Abstract
Early characterization of the immunostimulatory potential of therapeutic antisense oligonucleotides (ASOs) is crucial. At present, little is known about the toll-like receptor 9 (TLR9)-mediated immunostimulatory potential of third-generation locked nucleic acid (LNA)-modified ASOs. In this study, we have systematically investigated the TLR9-activating potential of LNA-modified oligonucleotides using different mouse and human cell culture systems. Although it has been reported that LNA modifications as well as cytosine methylation of 5'-cytosine-phosphate-guanine-3' (CpG) motifs can reduce TLR9 stimulation by phosphorothioate (PTO)-modified oligonucleotides, we identified CpG-containing LNA gapmers with substantial TLR9-stimulatory activity. We further identified immunostimulatory LNA gapmers without CpG motifs. Unexpectedly, methylation of cytosines only within the CpG motif did not necessarily reduce but could even increase TLR9 activation. In contrast, systematic methylation of all cytosines reduced or even abrogated TLR9 activation in most cases. Context dependently, the introduction of LNA-modifications into the flanks could either increase or decrease TLR9 stimulation. Overall, our results indicate that TLR9-dependent immunostimulatory potential is an individual feature of an oligonucleotide and needs to be investigated on a case-by-case basis.
Introduction
The field of oligonucleotide-based therapeutics has matured massively in the last years with an increasing number of approved drugs 1 and a multitude of projects in clinical development. While frequently chemically modified to enhance stability and efficacy, 1 oligonucleotide-based therapeutics are principally based on genetic material such as DNA or RNA. Genetic material is present extracellularly under pathological situations such as infection or immunogenic cell death. Higher organisms have hence developed sensors that recognize foreign DNA or RNA and trigger an immune response as a first line of defense against potential pathogens. 2 One of these sensors is the pattern recognition receptor (PRR) toll-like receptor 9 (TLR9), which, in humans, is expressed primarily on antigen presenting cells such as B cells and plasmacytoid dendritic cells. 3 TLR9 can be activated in a sequence-specific manner by single-stranded DNA.4,5 Therefore, single-stranded DNA-based oligonucleotide therapeutics such as antisense oligonucleotides (ASOs) have the potential to activate TLR9 resulting in immune-mediated adverse effects. Nonetheless, intentional stimulation of TLR9 can be exploited for treatment by synthetic DNA-based agonists and presently holds promise for the immunotherapy of cancer6,7 or infectious diseases.8,9 In all cases, a comprehensive characterization of the TLR9-stimulating properties of the compound is required.10,11
TLR9 agonists differ considerably in their potential to activate either murine or human TLR9.12–14 These differences hamper reliable testing of TLR9 stimulating potential of oligonucleotides intended for human use in the mouse. Therefore, preclinical characterization of TLR9-stimulatory potential relies on human cells expressing human TLR9.
Strategies to deal with unwanted TLR9 stimulation can be avoidance and early detection. TLR9 activation is associated with unmethylated CpG motifs, which are abundant in bacterial DNA and virus DNA but are underrepresented and selectively methylated in vertebrate genomes. 15 These are critical factors that limit TLR9 activation by endogenous DNA.16,17 It has been described that cytosine methylation in a CpG motif at the 5’ position of an oligonucleotide can lead to decreased unwanted immune stimulation. 12 Furthermore, selection of sequences without CpG motifs during the design of ASOs is a strategy to avoid unwanted TLR9 stimulation. However, CpG-independent TLR9 stimulation has been reported for some sequences, and thus, methylation or avoidance of CpG motifs does not guarantee the absence of unwanted TLR9 stimulation by an ASO.18,19
In this article, we have focused on the immunostimulatory potential of locked nucleic acid (LNA)-modified gapmer ASOs. Gapmer ASOs are designed to achieve a specific suppression of target gene expression in cells. Via base-pairing they bind to the target RNA in the nucleus and recruit the nuclear enzyme RNase H that cleaves the RNA, 20 initiating RNA degradation and suppression of gene expression. To enhance stability against nuclease-mediated degradation, they typically contain a phosphorothioate (PTO) 21 backbone. The flanks are chemically modified to enhance target affinity, leaving a central unmodified gap that is required for RNase H recognition. In case of LNA modifications, the ribose moiety is modified with a bridge connecting the 2' oxygen and 4' carbon which “locks” the ribose in the 3'-endo conformation22,23 resulting in a strongly enhanced binding affinity to the target RNA 24 and increased potency. 25 In contrast to earlier generations of oligonucleotides with lower affinity, good target knockdown can be achieved with LNA-modified ASOs in tissues with lower oligonucleotide biodistribution, broadening the range of targets and diseases that can be addressed with this therapeutic modality. In tissues with good biodistribution, doses can be reduced, lowering the risk of backbone-mediated side effects. Earlier studies reported that LNA modifications alter ASO properties besides their effect on target affinity. For example, LNA modifications reduced binding of the oligonucleotide to proteins associated with thrombocytopenia, thereby influencing their risk to cause this condition. 26 Moreover, incorporation of LNA modifications in CpG-containing, TLR9-stimulating oligonucleotides strongly reduced their potential to stimulate TLR9. 27 For these reasons, LNA-modified ASOs hold strong promise as a safe and efficacious therapeutic modality.
In the present study, we extensively investigated the effects of LNA-modifications on the TLR9-stimulatory potential of ASOs in the context of different sequences, as well as compared different cell-based assays that measure TLR9 activation. In addition, we challenged the concept that methylation of CpG motifs consistently abrogates TLR9 activation. We show that methylation of CpG motifs does not reliably reduce TLR9 stimulation in all cases and not all CpG containing oligonucleotides necessarily activate TLR9. Our results strengthen the prevailing necessity of experimental testing and early detection of TLR9-activating potential of ASOs in adequate cellular systems.
Materials and Methods
Antisense oligonucleotides
All ASOs were synthesized by and purchased from Ella Biotech, except for the negative control oligonucleotides, which were purchased from Axolabs. The sequences, targets, and chemistry of the used oligonucleotides are listed in Fig. 1. LNA Cs: always LNA 5-methylcytosines.
HEK-Blue cells
HEK-Blue reporter cells HEK-BlueTM hTLR9 (InvivoGen) and HEK-BlueTM mTLR9 (InvivoGen) were seeded in flat-bottom 96-well plates (Fisher Scientific) at a density of 40000 cells/well in DMEM (Thermo Fisher Scientific) supplemented with 1% GlutaMAX (Thermo Fisher Scientific), 1% antibiotic/antimycotic (Thermo Fisher Scientific), 10% heat-inactivated FCS (Thermo Fisher Scientific), and 1% sodium pyruvate (Thermo Fisher Scientific). Moreover, selection antibiotics were included in the media, HEK-BlueTM hTLR9 received 10 µg/mL blasticidin (InvivoGen) and 100 µg/mL Zeocin (InvivoGen); and HEK-BlueTM mTLR9 received 30 µg/mL blasticidin and 100 µg/mL Zeocin. Next day, ASOs were added to the cells at indicated final concentrations by exchanging half of the media. Approximately 20–24 h post-treatment, secreted alkaline phosphatase (SEAP) activity was measured in supernatants using the QUANTI-BlueTM solution (InvivoGen). To do so, 20 µL of supernatant per well was incubated with 180 µL QUANTI-BlueTM solution at 37°C for 1 h (HEK-BlueTM mTLR9) and 2 h (HEK-BlueTM hTLR9); plates were allowed to cool down for 20 min, and absorbance was measured at 620–655 nm with a Clariostar Microplate Reader (BMG Labtech).
RAW Dual cells
RAW DualTM reporter cells (InvivoGen) were seeded in flat-bottom 96-well plates (Fisher Scientific) at a density of 100000 cells/well in cell culture media consisting of DMEM (Thermo Fisher Scientific) supplemented with 1% GlutaMAX (Thermo Fisher Scientific), 1% antibiotic/antimycotic (Thermo Fisher Scientific), 10% heat-inactivated FCS (Thermo Fisher Scientific), and 1% sodium pyruvate (Thermo Fisher Scientific). After 2–3 h of incubation at 37°C and 5% CO2, cells were treated with ASOs and placed back at 37°C and 5% CO2 for 20–24 h. SEAP activity was measured as described for HEK-Blue cells, after incubation with QUANTI-BlueTM solution at room temperature (RT) for 4–4.5 h. Interferon regulatory factor (IRF) induction was determined by measuring luminescence immediately after mixing 20 µL of supernatant per well with 50 µL QUANTI-LucTM reagent (InvivoGen).
