Abstract
Hybridization-dependent off-target (OffT) effects, occurring when oligonucleotides bind via Watson–Crick–Franklin hybridization to unintended RNA transcripts, remain a critical safety concern for oligonucleotide therapeutics (ONTs). Despite the importance of OffT assessment of clinical trial ONT candidates, formal guidelines are lacking, with only brief mentions in Japanese regulatory documents (2020) and US Food and Drug Administration (FDA) recommendations for hepatitis B virus treatments (2022). This article presents updated industry recommendations for assessing OffTs of ONTs, building upon the 2012 Oligonucleotide Safety Working Group (OSWG) recommendations and accounting for recent technological advancements. A new OSWG subcommittee, comprising industry experts in RNase H-dependent and steric blocking antisense oligonucleotides and small interfering RNAs, has developed a comprehensive framework for OffT assessment. The proposed workflow encompasses five key steps: (1) OffT identification through in silico complementarity prediction and transcriptomics analysis, (2) focus on cell types with relevant ONT activity, (3) in vitro verification and margin assessment, (4) risk assessment based on the OffT biological role, and (5) management of unavoidable OffTs. The authors provide detailed considerations for various ONT classes, emphasizing the importance of ONT-specific factors such as chemistry, delivery systems, and tissue distribution in OffT evaluation. The article also explores the potential of machine learning models to enhance OffT prediction and discusses strategies for experimental verification and risk assessment. These updated recommendations aim to improve the safety profile of ONTs entering clinical trials and to manage unavoidable OffTs. The authors hope that these recommendations will serve as a valuable resource for ONT development and for the forthcoming finalization of the FDA draft guidance and the International Council for Harmonization S13 guidance on Nonclinical Safety Assessment of Oligonucleotide-Based Therapeutics.
Introduction
Oligonucleotide therapeutics (ONTs) are nucleic acid polymers with the potential to treat or manage a wide range of diseases. They can be divided into different classes, for example, antisense oligonucleotides (ASOs), small interfering RNA (siRNA), and steric blocking oligonucleotides (SBOs). While the various classes of ONTs share a common property of sequence-specific binding to the desired target RNA or protein, their respective mechanism of action can vary from recruitment of enzymes to induce degradation of mRNA transcripts (RNase H1 recruiting ASOs, and siRNA) to steric blocking that can alter splicing, antagonize microRNA (miRNA), and inhibit RNA translation (SBO or steric blocking ASOs). 1
While ONT molecules are designed to selectively hybridize with an intended, specific RNA transcript (i.e., on-target effect), it is crucial to consider the possibility of additional, unintended hybridization events [i.e., off-target (OffT) effects]. It is quite straightforward to avoid OffTs with perfect sequence homology, however with increasing counts of mismatches and insertions/deletions (a.k.a. edit distance), the number of false positive predictions can quickly increase. OffT effects can lead to reduced transcript levels, reduced translation rates, or aberrant splicing, utilizing similar molecular mechanisms and protein machineries as the intended on-target effects. ONTs can also trigger different safety effects that are not dependent on RNA hybridization.2–13 These hybridization-independent effects will not be further discussed in this article.
In 2012, a subcommittee of the Oligonucleotide Safety Working Group (OSWG) published considerations and recommendations on how to assess OffT concerns. 14 Still today, there are no formal regulatory guidelines addressing OffT effects apart from PMDA’s 2020 guideline 15 and the FDA’s 2022 recommendations specific to hepatitis B virus treatments 16 briefly addressing OffTs of ONTs. However, during the completion of this article, the US FDA published a draft guidance 17 covering OffT assessment, and the International Council for Harmonization (ICH) initiated work to develop nonclinical ONT-specific guidance (S13) 18 that will likely encompass OffT assessment.
Thanks to a better mechanistic understanding, improved in silico predictions, and increased use of RNA sequencing (RNAseq), an update of the 2012 recommendations is warranted. A new OSWG subcommittee was formed to update the recommendations and best practices for characterizing human hybridization-dependent OffT effects of RNase H1 and steric blocking ASOs, as well as siRNAs.
The risk to cause OffT effects is not unique to ONTs. All therapeutic modalities carry this risk, but the base-pairing mechanism for the ONT classes discussed herein makes this work to some extent predictable. Several preclinical studies have demonstrated the consequences of “promiscuous” ONTs silencing multiple OffT RNAs (e.g., liver toxicity of siRNAs and RNaseH-engaging ASOs).19–22 In addition, it has been demonstrated that minimizing broad, hybridization-dependent OffT effects for siRNAs correlates with reduced liver toxicity signals observed in Phase 1 clinical studies.21,23,24 Taken together with the long duration of antisense pharmacology effects of most current ONTs (weeks to months), understanding and minimizing potential safety risks associated with OffT effects remain a priority for both organizations developing these medicines and regulatory authorities.
In this work, we concentrate on the types of oligonucleotides designed to hybridize to endogenous RNA by means of predictable Watson–Crick–Franklin base pairing (ASO, siRNA, and SBO) and recommend a rational approach to identify and assess the risk of unintended hybridization events, building on the previous recommendations from OSWG. 14 Messenger RNA therapeutics replacing endogenous mRNA, DNA, or RNA editors will not be discussed in this article.
Although class-specific considerations are discussed, the core of the article is a proposed assessment framework built on shared principles applicable across these ONT classes with the following main steps (outlined in Fig. 1): (1) OffT identification via in silico homology prediction and transcriptomics analysis, (2) focus on cell types showing relevant ONT activity, (3) in vitro verification and margin assessment, (4) risk assessment based on OffT biological role, and (5) management of any remaining OffT.
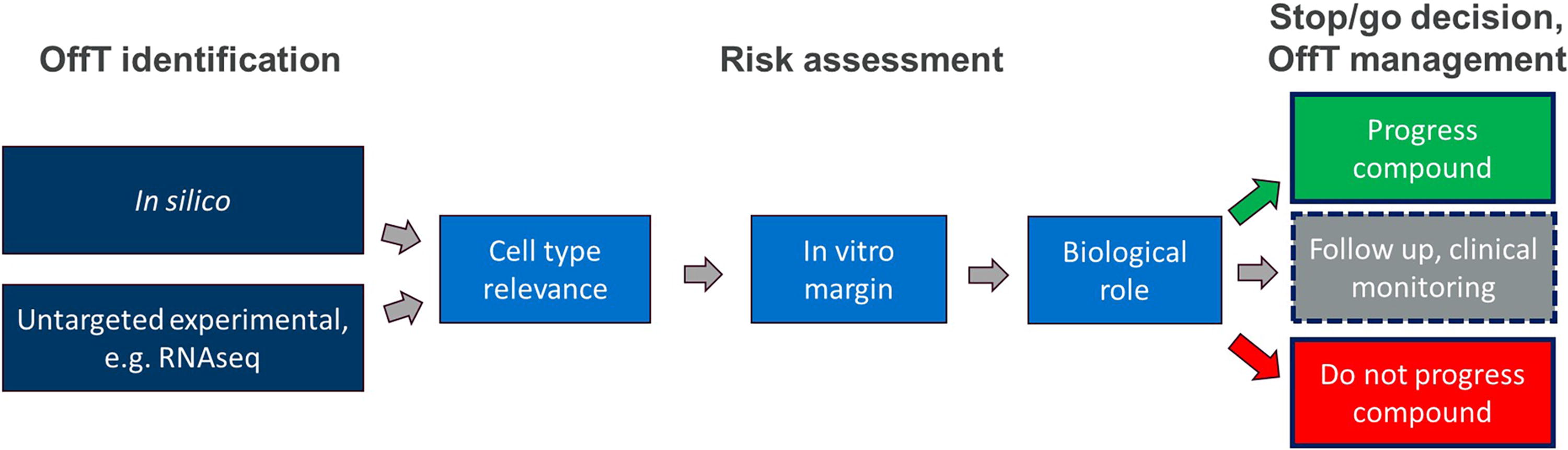
Recommended principles. Summary of the recommended principles and steps to identify and risk assess ONT hybridization-dependent OffTs. See Figure 4 for further details. OffT, off-target; ONT, oligonucleotide therapeutic; RNAseq, RNA sequencing.
