Lerebours et al. (2008) have shown that GADD45β expression was upregulated in inflammatory breast cancer using real-time quantitative RT-PCR. Engelmann et al. (2008) found an upregulation of GADD45β in NIH3T3 cells and showed that it could confer a resistance against death stimuli. After genistein treatment, we found a downregulation of GADD45β, suggesting a possible apoptotic property of this phytochemical. We also found an upregulation of the apoptosis-inducing gene CYCS expression after genistein treatment.
In our study, we reported a downregulation of CYP4B1 and CYP3A5 after genistein and daidzein exposures, respectively. Recent studies have associated CYP4B1 expression with inflammatory response and angiogenic activity (Mastyugin et al., 1999; Seta et al., 2007). However, Iscan et al. (2001) was the first to report that CYP4B1 is expressed in tumors and normal breast tissue. Our data may support the hypothesis that genistein could act as an anti-inflammatory agent.
In another study, Volak et al. (2008), demonstrated that curcumin inhibited CYP3A5 expression. Our data were consistent with the curcumin effects observed on CYP3A5 expression, and suggested that daidzein could act as an antigenotoxic molecule.
CYP1B1 expression was found upregulated after a 72-h treatment of daidzein. However, in a recent study, Wagner et al. (2008) examined the influence of E2, genistein, and daidzein on CYP1A1/1B1, COMT, and QR expression in MCF-7 cells by reverse- transcription polymerase chain reaction (PCR). They showed an insignificant change in CYP1B1 mRNA levels. Moreover, Nakajima (2007) reported that isoflavones could inhibit CYP2A6 catalysis. In our study, we observed an inhibition of CYP2A6 expression only in the MCF-7 cell line, thus suggesting an ER implication in the regulation of this gene. Higashi et al. (2007) examined the regulation of CYP2A6 gene by estrogen. Luciferase assays using MCF-7 cells revealed that the transcriptional activity of the CYP2A6 promoter was significantly activated by estradiol in an ER-dependent manner wherein ERE was responsible for the activation. Chromatin immunoprecipitation assays further verified the in vivo association of ER with the ERE on the CYP2A6 gene. They demonstrated that CYP2A6 is directly induced by estrogen in an ER-dependent manner, implying a biological role of CYP2A6 in estrogen-responsive tissues.
RAC2 has been implicated in cancer progression by promoting invasion and metastasis through its regulation of cell migration and actin spelling check. Mutations of RAC genes in cancers are rare, but upregulation has been identified in a range of human cancers such as breast cancer (Fritz et al., 1999). A recent study, showed that resveratrol, an estrogen-like phytosterol from grapes, acts in an antiestrogenic manner in MDA-MB-231 breast cancer cells to reduce cell migration (Azios et al., 2007). In our study, we found a lower expression of RAC2 in genistein-treated cells versus untreated cells, thus suggesting an anti-invasive effect of this phytochemical by migration modulations.
Recent studies have shown that an increased expression of VASP (vasodilator stimulated protein) is involved in the progression and invasion of adenocarcinomas and associated with tumor progression (Dertsiz et al., 2005; Turner et al., 2008). It has been demonstrated in vitro that upregulation of VASP resulted in neoplastic transformation of cell lines, and that normal cell growth may require the maintenance of VASP expression. After a 72-h treatment of genistein, we observed a downregulation of VASP, thus suggesting a possible antitumorigenesis potential of genistein.
We also observed modulation of signal transduction-related genes such as CDKN3 (or KAP), which is downregulated after a 72-h treatment of either genistein or daidzein. Lee et al. (2000) reported that breast and prostate malignancies are associated with high levels of CDKN3 expression.
AKR1C3, involved in complex lipid metabolism, was downregulated after a 72-h genistein exposure. Recent studies strongly associate the AKR1C3 elevated expression with prostate carcinomas and especially with biological aggressiveness (Fung et al., 2006; Lin et al., 2004). Thus, a repressive effect due to genistein might presumably be anticipated.
In conclusion, using pangenomic microarray technology, we showed that genistein and daidzein exposure differentially modulated a number of molecular pathways with considerable biological significance in the context of carcinogenesis. These pathways included, for example, apoptosis, xenobiotic metabolism, and cell communication as well as lipid metabolism and signal transduction. Moreover, these regulated pathways were found to be influenced according to the tumor status in breast cell lines, thereby lending evidence for the hypothesis that soy phytoestrogens could have a chemopreventive effect during the first phase of tumor development, that is, in the initiation phase. The molecular genomics data reported herein complement the previous clinical epidemiology studies on breast cancer prevalence and exposure to soy products in the diet. These observations need replication and evaluation in further translational clinical studies.
Footnotes
Acknowledgments
S. Satih was the thesis prize winner of La ligue contre le Cancer (comité du Puy de Dôme), France in 2009. N. Chalabi was a postgraduate of the CLARA “Cancéropôle Lyon-Auvergne-Rhône-Alpes,” France. N. Rabiau is the recipient of a grant “CIFRE” from Soluscience S.A., Saint-Beauzire, France. R. Bosviel is a recipient of a grant CPER, Crédits du Conseil Régional D'Auvergne/Axe Nutrition et Cancer, CR 3077, Secteur I, Convention 08/47/8221. We thank Timothy Gunnels for corrections to the English language in the manuscript.
Author Disclosure Statement
The authors declare that no conflicting financial interests exist.
