Abstract

The number of nucleic acid therapies in development, including oligonucleotide therapeutics (ONTs) and vaccines, increased rapidly since their inception in 1978 1 ; in particular, following the full approval of the first mRNA COVID-19 vaccine in 2021. 2 Today, ONTs comprise the third major class of drugs after small molecules and therapeutic antibodies, with many recent regulatory approvals and hundreds of ongoing clinical trials worldwide.3,4 ONTs, such as antisense oligonucleotides (ASOs) and small interfering RNA (siRNAs), are short fragments of nucleic acids complementary to a specific mRNA. Their function is to suppress, amplify, or alter the expression of a gene; by inhibiting, promoting or eliminating errors in RNA processing (e.g., RNA splicing or RNA degradation), ultimately influencing translation of the corresponding protein. The development, successes, and challenges of such ONTs have been reviewed recently.5–7 Given their genetically derived target, ONTs are applied across diverse disease areas, from oncology and neurology to infectious disease prevention.8–10 Importantly, however, a significant number of fundamental hurdles need to be overcome before an ONT drug can reach the market, including (1) poor stability and high sensitivity to endo- and exonucleases in tissues and fluids (e.g., plasma or cerebrospinal fluid), (2) inefficient uptake into target cells or tissues, (3) off-target effects due to interactions with unintended mRNA targets, and (4) unwanted immunogenicity 11 or immuno-stimulation. Furthermore, the development of ONTs as pharmaceutical drugs is hampered by limitations around their detection, including the quantitative methods available currently to demonstrate delivery, safety, and efficacy. 12 These limitations are echoed in current 2024 guidance documents issued by the US Food and Drug Administration (FDA) where it is recognized that the rapid evolution of ONTs, including new chemical modifications and delivery methods, can significantly impact immunogenicity risk and clinical assessment approaches. In its guidance, the FDA indicates that immunogenicity assays may be necessary to measure immune responses to different components of ONT drugs especially when the therapeutic includes carriers or conjugates.13,14 This guidance is concordant with earlier calls for better analytical tools to evaluate the drug metabolism and pharmacokinetics of ONTs and more broadly evaluate their absorption, distribution, metabolism, excretion, and toxicology properties. 15 Optimal methods for sample analyses should consider the advantages, disadvantages, and limitations of each assay, differences in the matrix (e.g., plasma, tissue, or urine), the sensitivity of detection required as well as the objectives for the analysis (e.g., quantitative PK measurement or qualitative metabolite identification). 16 While generally assessed by criteria established for small molecules, 17 ONTs should also be evaluated by bioanalytical assays, 18 for immunogenicity and anti-drug antibody (ADA) detection,11,19 for stereochemical control of chiral centers within certain ONTs18,20,21 and by methods to establish the drug release criteria; ideally without the use of radiolabeled tracers for ease of use, safety, and cost.17,22 We hypothesized that some of these needs might be addressed by deploying immunoassays, particularly if highly specific antibodies to ONTs could be generated and made available to researchers.
Existing Analytical Methods: Unmet Need for Alternatives
Many publications focusing on analytical assays for ONTs would arguably lead the reader to conclude that liquid chromatography combined with mass spectrometric detection23–25 is the method of choice to obtain structural information about ONTs directly, despite known sensitivity challenges associated with the assay.26,27 In Table 1 we present a summary of many existing analytical methods for ONTs. Absorption studies of ONTs mainly include the use of liquid chromatography-tandem mass spectrometry (LC-MS/MS)28,29 or ligand binding assays (LBA)31,32 to determine the concentration of ONTs in plasma, tissue homogenates, and other biofluids. 30 Distribution studies of ONTs, often using radiolabeling of the drug 33 or hybridization assays34,35 such as in situ hybridization (ISH) and more recently dual-ISH (DISH), 36 frequently rely on microscopy to determine tissue distribution and fluorescent imaging to determine intracellular localization. The latter sometimes requires fluorescent labeling of the ONT (e.g., Cy3) which may alter significantly cellular uptake and other pharmacodynamic properties of the drug. 30 Metabolism and excretion studies, including those designed to detect metabolites of ONTs, primarily use LC-MS/MS to analyze urine and other biofluids and may require the use of radiolabeling to isolate 3′ or 5′ “shortmer” sequences30,32,34 Sequential anion exchange and reversed-phase solid-phase extraction are also used for this purpose.28,37
Commonly Used Analytical Assays for ONTs
While LC-MS/MS, LBA, and HL are recognized as routine assays for ONT characterization, several other types of assays provide alternatives, in part, to overcome known limitations for core analytical assays. qPCR has the highest sensitivity and LC-MS/MS has the best specificity for measuring and monitoring ONTs.