ONT Class-Specific Mechanisms and Considerations
RNase H1-dependent ASOs
OffT mechanisms
RNase H1-dependent ASOs are designed to reduce target RNA expression. They typically consist of a central DNA region (the gap) flanked by 2′-ribose-modified RNA regions on the 5′ and 3′ ends (the wings or flanks), known as the gapmer design (Fig. 2A). When hybridizing to targeted RNA, the DNA (gap):RNA hybrid is recognized and bound by RNase H1. This endonuclease enzyme cleaves the RNA strand in the hybrid by hydrolysis, and the cleaved, unprotected RNA is rapidly degraded by exonucleases.25–27 Since the RNase H1 enzyme is present and active in both the nucleus and cytoplasm, 28 both primary and processed RNA may be targeted by ASOs and degraded in this manner.
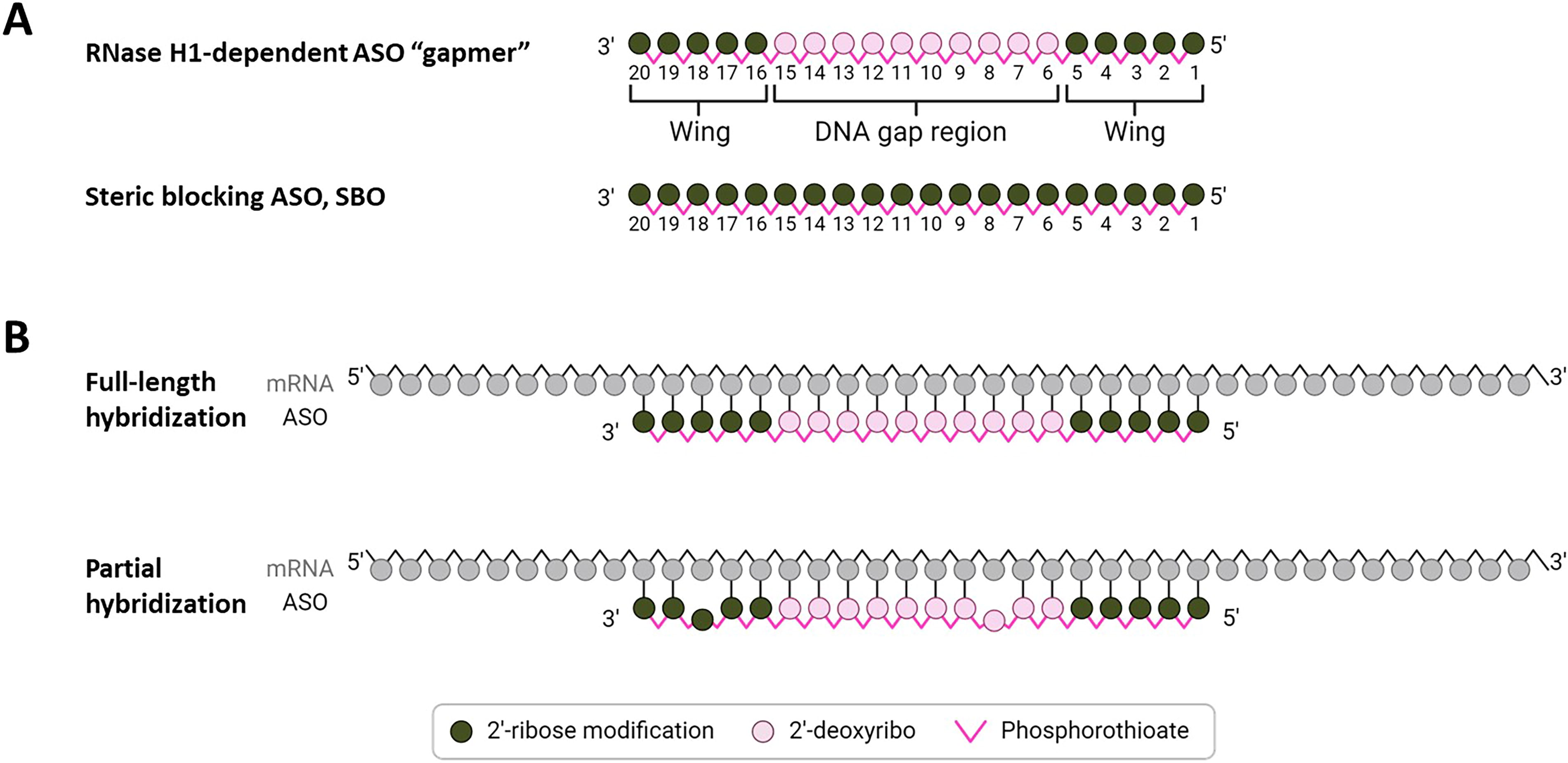
Schematic illustration of antisense oligonucleotides (ASOs) and examples of perfect and mismatched hybridization.
The potential for OffT RNase H1 recruitment depends on the nature of the hybrid formed between the ASO and its nonintended target. While RNase H1 works best with full sequence complementarity, it can tolerate some sequence divergence 29 (Fig. 2B). The number, type, and position of mismatches, as well as potential formation of bulge-loop structures, determine the extent of the reduction of substrate fitness for RNase H1 and thereby reduction in cleavage activity of the enzyme.30–34 Mismatches and bulges also lower the binding affinity between ASO and RNA and thereby reduce the amount of hybrid available as a substrate for RNase H1.
Numerous transcriptomic studies have explored the RNase H1-mediated OffT effects of RNase H1-dependent ASOs in vitro31,34–39 or in vivo.19,22,40,41 These studies as well as the authors’ experiences can guide OffT identification and management for this class of ONTs, which will be discussed next.
Class-specific considerations
The cut in the target or OffT RNA by RNase H1 leaving unprotected ends will lead to degradation of the cleavage products regardless of the position on the transcript where cleavage occurs. This allows the full length of the target sequence to be probed for accessible regions, providing great flexibility in identifying target sequences with potent on-target and minimal OffT activity.
Bioinformatic methods for computationally identifying on-target regions with minimal risk of OffT activity have been suggested.42,43 These methods typically catalogue all potential OffT regions up to a given number of mismatches and bulges. In general, the shorter the ASOs are, the higher the frequency of perfectly matching or nearly perfectly matching OffT regions. 44 On the contrary, longer ASOs may allow more mismatches while still establishing as many matching base pairs as shorter ASOs. In both cases this results in a loss of specificity. Furthermore, additional considerations such as pharmacokinetic properties and hybridization-independent toxicity not discussed herein, or the necessary compromise between affinity and potency, have resulted in practice to favor RNase H1-dependent ASO sequences between 16 and 20 nucleotides in length for clinical development.1,39,45,46 Irrespective of length, ASOs that only hybridize with perfect complementarity to the intended target will typically be preferred when identifying suitable on-target regions.
Based on the authors’ experience and published work, OffT effects are more often seen when mismatches are in the wing regions of ASOs than when mismatches are in the central DNA gap region14,31,47,48 (Fig. 2B). Consequently, when evaluating suitable target regions, one might consider mismatches in flanking regions of potential OffT RNA sequences as more important to avoid than mismatches in gap regions.
The binding affinity of RNase H1-dependent ASOs depends on their base sequences as well as chemical modifications. 49 In particular, the type and number of modified sugars in the flanks can change the binding affinity substantially. For high-affinity ribose modifications, such as locked nucleic acids (LNAs) or constrained ethyls (cEts), the binding affinity of the ASO may become so high that knockdown activity can be seen for more extensively mismatched regions than for, for example, 2′-O-methoxyethyl (MOE) ASOs with lower binding affinity. Historically, RNase H1-dependent ASOs with higher affinity modifications have tended to be shorter in order for optimal on-target potency to be achieved,1,50,51 increasing the likelihood of aligning to OffTs with (near-) perfect complementarity. Consequently, one strategy for minimizing OffT effects is to only select ASOs where OffT regions have multiple mismatches, and for those ASOs, adjust binding affinity to be just high enough for potent on-target effect, but not so high that the partially mismatched regions in other RNA sequences are bound and cleaved to an appreciable extent. This adjustment can be done by varying the number and placement of high-affinity (and lower-affinity) modifications in the flanks for a series of ASOs with the same base sequence and measuring on- and OffT effects, as discussed below. 40
Results from the development of allele-selective RNase H1-dependent ASOs have also increased our understanding of how to minimize OffT effects. When RNase H1-dependent ASOs target a region harboring a single nucleotide polymorphism (SNP) site, they are designed to lose most of their activity from a single mismatched nucleotide. Studies of such SNP-selective RNase H1-dependent ASOs show that the position of the SNP, as well as the types of chemical modifications in the flanks matter, as discussed above, but also, that shorter DNA gaps can result in more selective gapmers.48,52–55 Presumably, the longer the DNA gap, the more potential secondary sites are available for cleavage.
