Abstract
Nonclinical safety screening of small interfering RNAs (siRNAs) conjugated to a trivalent N-acetylgalactosamine (GalNAc) ligand is typically carried out in rats at exaggerated exposures in a repeat-dose regimen. We have previously shown that at these suprapharmacological doses, hepatotoxicity observed with a subset of GalNAc–siRNAs is largely driven by undesired RNA-induced silencing complex (RISC)-mediated antisense strand seed-based off-target activity, similar to microRNA-like regulation. However, the RISC component requirements for off-target activity of siRNAs have not been evaluated. Here, we evaluate the roles of major RISC components, AGO and TNRC6 (or GW182) proteins, in driving on- and off-target activity of GalNAc–siRNAs in hepatocytes, in vitro and in vivo. We demonstrate that knocking down AGO2, but not AGO1 or AGO4, is protective against GalNAc–siRNA-driven off-target activity and hepatotoxicity. As expected, knocking down AGO2, but not AGO1 or AGO4, reduces the on-target activity of GalNAc–siRNA. Similarly, knocking down TNRC6 paralogs, TNRC6A or TNRC6B, but not TNRC6C, is protective against off-target activity and hepatotoxicity while having minimal impact on the on-target activity of GalNAc–siRNA. These data indicate that while AGO2 is the only RISC component required for the on-target activity of GalNAc–siRNAs, the undesired off-target activity and hepatotoxicity of a subset of GalNAc–siRNAs are mediated via the RISC composed predominantly of AGO2 and TNRC6 paralogs TNRC6A and/or TNRC6B.
Introduction
RNA interference (RNAi)-based therapeutics have significant potential to address unmet medical needs. Over the last few years, several RNAi therapeutics have been approved for clinical use, including ONPATTRO® (patisiran), GIVLAARI® (givosiran), OXLUMO® (lumasiran), LEQVIO® (inclisiran), RIFLOZA® (nedosiran), and AMVUTTRA® (vutrisiran).1–7 RNAi is a highly conserved silencing pathway whereby 20–25 nucleotide-long double-stranded RNAs downregulate the expression of complementary target messenger RNA (mRNA). 8 Among these small RNAs are naturally occurring microRNAs (miRNAs) and synthetic small interfering RNAs (siRNAs) that can regulate gene expression by loading into the RNA-induced silencing complex (RISC). During the loading process, the antisense (guide) strand is retained in RISC, while the sense (passenger) strand is removed. 9 miRNAs recognize target mRNA via a partial sequence match primarily to the seed region of the antisense strand (nucleotides 2–8) leading to translational repression, mRNA destabilization, and degradation by 5ʹ and 3ʹ exonucleases, while siRNAs recognize target mRNA via a full sequence match leading to site-directed catalytic mRNA cleavage. 8
The argonaute (AGO) and trinucleotide repeat containing protein 6 (TNRC6, also known as GW182) are primary components of the RISC complex and play a central role in mediating gene repression via RNAi.10,11 There are four AGO proteins found in mammals, AGO1-4, of which AGO2 is thought to be primarily responsible for endonucleolytic cleavage of the fully complementary target transcript when bound to a guide RNA strand. 10 Emerging evidence suggests that AGO3 can also mediate slicer activity via short 14-nucleotide sequences.12,13 miRNAs can load into all AGOs to mediate post-transcriptional repression of partially complementary transcripts. In addition, gene expression regulation by miRNAs requires TNRC6 paralogs (TNRC6A, TNRC6B, and TNRC6C), which act as scaffolding proteins by binding to AGO and recruiting RNA degradation factors. 11 The role of the TNRC6 protein family in miRNA function is well understood, 11 but the individual roles of each TNRC6 paralog are still being elucidated. Using CRISPR knockout cell lines, Liu and colleagues demonstrated that TNRC6 paralogs have redundant functions in gene silencing by miRNAs. 14 Knocking out each paralog individually had no impact on miRNA activity, but knocking out TNRC6A and TNRC6B in combination was sufficient to partially inhibit miRNA activity. Importantly, while required for partially complementary miRNA-mediated activity, TNRC6 proteins are dispensable for fully complementary AGO2-mediated catalytic mRNA cleavage.14,15
Owing to key chemistry advances in the evolution of oligonucleotide therapeutics, siRNAs can now be effectively delivered to target tissues like the liver. Conjugating multivalent N-acetylgalactosamine (GalNAc) ligand to fully chemically modified siRNA allows targeted delivery to hepatocytes via highly conserved and abundant asialoglycoprotein receptor (ASGR). 16 Further refinement in recent years has led to enhanced stabilization chemistry (ESC) designs, which confer improvements in potency and duration of RNAi activity of GalNAc–siRNAs. 17 A small subset of ESC GalNAc–siRNAs have exhibited dose-dependent, asymptomatic, and transient increases in liver function tests (LFTs) in humans. 18 Subsequent mechanistic investigations indicated that the hepatotoxicity was largely driven by miRNA-like hybridization-based off-target effects, 19 whereby the seed region of the antisense strand of the siRNA can bind to complementary sites typically in the 3′UTR of undesired off-target mRNAs. This mimics miRNA-like post-transcriptional repression or noncatalytic degradation of the mRNA. 20 To mitigate these undesired off-target effects, we reported that seed-pairing destabilization through a single nucleotide substitution in the seed region with the (S)-isomer of glycol nucleic acid (GNA) could improve the mRNA target specificity and mitigate the hepatotoxicity of a GalNAc–siRNA in rats.19,21 Furthermore, this new design with improved specificity, termed ESC+, was found to be suitable for translation to humans, 18 and several ESC+ candidates are now advancing through clinical development. Here we assess the roles of AGO and TNRC6 protein families in off-target siRNA activity.