DISH, dual-ISH; ISH, in situ hybridization; ONTs, oligonucleotide therapeutics.
Bioanalytical methods, such as the enzyme-linked immunosorbent assay (ELISA), often rely on hybridization of the ONT to a complimentary oligonucleotide sequence where subsequent ligation of a tagged probe sequence is detected using enzyme-labeled antibodies, 18 or likewise in an electrochemiluminescent platform. 38 Variants of this method using branched DNA can increase sensitivity and accuracy.39,40 In addition, hybridization-ligation ELISA (HL-ELISA) and reverse-transcription quantitative polymerase chain reaction (RT-qPCR) have further improved the sensitivity of these assays and the ability to quantitate ONTs. 18 Others have developed assays for the bioanalysis of ONTs including capillary gel electrophoresis, 41 LC-fluorescence assays, and nanoscale secondary ion mass spectrometry (nanoSIMS), 42 as well as other chromatographic methodologies. 37 For measuring and monitoring ONTs, qPCR has the highest sensitivity and LC-MS/MS has the best specificity.45–48 Often bespoke antibodies are developed for specific ONTs themselves, where the resultant antibody may recognize sequence, conformation, modification, and other possible epitopes. As such, these antibodies are typically generated and used “in-house” and may not have utility for other ONT drugs or across drug platforms and may not be available for use except by their originators, or otherwise as gifts, or with limitations on use.43,44
However, all the above-listed methods have well-established pros and cons 15 and are highly dependent on length/size, sequence, and the chemical modification of the target oligonucleotide. For instance, hybridization assays, including ISH, have limitations that include lack of general utility (as unique probes must be designed individually for each ONT), the significant time and costs required to optimize probes, limitations for certain ISH probes to bind and detect short ONT drug sequences and difficulties encountered when repeated sequences are present within the ONT.45,49 Lastly, qPCR, arguably the gold standard for sensitive quantitation of DNA and mRNA molecules, has insufficient sensitivity for short or heavily modified ONTs. 50 Indeed, many existing assays suffer from a “general lack of sensitivity” needed for preclinical assays, creating an unmet need for alternatives. 18
Chemical Modifications of ONTs: Common Features to Exploit
Some of the practical limitations of ONTs as viable drugs have been at least partially addressed over several decades of development; specifically, strategies for tissue-specific drug delivery, such as GalNAc conjugates that bind to the asialoglycoprotein receptor for hepatic targeting, 51 or the use of lipid nanoparticles for drug encapsulation. 52 These strategies are expanding the potential clinical repertoire for ONTs. One further key development in ONTs has been the introduction of chemical modifications that significantly improve compound stability without modifying the ONT drug sequence.53,54 Medicinal chemists improved the “drug-like properties” of ONTs by incorporating modifications into the sugar-phosphate backbone and/or nitrogenous bases, 53 which are designed to increase ONT drug stability, uptake, and efficacy, but may increase the likelihood of toxicity. 55 Recent studies reveal that toxicity varies significantly between ONT sequences and impurities 56 although this can be modulated by adjustments to the ONT composition, such as the introduction of divalent cations. 57 First-generation chemical modifications (see Fig. 1) centered on phosphorothioate (PS) modification of the phosphate portion of the backbone58,59 whereas second-generation modifications included adding substituent groups to the 2′ position of the ribose portion of the backbone, specifically 2′-O-methyl (OMe), 2′-O-methoxy-ethyl (MOE), and 2′-fluoro modifications of RNA. 60 Third-generation modifications 61 include peptide nucleic acids, locked nucleic acids, morpholino phosphoramidate modifications, and others. 62 Of the ONT drug modalities, ASOs designed as “GapmeR”s are comprised of typically three to five 2′-modified nucleotides on either side of a central unmodified DNA “gap” sequence; this general structure can induce target mRNA degradation via endogenous RNAse H activity. 63 Drug developers can therefore integrate different chemical modification combinations around this basic structure while balancing both efficacy and tolerability. 55 Crucially, this has led to some convergence around optimal modifications across diverse clinical lead ONTs.
