Abstract
Functional sequestration of microRNA 122 (miR-122) by treatment with an oligonucleotide complementary to the miRNA results in long-lasting suppression of hepatitis C virus (HCV) viremia in primates. However, the safety of the constitutive miR-122 silencing approach to HCV inhibition is unclear, since miR-122 can modulate the expression of many host genes. In this study, a regulation system capable of specifically inhibiting miR-122 activity only upon HCV infection was developed. To this end, an allosteric self-cleavable ribozyme capable of releasing antisense sequence to miR-122 only in the presence of HCV nonstructural protein 5B was developed using in vitro selection method. The activity of the reporter construct with miR-122 target sequences at its 3′ untranslated region and the expression of endogenous miR-122 target proteins were specifically stimulated through sequestration of miR-122 only in HCV replicon Huh-7 cells, but not in naïve Huh-7 cells, when transfected with expression vector encoding the specific allosteric ribozyme. These findings indicate that miR-122 function can be specifically inhibited by the allosteric ribozyme only in HCV-replicating cells. Importantly, HCV replicon replication was efficiently inhibited by the allosteric ribozyme. This ribozyme could be useful for the specific, safe, and efficacious anti-HCV modulation.
Introduction
MicroRNAs (miRNAs) are endogenous, small noncoding RNAs that repress cellular gene expression in a sequence-specific manner, generally through degradation or suppression of the translation of target mRNAs (Chekulaeva and Filipowicz, 2009). MicroRNA 122 (miR-122), which is the most abundant miRNA in the liver (Chang et al., 2004), promotes HCV replication in an unusual manner through the stimulation of accumulation or translation of HCV RNA by interacting with the 5’ untranslated region (UTR) of the viral genome (Jopling et al., 2005; Henke et al., 2008; Jangra et al., 2010).
Functional sequestration of miR-122 using antisense oligonucleotides against the miRNA effectively reduces the abundance of viral RNA, implicating miR-122 as a potential target for anti-HCV therapeutics. Inoculation of chronically HCV-infected chimpanzees with a locked nucleic acid (LNA)-modified oligonucleotide complementary to miR-122 causes a long-lasting suppression of HCV viremia (Lanford et al., 2010). However, miR-122 can also regulate the expression of a large number of genes involved in cellular physiological function including lipid metabolism (Krützfeldt et al., 2005; Esau et al., 2006; Elmén et al., 2008a). Therefore, side effects cannot be completely excluded if the miR-122 silencing occurs independently of HCV infection.
In the present study, a HCV-dependent miR-122 silencing system was developed. The most effective antisense sequences against miR-122 were isolated with regard to the inhibition of miR-122 activity in cells. Then, an allosteric self-cleavable hammerhead ribozyme capable of specifically releasing the most efficacious antisense sequence to miR-122 (antimir-122), and hence deactivating miR-122 activity only in the presence of HCV nonstructural protein 5B (NS5B) RNA replicase, was created. HCV-dependent regulation of miR-122 activity by the allosteric ribozyme was validated in a human hepatoma cell line stably conferring the HCV subgenomic replicon replication (Lohmann et al., 1999; Krieger et al., 2001). Importantly, anti-HCV effect by the allosteric ribozyme was measured in the HCV replicon cell line.
Materials and Methods
Cells
HCV genotype 1b subgenomic replicon construct, pFK-I389neo/NS3–3′/5.1, carrying two cell-culture adaptive mutations in NS3 and one in NS5A (Krieger et al., 2001) was used. HCV replicon RNA was synthesized by in vitro transcription with the AseI and ScaI-digested replicon plasmid as described (Hwang et al., 2004; Shin et al., 2006). Huh-7 human hepatoma cell line was purchased from American Type Culture Collection and was cultured in Dulbecco's modified Eagle medium (Invitrogen) with 10% fetal bovine serum. Huh-7 cells highly express miR-122 and thus support HCV replication (Jopling et al., 2005). Huh-7 cells were electrophoretically transfected with the HCV replicon RNA using conditions of 950 μF and 250 V with a gene pulser system (Bio-Rad) and were cultured in the presence of 500 μg/ml G418 (Invitrogen) to isolate a stable HCV replicon cell line.
Reporter construct for evaluation of anti-miRNA activity
A reporter construct whose expression can be down-regulated in miR-122 harboring naïve Huh-7 and HCV replicon Huh-7 cells was created by inserting three copies of target sequence against miR-122 (5′-CAAACACCATTGTCACACTCCA) into the 3′-UTR of a vector (pTK-RL) encoding Renilla luciferase gene under control of the herpes simplex virus thymidine kinase promoter. PCR was performed with primers (5′-CCGCTCGAGACAAACACCATTGTCACACTCCACCGGACAAACACCATTGTCACACTC and 5′-CACTAGTTGGAGTGTGACAATGGTGTTTGTCCGGTGGAGTGTGACAATGGTGTTTG), and the amplified DNA was cloned into the XhoI and SpeI site of pTK-RL. The final construct was designated pTK-RL-3×miR122T. Reporter constructs lacking a target sequence against any miRNA or with three copies of the target sequence against miR-17-5p (pTK-RL-3×miR-17-5p) were produced as controls. A cytomegalovirus promoter-driven firefly luciferase gene construct (pCMV-F.luci) was used as an internal control.
Construction of expression vectors encoding anti-miRNA
To construct expression vectors encoding antisense sequence targeting diverse regions of miR-122 or miR-17-5p, sense and antisense oligonucleotides were synthesized (Bioneer), hybridized, and cloned into the SalI and XbaI site of U6+1 snRNA promoter or 7SL RNA promoter-driven expression cassettes (Paul et al., 2003). Antimir-122: (1) anti-Dicer star (targeting the Dicer processing site and part of the miRNA passenger strand); 5′-TCGACTGATAATGGCGTTTGATAGTTTAGAT (sense), 5′-CTAGATCTAAACTATCAAACGCCATTATCAG (antisense) (2) anti-miRNA Dicer (targeting the Dicer processing site and part of the mature miRNA); 5′-TCGACTTTGATAGTTTAGACACAAACACCATT (sense), 5′-CTAGAATGGTGTTTGTGTCTAAACTATCAAAG (antisense) (3) anti-Drosha star (targeting the Drosha processing site and part of the miRNA passenger strand); 5′-TCGACCCTAGCAGTAGCTATTTAGTGTGAT (sense), 5′-CTAGATCACACTAAATAGCTACTGCTAGGG (antisense) (4) anti-miRNA Drosha (targeting the Drosha processing site and part of the mature miRNA); 5′-TCGACTGTCACACTCCACAGCTCTGCTAT (sense), 5′- CTAGATAGCAGAGCTGTGGAGTGTGACAG (antisense) (5) anti-mature miRNA (targeting the mature miRNA); 5′-TCGACACAAACACCATTGTCACACTCCAT (sense), 5′-CTAGATGGAGTGTGACAATGGTGTTTGTG (antisense). Antimir-17-5p; (4) anti-miRNA Drosha; 5′-TCGACGTAAGCACTTTGACATTATTCTGAT (sense), 5′-CTAGATCAGAATAATGTCAAAGTGCTTACG (antisense) (5) anti-mature miRNA; 5′-TCGACACTACCTGCACTGTAAGCACTTTGT (sense), 5′-CTAGACAAAGTGCTTACAGTGCAGGTAGTG (antisense).