To identify AGO and TNRC6 paralogs that drive off-target activity and hepatotoxicity of GalNAc–siRNA, we selectively knocked down each protein in vitro in primary rat hepatocytes to assess the impact on siRNA transcriptional dysregulation and in vivo in rats to assess the impact on siRNA hepatotoxicity. We found that hybridization-based off-target activity and hepatotoxicity of a promiscuous GalNAc–siRNA were mediated via AGO2 but not AGO1 or AGO4. Similarly, knocking down TNRC6 paralogs TNRC6A or TNRC6B, but not TNRC6C, was protective against seed-based off-target activity and hepatotoxicity. These results indicate that a RISC complex composed of AGO2 and TNRC6A and/or TNRC6B is required for off-target activity of GalNAc–siRNAs in the liver.
Materials and Methods
Test materials and oligonucleotide synthesis
All GalNAc–siRNA conjugates were synthesized by Alnylam Pharmaceuticals (Cambridge, MA) according to previously published protocols.16,22,23 Three GalNAc ligands were covalently linked to the 3′end of the sense strand of the siRNA by a phosphodiester linkage between the pyrrolidine scaffold as described previously.16,22,23 All the information pertinent to the tested GalNAc–conjugated siRNAs can be found in Table 1.
Designs, Sequences, and Target mRNAs of the Small Interfering RNAs Used in This Study
S and AS represent sense and antisense strands.
Upper-case and lower-case letters indicate 2′-deoxy-2′-fluoro (2′-F) and 2′-O-methyl (2′-OMe) ribosugar modifications, respectively.
VP indicates vinylphophonate modification.
• indicates phosphorothioate (PS) linkage.
L indicates the trivalent N-acetylgalactosamine (GalNAc) ligand (structure above).
AGO1, argonaute RISC component 1; AGO2, argonaute RISC component 2; AGO4, argonaute RISC component 4; siRNA, small interfering RNA; TMP, transmembrane serine protease 6; TNRC6A, trinucleotide repeat containing adaptor 6A; TNRC6B, trinucleotide repeat containing adaptor 6B; TNRC6C, trinucleotide repeat containing adaptor 6C; TTR, transthyretin.
Care and use of laboratory animals
All studies were conducted using protocols consistent with local, state, and federal regulations, as applicable, and approved by the Institutional Animal Care and Use Committee at Alnylam Pharmaceuticals. The test articles were diluted with 0.9% NaCl to achieve appropriate dosing concentrations and dosed subcutaneously in the intrascapular region to male Sprague Dawley rats (6–8 weeks old) in a dose volume of 5 mL/kg with N = 3 animals/group. Eight-week-old rats were subcutaneously administered 0.9% saline or 10 mg/kg siAGO or siTNRC6, respectively, on Days 1 and 12. Each cohort that received siAGO or siTNRC6 also received three weekly injections of saline or 30 mg/kg siRNA2, respectively, on Days 8, 15, and 22. A separate cohort received only three weekly injections of siRNA2. Livers were collected for analysis 24 h after the last dose was administered. Randomization was performed using the partitioning algorithm in the Pristima® Suite (Xybion) that avoids group mean body weight bias.
Clinical pathology
Whole venous blood was collected into serum separator tubes (BD Microtainer) and allowed to clot at room temperature for 30 min prior to centrifugation at 3,000 RPM (1,489g) for 10 min at 4°C. Serum was then aliquoted and stored at −80°C until analysis. Serum chemistries were analyzed using the AU400 chemistry analyzer (Beckman Coulter-Brea, CA, USA), with reagents provided by Beckman Coulter, Randox, and Sekisui Diagnostics.
Histopathology
All animals were sacrificed as per Alnylam standard operating procedures, and livers were collected and fixed in 10% neutral buffered formalin (10% NBF) for 72 h prior to routine processing using TissueTek VIP 6A1 (Sakura). Tissues were trimmed, embedded into paraffin blocks, sectioned at 4 μm, stained with hematoxylin and eosin using TissueTek Prisma A1D (Sakura), and coverslipped using TissueTek Glass g2 (Sakura). Sections were examined microscopically from each liver in an unblinded fashion. The range of severity grade for each histological finding was graded on a scale of 0–4 with 0: no findings, 1: minimal, 2: mild, 3: moderate, and 4: marked severity.
Primary hepatocyte cell culture and treatment
Primary rat hepatocytes (BioIVT) were seeded in 96-well collagen I pre-coated plates (Gibco) at approximately 5 × 104 cells/well in INVITROGROTM CP Rodent Medium (BioIVT). At the time of seeding, the cells were transfected with 1 nM siAGO or siTNRC6 or 1X PBS (mock control) using Lipofectamine RNAiMAX (Thermo Fisher Scientific) according to the manufacturer’s instructions. The cells were incubated for 24 h at 37°C under these conditions. The media was then replaced, and cells were retransfected with 0, 0.0032, 0.0016, 0.08, 0.04, 2, 10, and 50 nM siRNA1 or 0.00001, 0.0001, 0.001, 0.01, 0.1, 1, and 10 nM siRNA2 or 1X PBS (mock control) for another 24 h at 37°C. The media was removed, and the cells were collected for RNA isolation.
RNA isolation
For the extraction of total RNA, frozen cells (5 × 104 cells/well) or powdered tissues (∼10 mg) were resuspended in 700 µL QIAzol and homogenized by vigorous pipetting. Samples were incubated at room temperature for 5 min, spun at 6,000g for 1 min 4°C, followed by the addition of 140 µL chloroform. Samples were mixed by shaking, followed by a 10-min incubation at room temperature. Samples were spun at 6,000g for 15 min at 4°C, the supernatant was transferred to a new tube, and 1.5 volumes of 100% ethanol were added. Samples were then purified using a miRNeasy Kit (Qiagen) according to the manufacturer’s instructions. The RNA was eluted from miRNeasy columns with 50–60 µL RNase-free water.