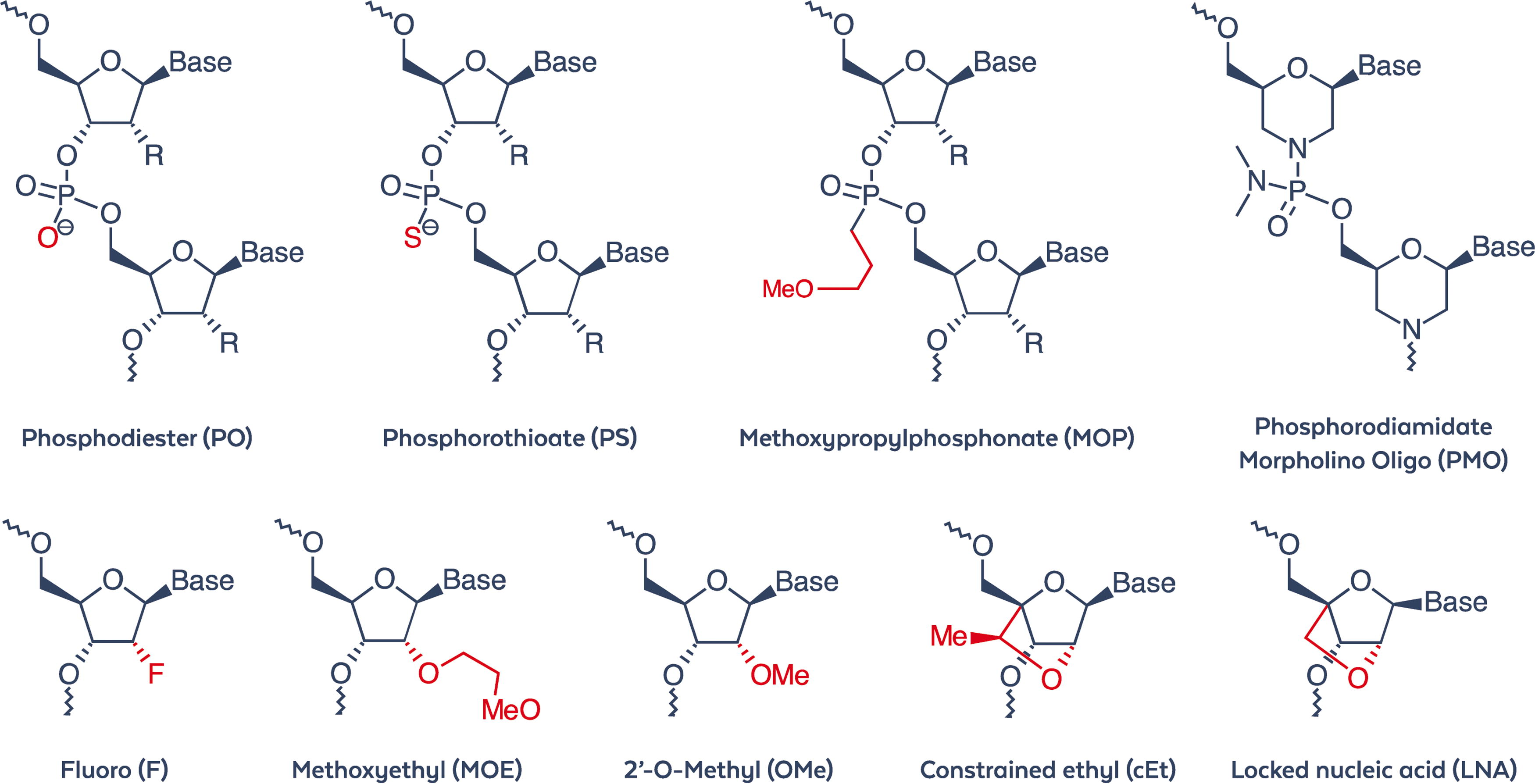
Partial structures of chemical modifications frequently used by medicinal chemists to enhance the “drug-like properties” of ONTs. ONTs, oligonucleotide therapeutics.