Selection of NS5B-dependent allosteric ribozyme
To construct the NS5B-dependent allosteric hammerhead ribozyme, in vitro selection was performed using a ribozyme library consisting of aptamer against NS5B of HCV type 1b (unpublished data), ribozyme catalytic core region, and 10-mer randomized communication module sequence as described previously (Kertsburg and Soukup, 2002; Roth and Breaker, 2004)
Construction of NS5B-dependent anti-miRNA releasing allosteric ribozyme (NDAR
)
To construct NS5B-dependent antisense sequence to mature miR-17-5p (antimir-17-5p NDAR) or to miR-122 releasing allosteric ribozyme (antimir-122 NDAR), PCR was performed using selected allosteric ribozyme clone No. 4 cDNA as template with following primer sets: antimir-17-5p NDAR: 5′-ACGCGTCGACGGGAGTGCAGCTGATGAGCCTCTCGCG-3′, 5′-CCCAAATTATTCGGTGTGTGCGCCCCTCACAATATTGCGCGAGAGGCTCATCAGC-3′ and 5′-GCTCTAGACAAAGTGCTTACAGTGCAGGTAGTTTGGTTTCCCAAATTATTCGGTGTG-3′; antimir-122 NDAR: 5′-ACGCGTCGACGGGAATGGTGCTGATGAGCCTCTCGCG-3′, 5′-TCCCAAATAATTCGGTGTGTGCGCCCCTCACAATATTGCGCGAGAGGCTCATCAG- 3′ and 5′-TGGAGTGTGACAATGGTGTTTGTTTGGTTTCCCAAATAATTCGGTGTG-3′. Each amplified DNA was digested with SalI and XbaI, and cloned into a pGEM T-easy vector (Promega). Each NDAR was generated by in vitro transcription using the cloned vector as the template. To construct the expression vector for each NDAR in cells, each vector containing NDAR cDNA was digested with SalI and XbaI and cloned into the 7SL RNA promoter-driven expression cassettes.
In vitro self-cleavage assay of allosteric ribozyme
The in vitro transcribed NDAR RNA was dephosphorylated with calf intestine alkaline phosphatase (Ambion) and 5′ end-labeled with [γ-32P]ATP (3,000 Ci/mmol, Amersham-Pharmacia Biotech) and T4 polynucleotide kinase (Ambion). The end-labeled NDAR RNA (20 fmol) was incubated with 3.2 pmol of BSA or HCV NS5B at 37°C for 1 hour in a reaction buffer (30 mM Tris-HCl, pH 7.5, 150 mM NaCl, 5 mM MgCl2, 2 mM dithiothreitol), and the reacted products were separated on a 12% polyacrylamide gel with 7 M urea. To identify self-cleavage sites by the ribozymes, a sequence ladder was generated and loaded on the gel as a size marker as follows. Alkaline hydrolysis was performed at 95°C for 15 minutes in a 20 μL reaction volume that contained 50 mM NaHCO3 (pH 9.0), 1 mM EDTA, 0.25 mg/mL tRNA, and 54 fmol of NDAR RNA. The cleaved RNA fragments were denatured at 80°C for 5 minutes and analyzed on the denaturing polyacrylamide gel. NDAR RNA (54 fmol) was also digested with RNase T1 (USB) at 37°C for 10 minutes in a reaction mixture (20 μL) containing 20 mM sodium citrate (pH 5.0), 7 M urea, 1 mM EDTA, 0.25 mg/mL tRNA, and 1 unit of RNase T1. The nuclease-treated reactions were stopped by the addition of 10 μL of RNA loading dye (10 mL formamide, 10 mM EDTA, 0.01% bromophenol blue, 0.01% xylene cyanol) and analyzed on the denaturing polyacrylamide gel.
To confirm the exact self-cleavage sites by the ribozymes, 3′-rapid amplification of cDNA ends (RACE) PCR of the cleaved products was performed. Self-cleaved RNA (30 pmol) was isolated from the polyacrylamide gel with urea and poly(A)-tailed using poly(A) polymerase (5 units, NEB) at 37°C for 30 minutes. Poly(A) RNA was reverse transcribed with 3′-RACE primer (5′-AAGCAGTGGTATCAACGCAGAGTAC(T)30VN, where N represents nucleotide equimolar incorporation of A, T, G, C and V denotes A, G, or C). The resulting cDNA was then amplified with a 5′ primer specific for the 5′ end of the ribozyme (5′-TAATACGACTCACTATAGGGAATGGTGCTG for antimir-122 NDAR, 5′-TAATACGACTCACTATAGGGAGTGCAGCTG for antimir-17-5p NDAR) and nested 3′ primer (5′-AAGCAGTGGTATCAACGCAGAGT) at 98°C for 30 seconds, at 58°C for 30 seconds, and at 72°C for 30 seconds for 25 cycles. The amplified products were cloned and sequenced.
Anti-miRNA reporter assay
For evaluation of anti-miRNA activity by antimir expression vectors, pCMV-F.luci (25 ng) and pTK-RL-3×miR122T or reporter construct lacking miR-122 binding sites [R.Luci(no target sites)] (100 ng) were cotransfected with 7SL- or U6+1-based anti-miRNA expression vector (400 ng) into 1×105 cells using LT1 transfection agent (Mirus Bio). After 24 hours of transfection, the cells were harvested and lysed, and reporter gene expression activities were determined by measuring relative light units using a TD-20/20 luminometer (Turner Designs Instrument) and dual-luciferase reporter assay system (Promega). For analysis of selective anti-miRNA activity by allosteric ribozyme expression vectors, after 48 hours of cotransfection of pCMV-F.luci (25 ng) and pTK-RL-3×miR122T (200 ng) with antimir NDAR expression vector (400 ng), reporter activities were assessed.
Real-time PCR
HCV replicon cells in a 12-well plate (each well containing 2×105 cells) were transfected with 7SL-driven NDAR construct (1 μg) using TransIT LT1 transfection reagent (Mirus Bio) according to the manufacturer's protocol. At 30 hours after transfection, total RNA was isolated and reverse transcribed with 3′-primer specific for the negative strand of HCV cDNA (5′-CGTAACACCAACGGGCGCGCCATG) or random primer for 18S cDNA. The resultant cDNAs were subsequently amplified with 2× real-time PCR (RT-PCR) premix (Solgent Co.) with forward (5′-CGTAACACCAACGGGCGCGCCATG) and reverse primer (5′-CTCGTCCTGCAGTTCATTCAGGGC) specific for the neomycin resistant marker gene of the HCV replicon construct. RT-PCR was performed using the Rotor-Gene (Corbett) and SYBR Green PCR Core Reagents (PE Biosystems), according to the manufacturer's protocol. The conditions for the PCRs were 95°C for 30 seconds, 58°C for 30 seconds, and 72°C for 30 seconds for 40 cycles. The threshold levels obtained from the HCV were adjusted to the threshold levels found in the 18S PCR reaction (forward primer; 5′-GTAACCCGTTGAACCCCATT, reverse primer; 5′-CCATCCAATCGGTAGTAGCG) to correct for minor variation in cDNA loading. Quantification of miR-17-5p and miR-122 was performed using TaqMan MicroRNA kit (miR-17-5p: 002308, miR-122: 002245; Applied Biosystem) according to the manufacturer's instructions. The 18S RNA was used as an internal control.