Real-time quantitative PCR
cDNA was synthesized using Applied Biosystems™ High-Capacity cDNA Reverse Transcription Kit (Applied Biosystems). Briefly, 10 µL of a master mix containing 1 µL 10X buffer, 0.4 µL 25X dNTPs, 1 µL 10X Random Primers, 0.5 µL MultiScribeTM Reverse Transcriptase, 0.5 µL RNase Inhibitor, and 6.6 µL of nuclease-free water per reaction was added to 10 µL of the isolated RNA. Plates were incubated in a Thermal Cycler programmed for four steps: Step 1: 25°C for 10 min, Step 2: 37°C for 120 min, Step 3: 85°C for 5 min, and Step 4: hold at 4°C.
cDNA was diluted with 130 µL of nuclease-free water, and 4.5 µL of the resultant mix was added to a master mix containing 0.5 µL of actin, beta (ACTB) or glyceraldehyde-3-phosphate dehydrogenase (GAPDH) TaqMan probe (ThermoFisher Scientific, Rn00667869_m1 or Rn01775763_g1, respectively), and 5 µL LightCycler® 480 probe master mix (Roche) per well in a 384-well plate (Roche) for each treatment. Similarly, in a separate set, 4.5 µL of cDNA was added to 0.5 µL of the respective target transcript probe (Table 2) for each GalNAc–siRNA treatment, along with 5 µL LightCycler® 480 probe master mix. qPCR was carried out with a LightCycler® 480 real-time PCR system (Roche) using the Ct(RQ) assay. Each treatment was tested in three independent replicates. To calculate relative fold change, real-time qPCR data were analyzed using the dCt method. On-target mRNA values were normalized to ACTB or GAPDH values for each treatment, and the data are presented as percent change relative to PBS-treated (mock) cells or saline-treated tissue samples.
Real-Time Quantitative PCR Primers and Probes for mRNA Knockdown Assessment
Quantification of total liver siRNA levels
Cohorts of rats were sacrificed 24 h after the last dose was administered, and livers were snap-frozen in liquid nitrogen and ground into powder for further analysis. Total liver siRNA levels were measured by stem-loop real-time quantitative PCR (SL-RT-qPCR) as previously described.23–26 RT-qPCR primers and probes used in this study are summarized in Table 3.
Real-Time Quantitative PCR Primers and Probes for Small Interfering RNA Quantification
RNA sequencing and bioinformatic analysis
RNA extracted with the miRNeasy kit (Qiagen) was used for cDNA library preparation with the TruSeq Stranded Total RNA Library Prep Kit (Illumina) and sequenced on the HiSeq or NextSeq500 sequencers (Illumina), all according to manufacturers’ instructions. Raw RNAseq reads were filtered with minimal mean quality scores of 25 and minimal remaining length of 36, using fastq-mcf. Filtered reads were aligned to the Rattus norvegicus genome (Rnor_6.0) using STAR (ultrafast universal RNAseq aligner) with default parameters. 27 Uniquely aligned reads were counted by featureCounts. 28 Differential gene expression analysis was performed by the R package DESeq2. 29
Results
Impact of AGO or TNRC6 knockdown on the on-target activity of siRNA in vitro
To investigate the role of the major components of RISC in siRNA activity, we designed ESC GalNAc–siRNA conjugates that specifically target each AGO (siAGO1, siAGO2, and siAGO4) or each TNRC6 (siTNRC6A, siTNRC6B, and siTNRC6C) transcript in rat (Table 1). AGO paralog expression is known to vary by cell type.30,31 We found that AGO1 and AGO2 mRNA were well expressed in the rat liver, while AGO3 and AGO4 mRNA were expressed at relatively lower levels than AGO2 (Supplementary Fig. S1). We were unable to identify a potent siRNA against AGO3, likely due to poor mRNA expression of AGO3 in rat liver; thus, we chose to evaluate AGO1, 2, and 4 in this study. On the contrary, the mRNA of all three TNRC6 paralogs was well expressed in rat liver (Supplementary Fig. S1).
The on-target activity of each siRNA against AGO and TNRC6 transcripts was assessed in primary rat hepatocytes by transfecting at 1 nM for 48 h as described in Figure 1A. AGO-targeted siRNAs showed ∼92%, ∼79%, and ∼88% mRNA knockdown of AGO1, 2, and 4, respectively, and were specific to each AGO transcript (Fig. 1B). Similarly, siRNAs against TNRC6 paralogs also showed robust and specific mRNA knockdown, ranging between ∼80 and 90% (Fig. 1C).
