Abstract
Development of antibiotic resistance can be achieved either by mutation or by acquiring a resistance gene from foreign sources, with some resistance genes likely originating in microbial populations to counteract antibiotics present in natural ecosystems. In this study, we describe the first report of a strain of nonclinical multidrug-resistant Stenotrophomonas sp. strain G4 with high-level resistance to colistin and meropenem, phylogenetically distinct from well-studied multiple drug-resistant species of Stenotrophomonas maltophilia. As the high-level colistin resistance of this strain was of great concern, the genome of this strain was completely sequenced. Only one chromosome was identified, and no plasmids were found. Chromosomal gene variants and other potential genetic determinants conferring resistance to colistin and meropenem were comparatively analyzed, and results showed that strain G4 harbored two putative colistin resistance determinants (named mcr-5.3 and mcr-8.2) and four extended-spectrum β-lactamase genes. In addition, 12 genes potentially encoding seven different types of efflux pumps were identified, which may have a major role in acquisition/transfer of colistin resistance. Our discovery of multiple antibiotic resistance determinants in this environmental strain extensively expands our understanding of the extent of dissemination of colistin and meropenem resistance.
The inappropriate use of antibiotics is a major factor in the rapid development of drug resistance and in therapeutic failure. Therefore, it is essential to consider, and screen for, the presence of multidrug-resistant (MDR) microbes before the formulation of appropriate antibiotic-based therapies. In general, carbapenems, which are β-lactam antibiotics, are used as a last line of defense against severe infections with MDR Enterobacteriaceae, and colistin (polymyxin E), a cationic antimicrobial polypeptide, is clinically regarded as the final defense line for treating lethal challenges with carbapenem-resistant pathogens. The coexistence of colistin resistance and extended-spectrum β-lactamase (ESBL) within individual variants of bacterial species isolated from humans, animals, and the environment is a strong reminder of the need to employ new approaches for combating MDR microbes.1–4
Unfortunately, the clinical use of carbapenems and colistin has been significantly challenged by the occurrence of the mobilized colistin resistance determinant (MCR-1) in clinical carbapenem-resistant isolates that produce New Delhi metallo-β-lactamase 1 (NDM-1) 5 or its variant NDM-5. 6 In addition to the intrinsic determinants of colistin resistance (e.g., PmrAB and PhoPQ mutations), transferable determinants, such as MCR-1, encoding phosphoethanolamine (PEA)-lipid A transferases have been discovered.2,7,8 The mechanism of action of MCR involves the covalent addition of PEA to the 4′-phosphate position of lipopolysaccharide lipid A moieties through a “ping-pong” trade-off.7–10 The increasing incidence of MCR-like determinants, including MCR-1, MCR-2, MCR-3, MCR-4, MCR-5, MCR-6, MCR-7, and the recently reported MCR-8, which are found in bacteria in patients, animals, and various environments, challenges the renewed interest in colistin for clinical therapies. In particular, MCR-1 and MCR-2 are commonly carried by plasmids, and >10 genetic variants of MCR-1/2 have been detected from these transferable reservoirs, suggesting the ongoing spread and dissemination of MCR-1/2. 9 The transferability of MCR-like polymyxin resistance is of social concern, as MCR-like mechanisms pose a serious threat to public health and the safety of the food chain. Furthermore, although rare, MCR has already been found to coexist with both ESBL 11 and NDM-1 12 (or its variants NDM-5 13 and NDM-9 14 ).