Strategies for the Development of Antibodies
Based on our experiences we recognize two challenges in generating antibodies to chemical modifications on oligonucleotides: nucleic acids are notoriously poor immunogens, and chemical modifications themselves represent inherently a minimal epitope for antibody detection. After analyzing common ONT drug structures we concluded that only monoclonal antibodies could effectively and reproducibly recognize minimal compositional changes, such as the substitution of an oxygen atom for sulfur within the chemical modification of PO to PS (e.g., phosphodiester to phosphorothioate). Additionally, selecting a cohort of clones from more than one splenocyte-myeloma fusion could improve the likelihood of obtaining fit-for-purpose antibodies capable of detecting chemical modifications in different types of ONT drugs (e.g., ASO, siRNA, mRNA, or aptamer), while ensuring the antibody’s utility across a range of immunoassays. This “panel” approach to develop monoclonal antibodies yielded the intended result for detecting PS, 2′-OMe, or 2′-MOE modifications with differential sensitivity and specificity independent of sequence or format (see Table 2).
Panels of Monoclonal Antibodies Sorted by Specificity for Chemical Modification, e.g., PS, 2′-MOE, or 2′-OMe
Data are shown for reactivity against fully modified ASO and a 5-10-5 GapmeR used for selection and screening clones and characterization of purified antibodies. The GapmeR used for screening contained both PS and 2′-MOE modifications but did not contain 2′-OMe, e.g., inotersen. See the text for a complete description of positive and negative controls used to develop panels of antibodies. Key: (−) negative; (+/−) weak; (+) positive; (++) strong; (+++) very strong; (ND) not determined.
Modified guanine nucleotide was excluded as a control.
MOEC is a cocktail of three unique clones mixed at a ratio of 1:1:1.
ASO, antisense oligonucleotides; PS, phosphorothioate.
The immunogens used to elicit the immune response in mice were either a modified nucleotide-carrier protein or a modified oligonucleotide-carrier protein conjugate, depending on which chemical modification was targeted for immune response. Upon analysis of our empirical data, we concluded that derivatized modified nucleotide/oligonucleotides conjugated to Keyhole limpet hemocyanin as a carrier protein in ratios ranging from 4:1 to 10:1 would serve as optimal immunogens. Likewise, the dosage used for the initial immunization of mice and subsequent booster injections, as well as the route of administration were carefully selected based on proprietary data.
Once generated, hybridomas were screened for sensitivity and specificity using ELISA and/or immunofluorescence (IF) microscopy as surrogate assays. We surmised that these assays could mimic in vitro and in vivo assays, including those assays designed to assess the biological effects of ONTs. Control oligonucleotides were synthesized and used to elucidate both the sensitivity and specificity of the antibody clones during the development phase. Negative control unmodified oligonucleotides possessed identical nucleotide sequences as the chemically modified PS, 2′-OMe and/or 2′-MOE positive control oligonucleotides. All oligonucleotide controls used in ELISA were conjugated to bovine serum albumin as a carrier protein to facilitate binding to the ELISA plate. In addition to indirect ELISA, a sandwich ELISA was employed where oligonucleotide controls were captured by candidate antibodies (capture antibody) and detected with an antibody carrying a quantifiable label (detection antibody), e.g., biotin or a fluorochrome.
Furthermore, modified oligonucleotide controls were synthesized with various incorporation levels of the selected modification, from 0% to 100%. For example, for the PS modification, controls were synthesized with 0% (naturally occurring PO backbone), 5%, 10%, 25%, 50%, 75%, and 100% PS incorporation. Additional negative control oligonucleotides were synthesized with 2′-MOE or 2′-OMe modifications but without PS modification. Additional positive control oligonucleotides were synthesized comprising the target chemical modification alongside additional modifications (e.g., as would be applied in GapmeR ASOs), and with varying nucleotide sequences. Unmodified nucleotides and nucleotides modified on the nitrogenous base rather than the internucleotide linkage were also used as negative controls.
Ultimately hybridomas were selected after iterative limiting dilution steps where the amount of screening oligonucleotide was varied from 0.1 µg to 0.5 µg to assess for sensitivity. Only hybridomas reactive to the positive controls and nonreactive to all negative controls were selected for further screening. Hybridomas showing sufficient titer, sensitivity, and specificity were selected for further development which included production in cell culture, cell banking, scale up, and purification using protein A chromatography. At each stage of development, the full complement of positive and negative control oligonucleotides was used to assess performance.