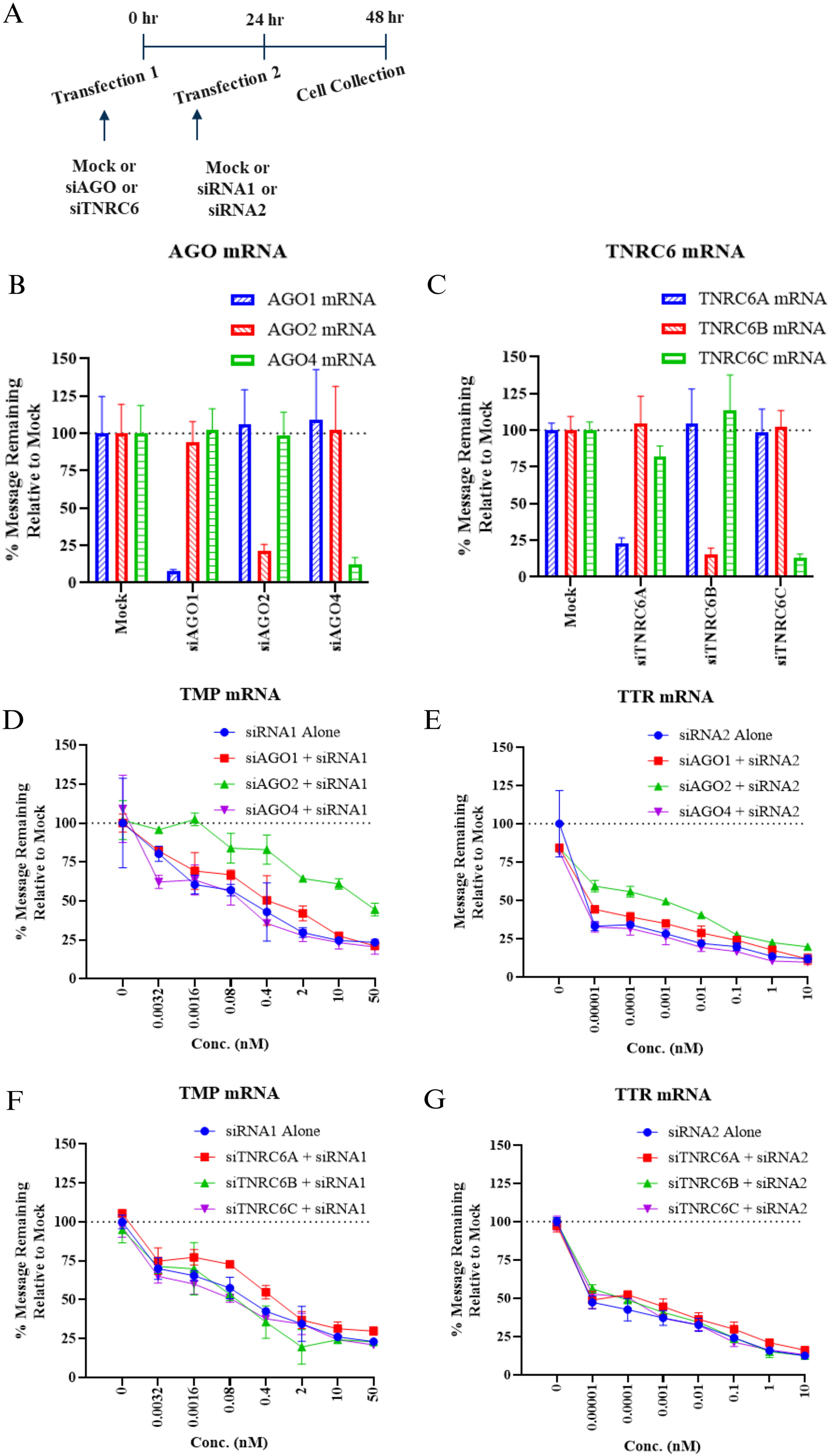
Impact of AGO or TNRC6 knockdown on siRNA on-target activity in primary rat hepatocytes.
Next, we sought to understand the impact of knocking down these RISC components on the on-target activity of unrelated siRNAs targeting rodent transcripts, siRNA1 targeting transmembrane serine protease 6 (TMP) and siRNA2 targeting transthyretin (TTR) (Table 1). As illustrated in Figure 1A, siRNAs against either AGO or TNRC6 paralogs were first transfected at 1 nM for 24 h. The media was then replaced, and siRNA1 or siRNA2 or mock control was transfected in a concentration-dependent manner (up to 50 or 10 nM for siRNA1 and siRNA2, respectively, based on their inherent potency) for an additional 24 h. We found that the concentration-dependent activity of siRNA1 and siRNA2 was not impacted when AGO1 or AGO4 was knocked down (Fig. 1D and E). However, when AGO2 was knocked down, there was a right shift in the concentration-dependent curves for both siRNAs, indicating a loss of activity (Fig. 1D and E). siRNA2 was inherently more potent than siRNA1 and thus showed loss of activity only at nonsaturating concentrations in the presence of siAGO2. This loss of siRNA on-target activity was not surprising since the catalytic activity of RISC-AGO2 is required for target transcript cleavage. In contrast, there was minimal to no impact on the on-target activity of siRNA1 or siRNA2 when any of the TNRC6 paralogs were knocked down (Fig. 1F and G). These results are consistent with published reports suggesting that TNRC6 paralogs are dispensable for siRNA-mediated mRNA cleavage. 14
Impact of AGO or TNRC6 knockdown on the off-target activity of siRNA in vitro
We have previously demonstrated that a subset of GalNAc–siRNAs can cause miRNA-like gene dysregulation mediated by the seed region of the antisense strand. 32 To demonstrate that siRNA1 can act in a miRNA-like fashion to downregulate seed-matched off-target transcripts, primary rat hepatocytes were transfected with 10 nM siRNA1 for 24 h, and the off-target activity was assessed by RNA sequencing (RNAseq). This resulted in downregulated transcripts that were enriched for perfect complementarity to the seed region of the antisense strand, and the magnitude of change generally did not exceed approximately 2-fold. Correspondingly, the magnitude of the cumulative distribution function (CDF) shift was increased across the three most common canonical seed matches (mer8, mer7-m8, and mer7-A1) in the 3′UTR relative to the background set of genes lacking 3′UTR seed matches in these categories (Fig. 2A). The shift in CDF was specifically observed for transcripts complementary to the seed region of the antisense strand, but not to the sense strand (Supplementary Fig. S3A and B).