In this study, we report on a nonclinical strain of MDR Stenotrophomonas sp., named G4, coharboring MCR- and ESBL-genetic determinants, which was isolated from sewage water of a poultry farm in Jinan, China, and likely developed its resistance pattern from antibiotic exposure and overuse in an agricultural setting. The sewage water was gradiently diluted with sterilized water, and then plated onto LB agar supplemented with 8 μg/mL colistin and 8 μg/mL meropenem, and incubated at 30°C for 24 hour to obtain single colonies; then one of the single colonies was picked out and streaked on the above LB agar to obtain a pure culture. After purification, a single colony (representing a pure culture) was named as strain G4. The G4 strain was cultured with LB supplemented with 8 μg/mL colistin and 8 μg/mL meropenem, and then the cells were harvested and the total DNA was isolated. The primer pairs 27F (5′-AGAGTTTGATCCTGGCTCAG-3′) and 1492R (5′-GGTTACCTTGTTACGACTT-3′) were used to amplify the partial fragment of 16S rDNA. Strain G4 was identified as Stenotrophomonas sp. based on 16S rDNA sequence analysis (GenBank Accession number: MK418754). A background survey found that amoxicillin and colistin had been regularly used in this livestock farm for poultry to prevent infection. To determine the minimal inhibitory concentrations (MICs) of strain G4 for different antibiotics, including ampicillin, cefixime, ciprofloxacin, florfenicol, meropenem, colistin, and tetracycline, the broth microdilution method was used, as described by the Clinical Laboratory Standard Institute (CLSI) guidelines. 15 Results showed that strain G4 has a striking resistance to the antibiotics tested (Table 1). Compared with the published MICs for colistin-susceptible Klebsiella pneumoniae, Escherichia coli, and Stenotrophomonas sp. strains,2,3,16–18 the MICs of colistin and meropenem for strain G4 were higher.
Minimal Inhibitory Concentrations of Antibiotics for Strain G4 and Escherichia coli Conjugants
Concentration units for the antibiotics tested: μg/mL