Immunoblot analysis
HCV replicon cells in 60 mm dishes (each containing 8×105 cells) were transfected with 7SL-driven NDAR construct (1 μg) using TransIT LT1 transfection reagent (Mirus Bio) according to the manufacturer's protocol. Cells were lysed 30 hours after transfection in a buffer consisting of 50 mM Tris-HCl (pH 8.0), 150 mM NaCl, 0.2% sodium azide, 0.1% SDS, 0.1% NP-40, and 0.5% sodium deoxycholate. The total proteins (50 μg) in the cell lysates were loaded into SDS-polyacrylamide gel and transferred on PVDF membrane (Bio-Rad) using electrophoresis in a condition of 300 mA for 3 hours at 4°C. The membrane was blocked with 5% (v/v) skim milk and immunoblotting was carried out using the following primary antibodies: mouse monoclonal anti-NS5A (#MAB8694; Millipore), goat polyclonal anti-Aldolase A (#sc-12059; Santa Cruz Biotechnology), rabbit polyclonal anti-Cyclin G1 (#C-18; Santa Cruz Biotechnology), and mouse anti-α-Tubulin (#DM1a; Santa Cruz Biotechnology). The following secondary antibodies were used: goat anti-mouse IgG-HRP (#sc-2005; Santa Cruz Biotechnology), mouse anti-goat IgG-HRP (#sc-23521; Santa Cruz Biotechnology), and goat anti-rabbit IgG-HRP (#sc-2004; Santa Cruz Biotechnology). The protein bands were visualized by an enhanced chemiluminescence protocol (Amersham-Pharmacia Biotech).
Results
Antimir-122 expressing vectors inhibit miR-122 activity in cells independent of HCV replication
To identify the most appropriate targeting region in miR-122, we designed diverse antisense sequences targeting miRNA precursor regions as well as mature miRNA (Fig. 1A), constructed U6+1 or the 7SL promoter-driven expression vectors for the anti-miRNAs, and compared their anti-miRNA activities using reporter genes with corresponding miRNA target sites at their 3′-UTR. The reporter gene construct for the assessment of miRNA activity regulated by each antimir expression vector is depicted in Fig. 1B. Reporter activity with constructs harboring each miRNA target site was efficiently reduced in Huh-7 cells as well as HCV replicon Huh-7 cells, when compared with reporter construct devoid of miRNA target sites (Supplementary Fig. S1; Supplementary Data are available online at www.liebertonline.com/nat).
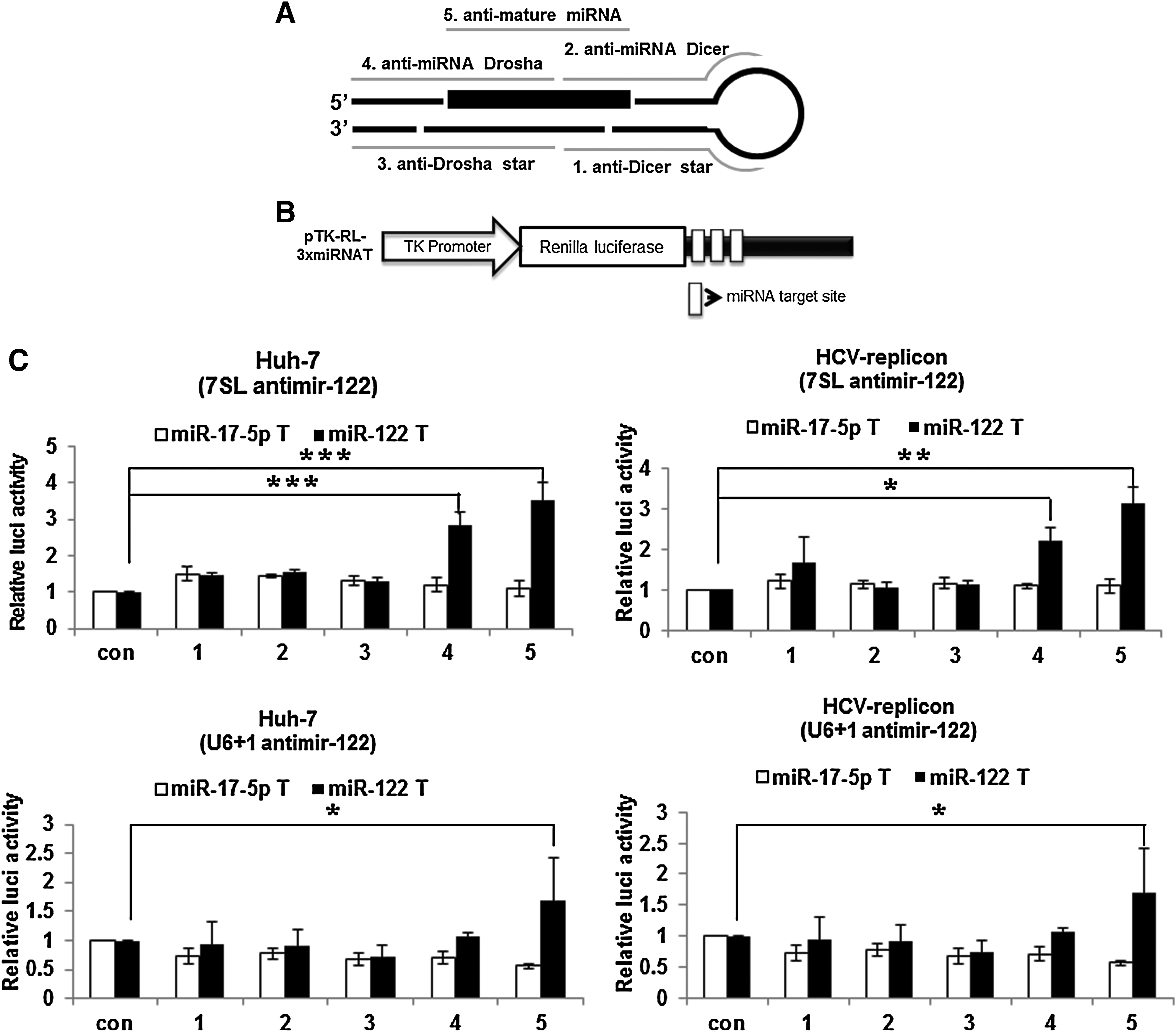
Effects of anti-microRNA (miRNA) expression vectors on miRNA activity in Huh-7 and hepatitis C virus (HCV) replicon cells.