Tumor growth in mice
All in vivo work was performed by Reaction Biology and mice were handled according to the German animal welfare law and the GV-SOLAS guidelines. This animal study has been approved by the Ethics Committee for Animal Experimentation and is registered by the regional board Freiburg. On Day 0, tumor cells (1.0 × 106 MC38-CEA cells in 100 μL PBS) were implanted into the left mammary fat pad of each mouse. Mice were randomized at a mean tumor volume of 58.1 mm3 and ASOs were injected intraperitoneally at a dose of 5 mg/kg dosing on days 5–9, 12, 14, and 16. Tumor growth monitoring was performed two times weekly after the start of therapy. Primary tumor volumes were determined by caliper measurement. Tumor sizes were calculated according to the formula W2 × L/2 (L = length and W = the perpendicular width of the tumor, L > W).
THP1-Dual hTLR9 cells
THP1-DualTM hTLR9 reporter cells were seeded in flat-bottom 96-well plates (Fisher Scientific) at a density of 100000 cells/well in RPMI (Thermo Fisher Scientific) supplemented with 1% GlutaMAX (Thermo Fisher Scientific), 1% antibiotic/antimycotic (Thermo Fisher Scientific), 10% heat-inactivated FCS (Thermo Fisher Scientific), and 1% sodium pyruvate (Thermo Fisher Scientific). Cells were then treated with ASOs and placed back at 37°C and 5% CO2 for 20–24 h. SEAP activity was measured as above after incubation with QUANTI-BlueTM solution at RT for 30 min. IRF induction was measured via luminescence immediately after mixing 10 µL of supernatant per well with 50 µL QUANTI-LucTM reagent (InvivoGen) (Supplementary Data S1).
Bjab cells
Bjab cells (DSMZ) were seeded in round-bottom 96-well plates (Fisher Scientific) at a density of 40000 cells/well in RPMI (Thermo Fisher Scientific) supplemented with 1% GlutaMAX (Thermo Fisher Scientific), 1% antibiotic/antimycotic (Thermo Fisher Scientific), 10% heat-inactivated FCS (Thermo Fisher Scientific), and 1% sodium pyruvate (Thermo Fisher Scientific). Cells were then treated with ASOs and placed back at 37°C and 5% CO2 for 48 h. For TNF-α measurements, cells were centrifuged for 5 min at 500g and supernatants were collected. Released TNF-α was measured via Enzyme-linked Immunosorbent Assay (ELISA) (BD Biosciences) according to the manufacturer’s instructions (Supplementary Data S1).
Peripheral blood mononuclear cells (PBMCs)
PBMCs were freshly isolated from buffy coats using SepMate-50 (IVD) tubes. Shortly, buffy coats (Donas GmbH) were diluted 1:1 in sterile PBS. Ficoll, 15 mL (Sigma), was added per SepMate-50 (IVD) tube (StemCell) and 17 mL of diluted buffy coat was added on top of the Ficoll layer. Tubes were centrifuged at 1200g at RT for 10 min with brake. The top layer was transferred to new 50 mL Falcon tubes and cells were washed twice with sterile PBS by centrifugation at 300g for 8 min. Finally, PBMCs were seeded in round-bottom 96-well plates (Fisher Scientific) at a density of 500000 cells per well in RPMI (Thermo Fisher Scientific) supplemented with 1% GlutaMAX (Thermo Fisher Scientific), 1% antibiotic/antimycotic (Thermo Fisher Scientific), 10% heat-inactivated FCS (Thermo Fisher Scientific), and 1% sodium pyruvate (Thermo Fisher Scientific). Cells were then treated with ASOs and placed at 37°C and 5% CO2 for 48 h. For interleukin-6 (IL-6) measurements, cells were centrifuged for 5 min at 500g and supernatants were collected. Released IL-6 was measured via ELISA (BD Biosciences) according to the manufacturer’s instructions.
Data presentation and statistical analysis
Statistical analysis was performed using GraphPad Prism 10. Data are presented as mean ± SD. Two-way ANOVA with Dunnett’s multiple comparison test was used for statistical comparison. Statistical significance has been indicated when three or more adjacent points in the curve had a P-value of P ≤ 0.05. *P ≤ 0.05, **P ≤ 0.01, ***P ≤ 0.001, and ****P ≤ 0.0001.
Results
Effects of LNA modifications and cytosine methylation on immunostimulatory activity of a mouse TLR9 agonist
ODN1668 is a CpG-containing immunostimulatory oligonucleotide that is stabilized with a PTO backbone. 28 It is a potent agonist of mouse TLR9 and has been shown to have a strong antitumor activity in mouse tumor models. 29 Therefore, we made use of its sequence as a basis to investigate the effects of LNA-modifications and cytosine methylation on TLR9 stimulation. Of note, we always used LNA 5-methylcytosines when replacing a cytosine by an LNA-C (all sequences are listed in Fig. 1).
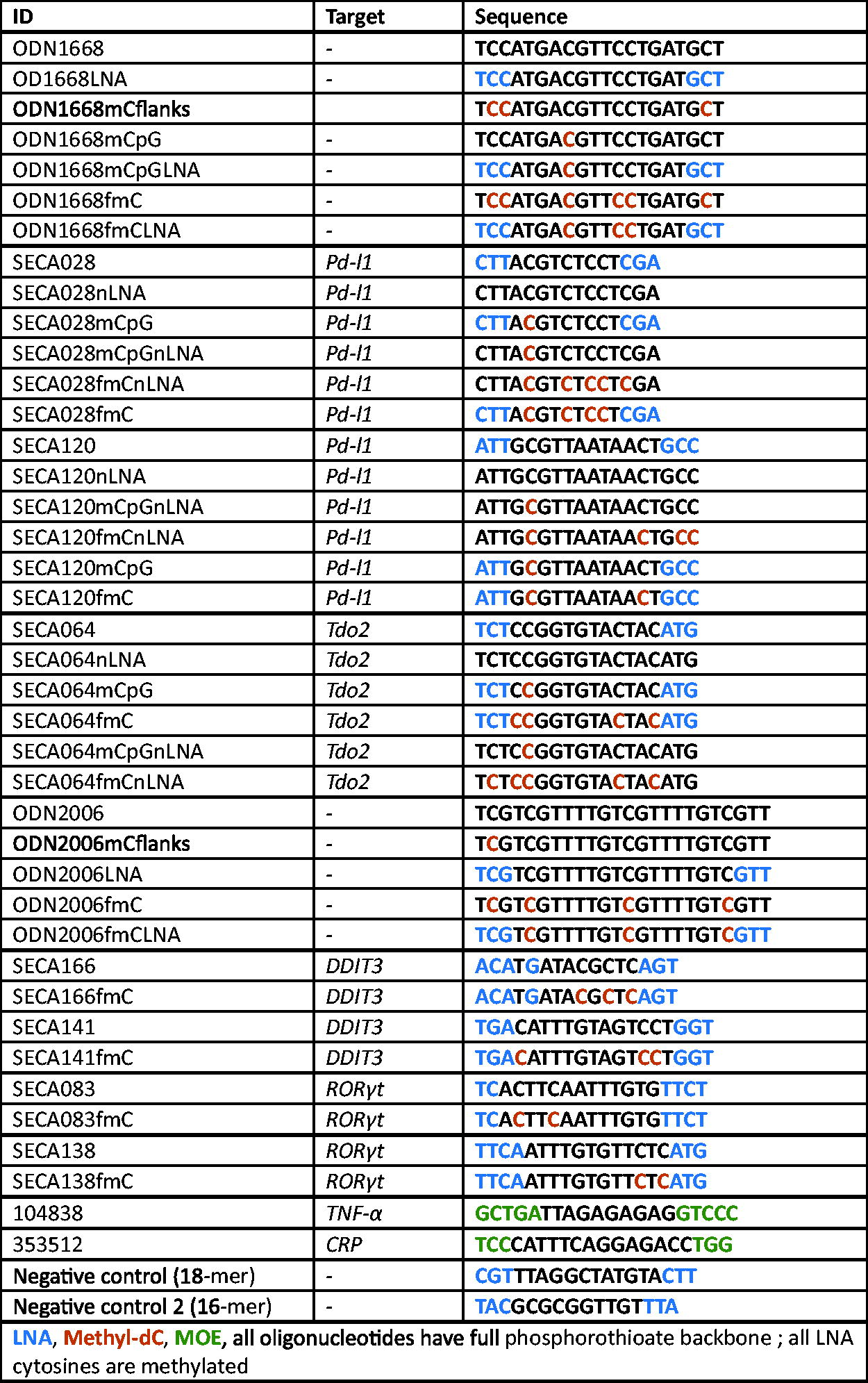
ASO sequences, targets, and chemical modifications.