Beyond its application to allele-specific RNase H1-dependent ASOs, chemical modifications can also be used to reduce OffT effects. Introduction of 2′-modifications or non-phosphorothioate (PS) nuclease-resistant linkages in the oligo-deoxynucleotide gap region of promiscuous PS backbone RNase H1-dependent ASOs can narrow the range of RNase H1 tolerant mismatch OffT sites while maintaining on-target potency. This will reduce the number of meaningful OffTs and increase OffT safety margins.32,52,56
In silico evaluation of ASOs for potential OffT effects attempts to balance these various observations with specific criteria being dependent on the stage of ASO development. Early on, when identifying suitable on-target regions, often only the most promiscuous regions with many perfect- or near-perfect OffT regions across the transcriptome are excluded. Later, after ASO hits have been identified and if RNAseq profiles have been generated, in silico evaluation of potential binding regions in genes with reduced expression after ASO treatment may help separate direct, hybridization-based OffTs from genes reduced by other effects. When the most likely regions of hybridization harbors three or fewer mismatches it is often judged as sufficient to conclude that RNase H1-binding and cleavage is the cause of the reduction. 40 Last, for genes where in silico analysis of potential OffT regions stand alone, and expression has not been evaluated experimentally, regions with four or more mismatches are typically not considered. For potential OffTs harboring regions with three or fewer mismatches, typically, the fewer the mismatches, the higher the likelihood that the region might support RNase H1-binding and cleavage. However, even for perfectly matching regions, only a fraction will support RNase H1-binding and cleavage.47,57 This may be attributed to only some target regions being unstructured and accessible to ASO binding.
Steric blocking oligonucleotides
OffT mechanisms
SBOs are single-stranded ASOs lacking a central DNA domain (Fig. 2A) that hybridize to RNA via Watson–Crick–Franklin base pairing at sites that regulate RNA processing or function. Unlike cleavage mechanisms, which result in RNA degradation, SBOs can regulate RNA via many different mechanisms including modulating the processing of intronic regions in the RNA, expression, and/or function. One of the most successful classes of SBOs is splice-switching oligonucleotides that target splicing of RNA transcripts. 58 Other alternative splicing mechanisms that SBOs can modulate via cis-acting effects include intron retention, 59 intron exclusion, 60 and alternative splice site selection, which includes exon extension or truncation. 61
Anti-microRNA (anti-miRNA) SBOs function via the blocking of miRNAs, small noncoding RNAs that regulate the degradation of many cellular mRNAs. Anti-miRNAs inhibit miRNAs binding to their mRNA targets, which can result in upregulation of the target mRNAs. Other types of SBOs may increase protein expression by blocking upstream open reading frames 62 or other 5′ UTR (untranslated region) inhibitory elements, such as RNA secondary structures and protein binding sites. 63 Protein expression can also be modulated by SBO hybridization to a polyadenylation cleavage site, thereby altering the 3′ UTR length and composition of regulatory elements such as miRNA binding sites. 64 Other regulatory elements residing on the mRNA transcripts amenable to SBO hybridization are miRNA, noncoding RNA (ncRNA), and RNA-binding protein target sites. 65
Class-specific considerations
SBOs lacking a central DNA domain do not support RNase H1 or RNA-induced silencing complex (RISC) dependent cleavage mechanisms. Even though a net effect on the levels of putative OffT RNA transcripts cannot be excluded and should be assessed, special attention should be paid to OffT matches that may target the same elements as the intended target, for example, matches within exon and near exon–intron junctions for splicing SBOs. Two recent studies have evaluated OffT effects of SBOs experimentally, either by qRT-PCR (quantitative reverse transcriptase polymerase chain reaction) 66 or RNA seq. 67 Both studies observed OffT splicing events close to some of the regions harboring near-complementarity to the SBOs.
The reduced sequence flexibility of SBOs leads to greater challenges in finding sequences that have minimal number of hits when screened for potential OffT hybridization. On the contrary, the greater flexibility in chemical modifications and oligo length 68 can be used to find optimal balance between on-target and OffT activity. From the OffT evaluation point of view, the potential SBO hybridization to other transcripts depends on multiple potential factors including the variable length and chemical modifications used. 69 While the specifics of the evaluated SBO should be considered, the experience of the authors is that putative OffTs with four or more mismatches are generally not considered for in silico prediction for SBOs of typical lengths around 20 nucleotides (nt), similarly to other ONT classes.
siRNA
OffT mechanisms
siRNAs are double-stranded oligonucleotides (Fig. 3A) that post-transcriptionally regulate gene expression via an RNA interference (RNAi) mechanism. 70 Following loading into the argonaute (Ago)-containing RISC, the sense (passenger) strand is removed and degraded, and the antisense (guide) strand directs the RISC complex to complementary transcripts.71,72 This loading and RISC activity is traditionally thought to occur in the cytoplasm, although nuclear localization of Ago2/RISC should also be considered.73,74 The seed region of siRNAs is located near the 5′ end of the antisense strand and generally encompasses nucleotide positions 2–6 or 2–8 75 (see Fig. 3B). The intended pharmacological effect of siRNAs is the result of seed region-mediated site recognition, hybridization of the full-length antisense strand, and Ago2-mediated cleavage of the on-target RNA transcript.76,77
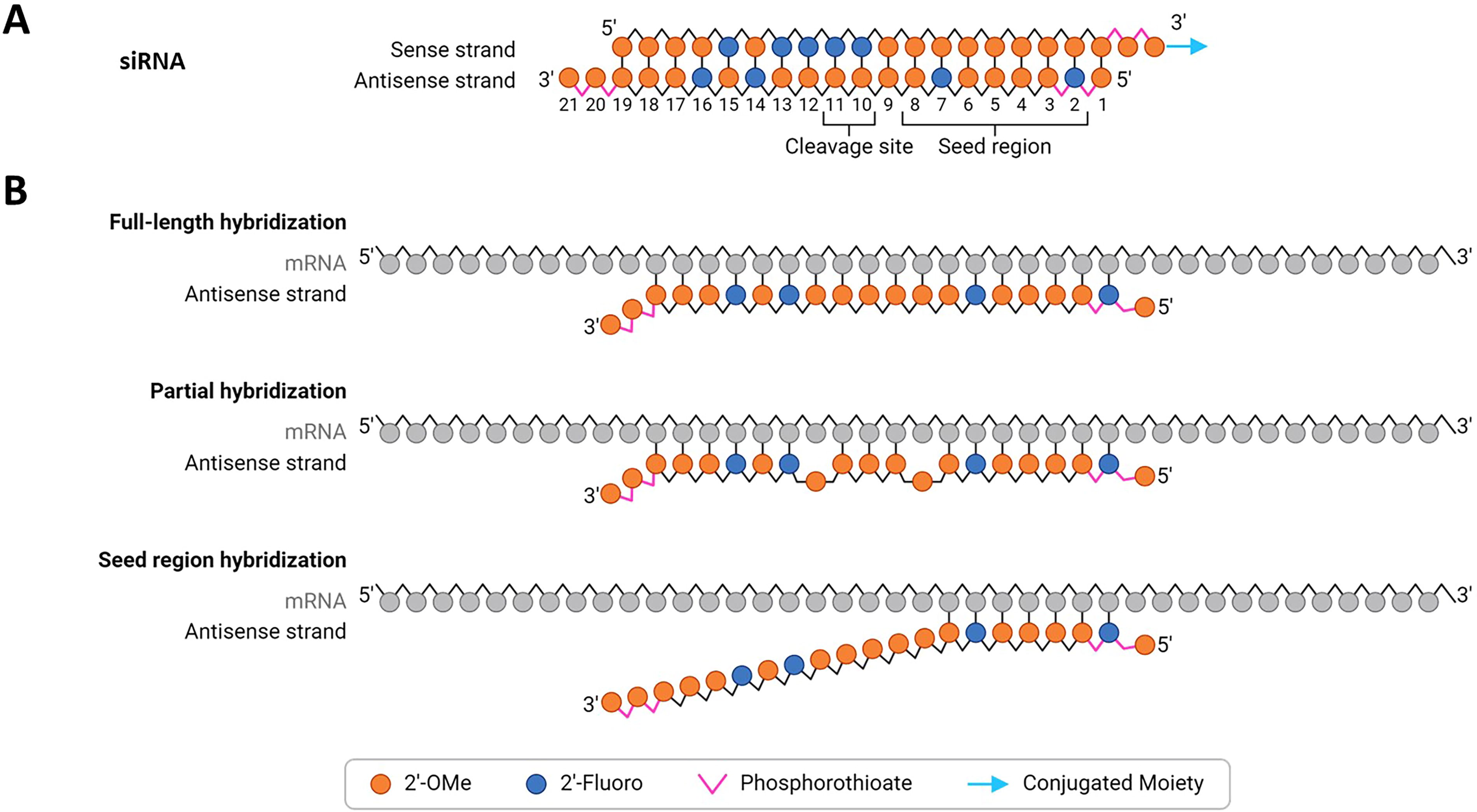
Schematic illustration of siRNA and examples of perfect and mismatched hybridization.