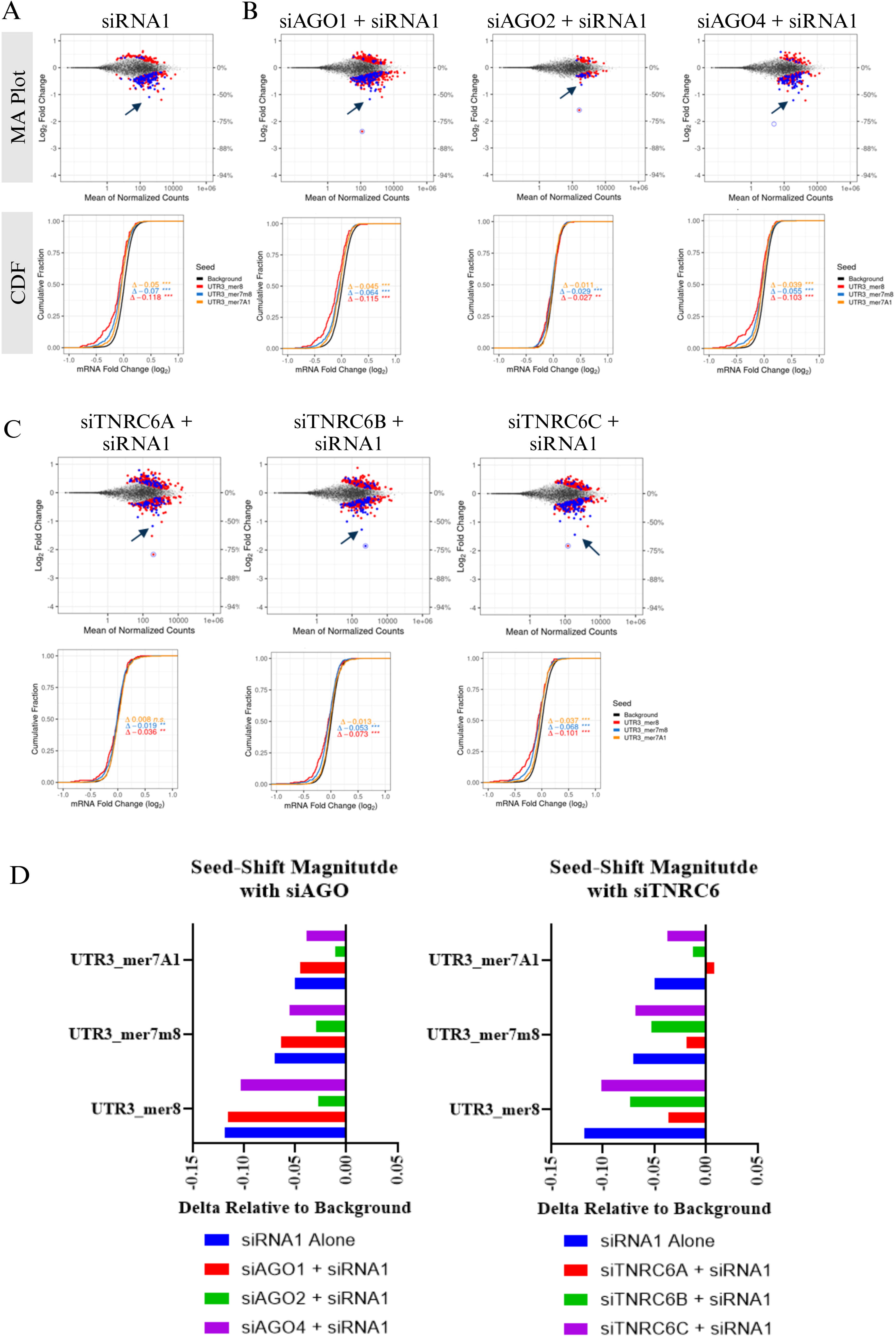
Impact of AGO or TNRC6 knockdown on siRNA seed-mediated off-target activity in primary rat hepatocytes. RNA sequencing was performed on RNA isolated from primary rat hepatocytes as described in Figure 1. Mock or siRNAs against AGO or TNRC6 paralogs were transfected at 1 nM for 24 h and mock or siRNA2 was transfected in the same wells at 10 nM for an additional 24 h. Upper: MA plots of log2 fold change vs abundance (average counts) of individual genes. Dots represent individual transcripts. Gray dots represent genes not differentially expressed after siRNA treatment relative to the control [measured against the antisense strand (AS) of the siRNA]; the blue and red dots represent differentially expressed genes (adjusted P < 0.05) with or without a canonical seed match, respectively. The P value associated with the log2 fold change of each gene was calculated by DESeq2 using the Wald test and was adjusted for multiple test correction using the method of Benjamini and Hochberg. On-target knockdown of siRNA1
To understand the role of AGO and TNRC6 paralogs in the off-target activity of siRNA1, each of the paralogs was first knocked down by transfecting respective siRNAs at 1 nM in primary rat hepatocytes. After 24 h, mock or siRNA1 was transfected at 10 nM as described in Figure 1A. There was variable transcriptional dysregulation when hepatocytes were only treated with siAGO or siTNRC6 (Supplementary Fig. S2A and B); however, the shift in CDF was less significant, relative to that of siRNA1 (Fig. 2A), indicating relatively minimal seed-mediated off-target activity. Moreover, there was minimal overlap in the downregulated differentially expressed genes (DEGs) between siAGO or siTNRC6 relative to the off-target signature of siRNA1 (Supplementary Fig. S2C). This suggests that when siAGO or siTNRC6 and siRNA1 were transfected together, the primary transcriptional dysregulation was driven by the seed region of antisense of siRNA1. As shown in the MA plots, the number of downregulated DEGs in the presence of siRNA1 alone was 257, while when AGO1 or AGO4 was knocked down first and then transfected with siRNA1, the downregulated DEGs were 456 and 211, respectively (Fig. 2A and B). In both conditions, the magnitude of dysregulation for canonical 3′-UTR seed match shown in CDF plots was minimally impacted. On the contrary, when AGO2 was knocked down, there was a notable reduction in the off-target activity of siRNA2. The number of downregulated DEGs was 65, and there was a significant reduction in the CDF shift (Fig. 2B and D). Together, these results suggest that the off-target activity of siRNA1 may be exaggerated when AGO1 is knocked down, greatly reduced when AGO2 is knocked down, and minimally impacted when AGO4 is knocked down, relative to the off-target activity of siRNA1 itself in rat hepatocytes.