25DN, sodium azide-resistant E. coli strain; 25DN-POL conjugant derived after colistin selection, with 25DN as recipient strain and G4 as donor strain.
To determine the genetic basis for the multidrug-resistance phenotype of this strain, whole-genome sequencing was performed using the Illumina NextSeq platform and the PacBio RS II platform. The resultant contigs were assembled by SOAPdenovo (soap.genomics.org.cn) and Velvet (ebi.sv.uk/∼zerbino/velvet) programs, generating a total genome sequence of 4,477,485 bp with an average GC content of 66.83%. The features of the complete genome sequence are shown in Supplementary Table S1. Whole genome alignment by BLASTp-based searches and a core genomic phylogeny established with CVTree 19 both grouped strain G4 with Stenotrophomonas species (Figs. 1 and 2), and strain G4 phylogenetic clade was close to Stenotrophomonas rhizophila and Stenotrophomonas maltophilia. S. rhizophila DSM14405 can facilitate plant growth and alleviate adversity. 20 Moreover, S. maltophilia pertaining to Stenotrophomonas species is the only phylogenetic clade known to cause human disease and prompt broad antibiotic resistance,17,21,22 and a recent study has described the presence of specific subgroups of S. maltophilia strains adapted to the human host resulting from their genomic diversity and variation. 22 As shown in Fig. 1, strain G4 was more closely related to S. rhizophila rather than S. maltophilia and Xanthomonas species, based on whole genomic level phylogeny with S. rhizophila DSM14405, four S. maltophilia and a variety of closely related Xanthomonas fully sequenced genomes in GenBank. However, the more accurate species assignment of strain G4 needs further clarified with the Biolog examination 23 and more fully sequenced Stenotrophomonas strains.
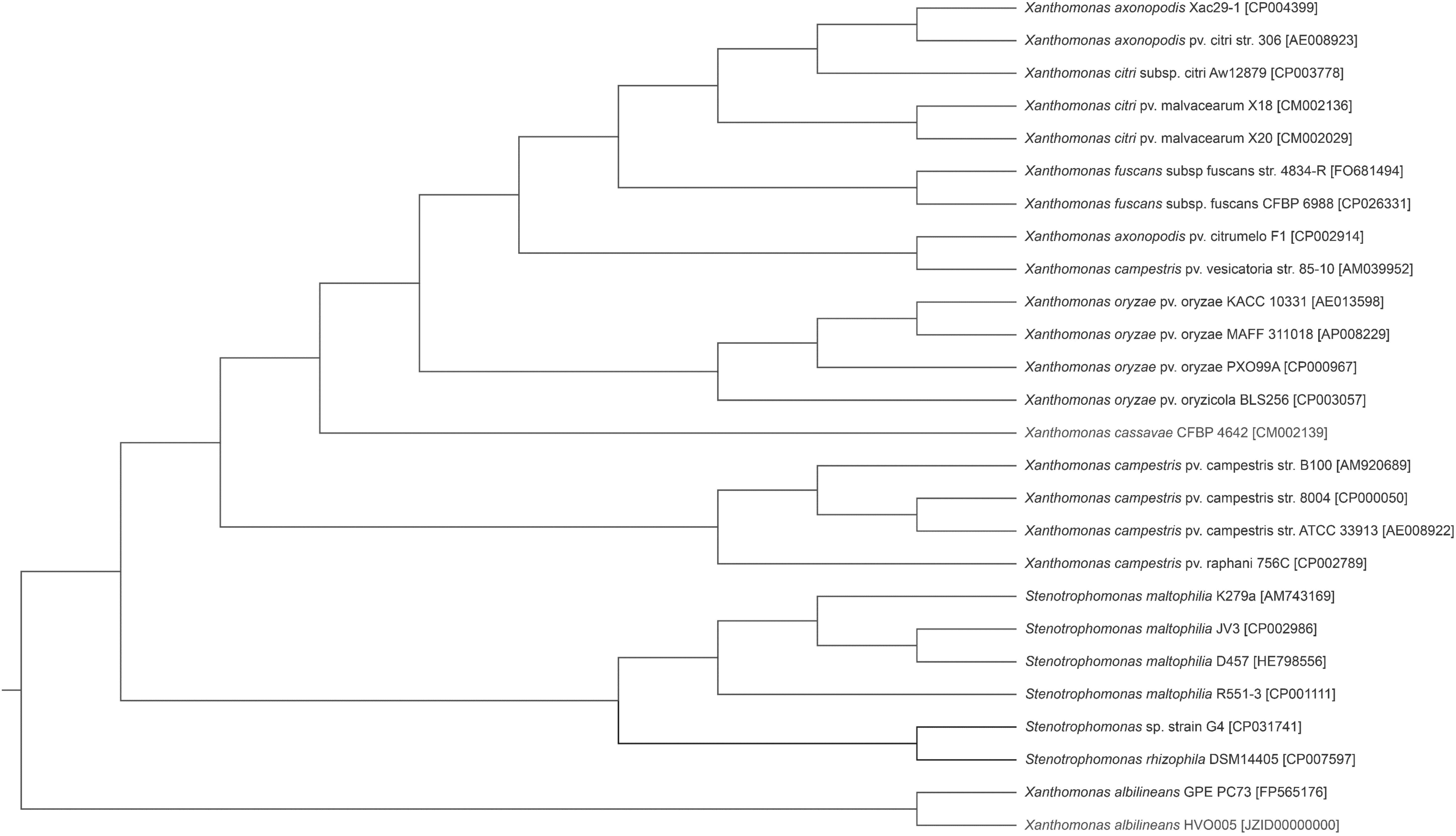

Phylogenetic relationships of strain G4. Whole-genome phylogeny was determined using CVTree3.0 and indicated that strain G4 is closely related to the genus Stenotrophomonas. Strain names are followed by their GenBank accession numbers.
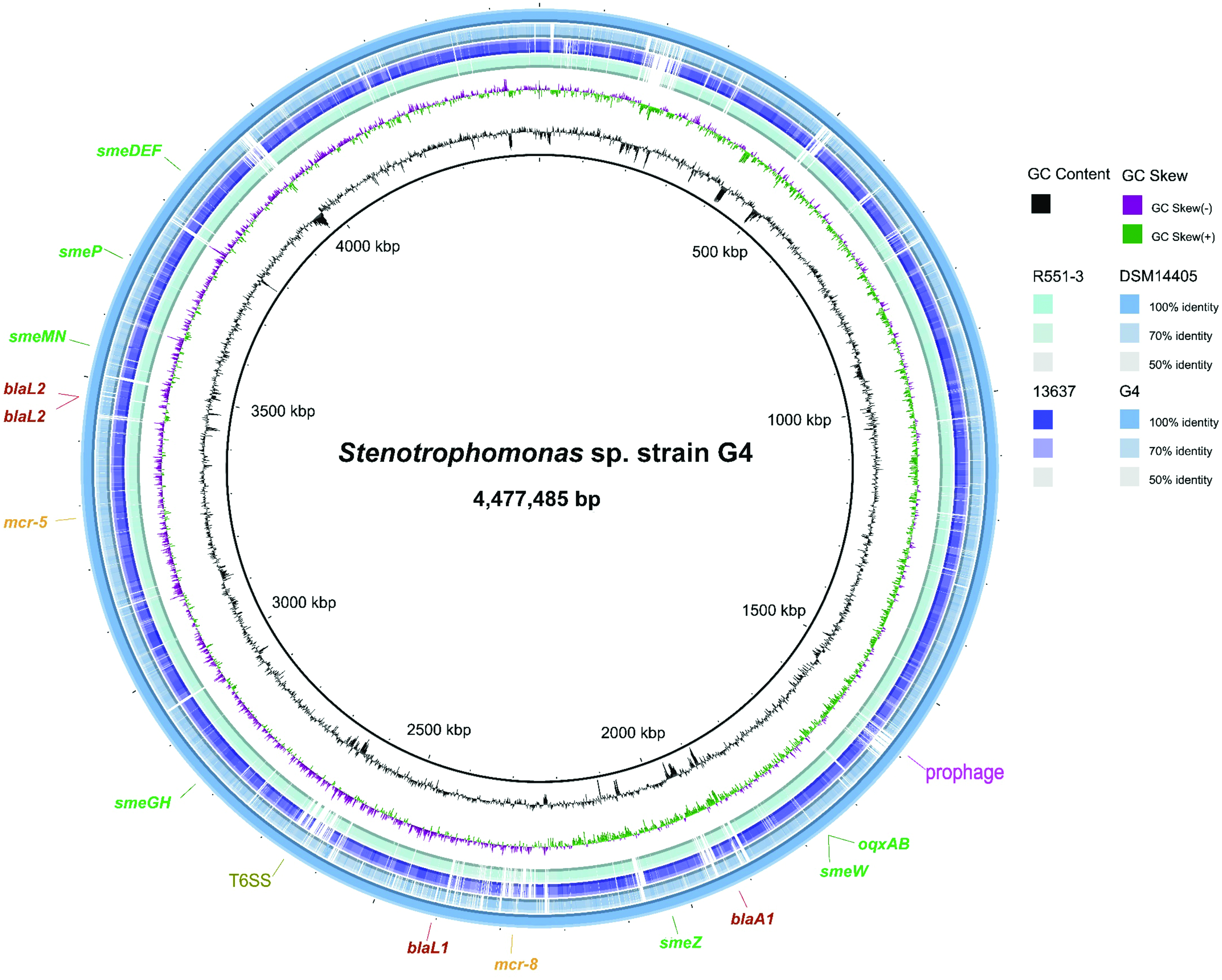