Orthogonal Approach: Antibodies Specific for Modification Independent of ONT Sequence
By using the strategies described above we exploited the common chemical features of modified ONTs to develop, optimize, and validate panels of monoclonal antibody reagents as analytical tools that precisely detect either PS, 2′-MOE, or 2′-OMe chemical modifications independent of nucleic acid composition, structure, strandedness, configuration or platform. 64 Example immunocytochemical data for anti-PS clones are shown here (Fig. 2) and preliminary studies using an anti-2′ MOE clone have been described elsewhere. 65 These “universal detection reagents” may be used directly in (1) immunohistochemistry to determine biodistribution, including co-localization with standard tissue/cell markers; (2) IF to determine intracellular localization, e.g., sequestration (Fig. 3); (3) ELISA to determine ADA binding, immunogenicity, and drug ranking; (4) immunoprecipitation to determine protein binding in serum and/or tissue lysates; (5) in vitro cell culture studies to support ONT potency assessments; (6) for quantification studies (Fig. 4); (7) to satisfy the unmet need to understand the cytosolic and nuclear penetration of ONTs, 66 (8) three-dimensional cell culture studies to determine the penetration of ONTs into culture systems such as spheroids, (9) FACS to identify and separate cells dissociated from tissue based on the presence or absence of intracellular ONTs within individual cells, (10) characterization of the stereochemistry of PS containing ONTs by assessing the chiral make-up of ONTs that may affect biological activity, and (11) other immunoassays. We summarize the potential uses of these novel analytical reagents in Table 3.
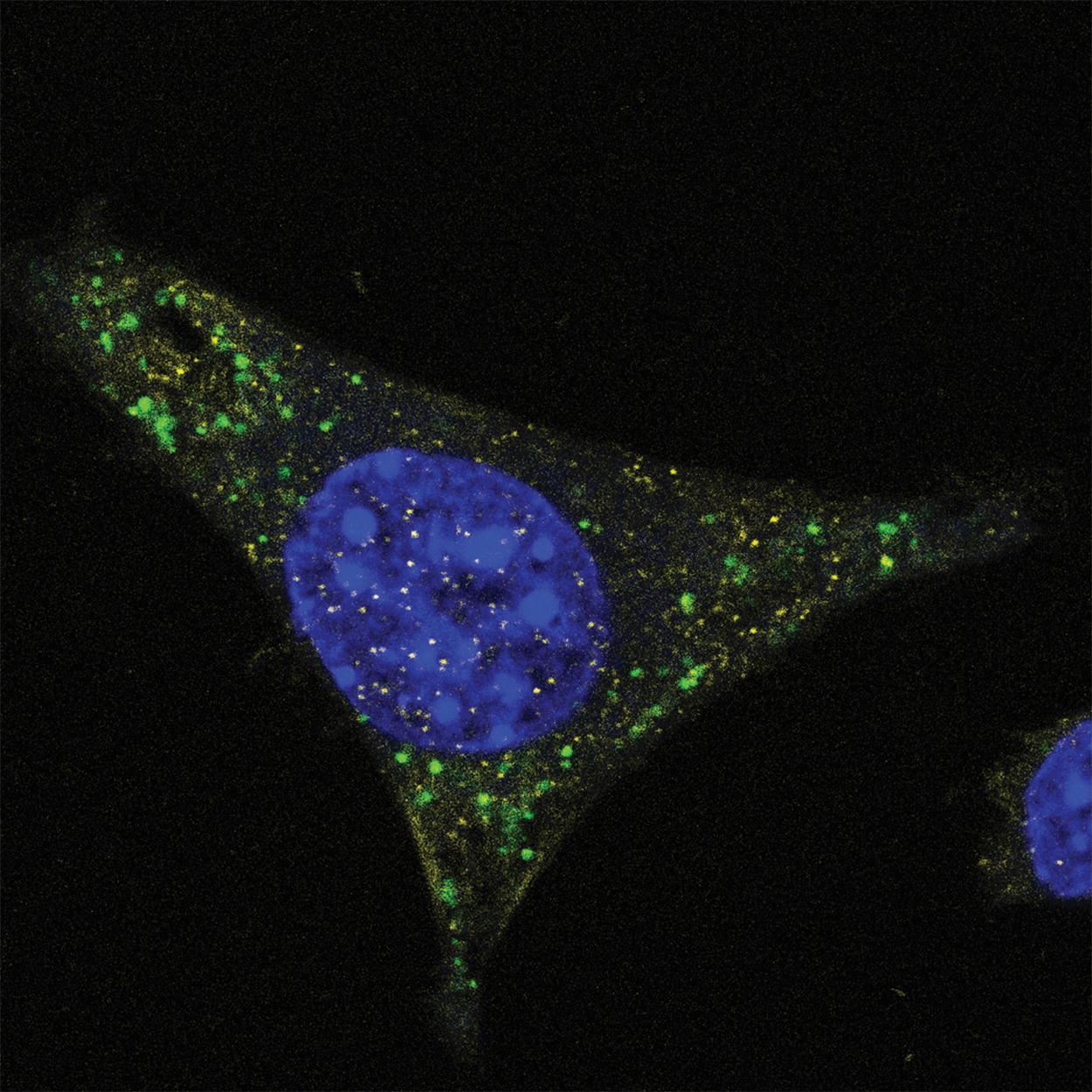
Immunofluorescence microscopy using anti-PS monoclonal antibody clone PS05. Mouse glioma cells derived from C57BL/6 mice were cultured and treated with fully PS-modified ASO drug. After fixation with paraformaldehyde, cells were incubated with a 1:2,000 dilution of anti-PS monoclonal antibody clone PS05 (green) in PBS followed by counterstaining of nuclei with DAPI (blue). Vehicle-only treated cells showed no staining (not shown). Image courtesy of CoSyne Therapeutics. ASO, antisense oligonucleotides; PS, phosphorothioate.