Mechanistically, RISC-dependent OffT activities of siRNAs can be divided into siRNA-like effects (whereby the OffT transcript is cleaved and degraded) and miRNA-like (where the exogenous siRNA behaves like a microRNA) (see section “Seed-mediated miRNA-Like OffT effects”). Seed-based OffT effects mediated by antisense strand-loaded RISC, which are complementary to the seed region but only partially complementary elsewhere in the sequence, representing the main mechanism of OffT activity. In some studies, this was associated with toxicity observed in vitro and in vivo.21,24,78 Importantly, reducing such OffT activity by chemically modifying the seed region of the antisense strand led to improved tolerability in rodents and in the clinic21,23,24; see section “Class-specific considerations” below. In addition to these RISC-dependent effects, other potential hybridization-dependent OffT mechanisms are possible and are discussed below.
siRNA-like effects
Transcripts with mismatches to the siRNA antisense strand (typically 19–23 nucleotides in length) may still be incorporated into RISC and cleaved by Ago2 79 (Fig. 3B). Analogous to RNase H1 recruiting ASOs, the position and nature of mismatches between the siRNA and OffT transcript are important and it is the authors’ experience that more than three mismatches greatly reduces activity. In particular, mismatches within the seed region (nt 2–8) and slicer site (nt 10–11) of the antisense strand are poorly tolerated and often lead to lack of activity.50,80 Additionally, the types of mismatches should also be considered. Purine:purine mismatches were shown to be deleterious to activity, 81 while G:U wobble pairing is tolerated and needs to be considered for OffT assessment.79,82 Notably, most studies describing the effects of mismatches were carried out with a small number of siRNAs, with limited chemistry modifications and patterns, and under diverse conditions (e.g., reporter assays, phenotypic screens). Thus, these specific position findings may not apply to current therapeutic siRNA designs, sequences, and modifications.
Seed-mediated miRNA-like OffT effects
siRNA binding via the seed region of the antisense or sense strand75,83,84 to unintended sites typically located in 3′ untranslated regions (3′UTRs) of nontarget mRNAs can lead to miRNA-like down-regulation of protein expression (Fig. 3B). These effects can be subtle (sometimes leading to <2-fold decrease in the mRNA and/or protein level, making detection challenging) and result from mRNA destabilization and/or translational repression of the OffT transcript.75,85,86 In contrast to catalytic Ago2-mediated mRNA cleavage, seed-based miRNA-like repression can be mediated by RISC ribonucleoprotein complexes containing any of the four Ago isoforms (Ago1, 2, 3, and 4). 87 Due to the stoichiometric nature of this process, such seed-mediated OffT effects are generally observed only at supratherapeutic siRNA doses in vivo 21 and at doses higher than the effective concentration in vitro for the intended on-target effect, 88 where sustained high levels of RISC-loaded siRNA are achieved. Owing to the short region of complementarity, and a lack of well-understood rules for this interaction, the accurate computational prediction of miRNA-like targets of therapeutic siRNAs remains a challenge.89,90
Class-specific considerations
For siRNAs, many OffT effects may be driven by the seed match, as discussed in section “Seed-mediated miRNA-like OffT effects,” while the on-target effect is driven by the full sequence match.20,91 To increase the specificity of binding and minimize the risks of OffT effects, it is recommended to employ strategies that selectively destabilize seed-based OffT binding while preserving on-target activity. For example, siRNA antisense strands with AU-rich seed regions can be prioritized. 92 Alternatively duplex-destabilizing chemical modifications, such as unlocked nucleic acid (UNA),93,94 glycol nucleic acid (GNA),21,23,24,95 threosefuranosyl nucleic acid 96 or 2′-deoxy-2′-α-F-2′-β-C-methyl pyrimidine nucleotides 97 can be introduced to the seed region to selectively disfavor seed-based hybridization. In addition, sense-strand RISC loading can be minimized by sequence selection (GC-rich 5′ end of the sense strand), use of chemical modification strategies (e.g., 2′-O methylation of specific sites), and/or 5′ modifications (e.g., morpholinos) that prevent phosphorylation of the sense strand. 98
OffT Assessment Workflow
In order for a hybridization-dependent OffT interaction to produce adverse effects, multiple criteria must be met. There must be sufficient ONT exposure to trigger an activity in the cells expressing the OffT RNA, the OffT RNA must be expressed at a time when the ONT is present, and the change in RNA function produced by the hybridization-dependent event must be anticipated to result in an adverse outcome.
Although technologies have improved the quality and speed of the assessment process, the workflow principles of characterizing potential OffT effects of a clinical candidate proposed by Lindow et al. 14 still hold. The proposed workflow is outlined in Figure 4 and includes the following steps:
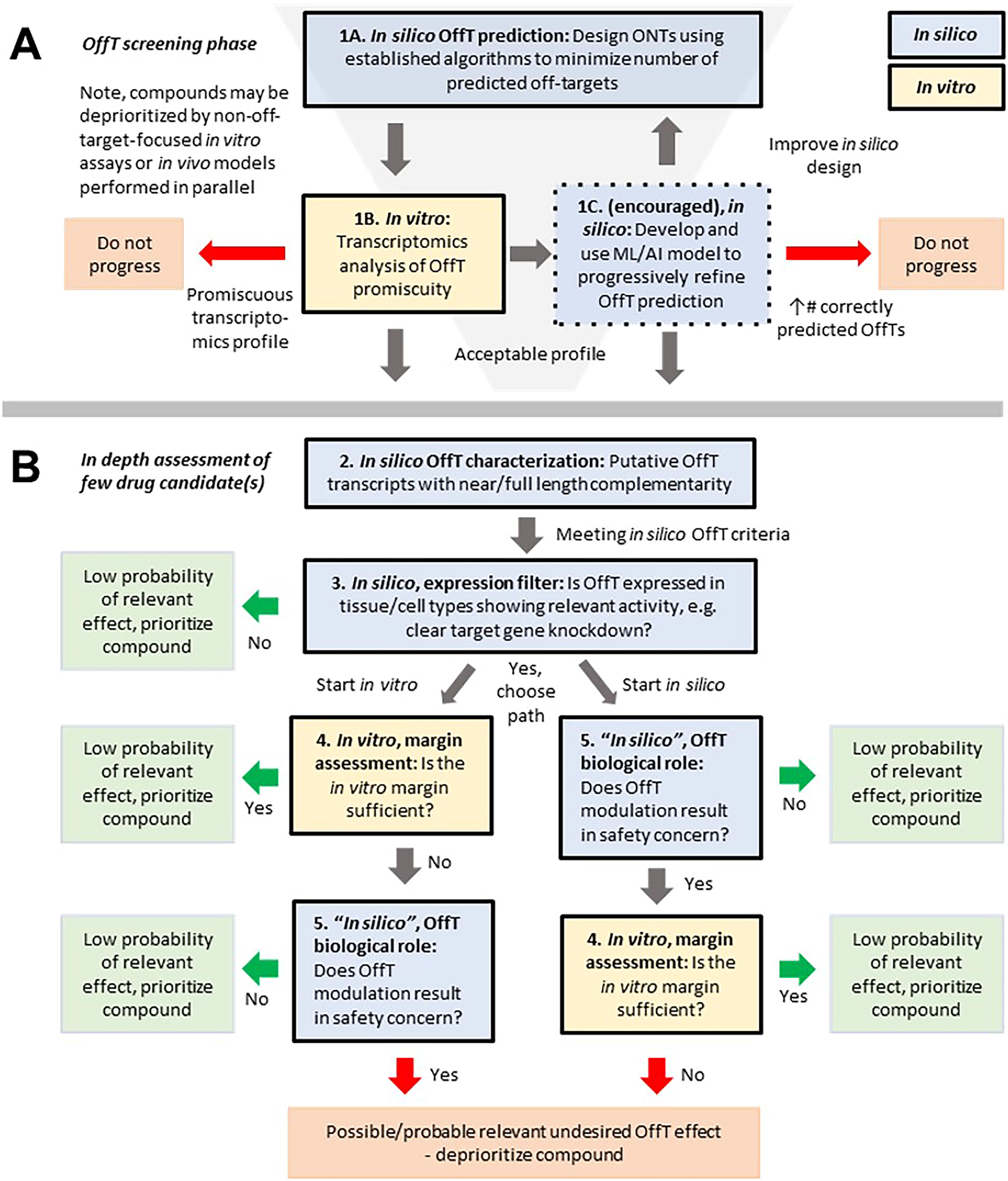
Proposed flowchart for systematic identification and assessment of potential hybridization-dependent OffTs for therapeutic oligonucleotides.