Circular schematics of the strain G4 genome and its BLASTn-based comparison with closely related bacterial genomes. Genomes of Stenotrophomonas strains DSM14405, R551-3, and 13637 were chosen for reference. Color images are available online.
As strain G4 has notably high levels of colistin and β-lactam resistance, we conducted comparative genomics to identify potential MCR- and ESBL-encoding genes, using the circular genome of strain G4 and closely related Stenotrophomonas spp. (Fig. 2). In addition, four genes coding for β-lactamases and two mcr-like genes were predicted (Table 2) using the ARG-ANNOT and VRprofile programs.24,25 We renamed the mcr-5-like gene, which contains mutations compared with the prototype mcr-5, as mcr-5.3, whereas the mcr-8-like gene was designated mcr-8.2 (Fig. 3 and Supplementary Fig. S1). The genomic contexts of the MCR-encoding genes were also assessed to investigate their transmission potentials and spread risk (Fig. 4). The mcr-pap2 unit was observed to be widely disseminated in species closely related to strain G4 (Fig. 4). However, no IS element, such as ISApl1, which is usually contained by a mcr-1/2 cassette [ISApl1-mcr-1-orf-ISApl1, presumptive Tn6330], 10 was found in the strain G4 genome. Therefore, strain G4 may comprise a reservoir of mcr genes to facilitate their spread and persistence. Similarly, four genes coding for ESBLs and which represent intrinsic antibiotic resistance capabilities were identified, highlighting the foreign and ancestral origins of ESBL- and MCR-related antibiotic resistance determinants. Group 3c β-lactamase (blaL1) and Group 2e β-lactamase (blaL2), responsible for the typical β-lactam resistance properties of S. maltophilia, were discovered in strain G4 (Table 2). In S. maltophilia, the blaL1-encoded broad-spectrum metallo-β-lactamase has been found to hydrolyze carbapenems. 26 In summary, the aforementioned β-lactamase antimicrobial resistance (AMR) determinants in strain G4 are likely intrinsic progenitors, which may have been widely disseminated to other species.