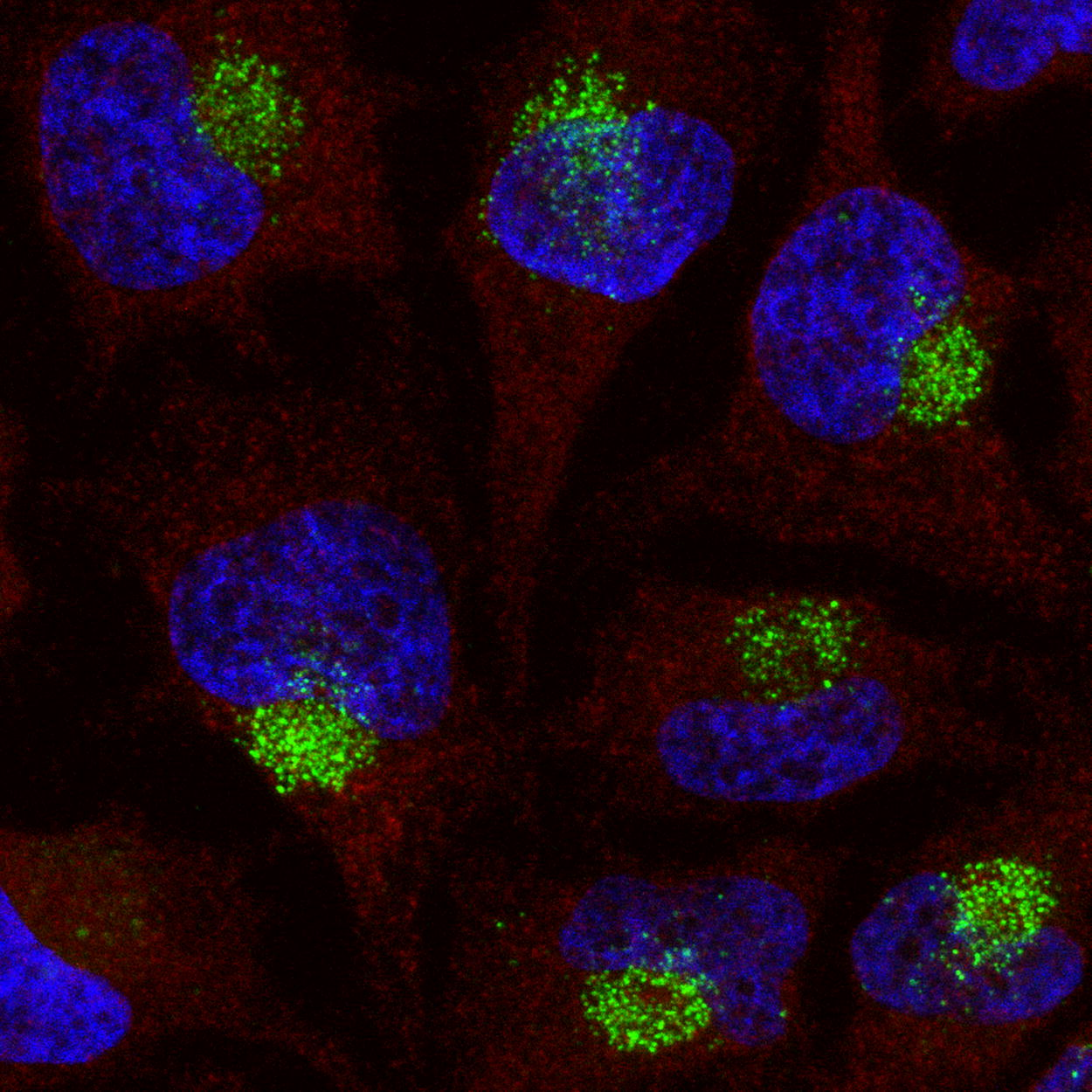
Immunofluorescence microscopy using anti-PS monoclonal antibody clone PS03. HeLa cells were cultured and treated with 100 nM of a fully-PS modified 16-mer MALAT-1 ASO by gymnosis for 72 h. After fixation with paraformaldehyde, cells were incubated with a 1:1,000 dilution of anti-PS monoclonal antibody clone PS03 (green) in PBS and an anti-alpha tubulin (red) antibody followed by counterstaining of nuclei with DAPI (blue). Cytoplasmic accumulation of signal indicative of endosomal compartmentalization is observed. Data generated by Inês Fial, Nucleic Acid Therapy Accelerator (NATA).