In a gapmer design, we introduced three LNA modifications into each flank (thereafter termed ODN1668LNA). There are three cytosines present in the ODN1668 flanks, which are methylated after introducing the LNA modifications, and thus, we also generated a corresponding non-LNA-modified version with methylation of these three cytosines, which was termed ODN1668mCflanks. We tested the modified versions’ TLR9 activating potential in comparison with ODN1668 in the mouse HEK-BlueTM mTLR9 assay over a wide range of concentrations. For these versions, we observed a concentration-dependent increase in TLR9 activity reaching a plateau around 0.25 µM. Methylation of the cytosines in ODN1668mCflanks and LNA modifications in the flanks (ODN1668LNA) slightly but nonsignificantly reduced the maximal TLR9 activation compared with ODN1668 (Fig. 2A).
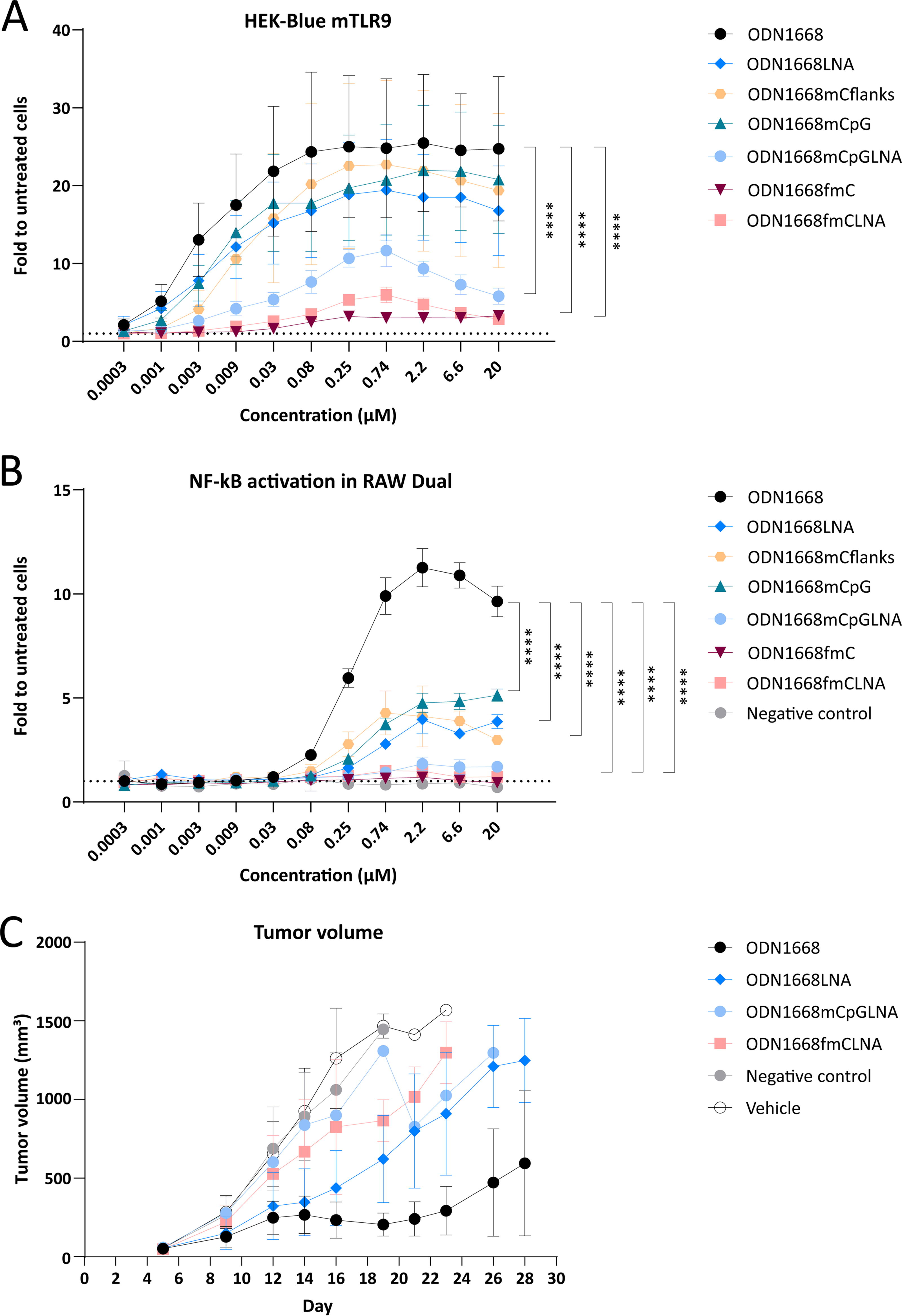
Effects of LNA modifications and cytosine methylation on immunostimulatory activity of ODN1668. HEK-BlueTM mTLR9 cells
It was previously described that TLR9 activation can be caused by unmethylated CpG motifs. 15 Surprisingly, methylation of the cytosine in the CpG-motif (ODN1668mCpG) only slightly and nonsignificantly reduced TLR9 activation, with peak TLR9 activation still >20-fold over untreated cells. TLR9 stimulation was, however, significantly reduced by further introduction of three LNA modifications (ODN1668mCpGLNA) into each flank. The concentration–response curve was more bell-shaped with a peak at 0.74 µM and decreasing TLR9 activation at higher concentrations. The strongest significant reduction was achieved by methylation of all 6 cytosine residues, both in the non-LNA-modified (ODN1668fmC) and LNA-modified (ODN1668fmCLNA) versions (Fig. 2A).
The HEK-BlueTM mTLR9 assay is a reporter cell system, in which the mouse TLR9 (mTLR9) is overexpressed in the human HEK293 kidney cell line together with a reporter for TLR9 signaling. We sought to confirm our findings in a physiologically more relevant test system without TLR9 overexpression. We selected the RAW DualTM cell system, which is a murine macrophage cell line with endogenously expressed TLR9 and expression of other PRRs such as the toll-like receptors TLR2 and TLR4, the cytosolic DNA sensor cGAS, and the cyclic dinucleotide sensor STING. Moreover, the stable expression of two reporter genes encoding for SEAP (secreted embryonic alkaline phosphatase), which is regulated by the NF-κB pathway, and Lucia luciferase, which is under the control of the IRF pathway, enables independent monitoring of both pathways. ODN1668 potently stimulated the NF-κB arm with a maximal ∼12-fold induction at a concentration of 2.2 µM. LNA modification of the flanks (ODN1668LNA) as well as methylation of the three cytosines in the flanks (ODN1668mCflanks) or of the CpG motif only (ODN1668mCpG) significantly reduced activity, although the maximal stimulation by these variants was still approximately fivefold over untreated cells. Methylation of the CpG-motif in combination with LNA-modified flanks (ODN1668mCpGLNA) further reduced activation of the TLR9- NF-κB arm. As in the HEK-BlueTM mTLR9 assay, methylation of all cytosine residues with (ODN1668fmCLNA) and without (ODN1668fmC) LNA flanks mostly abrogated the TLR9-stimulatory potential measured by the NF-κB arm. The negative control oligonucleotide, which is a nontargeting LNA gapmer 18-mer oligonucleotide, did not show any activity on this readout (Fig. 2B). The IFN-stimulated response arm of TLR9 was not activated by any of these oligonucleotides (Supplementary Fig. S1A), although it could be stimulated by the STING activator 2′3′-cGAMP (Supplementary Fig. S1B).
Overall, both the HEK-BlueTM mTLR9 and the RAW DualTM assays showed that LNA-modified oligonucleotides are still capable of stimulating TLR9 signaling. Cytosine methylation of CpG motifs only had a limited effect on reducing TLR9 stimulation. In contrast, methylation of all cytosines resulted in a strong reduction of TLR9 activation in both assays.