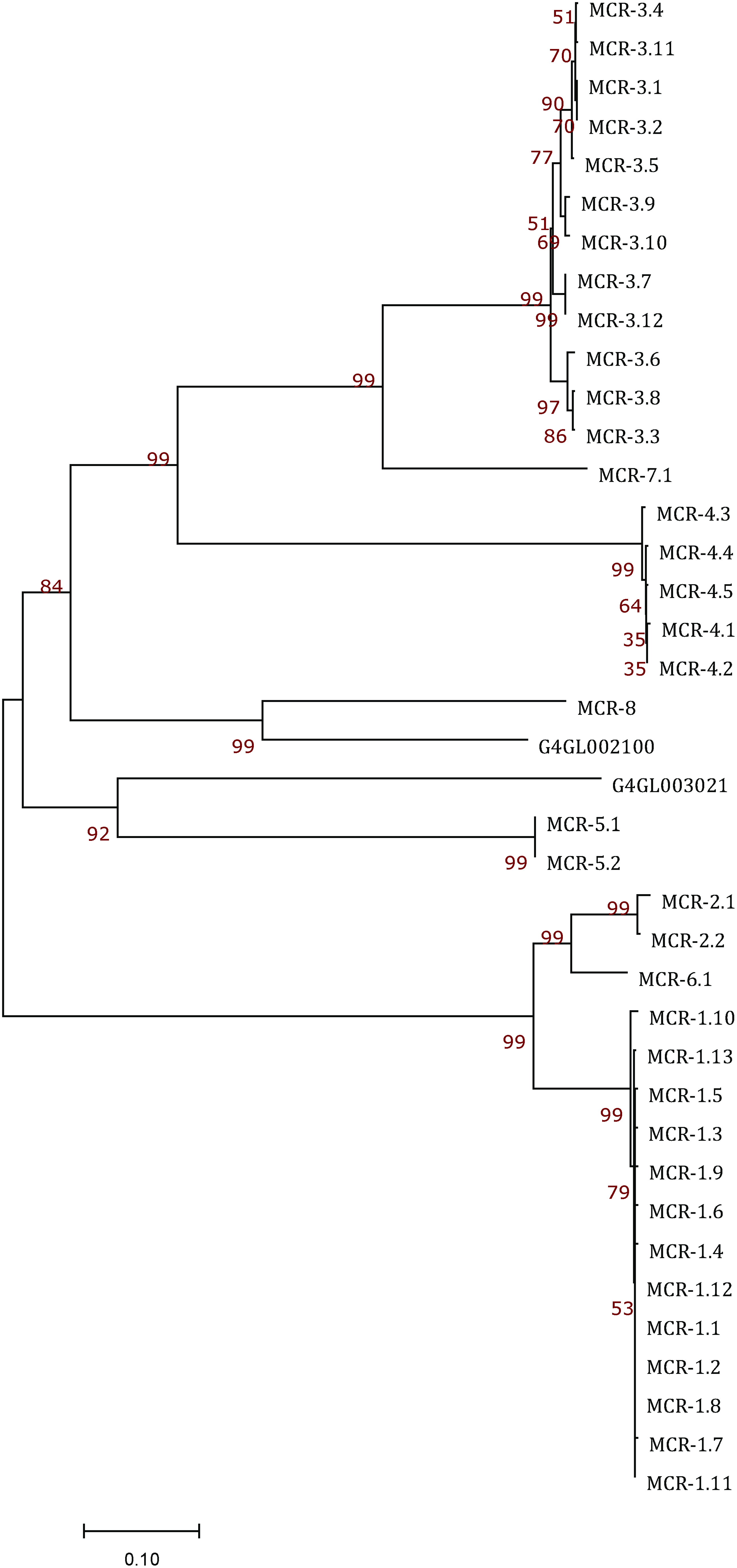
Phylogenetic relationships of mcr genes identified in this study. G4GL002100 and G4GL003201 in strain G4 were identified as the mcr-8 and mcr-5 homologs, respectively. Other MCR-like proteins were curated from two publications [9,10]. The 37 manually collated sequences and two sequences for the two MCR-like products of strain G4 were included in the Supplementary Figure S1. The MCR phylogeny was conducted through the MEGA7 program (megasoftware.net), and WAG model with 1,000 bootstrap. MCR, mobilized colistin resistance. Color images are available online.