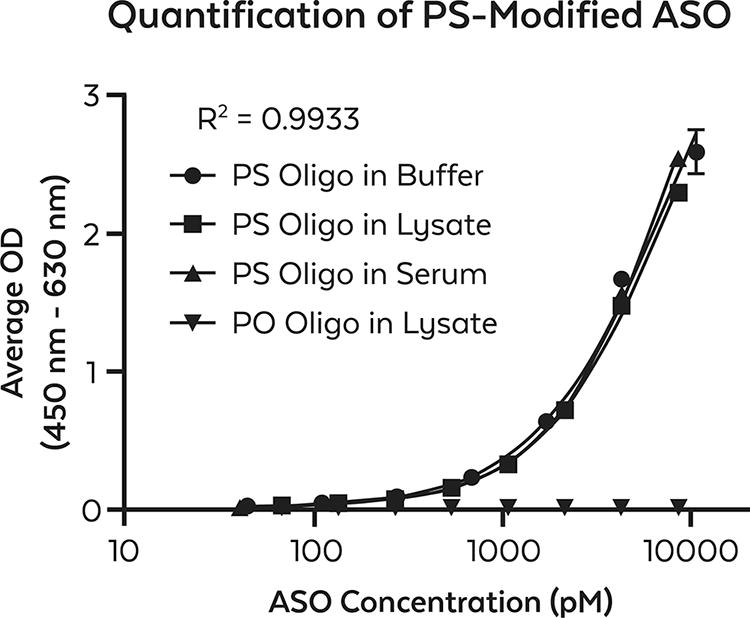
Quantification of fully PS-modified ASO by ELISA using anti-PS monoclonal antibody (clone PS04). The ability to quantify PS-modified ASO diluted in tris-based buffer (●) over a range from 44 pM to 11 nM concentrations is shown in the curve as indicated by spiking known amounts of the ASO from a 100 nM stock by 2.5-fold serial dilution and subsequently detected using anti-PS monoclonal antibody. Samples were run in triplicate. A standard curve was plotted as the average OD result vs. the log of ASO concentration in pM using a 4PL best-fit formula. The effect of matrices was investigated by performing similar experiments replacing the tris-based buffer with either HeLa whole cell lysate (■) or mouse serum (▲) at initial protein concentrations of 0.3 µg/mL and 2.5 mg/mL, respectively. Nonmodified PO oligonucleotide of the same sequence was used as a negative control (PO Oligo) and was similarly prepared in HeLa whole cell lysate (▼). Reproducibility was confirmed by repeating the assay twice (data not shown). For PS-modified ASO diluted in tris-based buffer, the LLOQ defined as the lowest standard detected is 44 pM. The LOD, defined as the average signal of background + 3× standard deviation) was determined to be <33pM. The ULOQ as shown is 11 nM. The LLOQ/ULOQ is defined as the lowest/highest standard concentration meeting the acceptance criteria of calculated/theoretical × 100% recovery between 70% and 130% and CV ≤25% for LLOQ and between 80% and 120% and CV ≤20% for ULOQ. Further optimization and sensitivity enhancement will likely decrease the LLOQ by one log unit. ELISA, enzyme-linked immunosorbent assay; LLOQ, lower limit of quantitation; LOD, limit of detection; ULOQ upper limit of quantitation.