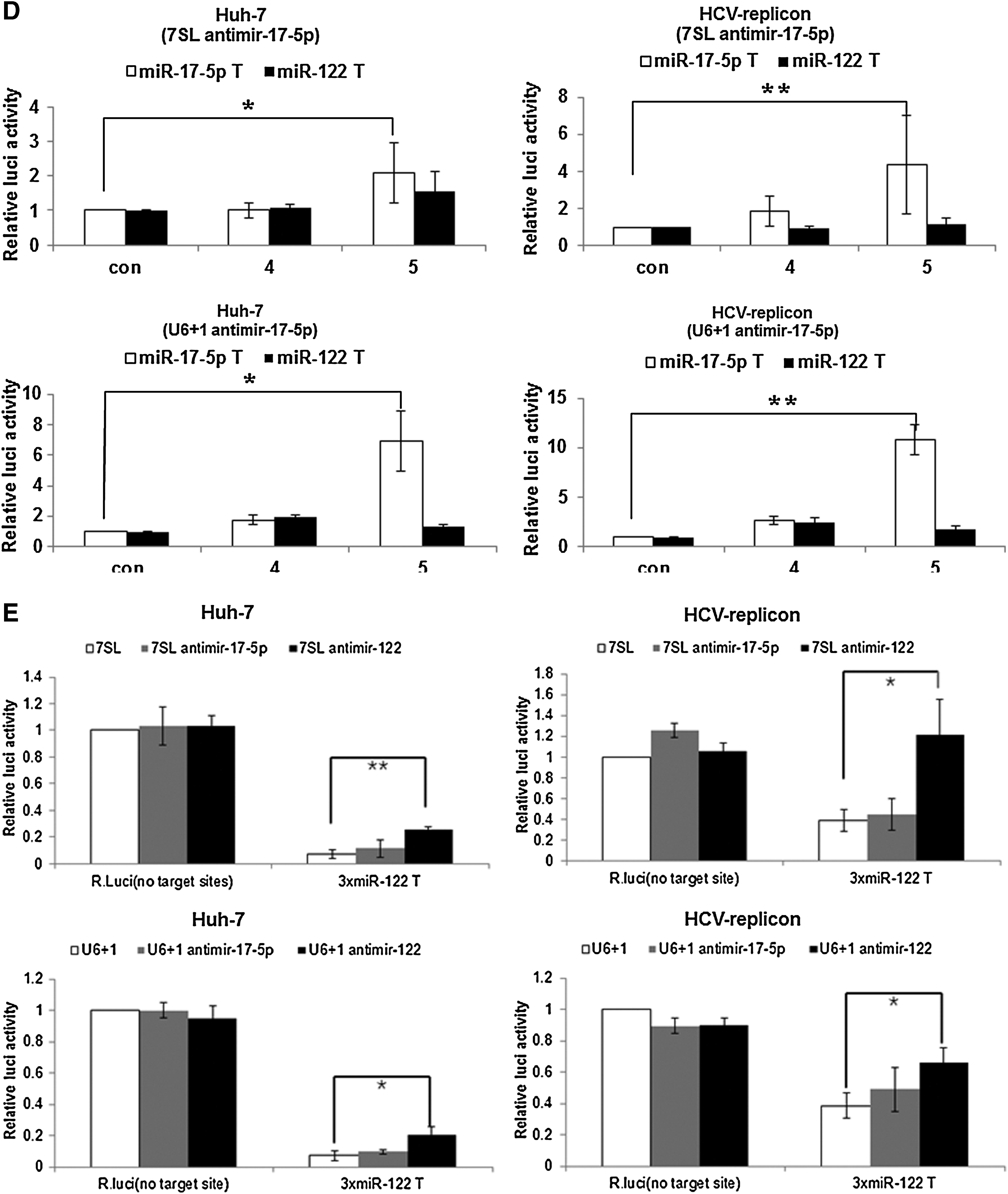
Transfection with expression vectors for either anti-mature miRNA or anti-miRNA Drosha derepressed reporter activity in both Huh-7 and HCV replicon cells (Fig. 1C). The derepression levels of the reporter activity through the expression of the 7SL promoter-driven antimir-122 were two-fold higher than those by U6+1 promoter-driven antimir-122. However, no derepression of reporter activity was evident in constructs harboring miR-17-5p target sites by transfection of any of the antimir-122 expression vectors, indicating target specificity of antimir-122. Taken together, the intracellular expression of an antisense sequence targeting full or seed sequences of mature miR-122 could efficiently and specifically inhibit miR-122 activity. Correspondingly, treatment with chemically synthesized antagomir targeting the full or seed region of mature miR-122 inhibited miR-122 activity in cells (Supplementary Fig. S2). A similar analysis was performed for miR-17-5p, another miRNA expressed in Huh-7 cells (Fig. 1D). Intracellular expression of an antisense sequence targeting the full region of mature miR-17-5p efficiently and specifically inhibited miR-17-5p activity. Specific anti-miR-17-5p activity was shown in both naïve Huh-7 and HCV replicon Huh-7 cells, and was much higher when antimir was expressed from U6+1 promoter than from the 7SL promoter.
Intracellular expression of the anti-mature miR-122 resulted in derepression of reporter gene expression more highly in HCV replicon cells than in Huh-7 cells (Fig. 1E). Of note, miR-122 inhibition through expression of anti-mature miR-122 with 7SL promoter in HCV replicon cells was most effective, resulting in derepression at a level of reporter gene expression comparable to reporter gene lacking miR-122 binding sites in the cells.
Overall, although the relative fold derepression of the reporter activity indicating effectiveness of anti-miRNA activities is different between expression systems depending on the antimir and miRNA target/reporter gene, the effect of different antimir sequences is comparable. Conclusively, intracellular expression of antisense sequence targeting the full sequence of mature miRNA was most effective with regard to miRNA silencing, however, independent of HCV infection.
Strategy of HCV-specific miR-122 inhibition with HCV NS5B-dependent antimir-122 releasing allosteric ribozyme (antimir-122 NDAR)
A modular rational design was used to create artificial ribozymes that could be activated by the binding of HCV-specific ligand, and so could deactivate miR-122 specifically in HCV-replicating cells. These allosteric ribozymes consisted of 2 independent structural domains (the first was a self-cleavable hammerhead ribozyme and the second was an aptamer for HCV NS5B that is a HCV-encoding regulatory protein critical for HCV replication (Moradpour et al., 2002) and a communication module sequence that connected the domains (Fig. 2A). The conformational changes upon binding of the aptamer with the ligand can trigger the kinetic activation of the adjoining catalytic domain mediated by the communication module sequence (Soukup and Breaker, 1999), resulting in the specific release of an antisense sequence against miR-122 only in the presence of HCV NS5B (Fig. 2A).
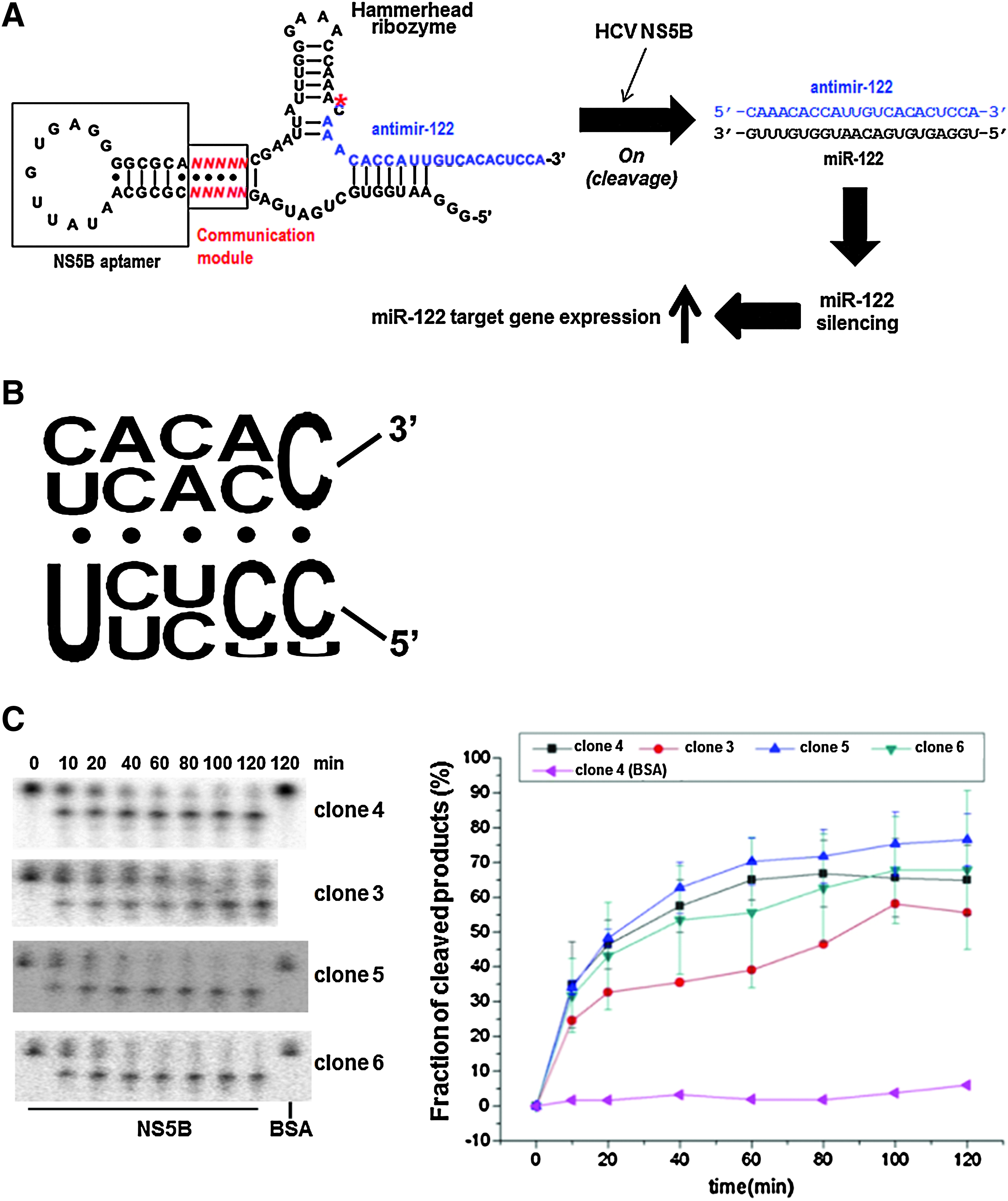
Selection of HCV nonstructural protein 5B (NS5B)-dependent allosteric hammerhead ribozyme.