OffT identification, in silico and experimental—Boxes 1–2 in Figure 4.
Focus on cell types showing relevant ONT activity—Box 3.
Experimental verification and margin assessment between on-target and OffT effects—Box 4.
Risk assessment of experimentally confirmed OffTs not possible to avoid—Box 5.
Risk management in defining a clinical path.
Step 1—Identification of potential OffTs
OffT identification via in silico prediction approaches
The proposed OffT assessment workflow starts with an in silico step to focus the assessment on potential OffT transcripts with sufficient ONT sequence homology and likelihood of being a hybridization-dependent OffT. Although straightforward in theory, the majority of in silico predicted OffTs are not downregulated when experimentally assessed, illustrating a high frequency of false positive predictions. Considering that screening for active sites is also required for intended ONT candidates with perfect sequence complementarity,47,57,99,100 this high rate of false positives for sequences with imperfect complementarity is not surprising.
In silico prediction takes the sequence of the oligonucleotide of interest and aligns it against full transcriptomes to identify regions of sequence complementarity. There are many possible algorithms to choose from when performing similarity searches, see the examples listed in Table 1. ONT class-specific considerations are outlined below. Sponsors wanting to minimize risk for species-specific OffT effects in toxicology studies can also consider homology searches in relevant animal transcriptomes.
Examples of Tools for In Silico Predictions of Hybridization-Dependent OffTs
FISH, Fluorescence in situ hybridization.
The result of the homology prediction is then filtered on expression in tissues and cell types where the ONT class being studied shows relevant activity (see section “Step 2—Focus on cell types with relevant productive uptake for verifying OffTs”), using RNA expression databases, for example, listed below in Table 2.
Resource Examples for Assessing OffT Biological Role
Examples of publicly available resources that are useful for identifying and understanding potential on- and off-target related hazards.
CRISPR, clustered regularly interspaced short palindromic repeats.
Evidence provided by the Kamola et al. 31 emphasizes the importance of searching for putative OffTs not only in mature mRNA but also in nonprocessed pre-mRNA and noncoding RNAs.
Although comparably fast and computationally practical, simple alignment tools are poor at predicting ONT hybridization. This poor sensitivity and/or specificity applies to short-read alignment tools (e.g., Bowtie2, STAR, BWA, GGGenome; see Table 1) and BLAST-like tools as well, and requires parameter optimization to improve. Models based on thermodynamic simulation, dynamic programming, or nearest-neighbors can be more representative of ONT-RNA occupancy than edit distance. A practical strategy is to first run a sequence similarity search (e.g., BLAST-like search) in an overly sensitive configuration (suffering low specificity) and consider further evaluation of the remaining candidates with more complex search models like energy-based models for ASOs or seed-region searches for siRNAs.
For all the described in silico models above, potential false positives and false negatives must be considered carefully. On the one hand, relaxed filtering criteria may result in a large number of false positives (i.e., predicted OffTs that are not validated experimentally), and candidate ONTs may be removed prematurely during in silico design for the wrong reason. On the other hand, criteria that are too narrow may result in false negatives (i.e., true OffTs not predicted by in silico methods). To account for the latter case, experimental screening using an open-ended technology platform like RNAseq is highly recommended, see section “OffT identification via transcriptomics approaches.”
siRNA considerations
OffT siRNA searches should focus on potential RNAi transcripts that require near full-length complementarity to the antisense strand. Sequence complementarity searches for siRNA-like OffT effects are frequently conducted for nucleotides 2–19 of at least the antisense strand, as position 1 and the 3′ overhangs of the antisense strand interact with Ago2, 112 and do not contribute to base pairing with mRNAs (see Fig. 3B). The number of edits (e.g., mismatches, insertions, deletions, G:U wobbles) can be changed in the search tools to suit the requirements of the user; increasing the number of edits will generally decrease the false negative rate but also increase the false positive rate. siRNA sequences and chemical modification patterns are typically designed to strongly favor antisense strand loading.113–115 Although it may be a rare event, the sense strand may be loaded into RISC or result in steric blocking effects so OffT complementarity searches should include the sense strand. For the reasons mentioned above in section “siRNA”, the in silico identification of all potential seed-based (miRNA-like) OffTs is not feasible, and investigators may consider experimentally identifying siRNA for seed-based regulation and rank the sequences based on lower miRNA-like activity.
Application of machine learning models to the identification of putative OffT transcripts
The systematic implementation of RNAseq techniques in the early stage (in vitro) identification of active and specific ONTs will enable the progressive accumulation of a massive amount of wide-scale transcriptomic data across a broad range of ONTs. Properly curated, this wealth of data will provide the opportunity to validate and refine in silico algorithms for identifying putative OffT effects. Moreover, the availability of such data will also support the development of new, proprietary machine learning models to predict on-target activity as well as OffT specificity (Fig. 4, Box 1C).
Unfortunately, several challenges hinder the systematic deployment of these models. For instance, with most RNAseq and other relevant data being owned by individual sponsors, obtaining large enough training data sets for comparable ONTs takes time and is expensive. Nevertheless, organizations should remain proactive in developing and testing such machine learning models, as they have the potential to significantly improve the identification of active and specific ONTs in silico. Ultimately, these models could prove valuable during the in silico design phase, enhancing the quality of initial screening libraries.
OffT identification via transcriptomics approaches
In silico predictions will identify a high number of putative OffTs that will not show differential expression when tested, resulting in a high rate of false positives. It is therefore crucial to experimentally verify which in silico predicted OffTs are modulated, ideally in a cell line that also expresses the on-target as a positive control for active ONT uptake.
Early in the discovery phase, when a large number of sequences are being considered, the use of technologies assessing global transcriptome changes (e.g., RNAseq) can be beneficial. This type of global approach can help assess the overall promiscuity of an ONT sequence, as well as be an efficient way to confirm OffTs identified by in silico approaches (Fig. 4, Box 1B).
Different approaches, such as microarray expression profiling, RNAseq, and real-time quantitative PCR (RT-qPCR), have been employed in previous research to explore hybridization-dependent OffT effects. 40 Among these methods, RNAseq is arguably more effective, especially during early ONT prioritization, as it offers unbiased and comprehensive robustness, enabling more accurate quantification of transcripts with low expression across the transcriptome. Regardless of methodology, expression changes will only be detected from genes expressed in the cell/tissue system being assessed. For practical and cost reasons, it is not possible to use enough models to cover the entire human transcriptome throughout the myriad of physiological conditions, and we argue that the use of one or a few in vitro human models using an unbiased approach can be useful to rank ONTs early in a program.