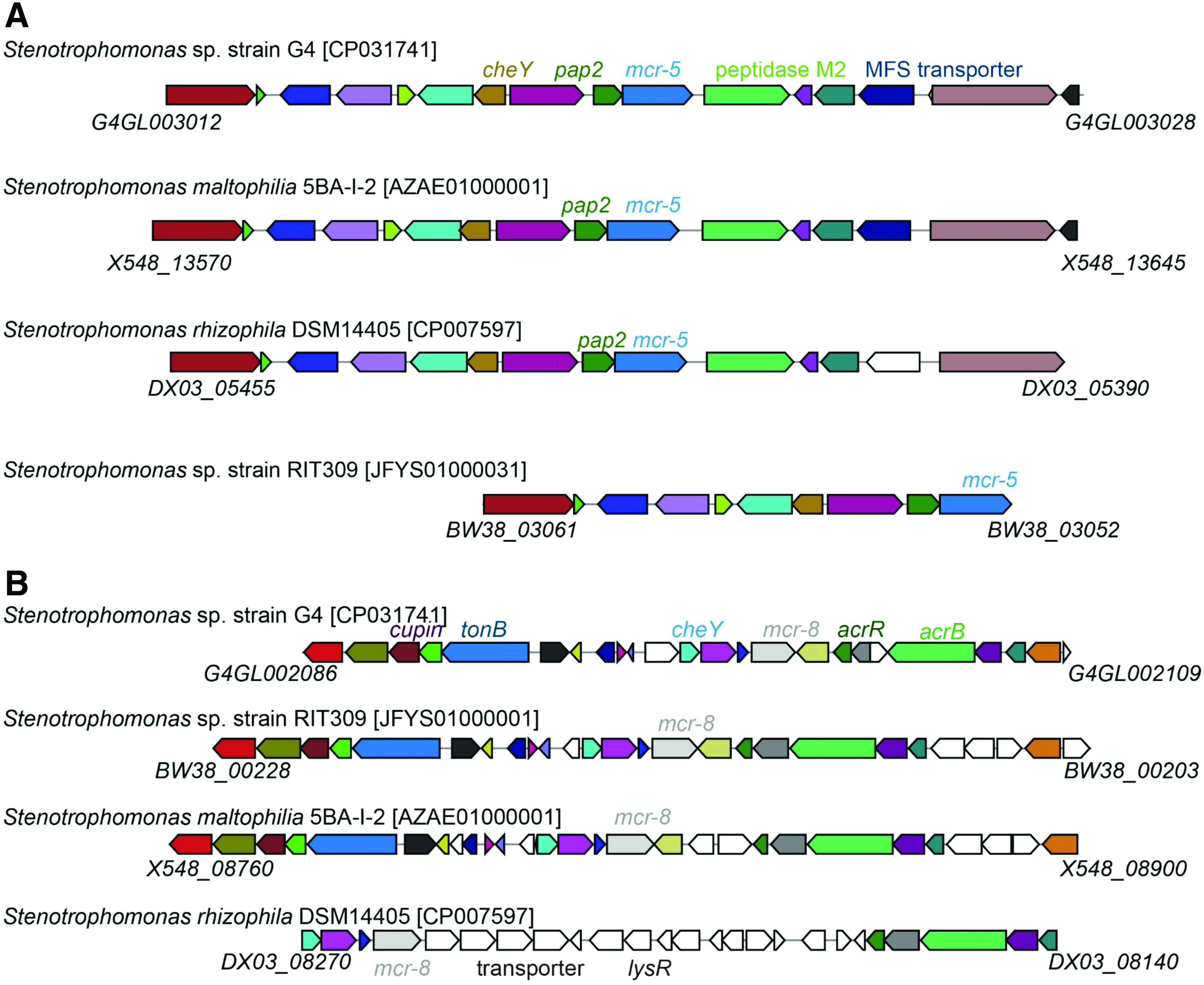
Genomic contexts of mcr-like genes in the strain G4 genome and closely related strains.