Isolation of HCV NS5B-dependent self-cleavable allosteric hammerhead ribozyme
In vitro selection was used to identify effective communication module sequences of allosteric ribozymes that function as precise molecular switches to specifically release antimir-122 and turn off miR-122 activity only in HCV-infected cells. To this end, we generated a population of more than 106 variant RNAs consisting of HCV NS5B-binding aptamer (Lee et al., unpublished) and hammerhead ribozyme domains connected by a random-sequence communication module. The communication module resides in the stem II motif of the hammerhead ribozyme, which is a critical structural element for the ribozyme activity (Tuschl and Eckstein, 1993; Long and Uhlenbeck, 1994). RNA pools isolated after 4 rounds of selection displayed HCV NS5B-dependent self-cleavage, but not in the presence of buffer only or BSA (Supplementary Fig. 3). This allosteric cleavage was due to ribozyme activity, based on the observation that no cleavage reaction was shown by inactive ribozyme version even in the presence of HCV NS5B (Supplementary Fig. S3).
Ultimately, 25 different ribozyme clones were isolated and sequenced (Supplementary Fig. S4). Twenty-two different RNAs were selected with pyrimidine-rich communication module sequence (Fig. 2B), some of which were found in multiple clones. Several ribozyme clones were then analyzed for their time-dependent self-cleavage reaction in the presence or absence of HCV NS5B to determine NS5B specificity and efficacy of the selected allosteric ribozymes (Fig. 2C). The presence of BSA did not induce or suppress the self-cleavage activity of the ribozyme, even after an extended incubation. On the contrary, all tested allosteric ribozymes were efficiently inducible in their self-cleavage activity in the presence of HCV NS5B with similar catalytic efficacy with a rate constant (kmin) of approximately 0.15 min−1. Thus, ribozymes whose self-cleavage activity can be allosterically inducible by HCV NS5B were isolated using the presently designed in vitro selection method. The mechanism of the allosteric function was not analyzed. A slip-structure mechanism is one of possibility for the allosteric conformational changes in the ribozyme upon aptamer binding (Soukup and Breaker, 1999; Kertsburg A and Soukup, 2002).
Activity of antimir-releasing self-cleavable allosteric ribozymes depends on HCV NS5B
To construct NS5B-dependent antimir-releasing allosteric ribozyme (antimir NDAR) that can switch off targeted miRNA activity only in the presence of HCV NS5B, the sequence of the released part of the selected ribozyme was replaced with the most efficacious antisense sequence against miRNA (antisense against mature miRNA) (Fig. 3). A self-cleavage reaction was clearly manifested with the antimir-122 NDAR (Fig. 3A) and antimir-17-5p NDAR (Fig. 3B) in the presence of HCV NS5B, but not in the presence of BSA, indicating NS5B-dependency for antimir NDAR activity in vitro. Analysis of the size of the cleaved RNA products in the sequencing gel suggested that self-cleavage reaction occurred at sites around the predicted cleavage regions in the NDAR, indicating that antisense sequence targeting mature miRNA can be released by antimir NDAR. Sequence analyses of the 3′-RACE amplified cDNA products of the self-cleaved RNAs also showed that the antimir NDARs were self cleaved at the sites around sequences jointed with antisense sequence against each targeted mature miRNA (data not shown). Both NDARs were heterogeneously self-cleaved