This type of global assessment will capture changes in gene expression due to all aspects of treating the cell with the ONT and does not necessarily confirm a hybridization-dependent effect. For initial ONT ranking for promiscuity, the count of the statistically significant, downregulated genes is commonly used to help prioritize sequences to move forward, capturing both hybridization and hybridization-independent ONT effects. Examples of hybridization-independent effects include downstream effects of certain ASO sequences inadvertently inducing proinflammatory responses.116–120
Overlaying the in silico predicted putative OffTs can help distinguish between the hybridization-dependent and -independent types of effects. In addition, an overlapping set of differentially expressed genes across multiple ONTs with different sequences for the same target can assist in identifying expression changes resulting from modulation of the primary target transcript. By subtracting such on-target transcript sets from an individual ONT’s in silico-predicted, downregulated set of genes, confidence can be increased that the remaining signal is due to sequence-specific hybridization OffT effects.121,122 The purpose of this early in vitro assessment is to prioritize sequences and ONT designs selective and active for the target gene with minimal OffT effects. It also helps identify true OffTs for progressing ONTs to further verify with a dose-response assay.
To accurately assess OffT effects and ensure the reliability of experimental results, it is essential to consider various factors, including the selection of appropriate cell line(s), time points, and concentrations for each ONT platform and experimental system, see section “Step 3—Experimental verification of potential OffTs.” By carefully optimizing these parameters, researchers can minimize confounding variables and obtain more reliable data on the OffT effects of ONTs.
Step 2—Focus on cell types with relevant productive uptake for verifying OffTs
The distribution and activity of a given ONT differ significantly between tissues and cell types. This is further influenced by the choice of administration route (e.g., systemic or local such as inhalation, intrathecal) and the ONT chemistry and design. Another factor that significantly impacts distribution and productive uptake (i.e., uptake leading to ONT activity, e.g., RNA degradation or splice modulation) is the choice of ONT delivery system (e.g., conjugates or lipid nanoparticles, LNP). Examples of targeting conjugates are N-acetyl galactosamine (GalNAc)-conjugated oligos that are preferentially taken up by hepatocytes via the asialoglycoprotein receptor ASGR.123–125 Targeted uptake into muscle and across the blood–brain barrier can be achieved via Transferrin receptor conjugation, 126 whereas lipid nanoparticles can be used to deliver siRNA to hepatocytes. 127
All biological systems show some degree of redundancy, and a certain level of change is required to exert an intended or unintended perturbation. Given that hybridization-dependent on- and OffT effects act via the same molecular mechanisms, tissues, and cell types with a low degree of intended on-target ONT activity will also show a low degree of OffT activity. Further analysis of potential OffTs identified in silico or via transcriptomics methods, should focus on putative OffTs expressed in tissues and cell types with high enough productive uptake allowing ONT activity to result in a biologically relevant change. Conceptually, putative OffTs that are only expressed in tissues and cell types with a low degree of ONT productive uptake and activity are of minimal concern and should not need to be further assessed.
This approach raises at least two key questions: (1) What is regarded as a relevant level of productive uptake and ONT activity? and (2) Which tissues and cell types show relevant ONT productive uptake and activity? The answer to the first question may depend on the ONT mechanism. For RNase H1-dependent and siRNA-triggering degradation of a target RNA, gene dosage analyses and experience from ONT pharmacological studies indicate that many biological processes are buffered to resist some degree of reduced expression before a relevant biological effect is seen. 128 However, there is no general gene expression modulation threshold that can be used for all OffT genes given the complexities of ONT activity, specific gene modulation, and their phenotypic expression as discussed in section “Assessment of OffT biological role”. Nevertheless, small-expression modulations that are unlikely to meaningfully impact gene function indicate no need for further OffT assessment.
To answer the second question on which tissues and cell types show relevant ONT productive uptake, a platform approach could be used, i.e., run an ONT activity/target engagement study in a relevant animal model using a tool ONT. To be relevant, this tool ONT should be of the same chemistry, and design and use the same delivery system and administration route as the ONT under study. This approach would need to be repeated for novel chemistries or delivery systems. The tool ONT must be active and sufficiently tolerated in the species of choice and should ideally target an RNA expressed in most cell types. Malat-1 (metastasis-associated lung adenocarcinoma transcript 1), a long noncoding RNA expressed at high levels in almost all cell types in most species, is a good example of a tool target that has been used to assess productive uptake and ONT activity for RNase H1-dependent ASOs in mice and NHPs. 129 Since Malat-1 is localized to the cell nucleus, it is not useful for the study of siRNA activities in the cytosol, but other targets showing broad expression profiles have been described that can be used with siRNA, for example, cyclophilin B (PPIB)130,131 or superoxide dismutase 1 (Sod1). 132 When designing the “productive uptake distribution” experiment, we propose to use the same administration route and delivery system as intended in the clinic. The dose level(s) and exposure time should at least lead to the pharmacological activity level (e.g., target RNA knockdown) anticipated in the intended target organ(s) in clinical studies. The selection of test species for these experiments should consider species differences in the activity of any targeting moiety.
If utilizing a conjugate delivery approach, the tissues to analyze could partly be guided by the expression pattern of the receptor to which the conjugate binds but the possibility for relevant productive uptake in other cell types and tissues should be assessed. If the delivery system is less specific, for example, a lipid conjugate or unspecific LNP, a broader set of tissues may be explored. To assess ONT activity, qRT-PCR on tissue homogenate is a straightforward method. However, given that productive uptake can differ significantly between cell types in the same tissue, more granular analysis may be needed. To understand active uptake in both tissue and cell types in vivo, methodologies such as in situ hybridization (ISH) or single cell/nucleus RNAseq can be used. Indeed, recent progress offering several flexible options for sample preparation supports this need. 133
As long as key properties influencing productive uptake of the ONT cargo are kept the same, results from a carefully designed study as outlined above should be able to inform productive uptake distribution for subsequent candidates using the same delivery system and administration route, thereby guiding which cell types and tissues are most relevant for OffT assessment. In other words, the focus of the OffT assessment should be on OffTs expressed in cell types and tissues showing enough productive uptake to result in a biologically relevant level of RNA modulation (Fig. 4, Box 3). This does not imply that all cell types showing productive uptake need to be covered in the experimental verification phase; experimental verification in one or a few relevant cell model expressing a given OffT should be sufficient.
Step 3—Experimental verification of potential OffTs
Due to the high rate of false positive OffT predictions using in silico models (see section “OffT identification via in silico prediction approaches”), experimental verification is required for risk assessment (Fig. 4, Box 4). We strongly recommend focus on human model systems. If an RNAseq analysis has been run (see section “OffT identification via transcriptomics approaches”) these results may be used for initial verification. If the concentration(s) selected in the transcriptomics experiment result in an on-target ONT activity level clearly above the highest activity anticipated in patients (e.g., 70% target knockdown), in silico predicted OffTs expressed in the model system that do not show relevant change in expression levels can be regarded as inactive and do not need further assessment.
For more granular and quantitative analyses, we recommend dose-response experiments to generate an ICX for each OffT (e.g., IC20, IC50, IC90). In the ideal case where the in vitro system expresses the on-target and putative OffT, an in vitro margin can be calculated to inform the likelihood of relevant activity of the OffT. Multiple human cell systems may be used to cover the OffTs identified via in silico prediction or transcriptomics analysis and expressed in relevant cells/tissues (see sections “Step 2—Focus on cell types with relevant productive uptake for verifying OffTs” and “Assessment of OffT biological role”). Depending on the number of putative OffTs identified in silico, this analysis can be performed using qRT-PCR, global transcriptomic technologies (e.g., RNAseq), or other molecular techniques.
Compared to RNAseq, qRT-PCR has limitations in assessing the activity of a large number of OffTs due to its lower throughput and the need for a priori knowledge of target sequences with primer:probe sets to be designed and validated for each potential OffT. PCR-based approaches might be better suited for focused investigation discussed in “Step 4—Management of remaining OffTs” section.