Potential Use of Monoclonal Antibodies Specific for Chemical Modifications to ONTs Independent of Nucleic Acid Sequence in Various Immunoassays and Their Intended Purpose
Assays may be considered either to be primary assays or constitute an orthogonal approach to the collection of data intended to ensure the safety and efficacy of ONTs.
Conclusion
There is an ongoing and significant need to develop new analytical methods for ONTs and improve existing localization and quantification methodologies. 18 Beyond fundamental analysis (e.g., localization and quantification in tissue), there are additional needs “to study the metabolism and biotransformation of these molecules,” 67 including better methods to identify oligonucleotide metabolites and the ability to quantify ONTs in extrahepatic tissues, especially when low concentrations of these drugs are likely present due to their relatively long drug half-lives and infrequent administration. For instance, within the oligonucleotide field, it is certain that quality control programs require “differential” analytical expertise and technologies. 68 While LC-MS/MS, HI-ELISA, ISH, qRT-PCR, and 31P NMR-SAX/HPLC 69 have become accepted methods for analysis and quantification of ONTs, the novel universal detection reagents we describe present opportunities to expand the analysis of ONTs to support PD/PK studies intended to demonstrate their efficacy and safety. Ultimately, we envision this technology accelerating the rate at which ONT drugs advance through clinical trials and regulatory approval.
Here, we opine that highly specific antibodies to ONT modifications and the immunoassays they enable may potentially enhance the development and production of ONTs, expand studies to determine the safety and efficacy of this class of drugs, and serve as orthogonal assays to supplement data collected using established assays for the analysis of ONTs. Although immunoassays were not specifically named in the recently issued European Medicines Agency draft guidelines, the use of “orthogonal analytical assays” for certain purposes was encouraged. 70 Immunoassays based on the use of precise antibody-based analytical tools that only recognize chemical modifications, independent of nucleic acid sequence, would be an added tool in the toolbox of analytical assays. Commonly, immunoassays are well-established as a rapid and highly reproducible analytical tool. 71 Thus, novel, well-validated monoclonal antibodies that detect unique chemical modifications in ONTs may have broad applications to preclinical, nonclinical, and clinical studies involving ONTs. This is especially true now as the diversity of chemical modifications, the length of ONTs, and emerging backbone chemistries continue to expand creating even greater challenges to existing assays in this field. 72
Identifying ONTs within the cytosol is quite difficult and has been described as a significant barrier to drug development. 66 Our unpublished immunochemical studies to date have shown accuracy and ease of use to localize and quantitate ONTs in cells and tissue and specifically determine their subcellular localization. Furthermore, as siRNA and potentially other RNA modalities are considered a platform technology by the FDA, immunoassay-based analytical tools may facilitate the identification of targeting moieties and aid in quality control assessments. These reagents, binding independently of sequence, facilitate the efficient comparative assessment of how sequence modification affects biological activity and safety, and therefore constitute a solution to problems identified by the FDA. 73 In a 2020 review article, Sutton et al. concluded that “there are still many advances to be made in the field of oligonucleotide analysis.” 26 We agree and believe that highly specific antibody-based immunoassays capable of detecting as little as a single modified internucleotide linkage in 20 may represent a significant advance and should be thoroughly explored for utility by the oligonucleotide community.
To validate our hypothesis, we encourage researchers to use and interrogate this novel technology. Our data, while limited, is highly suggestive of the broad suitability of these immunoassays to collect meaningful analytical data that has relevance to preclinical, nonclinical, and clinical studies. If this technology truly signifies a breakthrough in the analysis of ONTs, this can only be verified by rigorous assessment of the reagents and methods. Such an assessment should consider various aspects of ONT quality including pharmacological properties, safety, and efficacy by utilizing the immunoassays as described herein. Ideally utilizing this technology to collect data by an orthogonal approach could go far in demonstrating the value of this technology.
Footnotes
Acknowledgments
The authors acknowledge critical comments from P. Oliver and A. Krainer. The authors also thank A. Gilmore for the careful editing of the document and T. Giardiello and S. Farrier for assistance in the preparation of figures.
Author Disclosure Statement
D.P.C., A.L.A., and C.A.A. are employees of Rockland Immunochemicals, Inc. I.F. has no disclosures.
Funding Information
NATA is supported by the Medical Research Grant MC PC 20061. The information presented is covered under U.S. Provisional Patent Application No.: 63/547,067.