at approximately 2 major sites, but all of the cleaved and released products covered most of the full sequence of targeted mature miRNA, including seed region. It is unexpected that antimir NDARs yielded multiple cleavage products. This suggests some considerable and unanticipated flexibility in the stem I and/or stem III of the ribozymes upon binding to HCV NS5B. Extensive analysis of conformational features of the ribozymes in the presence or absence of HCV NS5B will be needed in order to elucidate what causes this phenomenon.
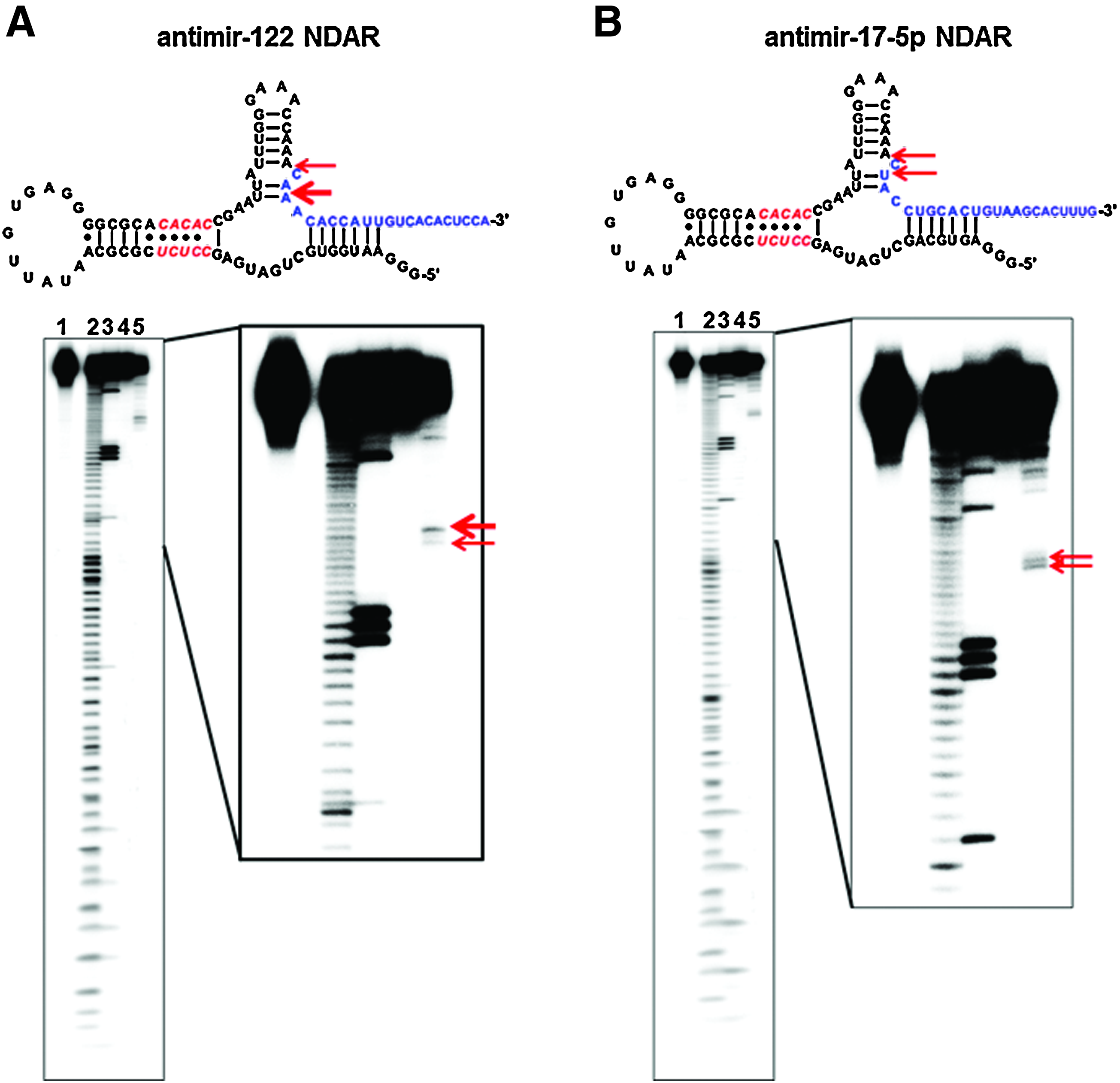
HCV NS5B-dependent in vitro self-cleavage reaction of antimir NDAR. (Upper) Scheme of antimir-122 NDAR
Allosteric ribozyme inhibits miR-122 activity in cells, which is dependent on HCV replication
Once the in vitro HCV NS5B-selectivity of self-cleavable antimir-122 NDAR had been determined, the next experiment validated the selectivity of antimir-122 NDAR in cells in terms of selective inhibition of miR-122 activity in HCV-replicating cells (Fig. 4). To this end, a 7SL-based expression vector encoding antimir-122 NDAR was constructed and cotransfected into naïve Huh-7 or HCV replicon Huh-7 cells with a reporter construct harboring miR-122 target sites (Fig. 4A). Consistent with the results shown in Fig. 1C, expression of the antisense sequence against mature miR-122 inhibited miRNA activity and resulted in derepression of the reporter activity in Huh-7 cells, regardless of HCV replication. In sharp contrast, expression of antimir-122 NDAR highly derepressed reporter gene activity in HCV replicon Huh-7 cells, but not in naïve Huh-7 cells. Of note, the expression of antimir-122 NDAR selectively increased reporter gene activity in HCV replicon cells with efficiency comparable to that associated with the expression of antimir-122. Several variants of antimir-122 NDAR were constructed such as to have extended base pairing at stem I of the ribozyme to exclude the possibility of nonspecific binding of antimir sequence in the NDAR with the target miRNA (Supplementary Fig. S5). Antimir-122 NDAR variant 4 (V4) harbored HCV NS5B specificity in vitro with regard to its HCV NS5B-dependent self-cleavage activity and in cells also in terms of selective derepression of reporter gene activity, but with less inducibility of antimir-122 activity compared with unmodified antimir-122 NDAR (Fig. 4A).
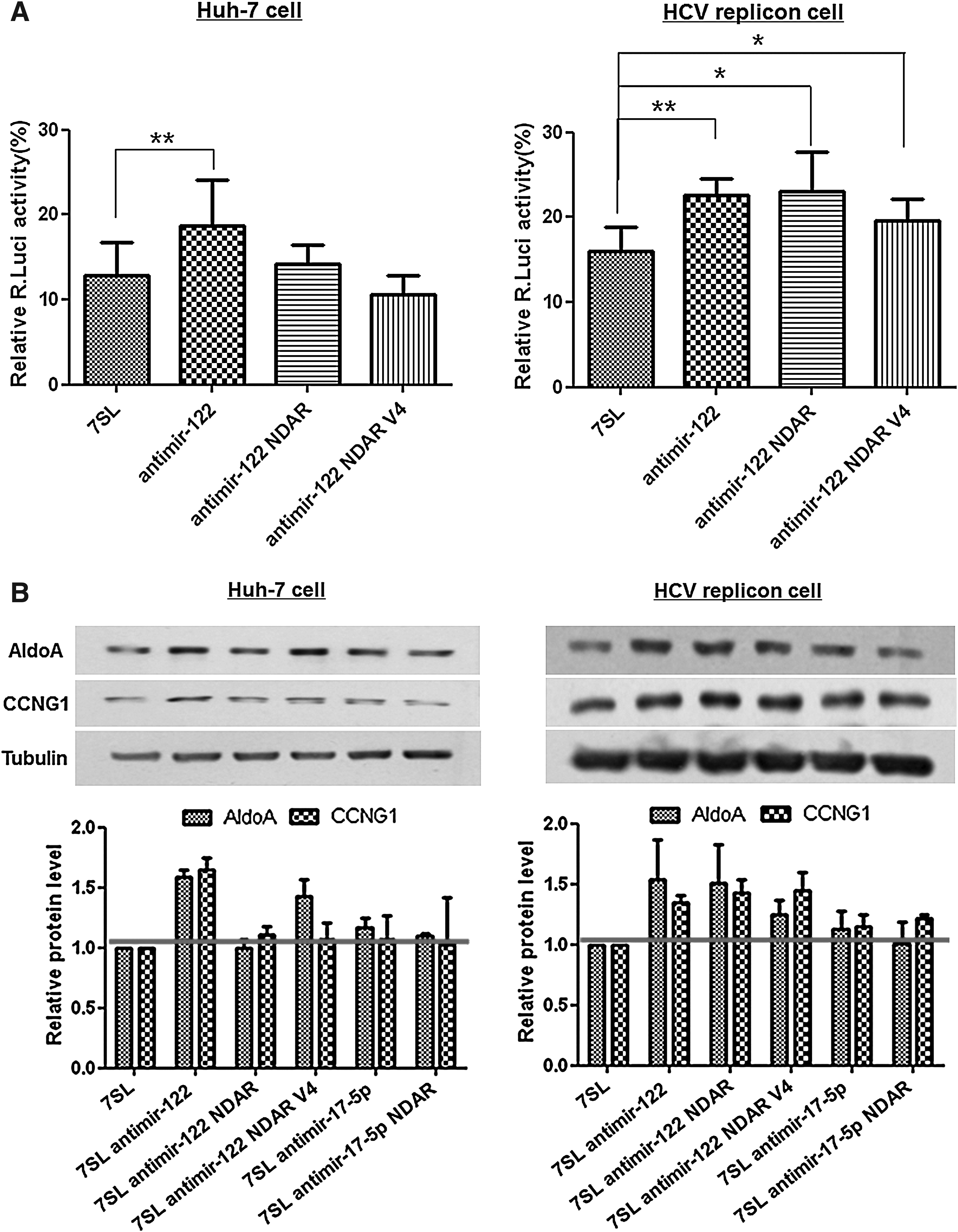
HCV-selectivity of antimir-122 NDAR in cells.
HCV-selective inhibition of miR-122 activity by antimir-122 NDAR was confirmed by western analysis of endogenous miR-122 target proteins, Cyclin G1 (Gramantieri et al., 2007) and Aldolase A (Elmén et al., 2008b) (Fig. 4B). Expression of antimir-122 derepressed the expression of both miR-122 target proteins, independent of HCV replication. Noticeably, consistent with Fig. 4A, expression of antimir-122 NDAR increased the expression of the cellular miR-122 target proteins selectively in HCV replicon Huh-7 cells. However, expression of the miR-122 target proteins was not affected by expression of either antimir-17-5p or antimir-17-5p NDAR, indicating target miRNA specificity of the corresponding antimir. These findings suggest that the intracellular expression of the mature miRNA-targeting antimir-122 releasing allosteric ribozyme specifically activates ribozyme activity and hence releases an antisense sequence against the miRNA in the presence of HCV NS5B, resulting in the efficient and specific inhibition of miR-122 activity only in HCV-replicating cells.