As mentioned above, the choice of cells, time point, delivery method, and concentration range are important considerations. For human risk assessment, in vitro models of human origin are key but sponsors wanting to de-risk species-specific OffT effects in toxicology studies could consider in silico and in vitro assessment also for the species to be used for nonclinical toxicology studies. When selecting the cell systems/tissues for in vitro experiments, we recommend focusing on experimental models representing cell types showing relevant activity (“productive uptake”) of the ONT type under study and the intended clinical route of administration (see section “Step 2—Focus on cell types with relevant productive uptake for verifying OffTs”). This should increase the likelihood of covering the in silico predicted OffTs of most relevance from an expression pattern perspective. The preferred in vitro ONT delivery method is with free/gymnotic uptake if possible, to minimize effects from non-physiological uptake. When other delivery methods are needed, for example, lipid transfection or electroporation, ensure appropriate controls are included in the study design. Optimal timing for robust knockdown should be established, ideally guided by the time course of primary target knockdown using the same ONT format. This is typically 24–48 hours in most cell systems.
Following initial untargeted in vitro experiments (e.g., RNAseq) to identify which in silico predicted OffTs are modulated by an ONT, a more targeted experiment is conducted to identify in vitro margins of modulation for these experimentally verified OffTs (see “Box 4, in vitro margin assessment” in Fig. 4). This is a human in vitro experiment using, for example, RT-qPCR or RNAseq analysis in preparation for candidate selection and generating margin assessment for regulatory filings. It is recommended to include multiple concentrations up to concentrations that exceed saturated on-target activity. A dose-response of the OffT can then be compared to the on-target to establish an in vitro OffT margin to efficacious on-target knockdown, for example, IC50 on-target versus IC50 of potential OffT. The safety margin needed is highly context-dependent so the authors cannot recommend a single default margin to be used.
Challenges with in vivo assessment
As described above, for human risk assessment, we advocate the use of human in vitro models to experimentally verify potential OffTs identified by in silico methods. The main reasons include analysis of transcriptome from the intended target species (i.e. human), being cost-effective and amenable to apply relatively early in the compound selection process. In vivo animal models can be useful for some aspects of the OffT assessment process like understanding the potential consequences of affecting an OffT not possible to avoid (see section “Step 5—Management of remaining off-targets”); however, we do not recommend animal models for the identification or verification of potential human ONT OffTs.
Sequence differences can have a substantial impact on ONT activity, with very few mismatches leading to significantly reduced potency between species. Even in cases of perfect sequence homology, the activity can vary from full activity to inactive regardless of ONT class as observed during the screening phase for on-target activity.47,57,99,100 Similar variability in activity for perfect matches will also be the case for predicted OffTs. Thus, the absence of activity on a potential OffT in animal species cannot be interpreted as no risk for effect in humans. Also, lack of toxicity beyond expected class effects in animal models cannot be taken as an indication for lack of OffT effects in humans. To focus efforts on the most promising clinical candidate(s), identification of experimentally verified OffTs of concern is ideally done as early as possible in the ONT discovery process. Since studies in animal models of highest likelihood for homology (e.g., NHP) are run relatively late in the sequence selection process, any data that could be relevant would likely be generated too late to make an impact on compound selection. That said, OffT effects experimentally confirmed in nonclinical toxicology studies can provide valuable insight to understand potential consequences and inform overall OffT risk assessment (see section “Step 5—Management of remaining OffTs”).
Step 4–Assessment of OffT biological role
Risk assessment of relevant OffT hits follows the same principles as assessment of potential on-target safety concerns, a.k.a. exaggerated pharmacology. 134 This starts with identifying potential hazards (i.e., adverse effects that could occur in the intended or nonintended organ). Valuable information for this assessment includes expression levels and tissue distribution of the target RNA, temporal expression, as well as phenotypes associated with known human mutations and polymorphisms and genetically modified animal models. Data from other compounds modulating the same target can also be informative, but modality-specific aspects need to be considered. Examples of useful, publicly available databases for expression and phenotypes are listed in Table 2.
This hazard identification should be followed by a quantitative theoretical risk assessment defining the likelihood of these potential adverse effects to occur, accounting for cell type-specific drug activity, effect duration, and species differences in biology. This may require dedicated experiments with a specific study design with readouts reflecting the biology being assessed. Compared to small molecule and antibody drugs, other considerations are needed for potential on-target safety risk for ONTs 135 that are also relevant for OffT risk assessment.
The amount of information available relevant for hazard identification and risk assessment of a given OffT can differ significantly. For some potential OffTs, information exists describing phenotypes of human mutations and genetically modified mice that may be useful in identifying potential OffT hazards. When using such information, it is important that the “functional direction” of the genetic variant is understood, that is, does it result in reduced expression or disrupted activity or the other way around. One should also remember that ONTs rarely result in activity levels mimicking the situation of homozygous gene deletion often reported for mouse knockout models or phenotypes resulting from human autosomal homozygous or X-linked loss of function mutations. Special attention should be paid to dosage-sensitive (e.g., haploinsufficient or triplosensitive) genes that are sensitive to a single copy gain or loss, 136 which is analogous to a modest change in gene expression at the RNA level. Phenotype data from heterozygous subjects can therefore be particularly valuable. This can provide information about which level of change for a given OffT transcript is needed for a biologically relevant effect to occur.
The nature and severity of the disease that may be linked to variants of the OffT transcript is an important part of the risk evaluation exercise. Potential OffT modulation of cancer drivers such as oncogenes and tumor suppressors is of particular concern since a single transformed cell can initiate a lethal disease even if all other cells in the tissue are healthy. The lack of reported human phenotypes associated with a specific gene should not be automatically considered as an indication of low risk since inactivation of essential genes is often lethal in early embryogenesis. Investigation of available mouse data such as the International Mouse Phenotyping Consortium (IMPC) or MGI is essential to better understand a gene’s role in early embryogenesis.
Evaluation of the potential OffT sequence variation is necessary to assess the risk in different human populations. The presence of polymorphic variants such as SNPs within a confirmed OffT sequence may add or remove mismatches between the ONT and OffT transcripts and impact potential modulation. If there is any SNP with relevant frequency detected within the OffT sequence (e.g., >1%), the likelihood for potential differential OffT hybridization due to different edit distance in human subpopulations should be discussed.
Step 5—Management of remaining OffTs
For some confirmed (based on the above recommendations) OffTs, there might be insufficient or conflicting information and data available to allow a clear stop/go decision, thus requiring additional experimental evaluation to assess hazard and quantify risk margins. Understanding and quantifying potential consequences of an OffT effect may require a model system expressing the OffT with relevant “downstream” function retained. For functional consequences not possible to study in simple cell lines, primary cells, and different 3D in vitro models, including microphysiological systems may be useful in mimicking OffT in vivo function, ONT uptake, and complex readouts. 137
We do not recommend animal toxicity studies for OffT identification and verification. However, under certain conditions, regular animal toxicity studies can provide additional weight of evidence to understand potential consequences of an OffT effect. For this to be relevant, the anticipated OffT effect (e.g., target RNA knockdown) must be experimentally verified in the study referred to. For example, if in vitro experiments indicate that an RNase H1-dependent ASO or siRNA may cause 30% OffT knockdown in a clinical setting, the magnitude of OffT knockdown in the in vivo toxicology study should reach the same level or ideally higher level of activity. Known species differences in OffT function should also be considered during the integrated risk assessment.
For organ systems or readouts lacking relevant human in vitro models, rodent active surrogate ONTs against the OffT can be generated and used for quantitative in vivo risk assessment. 44 When designing OffT species-specific surrogate ONTs several aspects need to be considered to mimic the human situation as much as possible. The animal active surrogate ONT should have as similar chemistry, design, and tissue distribution as the clinical candidate as possible. Such species-specific OffT surrogate ONTs should be selected with similar stringency as the human candidate. This would ideally include having comparable OfffT potency in the animal model as would be expected from the clinical candidate and with minimal animal-specific OffTs. Should a compound with remaining OffT liability be progressed to clinical development, such OffT surrogates could be useful (and perhaps requested) to be included in safety studies run in that species.
Discussion and Conclusions
Hybridization-dependent OffTs remain an important area for safety risk assessment of ONT clinical candidates. Besides the correlation between broad OffT effects and liver toxicity described for some siRNAs,21,23,24 the authors are not aware of any other case of toxicity observed in clinical studies that could directly be attributed to a hybridization-dependent OffT effect. This suggests that the approaches to minimize relevant OffTs have been successful.