To investigate whether the in vitro results are of relevance in vivo, we utilized the finding that TLR9 agonists such as ODN1668 can have profound antitumor effects in the mouse. 29 We performed our study in the syngeneic MC38 colorectal cancer mouse model, since it is reported to respond to systemically administered TLR9 agonists. 6 Tumor-bearing mice were treated with ODN1668 and selected variants, specifically ODN1668LNA, ODN1668mCpGLNA, and ODN1668fmCLNA by intraperitoneal injection. In mice treated with ODN1668, we observed a strong tumor growth reduction compared with vehicle-treated mice or mice treated with the nontargeting and non-TLR9-stimulating control oligonucleotide. ODN1668LNA also considerably reduced tumor growth, however, to a lesser degree. ODN1668mCpGLNA and ODN1668fmCLNA further diminished antitumor activity (Fig. 2C). Together, these results show that—in the mouse—some antitumor activity could be achieved by LNA-modified ASOs and that the antitumor activity of ODN1668 variants at large correlated with their TLR9 activity in vitro.
Stimulation of mouse TLR9 by LNA gapmers designed to suppress gene expression
ODN1668 was designed to stimulate immune responses and not to suppress gene expression. As a next step, we investigated the immunostimulatory potential of LNA gapmers that were designed to suppress gene expression in the mouse. We used the mouse HEK-BlueTM assay, shown above to be a sensitive in vitro assay for TLR9 stimulation detection as well as to correlate with antitumor activity in vivo in the mouse (Fig. 2). SECA028 is an LNA gapmer targeting mouse Pd-l1 that has been reported to stimulate TLR9 in the mouse. 6 Its sequence contains three LNA modifications in each flank, an unmethylated CpG motif in the gap, as well as three additional unmethylated cytosine residues in the gap (Fig. 1). Figure 3A shows that it induced an ∼4.5-fold increase in TLR9 activity over a broad concentration range. Removal of the LNA-modifications (SECA028nLNA) did not overtly change the TLR9-stimulating properties of the ASO. To reduce the TLR9-stimulating properties of these ASOs, we methylated the CpG motif. To our surprise, TLR9-stimulation was significantly increased in the LNA-modified version, reaching levels >10-fold over untreated cells, whereas in the non-LNA-modified version (SECA028mCpGnLNA), CpG methylation resulted in complete abrogation of TLR9 stimulation. There was also no detectable TLR9 activity of the non-LNA-modified version after methylation of all cytosine residues (SECA028fmCnLNA), whereas the LNA-modified fully methylated counterpart SECA028fmC still had TLR9-stimulating activity, although very low (Fig. 3A).
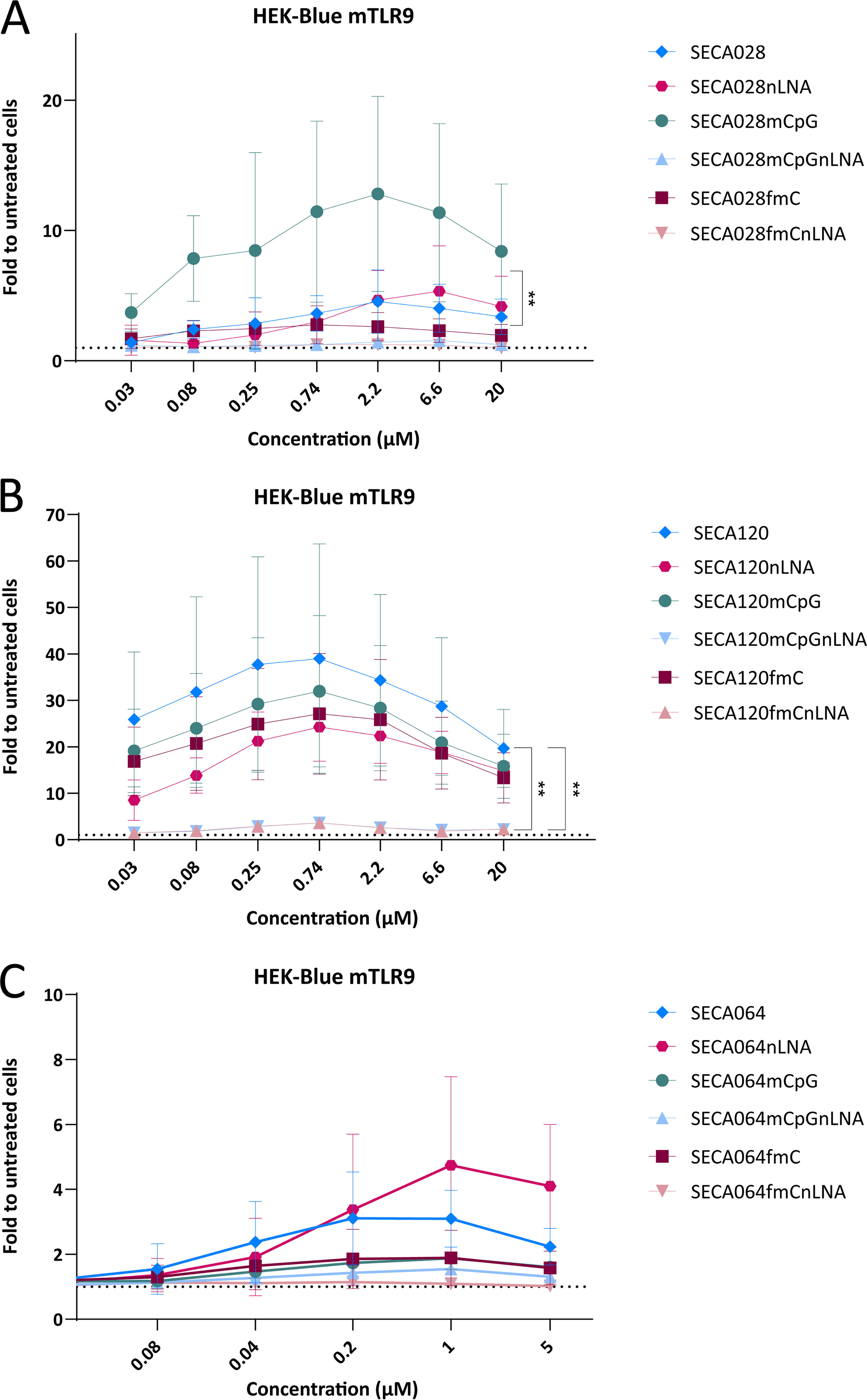
Stimulation of mouse TLR9 by gene-targeting LNA gapmers. HEK-BlueTM mTLR9 cells were treated with a concentration range of SECA028 and its variants
Altogether, these results not only indicate that, for some LNA-modified sequences, methylation of cytosines only in CpG-motifs could even increase TLR9 stimulation, while methylation of all cytosines resulted in robust reduction of TLR9 stimulation. They also show that, in certain instances, LNA-modified ASOs could have a higher immunostimulatory potential than their non-LNA counterparts.
We were interested in whether these findings could be reproduced with other sequences and tested SECA120, another mouse-specific LNA gapmer designed to suppress Pd-l1 expression. Its sequence contains two unmethylated cytosines in the gap, one of them within a CpG motif. There are two additional methylated cytosines in the LNA flanks (Fig. 1). SECA120 resulted in robust TLR9 stimulation with a bell-shaped concentration–response curve and an almost 40-fold induction at 0.74 µM. The non-LNA-modified version (SECA120nLNA) had a comparable profile although its maximal TLR9 stimulation was reduced compared with SECA120, however, not significantly. Interestingly, methylation of either the CpG-motif (SECA120mCpGnLNA) or of all cytosine residues (SECA120fmCnLNA) mostly abrogated the immunostimulation only in the non-LNA-modified versions, while they only had minor effects in the LNA-modified versions SECA120mCpG and SECA120fmC, respectively. While methylation of only the CpG motif in SECA120 did not increase TLR9 stimulation as observed for SECA028, methylation had a more pronounced effect on the reduction of TLR9 stimulation of non-LNA-modified ASOs than on LNA-modified counterparts in both sequences (Fig. 3B).
As a third example we tested SECA064, an ASO targeting mouse Tdo2. Its sequence contains one CpG motif in the gap, three further unmethylated cytosine residues in the gap, and one methylated cytosine in the LNA-flank. Its TLR9-stimulatory potential was low when compared with the Pd-l1 ASOs (approximately threefold induction compared with untreated cells). For this example, the non-LNA-modified version SECA064nLNA achieved a stronger activation of TLR9 than the LNA-modified ASO SECA064. Methylation of cytosines either in the CpG motif or of all cytosines strongly downregulated TLR9 activity both in the LNA and in the non-LNA-modified versions (Fig. 3C).