Identified Antibiotic Resistance Genetic Determinants and Their Locus Tags in the Strain G4 Genome
Gene locus tags: ORF_No. for the abbreviation of G4GL00(ORF_No.), such as 3512-4 indicating ORFs from G4GL003512 to G4GL003514.
RND- and AcrAB-type efflux pumps that increase resistance to antimicrobial agents have been recently reported and are also involved in colistin and β-lactam resistance.17,27,28 The strain G4 chromosome encodes seven efflux pumps, including SmeDEF, SmeMN, SmeGH, SmeP, SmeW, SmeZ, and OqxAB (Table 2). In Stenotrophomonas species phylogenetically related to strain G4, the overproduction of efflux pumps, such as SmeDEF, confers high-level antibiotic resistance; moreover, antibiotic extrusion and colonization are both likely related to SmeDEF. 17 The expression and activity of efflux pumps are usually inhibited, but SmeDEF expression could be induced by plant-produced flavonoids and, therefore, impact antibiotic resistance. 17 Since SmeDEF expression can be triggered by plant-produced flavonoids in Stenotrophomonas, we hypothesized that this efflux pump might have a role in antibiotic resistance and the colonization of plants by strain G4.
The strain G4 chromosome contains an intact prophage (T6SS) as well as one type VI secretion system (Fig. 5). Prophages have been previously determined to contribute to host specificity of pathogens and growth fitness.25,29 Of note, a variable region of strain G4 was comparatively identified in the prophage, and this unstable region and its vicinity are proposed to correlate with strain G4 environmental adaptability (Fig. 5A). The genetic arrangement of the T6SS of strain G4 is almost identical to that of its counterparts in other closely related strains and to that of the type i4b T6SS in Xanthomonas oryzae 30 (Fig. 5B). Type i4b T6SSs have been experimentally verified to provide virulence traits and alternative adaptabilities in other strains.31–33 Given that the activation of T6SS renders Acinetobacter baumannii more susceptible to antibiotics,34,35 it will be important to elucidate the role of T6SS in the antibiotic resistance of the MDR strain G4.
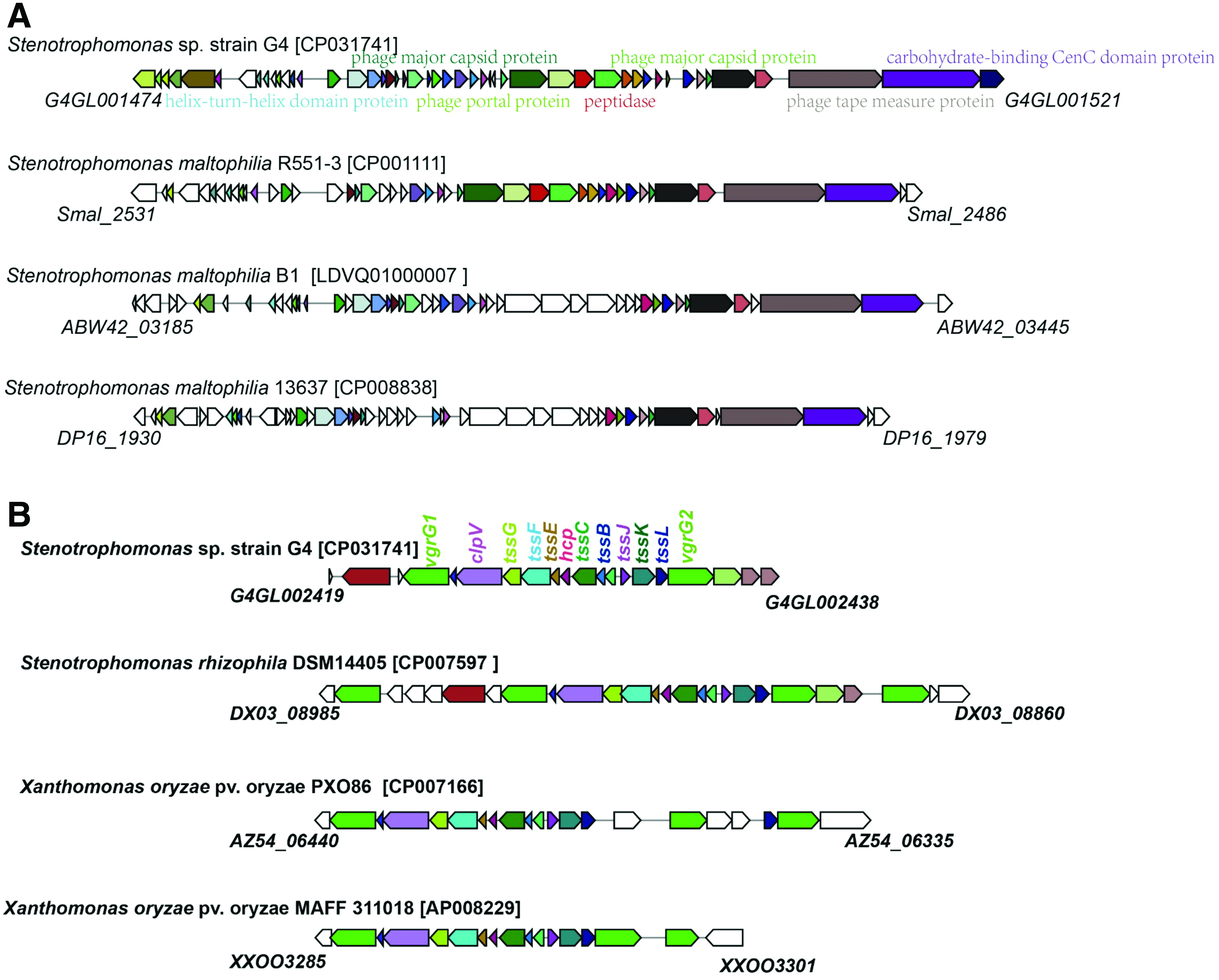
Synteny analyses of the gene clusters coding for
To determine if the antibiotic resistance genes could be transferred from strain G4 to other bacteria by horizontal gene transfer, the sodium azide-resistant E. coli 25DN strain (derived from E. coli 25922) was used as the recipient strain for examining conjugal transfer of resistance. Colistin and meropenem were used to select the resistant conjugants, as described previously.