Since the last OSWG recommendation, 14 experience has increased and in silico and experimental methodologies have further improved. This article provides updated and pragmatic recommendations on how to identify, characterize and risk assess human OffT effects of RNase H1-dependent and steric blocking ASOs and siRNAs. During the revision of this article, FDA published a draft version of Nonclinical Safety Assessment of Oligonucleotide-Based Therapeutics 17 and ICH has formally initiated work to develop a nonclinical ONT-specific guidance (S13) 18 that will likely encompass OffT assessment. It is recognized that these formal guidances will take priority over recommendations discussed in this article.
While applying ONT class-specific considerations described in more detail above, the authors recommend conducting all steps to best identify and assess potential relevant OffTs. In our flowchart of recommended activities (Fig. 4), two paths are outlined for steps 3 and 4 (boxes 4 and 5). Both activities (in vitro experiments vs. understanding OffT biology), are different in nature and quite resource demanding. Since established expertise and capabilities may differ, sponsors can choose in which order they perform these steps. If a sponsor chooses to start the process with in vitro verification (Box 4) or OffT biology understanding (Box 5), the resulting risk assessment for the prioritized compounds will still be the same. However, for many OffTs the information available may be insufficient to allow a conclusive risk assessment based on biological role (step 4, box 5), in practice forcing step 3 (box 4) to be run first.
In silico OffT identification
In silico prediction includes alignment of the ONT sequence of the oligonucleotide(s) of interest to transcriptome databases. There are many search models, possible algorithms, and important considerations when performing similarity searches (Table 2). In general, we suggest the in silico search be performed on entire primary and mature transcripts (including both introns and exons as well as exon:exon junctions, nuclear, noncoding, and mitochondrial sequence 31 ). Other considerations are specific for the ONT classes.
All in silico models described to date tend to return a high number of false positive predictions, thereby requiring experimental assessment to support OffT predictions. Depending on the prediction model, false negative predictions can still occur but can be assessed using open-ended transcriptomic methodology such as RNAseq. Synthesizing compounds with a large number of unverified OffT predictions can still be informative during the initial design and screening phases (Box 1A) when identifying active hot spots for on-target activity on the target transcript. Stricter progression criteria are normally warranted for the selection and prioritization of shortlisted compounds that will undergo more in-depth analysis. This is depicted by Box 2 in the flowchart. The same alignment model as in Box 1 can be used but with stricter acceptance limits.
In addition to sequence homology, the likelihood of a relevant OffT-driven effect depends on the OffT expression distribution pattern. Due to a low level of productive uptake in most tissues and cell types, the tissues and cell types amenable to relevant OffT effects are also restricted. For the next steps in the assessment cascade, we recommend focusing on those OffTs that are expressed in cell types in which relevant activity can be expected for the ONT class being studied, taking into account delivery and targeting chemistry (e.g., GalNAc conjugates).
In vitro verification
As mentioned in “OffT identification via in silico prediction approaches” section, in silico predictions using permissive settings are likely to return a high number of false positives, that is, sequences meeting the in silico criteria but not showing differential expression when experimentally tested. It is therefore crucial to experimentally verify which in silico hits are true OffTs. This should be done in vitro, using human cell types that ideally express both the primary target and the OffT(s) of interest.
The choice of cells, time point, delivery method, and concentration range are important considerations. Ideally, the cell model(s) used has the same origin as the target tissue of concern, but the primary selection criteria should be expression of the OffT. Expression of the on-target transcript in the same model system allows direct assessment of in vitro margins. To experimentally cover the in silico predicted OffTs, different in vitro models may be required. Besides the choice of cell model, the selection of a relevant concentration range and study duration need to be empirically determined for the specific ONT platform and experimental system. For ONTs aimed to induce degradation or increase expression of RNA, different methods exist to assess OffT effects. During the screening phase, when there are many ONT candidates to evaluate, the use of RNAseq or other transcriptomics technologies is recommended (Fig. 4, Box 1B). In theory, this allows the assessment of all potential OffTs expressed in that model system for a given ONT. Cost may prevent full concentration-response curves, but the comprehensive coverage allows early de-selection of promiscuous ONTs and selection of in silico predicted OffTs to further characterize the ONT compounds selected for progression. Properly curated, this wealth of data presents the opportunity to validate and refine in silico algorithms for identifying putative OffT effects. Moreover, the availability of such data allows the development of new machine learning models (Fig. 4, Box 1C) that can be used to improve the design of compound libraries to be synthesized for initial testing.
In contrast to the “open-ended” transcriptomics methodologies, RT-qPCR is a directed analysis that can be used as an alternative or complement to RNAseq methodology. A main advantage of this method may be faster delivery of results and the ability to run in a standard in vitro lab. This makes RT-qPCR an attractive technology to use to verify and establish granular in vitro margins when fewer candidates remain at the shortlist stage (Fig. 4, Box 4).
Integrated risk assessment
With the common “funnel” approach to identify the ONT compound with the best overall properties for OffT risk assessment, stop/go decisions should be made at several steps throughout the ONT development process. At the early design stage, the risk assessment and decision process is relatively simple with sequences having few in silico predicted OffTs being favored over those that have many. Further down the funnel, additional parameters become important for understanding potential risk. These include tissue/cell type expression pattern (Fig. 4, Box 3), the in vitro margin to the on-target activity (Fig. 4, Box 4), and an understanding of the biological role of the potential OffT (Fig. 4, Box 5).
Together with the expression pattern information, the result from the in vitro margin assessment shows the likelihood/probability of a relevant effect of a given OffT at therapeutic dose/exposure levels. This information needs to be combined with an assessment of the potential biological consequence, should relevant effects on the OffT RNA occur. This follows the same principles as assessing potential risk for exaggerated pharmacology of the intended on-target 134 (i.e., compiling available information on the biological role of the OffT). Key parameters are phenotypes associated with known human mutations and polymorphisms and genetically modified animal models. Data from other compounds modulating the same target can also be informative, but modality-specific aspects like selectivity, duration of effect, and distribution pattern should be considered when interpreting safety data from other drug modalities.
As the last step of this risk assessment, the identified likelihood and potential biological consequence need to be discussed in light of the project context. That is, assessing the risk-benefit for the intended treatment situation, considering, for example, severity of the disease and age of the typical patient to treat and if treatment alternatives are available. Some OffTs may only play an important role during a certain stage of human life, for example, fetal development or adulthood so understanding the temporal and spatial expression of the OffT in relation to intended patient population is also critical.
Until the FDA draft guidance 17 is finalized and the future oligonucleotide-specific ICH S13 guidance 18 is published, the authors of this article anticipate the considerations and recommendations for OffT evaluation outlined here will be valuable for the development of ONTs and that it may serve as a complement to the new guidance after their finalization. Our aim has been to provide principles and recommendations that are specific enough to provide helpful guidance, but also flexible enough to be applicable to the wide range of ONT platforms available today and in the future.
Footnotes
Acknowledgments
The authors would like to express their gratitude to the following fellow scientists who kindly provided valuable comments on a mature version of the article. Maja Janas (Alnylam), Fredrik Karlsson, Nicola Guzzi, Sebastian Prill (AstraZeneca), Kari Neier (Boehringer Ingelheim), Chris Hart (consultant), Jibo Wang (EliLilly), Joel Parry and Piotr Kamola (GSK), Morten Lindow (F. Hoffmann—La Roche), Tim MacLachlan, Megumi Onishi-Seebacher, Stefan Haemmig (Novartis), Yann Tessier (OligoSafety), Shan Gao, Hong Yu (Ribocure), Marie Wikström Lindholm, Steffen Schubert, and Alexander Bird (Silence Therapeutics).
Author Disclosure Statement
The authors declare no conflicts of interest given the precompetitive nature of this work. Their respective employers at the time of publication are detailed in the affiliations.
Disclaimer
The views and opinions expressed in this article are those of the authors and do not necessarily represent the views, opinions, and/or policies of their employers or OSWG.
Funding Information
No funding was received for this work, other than the publication fees, which were covered by AstraZeneca R&D (employer of the subcommittee Chair).