Taken together, these results show that also LNA-modified gapmers can have substantial mouse TLR9-stimulatory activity and that methylation of cytosines in CpG motifs only does not always maximally reduce TLR9 stimulation, in the LNA context it can even have the opposite effect.
Effects of LNA modifications and cytosine methylation on immunostimulatory activity of a human TLR9 agonist
ODN2006 (also known as agatolimod or ODN7909) is a CpG-containing agonist for human TLR9, which was early reported to be immunostimulatory in primary human B cells 30 and has recently been approved by the Food and Drug administration (FDA) as an adjuvant for a new anthrax vaccine. Characterization of immunostimulatory effects mediated by human TLR9 activation is crucial during the development of ASOs as medicines. Therefore, in a complementary approach to our investigations regarding mouse TLR9, we made use of the ODN2006 sequence as a basis for investigation of the effects of LNA modifications and cytosine methylation on human TLR9 stimulation.
First, we tested different ODN2006 variants in human TLR9 reporter HEK293 cells (HEK-BlueTM hTLR9), which are based on the same reporter system as the murine version but overexpress the human TLR9 (hTLR9) sequence instead. ODN2006 led to an ∼15-fold increase in TLR9 activation over untreated cells and as observed with ODN1668, partial methylation of cytosines (ODN2006mCflanks) slightly reduced but did not abrogate TLR9 activation. In contrast, introducing three LNA modifications into each flank (ODN2006LNA), methylating all four cytosines in the sequence (ODN2006fmC), or a combination of both LNA modifications with full methylation (ODN2006fmCLNA) resulted in complete abrogation of TLR9 activation levels (Fig. 4A).
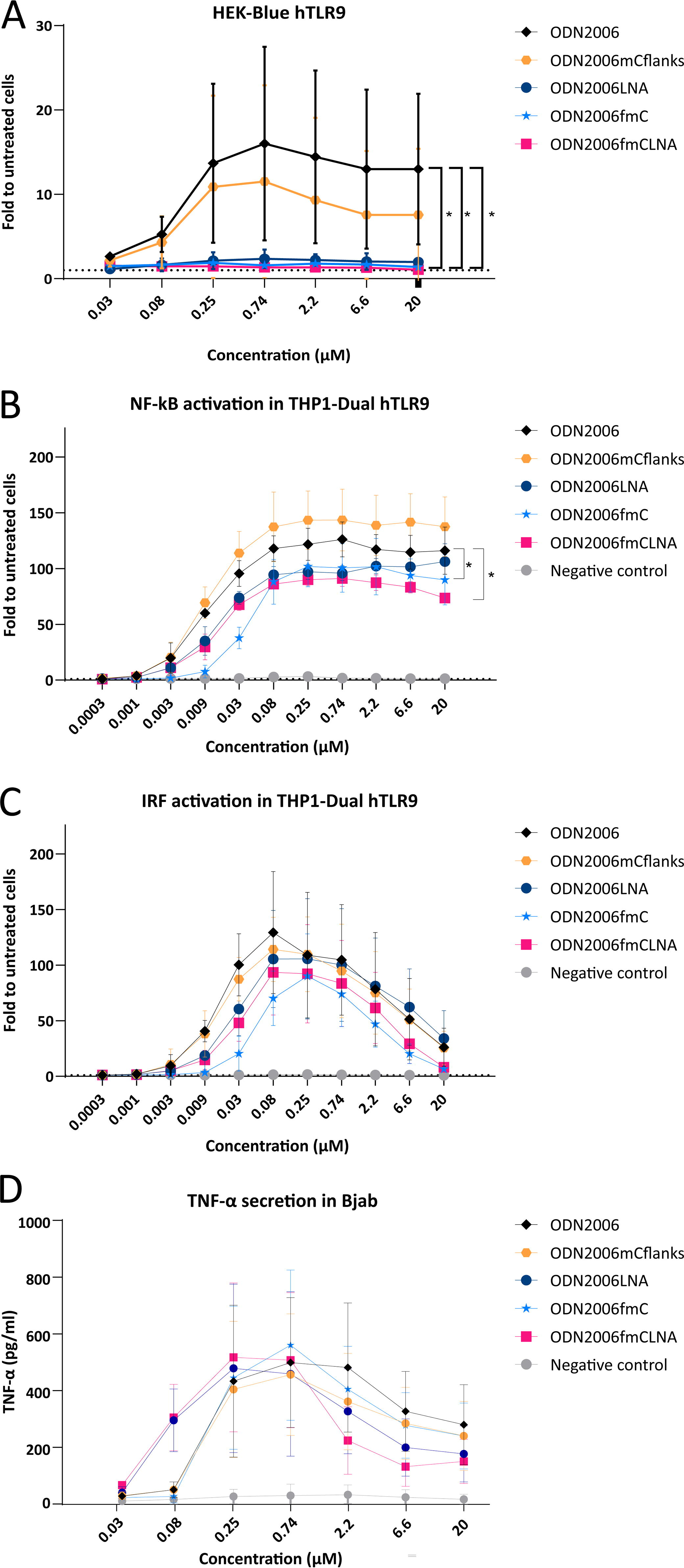
Effects of LNA modifications and cytosine methylation on immunostimulatory activity of ODN2006. HEK-BlueTM hTLR9 cells
To further study the effects of LNA and cytosine modifications on human TLR9 activation, we used two additional human cell lines. The first one was the THP1-DualTM hTLR9 cell line. These cells overexpress the human TLR9 gene, as well as two TLR9-regulated reporter genes, which allow the investigation of the TLR9-mediated activation of the NF-κB and IRF pathway. Treatment with ODN2006 for one day resulted in pronounced activation of both pathways under the tested conditions with maximal activity at a concentration of ∼0.08 µM, while the negative control oligonucleotide did not show any effect. Methylating Cs in the flanks (ODN2006mCflanks) did not ameliorate TLR9 induction. Introduction of LNA modifications (ODN2006LNA) slightly, but not significantly, reduced the activation of the NF-κB pathway, but had no clear effect on the IRF pathway. By contrast, methylation of all cytosines (ODN2006fmC), or a combination of both LNA modifications with full methylation (ODN2006fmCLNA), significantly reduced the activation of the NF-κB pathway and had minor nonsignificant effects on the ability of ODN2006 to stimulate the IRF pathway (Fig. 4B, C).
In addition, we investigated which effects the mouse TLR9 agonist ODN1668 and its variants have on THP1-DualTM hTLR9 reporter cells. Notably, ODN1668 robustly activated both the NF-κB and the IRF pathways in these cells. However, maximal stimulation was reached at higher concentrations and was lower compared with the human agonist ODN2006 (Supplementary Fig. S1C, D). As previously observed (Fig. 2), partial methylation of cytosines (ODN1668mCflanks, ODN1668mCpG) led only to a moderate reduction in TLR9 activation. Importantly, introducing LNA modifications in the flanks or combining the LNAs with partial or full cytosine methylation (ODN1668LNA, ODN1668mCpGLNA, ODN1668fmCLNA) led to abrogation of activity in both pathways downstream of TLR9 (Supplementary Fig. S1C, D).
The other cell line we used to explore the effects of the different ODN2006 variants was the Bjab cell line. These cells are derived from a Burkitt lymphoma and are a human B cell line, which has recently been described as a potentially useful predictor of innate immune responses in humans (via analysis of CCL22 mRNA expression, as well as TNF-α and IL-10 secretion). 31 Interestingly, although treatment of Bjab cells with ODN2006 resulted in high levels of TNF-α secretion, no significant differences in secretion were observed between ODN2006 and its different variants in these cells (Fig. 4D).