Allosteric ribozyme reduces miRNA amplification by miRNA-specific quantitative RT-PCR selectively in HCV-replicon cells
In order to figure out how antimir-122 NDAR inhibits miR-122 function selectively in HCV-replicating cells, the level of miRNA in the cells was assessed and compared after transfection of expression vector encoding the NDAR. The transcript levels of mature miRNA or pre-miRNA were only marginally affected by transfection of any of the antimir expression vectors in northern blot analysis (data not shown). In contrast, miRNA-specific quantitative RT-PCR analyses (qRT-PCR) showed that expression of the antisense sequence against mature miRNA specifically reduced targeted miRNA level, regardless of HCV replication (Fig. 5A and B). This discrepancy is probably due to interference of qRT-PCR by antimir through prevention of the binding of miRNA-specific looped RT primer and following suppression of the reverse transcription reaction. Scherr et al. (2007) also showed reduction of miRNA amplification by qRT-PCR with lentivirus-mediated antagomir expression. Much higher inhibition of miR-122 amplification by expression of the antisense sequence against mature miRNA was shown in HCV replicon cells than Huh-7 cells. This is partly because of the much lower amount of steady-state miR-122 in HCV replicon cells (Supplementary Fig. S6).
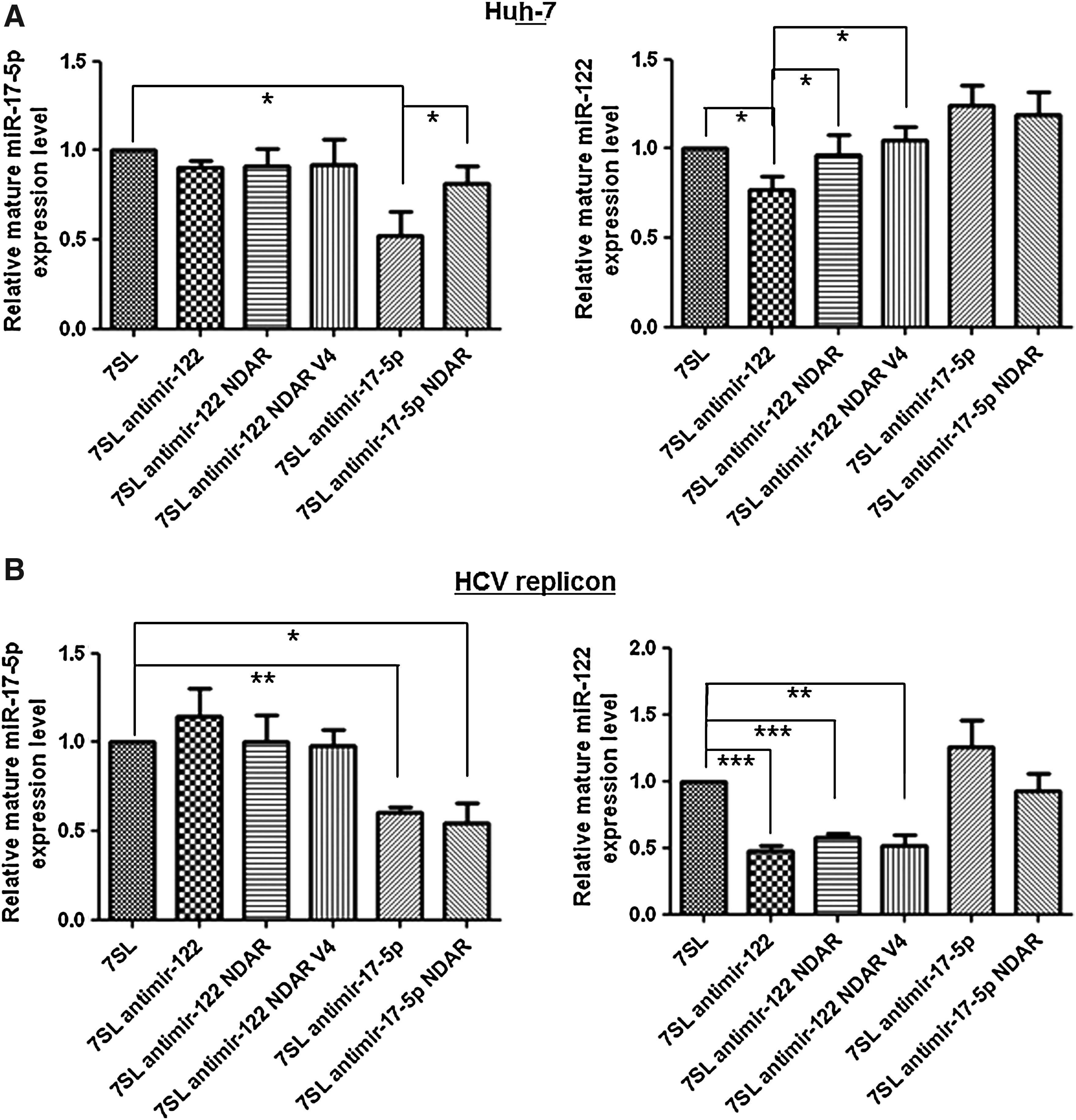
HCV-specific inhibition of miRNA amplification by antimir NDAR. Huh-7
Of note, expression of antimir NDAR selectively suppressed targeted miRNA amplification only in HCV replicon cells, but not in naïve Huh-7 cells, with efficiency comparable to that associated with the expression of the antisense sequence against mature miRNA (Fig. 5A and B). Taken together, selective inhibition of miR-122 function by antimir-122 NDAR could be due to sequestering and saturating of available miR-122 selectively in HCV-replicating cells.
Allosteric ribozyme efficiently inhibits HCV replicon replication
Once the HCV-selective silencing of miR-122 activity by the NDAR had been observed, we next evaluated anti-HCV activity of the ribozyme in human cells stably supporting HCV subgenomic replicon replication (Fig. 6). To this end, the level of HCV negative (−) strand RNA and HCV NS5A protein was quantified in the HCV replicon cells transfected with the expression vector encoding antimir-122 NDAR. Expression of antimir-122 inhibited HCV replication by up to 26%. In contrast, HCV replication was not affected by unrelated antimir-17-5p, indicating specific up-regulation of HCV replication by miR-122 (Jopling et al., 2005). Expression of antimir-17-5p NDAR also inhibited HCV replication by up to 30%. This inhibition was probably due to sequestration of HCV NS5B protein by aptamer. Noticeably, expression of antimir-122 NDAR suppressed HCV replication the most, by up to 47%, indicating that the intracellular expression of the allosteric ribozyme deactivates miR-122 activity simultaneously with inhibiting HCV NS5B function and hence induces synergistic effects on the HCV replication.