36
For example, after mixing the donor strain G4 and recipient strain 25DN, the mixture was spread on LB agar medium with X-Gluc (5-bromo-4-chloro-3-indolyl-β-
Antibiotic resistance genes and their transfer are an urgent risk to human health.37,38 It has been frequently reported that mcr-like genes could cotransfer with other antibiotic resistance genetic determinants, such as NDM-1 gene and/or its derivates (NDM-5 and NDM-9).5,39 This genetic colocalization of a variety of AMR genes raises the possibility that bacteria could acquire resistance to both colistin and carbapenem, two categories of antibiotics that are both considered the last resorts against MDR bacteria. It is likely that Enterobacteriaceae could predominantly account for the mcr-like genes in commensal and gut microbe.9,10,40 However, our discovery of a Stenotrophomonas strain possessing two MCR-like determinants and several ESBL genes implies that the sources and distribution of AMR genes may be substantially complex. Presently, the emergence and transmission of MCR and ESBL pose great risks to public health through the food chain and environmental exposure. Based on this scenario, the proposed “one health” approach 10 might benefit the effective control of worldwide AMR problems and assist combating the stringent challenges from MDR superbugs, which requires the systemic coordination of three sectors: environment, animals, and humans.
Data Availability
Complete sequence of the strain G4 genome was deposited into GenBank under accession No. CP031741, and 16S rRNA sequence of the strain G4 was submitted to GenBank assigned with accession No. MK418754.
Footnotes
Acknowledgments
We thank Dr. Susan T. Howard for critical reading of the article. This study was supported by the National Natural Science Foundation of China (31800118 and 31500094), China Postdoctoral Science Foundation (2017M621965), and the Innovation Project of Shandong Academy of Medical Sciences.
Author Contributions
J.L. and G.C. designed and supervised this project; S.L., J.Z., and J.F. performed experiments; J.L., J.Y., C.Z., and G.C. analyzed the data; J.L., C.Z., and G.C. drafted this article.
Disclosure Statement
Authors declared no potential conflict of interest.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