Stimulation of human TLR9 by LNA gapmers designed to suppress gene expression
To complement our investigations on the effects of ODN2006 variants in the abovementioned cellular assays, we selected a variety of LNA gapmers that were designed to suppress gene expression in humans. SECA166 is a 16-mer LNA gapmer targeting DNA damage-inducible transcript 3 (DDIT3) and its sequence contains an unmethylated cytosine in a CpG motif in the gap, as well as two additional unmethylated cytosine residues in the gap. SECA141 is a 19-mer, which also targets DDIT3, its sequence contains three unmethylated cytosine residues in the gap, but no CpG motif. SECA166 leads to ∼60% DDIT3 knockdown and SECA141 leads to ∼80% knockdown in EFO-21 cells (Supplementary Fig. S2A). SECA083 and SECA138 are LNA gapmers targeting the second isoform of retinoic acid receptor (RAR)-related orphan receptor gamma (RORγt) and contain two unmethylated cytosine residues in the gap. SECA083 and SECA138 are partially overlapping 19-mers, with no CpG motif, that share 15 of their 19 residues (Fig. 1). Both LNA gapmers lead to ∼50–60% knockdown in SUP-M2 and HDLM-2 cells (Supplementary Fig. S2B, C).
Importantly, none of the tested LNA gapmers led to TLR9 activation in HEK-BlueTM hTLR9 cells, neither after a single concentration treatment (Supplementary Fig. S3A, B) nor after treatment with a range of different concentrations (Fig. 5A), whereas the positive control ODN2006 did achieve an ∼20-fold activation compared with untreated cells. The LNA-modified version of ODN2006 did not lead to strong activation of TLR9 in those reporter cells either (Fig. 4A). NFkb or IRF activation in THP1 Dual cells without hTLR9 overexpression after treatment with SECA166 and SECA141 (Supplementary Fig. S3C-E). Thus, we wondered whether these cells might not be sensitive enough to measure TLR9 activation in the LNA context. Indeed, when tested in THP1-DualTM hTLR9 cells, SECA141 and SECA138 resulted in robust activation of both the NF-κB and the IRF pathways (Fig. 5B–E). Interestingly, SECA166 did not have any activating effects on TLR9 in any assay, although this oligonucleotide contains an unmethylated CpG sequence in the gap (Fig. 5B, C). SECA083 did not stimulate TLR9 either, although it shares most of its sequence with SECA138 (Fig. 5D, E). Moreover, cytosine methylation did not reduce nor increase the TLR9 activation of the tested ASOs in THP1-DualTM hTLR9 cells.
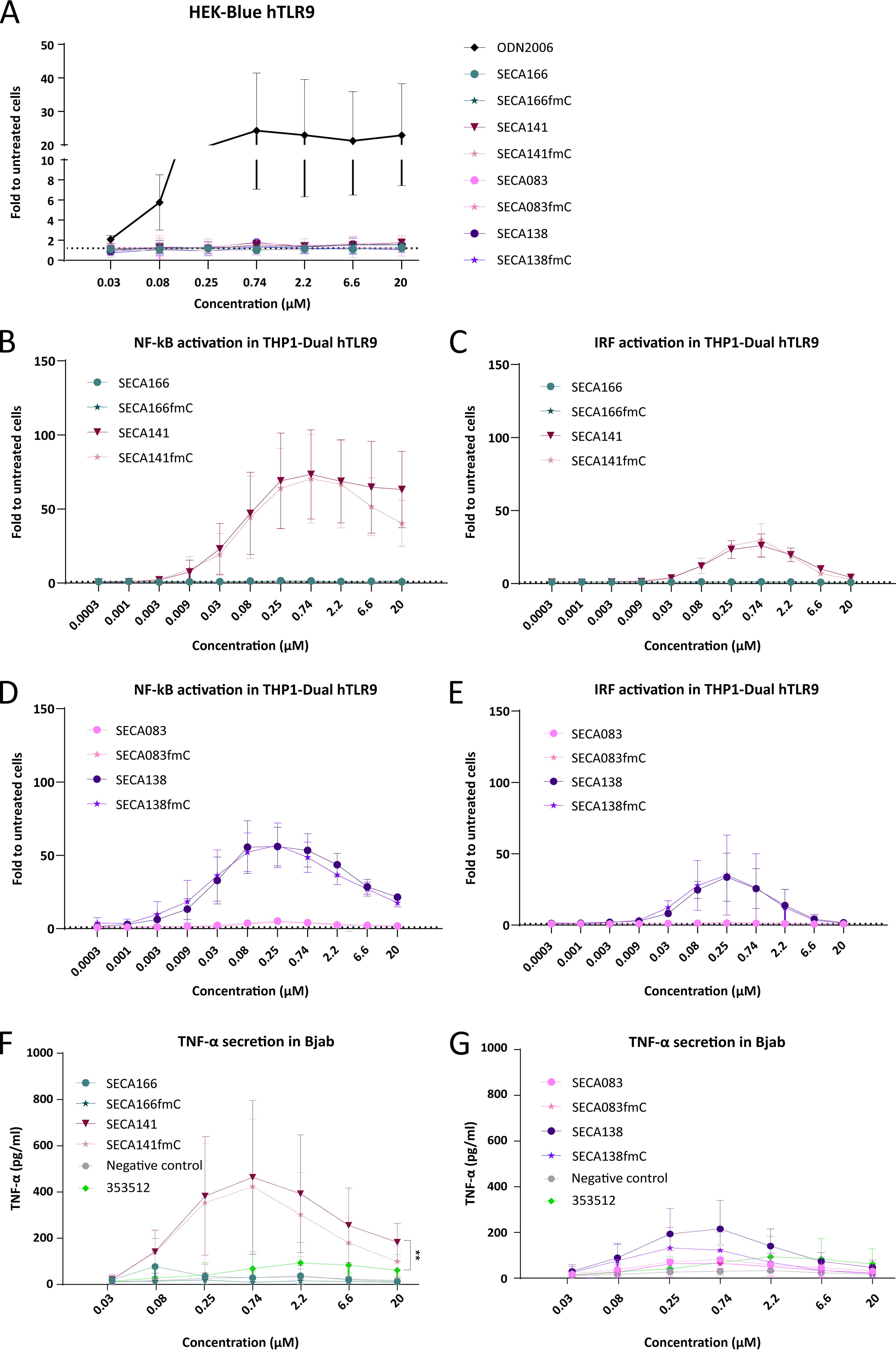
Stimulation of human TLR9 by gene-targeting LNA gapmers. HEK-BlueTM hTLR9 cells
Next, we tested the ASO effects on TNF-α secretion in Bjab cells. For these experiments, we included the 18-mer LNA gapmer negative control oligonucleotide, which was used in previous experiments, and as positive control, we included the 2′-O-methoxyethyl (MOE)-modified oligonucleotide 353512. This ASO caused inflammatory reactions in healthy human volunteers (clinical trial identifier NCT00734240) and was used as benchmark in the establishment of Bjab cells as a screening tool to detect immunostimulatory ASOs. 31 SECA141 resulted in significantly higher TNF-α release than 353512 in Bjab cells, whereas SECA166 had only a low effect on TNF-α release (Fig. 5F). SECA138 resulted in higher, but not significant TNF-α release than 353512, while SECA083 led to intermediate TNF-α levels between the negative control oligonucleotide and 353512 (Fig. 5G). In this assay, methylation of all cytosines slightly decreased TNF-α release both for SECA141 and SECA138 (Fig. 5F, G). Taken together, SECA141 and probably SECA138 (as well as their corresponding methylated versions) would be flagged as potentially immunostimulatory ASOs. Similar results have been observed regarding CCL22 mRNA expression in BJAB cells (Supplementary Figure 3F, 3G).
Overall, our results indicate that HEK-BlueTM hTLR9 cells might not be sufficiently sensitive to identify potentially immunostimulatory LNA-modified ASOs. The assay results in THP1-DualTM hTLR9 and Bjab cells show similar trends and might be better suited for in vitro screening and exclusion of TLR9 activating LNA-modified oligonucleotides.
Immunostimulatory effects in human peripheral blood mononuclear cells (hPBMCs)
Finally, we sought to confirm our findings in a more physiological cellular system and treated freshly isolated hPBMCs with the different ODN2006 variants and LNA-modified gapmers. In these experiments, we treated cells from three different blood donors and, based on a recent report on proinflammatory responses in hPBMCs, 18 we assessed IL-6 secretion as a measure for immunostimulation.
Strong donor variability in IL-6 responses after treatment with ASOs in hPBMCs has previously been reported 18 and here we also observed this phenomenon. hPBMC responses to treatment with ODN2006 and its variants showed remarkable variability between different donors. In two out of the three donors, the basic ODN2006 sequence led to the highest IL-6 secretion levels (Fig. 6A, B) and methylation of all Cs (ODN2006fmC) resulted in the strongest reduction in IL-6 secretion in two donors (Fig. 6A, C).