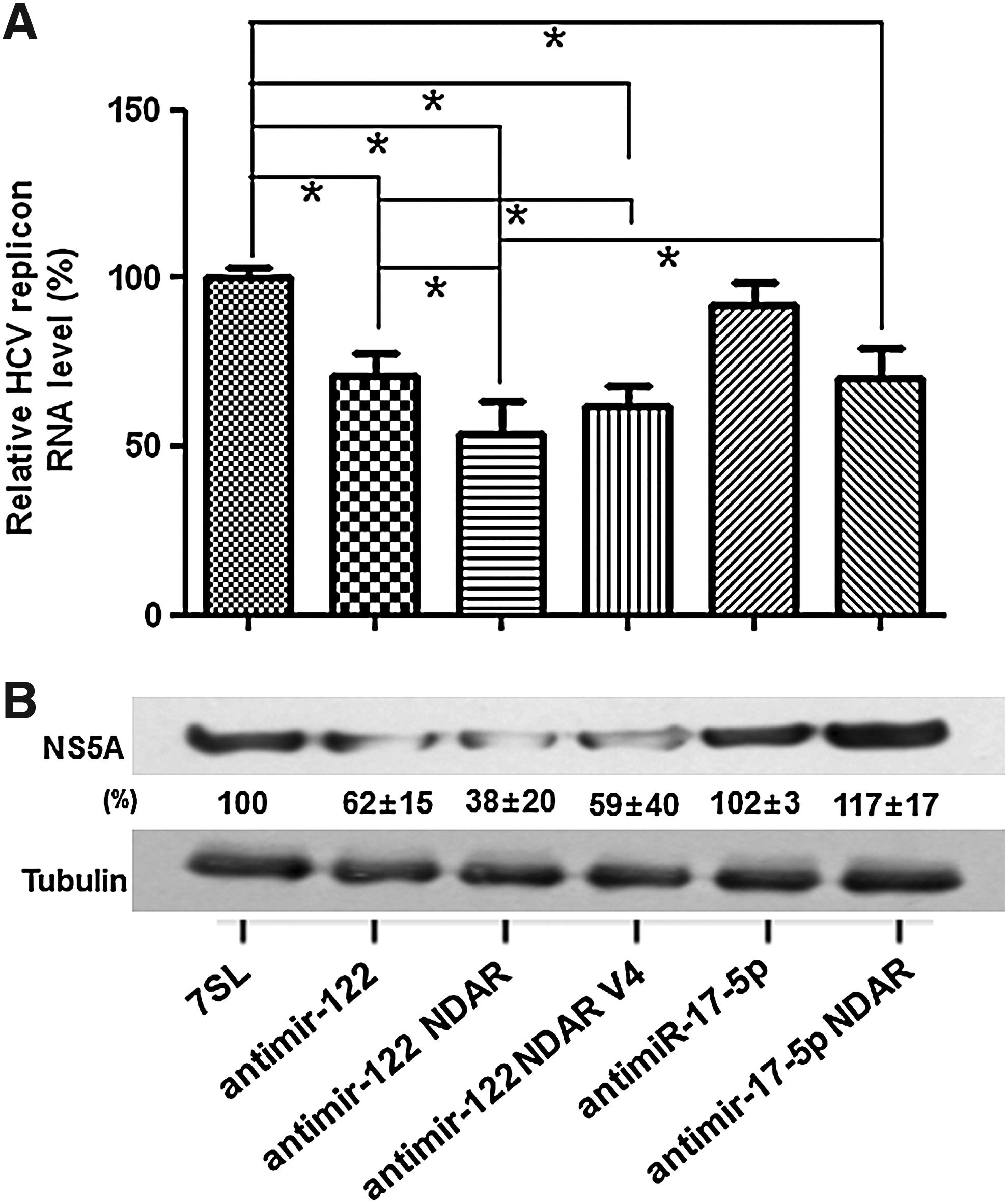
Inhibition of HCV replicon replication by antimir-122 NDAR. HCV replicon cells were transfected with control 7SL vector or 7SL-driven expression vector for antimir-122, antimir-122 NDAR, antimir-122 NDAR V4, antimir-17-5p, or antimir-17-5p NDAR.
Discussion
In this study, a genetic regulation system specifically capable of inhibiting miR-122 activity only in HCV-replicating cells was developed to overcome any possibility of nonspecific side effects due to miR-122 silencing in the normal liver. To this end, allosteric hammerhead ribozyme (antimir-122 NDAR), which can be specifically activated to release antisense sequence against mature miR-122 only in the presence of HCV NS5B, was constructed. The intracellular expression of the antimir-122 NDAR silenced miR-122 activity, probably through sequestering of miR-122, selectively in HCV-replicating liver cells.
Selective silencing of miR-122 could be achieved by the specific introduction of antimir-122 into HCV-infected cells. However, no systems capable of delivering genes or oligonucleotides selectively into HCV-infected livers have been reported. Another approach to selective silencing of miR-122 is to utilize tissue-specific promoter systems to express antimir only in HCV-infected cells. However, no specific promoter systems induced only in HCV-infected livers have been described to date. Extensive analysis of molecular changes including both coding and noncoding RNA expression profile changes in liver tissue when infected by HCV (Ciccaglione et al., 2008; Murakami et al., 2009) should be performed first to develop the HCV-specific delivery or expression system. To our knowledge, the allosteric ribozyme system developed in this study is the first system that selectively turns off miR-122 activity only in HCV-infected cells.
Since the HCV genome is present exclusively in RNA form during replication, various RNA-based therapeutic strategies targeting the HCV genome including trans-cleavage ribozyme, trans-splicing ribozyme, antisense RNA, or small interfering RNA have been proposed as means of halting HCV replication (Hugle and Cerny, 2003; Ryu et al., 2003; Khaliq et al., 2010). However, the rapid emergence of HCV escape mutants could limit such HCV genome-targeting RNA approaches to HCV treatment. Targeting host factors critical for HCV replication instead has emerged as an alternative strategy to circumvent the occurrence of drug-resistant escape variants (Ye et al., 2003; Klebl et al., 2005; Watashi et al., 2005; Okamoto et al., 2006; Kim et al., 2009). MiR-122 also has been included as a targetable host factor. However, miR-122 is significantly down-regulated in human primary hepatocellular carcinoma and negatively regulates tumorigenesis (Gramantieri et al., 2007; Coulouarn et al., 2009; Tsai et al., 2009). Therefore, the possibility of nonspecific toxicity or pathogenesis due to host factor silencing in normal livers should be considered. A strategy for the selective inhibition of the host factor activity only in HCV-replicating cells may be valuable as a host factor-targeting approach to HCV treatment. NDAR, which can release antisense sequence against any kind of target host factors and thus inhibit the activity of wide range of targeted host factors specifically in HCV-replicating cells, could be easily constructed with the backbone of NDAR selected in this study and applied for selective host factor-targeted HCV modulation.
In addition to specificity, NDAR could be more efficacious for HCV inhibition than only the miRNA silencing approach, because it contains an aptamer domain that can specifically bind and sequester HCV NS5B protein. Indeed, NDAR was shown to inhibit HCV replicon replication more efficiently than either antimir-122 or NS5B aptamer in this study. We have shown that intracellular introduction of aptamer against HCV regulatory proteins inhibits the replication of HCV replicon (Hwang et al., 2004; Shin et al., 2006). Combined with activities of selective miR-122 silencing and specific sequestering of HCV NS5B, antimir-122 NDAR could be a more efficient and specific anti-HCV agent.
Footnotes
Acknowledgments
We thank Ralf Bartenschlager (Heidelberg University, Germany) for supplying the HCV replicon construct and David R. Engelke (University of Michigan) for the U6+1 snRNA- and 7SL RNA-based expression vectors. This study was supported by grants from the Korea Research Foundation funded by the Korean Government (KRF-2008-313-C00668) and from the Korea Healthcare Technology R&D Project by the Korean Ministry for Health, Welfare, and Family Affairs (A080173). C.H.L. and J.H.K. are recipients of Brain Korea 21 fellowship.
Disclosure Statement
No competing financial interests exist.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