The knockdown of TNRC6A, TNRC6B, or TNRC6C did not greatly impact the overall transcript dysregulation following siRNA1 treatment. The number of downregulated DEGs was 241, 305, and 274 for TNRC6A, TNRC6B, and TNRC6C, respectively, compared to 257 with siRNA1 alone, as shown in the MA plots (Fig. 2C). However, the magnitude of dysregulation of transcripts with a canonical 3′-UTR seed match to the antisense strand of siRNA1, shown in CDF plots, was reduced when TNRC6A or TNRC6B was knocked down, relative to siRNA1 alone (Fig. 2C and D). This shift in CDF was specifically observed for transcripts complementary to the seed region of the antisense strand, but not to the sense strand (Supplementary Fig. S3A and C). Previous work has demonstrated that miRNA activity was almost completely inhibited when all three TNRC6 paralogs were knocked out, partially inhibited when TNRC6A and TNRC6B were knocked out, and had no impact when they were individually knocked out. 14 Here we find that TNRC6A or TNRC6B, individually, may be sufficient to have an impact on antisense strand seed-mediated activity, as evident by CDF shifts, while TNRC6C has a minimal impact. It is likely that depleting both TNRC6A and TNRC6B paralogs will lead to a greater impact based on the redundancies in their function. Together these data indicate that the AGO2, TNRC6A, and TNRC6B components of RISC contribute to miRNA-like antisense strand seed-mediated off-target activity of GalNAc–siRNA in rat hepatocytes.
Impact of AGO or TNRC6 knockdown on GalNAc–siRNA-mediated hepatotoxicity in vivo
To investigate whether our in vitro findings in primary rat hepatocytes translate in vivo, rats were subcutaneously administered 10 mg/kg GalNAc–siRNA against each AGO (siAGO1, siAGO2, and siAGO4) or TNRC6 paralog (siTNRC6A, siTNRC6B, and siTNRC6C) on Days 1 and 12 and sacrificed on Day 23 (Table 1 and Fig. 3A). Consistent with the in vitro data, there was ∼60%–75% mRNA knockdown of AGO proteins (Supplementary Fig. S4B) and ∼55%–70% TNRC6 paralogs in rat liver (Supplementary Fig. S4C). Next, we sought to characterize the impact of AGO and TNRC6 knockdown on the safety profile of a GalNAc–siRNA known to mediate hepatotoxicity in rats.
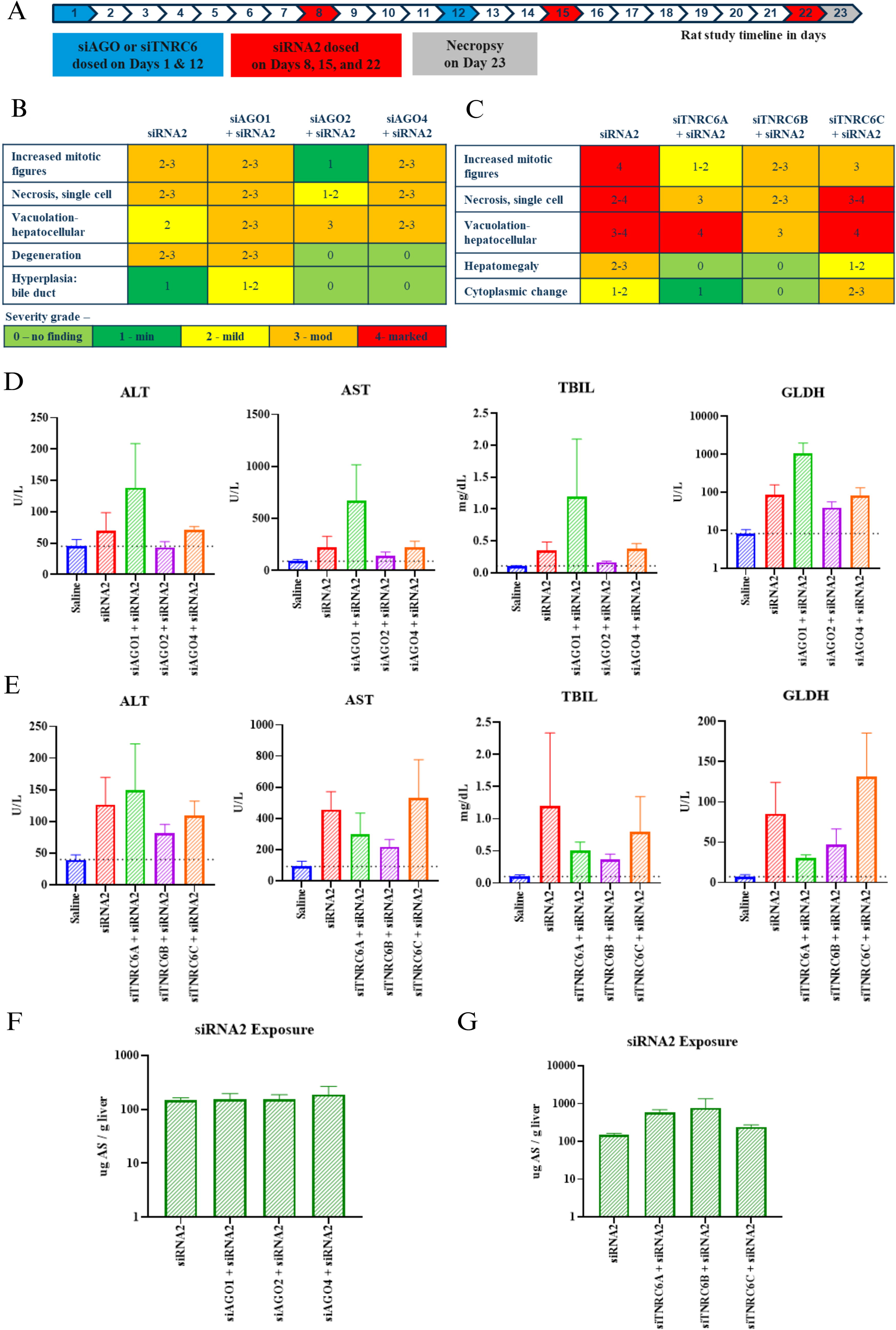
Impact of AGO and TNRC6 knockdown on siRNA-mediated hepatotoxicity in rats.