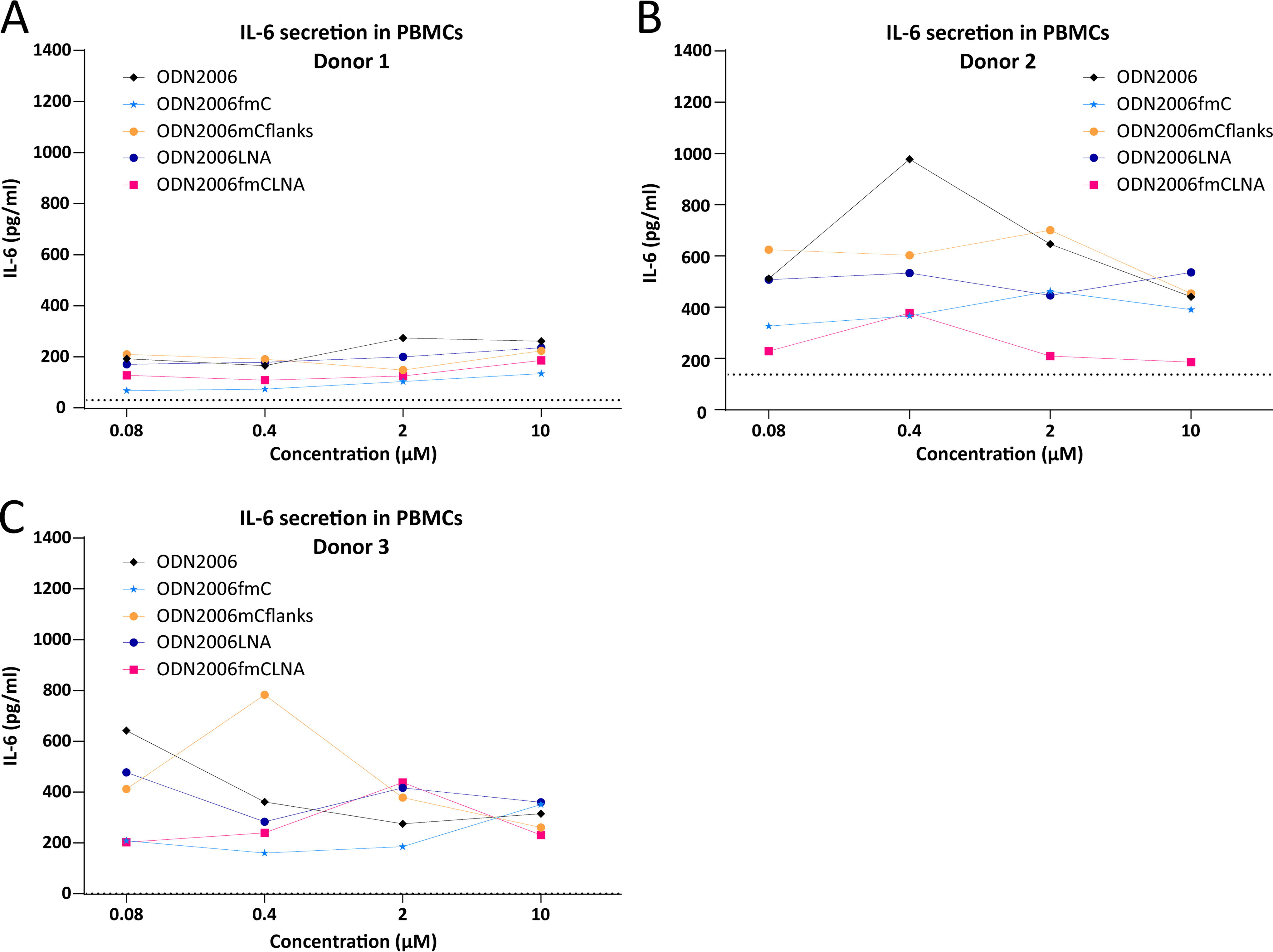
Immunostimulatory effects of ODN2006 variants in hPBMCs.
Next, we tested LNA-modified gapmers in hPBMCs. We again included the oligonucleotide 353512 as an immunostimulatory positive control. As second control we selected another MOE-modified oligonucleotide, 104838, which was reported to be less immunostimulatory than 353512 and recently used as benchmark in Bjab cells 31 and in hPBMCs. 18 Despite high variability across donors, SECA141 consistently resulted in higher IL-6 secretion than SECA166 and the two reference MOE-modified controls (Fig. 7A–C). These results are consistent with the high TLR9 activation observed in THP1-DualTM hTLR9 cells and high TNF-α secretion in Bjab cells observed after SECA141 treatment (Fig. 5). Conversely, results with SECA138 and SECA083 in hPBMCs were not as clear. SECA138 treatment resulted in higher IL-6 release than SECA083 only in two of the three donors (Fig. 7D–F), and the IL-6 values after treatment with SECA138 were only clearly higher in one donor compared with the two MOE-modified controls (Fig. 7F). SECA138 had shown lower stimulation in THP1-DualTM hTLR9 and Bjab cells than SECA141 (Fig. 5). These results highlight the complexity of determining the immunostimulatory potential of oligonucleotide gapmers based on in vitro cellular assays.
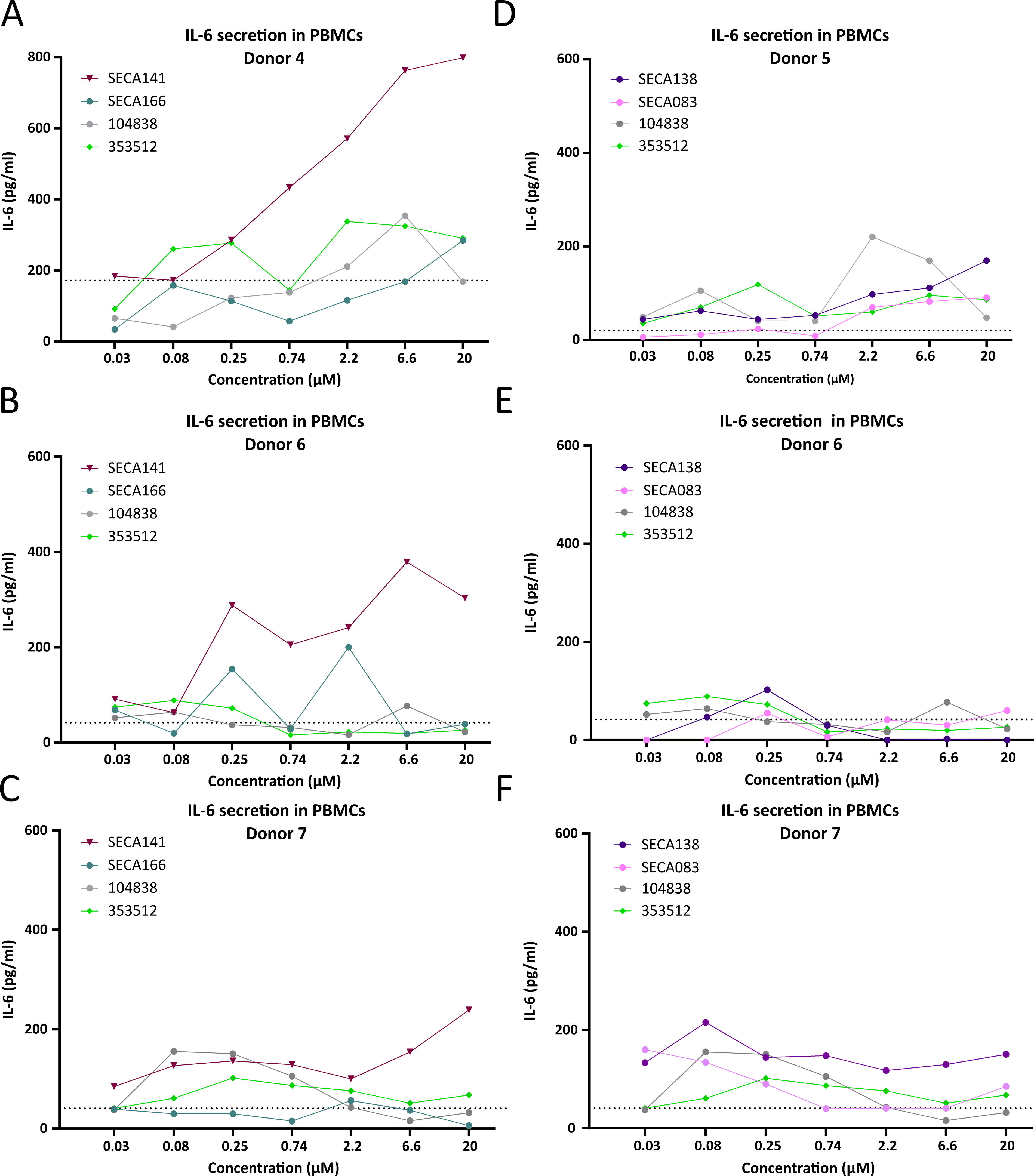
Immunostimulatory effects of LNA gapmers in hPBMCs. IL-6 secretion by hPBMCs after 48 h of treatment with a concentration range of DDIT3-targeting LNA-modified ASOs
Altogether, these results suggest that THP1-DualTM hTLR9 and Bjab cells may detect the immunostimulatory potential of human ASOs with higher sensitivity than the HEK-BlueTM hTLR9 assay, while showing more consistent results than more physiological and complex systems such as hPBMCs.