Nonclinical safety screening of GalNAc–siRNAs is typically carried out at exaggerated exposures in a repeat-dose regimen, and under these conditions, we have previously shown that hepatotoxicity observed with a subset of GalNAc–siRNAs is largely driven by undesired antisense strand seed-mediated off-target activity. 19 Following this paradigm, a cohort of rats received three weekly exaggerated doses of 30 mg/kg of a known hepatotoxic sequence, siRNA2, targeting rodent transthyretin (TTR) (Table 1), on Days 8, 15, and 22, and sacrificed 24 h after the last dose on Day 23. This represents a 2–3 log exaggeration of the pharmacological dose range. Resultant data from animals treated with siRNA2 were compared to another cohort of rats treated with two injections of siAGO or siTNRC6, respectively, on Days 1 and 12, and three weekly injections of siRNA2, on Days 8, 15, and 22, as described in Figure 3A. Unlike the in vitro data, there was no impact on the on-target activity of siRNA2 after knocking down any of the AGO or TNRC6 paralogs (Supplementary Fig. S4D and E), with ≥95% mRNA knockdown of the siRNA2 target, TTR. This is likely due to the exaggerated dose and repeat dosing regimen leading to saturated on-target activity of siRNA2.
Terminal rat liver samples were harvested at the end of the study on Day 23, 24 h after the last dose, for microscopic evaluation, and serum samples were evaluated for clinical pathology (Fig. 3). The impact of AGO and TNRC6 knockdown on siRNA2-mediated hepatotoxicity was evaluated in two separate studies. As expected, in both studies, rats administered once weekly (qw) × 3 doses of 30 mg/kg of siRNA2 alone had considerable hepatotoxicity, although the severity of findings was not identical between studies. In the first study, treatment with siRNA2 led to mild hepatocellular vacuolation, mild to moderate single-cell necrosis, increased mitosis, mild to moderate hepatocellular degeneration, and minimal bile duct hyperplasia (Fig. 3B). In agreement with trends observed in vitro, knocking down AGO1 or AGO4 and then treating with siRNA2 did not lead to any substantial change in microscopic findings in the liver, relative to siRNA2 alone. However, knocking down AGO2 either eliminated or significantly reduced the severity of most microscopic findings with the exception of hepatocellular vacuolation, relative to siRNA2 alone. Three weekly doses of siRNA2 alone also led to substantial increases in multiple liver function tests (LFTs), such as alanine aminotransferase (ALT), aspartate aminotransferase (AST), total bilirubin (TBIL), and glutamate dehydrogenase (GLDH) (Fig. 3D). siRNA2-mediated increases in LFTs were reduced when AGO2 was knocked down, but not when AGO1 or AGO4 were knocked down. Treatment of each of siAGO, in the absence of siRNA2, did not lead to significant microscopic changes in the liver or in LFTs (Supplementary Fig. S4F and G). Interestingly, in the case of treatment with siAGO1 and siRNA2, there were further increased LFTs relative to siRNA2 alone (Fig. 3D). Consistent with the in vitro data, this may suggest that knocking down AGO1 exaggerates the impact of siRNA2 on liver toxicity, although LFTs in this case did not correlate with microscopic findings. A potential explanation for this outcome may be that knocking down AGO1 leads to the redistribution of siRNA into AGO2, leading to an exaggeration of off-target activity and liver toxicity.
In the second study, three weekly 30 mg/kg doses of siRNA2 led to moderate to marked hepatocellular vacuolation, mild to marked single-cell necrosis, marked increase in mitosis, mild to moderate hepatomegaly, and minimal to mild cytoplasmic change (Fig. 3C). Knocking down TNRC6B led to the most substantial reduction in the severity of microscopic findings, followed by TNRC6A, while knocking down TNRC6C had similar results as treatment with siRNA2 alone. Consistent with the microscopic findings, LFT increases driven by siRNA2 were reduced when TNRC6A or TNRC6B was knocked down (Fig. 3E). Knocking down TNRC6C did not have a significant impact, relative to siRNA2 alone. Treatment of each siTNRC6 alone did not lead to significant microscopic changes in the liver or LFTs (Supplementary Fig. S4H and I).