Discussion
Characterization of the immunostimulatory potential of oligonucleotides such as ASOs is critical for many different applications. For most therapeutic applications in humans, TLR9-mediated immune stimulation is an unwanted side effect and needs to be avoided, unless TLR9-stimulation is a desired effect, for example, to boost the immune system for treatment of cancer 7 or infectious diseases.8,9
Proof-of-concept investigation of human therapeutic ASOs using mouse surrogates, as well as target validation studies, and gene function assessment using ASOs is common practice. Underlying immunostimulation may affect the outcome of studies, especially in oncological research, for which tumor models in immune-competent mice are used. TLR9 activity can have a profound effect on tumor growth independently from target knockdown (Fig. 2C) and may result in misinterpretation of the results. Therefore, there is a strong need to not only characterize TLR9 activation by therapeutic ASOs, but also by antisense oligonucleotides that are being used for in vivo proof-of-concept studies in animal models.
To avoid undesired TLR9-mediated immunostimulation by ASOs, certain motifs, such as CpG motifs, can be filtered out during sequence selection and chemical modifications, such as cytosine methylation, can be introduced. However, even with these preventive measures, TLR9 activation cannot be completely avoided. 32 Bauer et al. reported that the optimal sequence motif for activation of mouse TLR9 is GACGTT, while for human TLR9 is GTCGTT. 12 Indeed, the mouse TLR9 agonist ODN1668 contains a GACGTT motif and the human TLR9 agonist ODN2006 contains a GTCGTT motif. However, none of the other ASOs tested in this study contained a GTCGTT or a GACGTT motif (Fig. 1). The findings described in this study highlight the need for identification and deselection of immunostimulatory oligonucleotides using adequate test systems before they are tested in vivo in animal models or even in patients.
While efforts to identify and characterize the effects of immunostimulatory ASOs have been recently described for second-generation chemistries,31,32 little is known about the immunostimulatory effects of third-generation LNA-modified ASOs. In this study, we extensively investigated the effects of introducing chemical modifications such as cytosine methylation and LNAs and moreover compared different assays to screen for mouse and human TLR9 activation.
Earlier investigations attributed TLR9 stimulation mostly to unmethylated CpG motifs. 15 It was, for example, published that for human TLR9 at least two CpG motifs in close vicinity were required. 13 ODN2006 having in total four CpG motifs fulfills this requirement. 30 Alternatively, for mouse TLR9 it was shown that one CpG motif at 4–6 nucleotides from the 5′ end was sufficient. 14 The CpG motif of the mouse TLR9 agonist ODN1668 does not follow this rule, since it is located further apart from the 5′ end. It contains, however, a TCC motif close to the CpG motif, which is reported to enhance activity. 14 SECA028 has both a CpG motif close to the 5′ end and a TCC motif in close vicinity to the CpG motif. SECA120, which had more potent mouse TLR9 activity compared with SECA028, has a CpG motif close to the 5′ end, however, no TCC motif (Fig. 1). It could therefore be assumed that avoidance of CpG motifs would be a viable option to reduce the risk of TLR9 activation. However, it has been postulated for second-generation ASOs 18 and confirmed here for LNA-modified ASOs that sequences without CpG motifs can activate TLR9 (SECA141 in Figs. 5, 7). Moreover, sequences without CpG motifs have been reported by others to have adverse immune effects in humans [clinical trial identifier NCT00734240]. 33 In addition, excluding all CpG containing sequences during the in silico selection of ASOs, without testing their immunostimulatory potential, might lead to the loss of potentially active and safe sequences.
Cytosine methylation is another measure to reduce TLR9 activation. 34 Our results demonstrate that methylation of only the cytosine within CpG motifs (or methylation of only some cytosines in the sequence) is mostly not sufficient to robustly reduce TLR9 activation (Fig. 2). In some cases, it can even have the opposite effect, particularly in combination with LNA modifications (Fig. 3A). Methylation of all cytosine residues leads to a more profound reduction of TLR9 activity. In some cases, it can completely abolish measurable TLR9 activity (Fig. 3), in other cases it has a moderate or no effect, especially on activation of human TLR9 (Figs. 4, 5). This disparity may be related to functional differences between mouse and human TLR9. Moreover, in combination with LNA modifications, the reduction of TLR9 stimulation by cytosine methylation can be less pronounced compared with the non-LNA-modified counterparts (Fig. 3B).
It was reported that LNA modifications reduced the immunostimulatory potential of known TLR9 agonists. 27 It needs to be mentioned here that Vollmer and colleagues used a mixed phosphodiester/PTO-backbone (only the gap was stabilized by PTO modifications). Therefore, it is unclear whether the findings can be extended to oligonucleotides with a full PTO backbone as they are commonly used for LNA-gapmers and in the present study. We show that LNA modifications can reduce TLR9 activity for some sequences (Fig. 3C) but result in higher TLR9 stimulation for other sequences (Fig. 3B), especially in the context of methylated cytosines. In addition, the observation that introducing LNA modifications in the mouse TLR9 agonist ODN1668 (Supplementary Fig. S1 C, D) had a stronger effect on the reduction of TLR9 activation in THP1-DualTM hTLR9 than introducing them in the human TLR9 agonist ODN2006 (Fig. 4B, C) is particularly interesting. These results suggest that the effect of LNA modifications on TLR9 activity might be sequence- and context-dependent, even in the case of CpG-containing oligonucleotides having a full PTO backbone. Since TLR9 responses are different between mouse and human, testing of human therapeutic drug candidates for their immunostimulatory potential in the mouse is not a reliable option. Mouse surrogates can nevertheless be investigated in vitro in systems expressing mTLR9. We tested a widely used mTLR9 agonist as well as several mouse surrogate ASOs, including ASOs with LNA modifications and methylated cytosine residues, in HEK-BlueTM mTLR9 and RAW DualTM reporter cells and confirmed a wide range in mTLR9 responses (Figs. 2, 3). The degree of TLR9 activation in HEK-BlueTM mTLR9 cells by different variants of the mouse TLR9 agonist ODN1668 mostly correlated with the antitumor activity in a syngeneic mouse tumor model. In contrast, testing of LNA-modified gapmers in the widely used HEK-BlueTM hTLR9 cells revealed poor sensitivity of these reporter cells for ASOs with such chemical modifications (Figs. 4A, 5A). Indeed, evaluating the same ASOs in other cell systems (THP-1 Dual hTLR9 and Bjab cells) uncovered important TLR9-mediated immunostimulatory responses (Fig. 5), which proved to be physiologically relevant when compared with clinical reference compounds in primary PBMCs (Fig. 7). It has been described that extracellular proteins such as HMGB1, S100A8, and HRG can enhance the TLR9 stimulatory capacity of ASOs in vitro. 32 It is likely that HEK293 cells secrete different proteins compared with THP-1 or Bjab cells. Extracellular proteins that increase the immunostimulatory potential of ASOs may be lacking in the supernatant of HEK293 cells, explaining their lower sensitivity to ASO-mediated TLR9 stimulation.
To conclude, our data show that it is possible to identify ASOs against different targets achieving potent target knockdown without TLR9 stimulation. Nevertheless, the immunostimulatory properties of an oligonucleotide can hardly be predicted and our results highlight the need to test ASOs in sensitive and relevant in vitro systems to select nonimmunostimulatory candidates for clinical testing.
Footnotes
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