To ensure that reduced hepatotoxicity was not simply due to reduced exposure to siRNA2, we evaluated the total amount of the antisense strand of siRNA2 in the terminal rat livers. The concentration of siRNA2 in the liver was measured by Stem-Loop Real-Time Quantitative PCR (SL-RT-qPCR) using the primers and probes detailed in Table 3. Indeed, we found that the levels of antisense strand of siRNA2 were maintained after knocking down any of the AGO proteins (Fig. 3F) or TNRC6 paralogs (Fig. 3G), indicating that any differences that were observed in hepatoxicity driven by siRNA2 were not related to the difference in total exposure in hepatocytes. Together with the in vitro data, these results suggest that AGO2, TNRC6A, and TNRC6B components of RISC contribute to GalNAc–siRNA-mediated hepatoxicity in rats at exaggerated doses.
Discussion
The specificity and safety of siRNAs are key factors for the development of RNAi therapeutics. We have previously demonstrated that hybridization-based off-target effects driven by the seed region of the antisense strand are the major cause of the rat hepatotoxicity observed with a subset of GalNAc–siRNA conjugates 19 and outlined a possible mitigation strategy using GNA to destabilize seed-mediated pairing to off-target transcripts. 21 In this report, we aimed to further expand our mechanistic understanding of siRNA-mediated off-target activity by assessing the role of major RISC components, AGO1-4 proteins, and TNRC6A-C paralogs. We selectively knocked down each AGO protein and TNRC6 paralog in primary hepatocytes or rats and asked which of these RISC components are needed to drive the off-target activity and hepatotoxicity of GalNAc–siRNA at exaggerated doses. We found that hybridization-based off-targets driven by the seed region of the antisense, and consequent hepatotoxicity of a noisy GalNAc–siRNA, were mediated by AGO2, not AGO1 or AGO4. Similarly, knocking down TNRC6 paralogs TNRC6A or TNRC6B, but not TNRC6C, was protective against seed-based off-target activity and hepatotoxicity of GalNAc–siRNA. The on-target activity of GalNAc–siRNA was only impacted by the knockdown of AGO2, as expected. Together, these results confirm that undesired off-target effects and hepatotoxicity of a subset of GalNAc–siRNAs are driven by miRNA-like seed-mediated activity and that this is carried out via the RISC complex composed of AGO2 and TNRC6 paralogs TNRC6A and/or TNRC6B.
In humans (in contrast to flies33,34), there is no known sorting system that exists to selectively load miRNA or siRNAs into the different AGOs, and thus, non-cleavage-based off-target activity may be mediated via all AGO1-4 paralogs. 35 In the present report, a question arises whether GalNAc–siRNA loaded into AGO1, AGO3, or AGO4 could lead to miRNA-like off-target activity and potentially contribute to hepatotoxicity. However, even when AGO2 was depleted, any siRNA loaded into AGO1, AGO3, or AGO4 did not appear to be sufficient to drive miRNA-like off-target activity or hepatotoxicity, at least not to the extent of AGO2. While AGO2 can cleave the sense strand to facilitate antisense strand loading, AGOs 1, 3, and 4 likely have to rely on sense strand unwinding, which is impacted by the thermodynamic stability of the siRNA duplex. 35 Since the siRNAs used in this study had relatively high melting temperatures due to full chemical modification, it is possible that only cleavage-competent AGO2 formed sufficiently high amounts of active RISC to mediate noncatalytic seed-based off-target activity.
Advances in chemistry in the evolution of oligonucleotide therapeutics have enabled the effective delivery of siRNAs to target tissues like the liver. Beyond the liver, there is a significant ongoing effort to deliver siRNAs in tissues such as the central nervous system, muscle, eye, skin, and lung, to name a few. 36 The expression of each AGO1-4 and TNRC6A-C protein can vary between cell types in each tissue, between various tissues, and between species (Supplementary Fig. S5; GTEx Consortium, 2024). For example, the cerebellum appears to have higher expression of each of the AGO and TNRC6 proteins relative to the rest of the brain regions, such as the frontal cortex, hippocampus, or the spinal cord (Supplementary Fig. S5; GTEx Consortium, 2024). It is conceivable that certain tissues may be more sensitive to the miRNA-like off-target activity of a subset of siRNAs due to the AGO and TNRC6 expression differences. In future studies, it would be interesting to ask whether there is greater propensity of seed-mediated off-target activity in the cerebellum, relative to other regions of the brain, potentially driven by the high expression of AGO proteins and TNRC6 paralogs. Similarly, it is possible that tissues where endogenous expression of these RISC components is low would be more resistant to siRNA toxicity driven by seed-mediated off-target activity.
In conclusion, we built on our previous work that demonstrated hybridization-based off-target effects are driven by the seed region of the antisense strand and are the major cause for the rat hepatotoxicity observed with a subset of GalNAc–siRNA conjugates. Here we demonstrate that this off-target activity is carried out via the RISC complex composed of AGO2 and TNRC6A and/or TNRC6B, which further confirms the mechanism of GalNAc–siRNA-mediated hepatoxicity.
Footnotes
Acknowledgments
The authors thank Alnylam’s chemistry team for all siRNA synthesis; Elena Stamenova, Alexa Bacha, and Jessica Grant for RNAseq library preparation; Lauren Moran, Sara Woldemariam, Antoinette Hayes, Linhong Sun, Christine Fregeau, and Matthew Whittaker for in vivo study support; Kellie D’Angelo and Bangyi Ma for clinical pathology support; Paul Gedman, Kimberly Harding, and Brenda Carito for histology support; and Julia Varao, Michael Regan, and Yesenia Manche for sample management.
Author Disclosure Statement
During the time this work was conducted, all authors were employees of Alnylam Pharmaceuticals with salary and stock options.
Funding Information
This work was funded by Alnylam Pharmaceuticals.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
