Abstract
The cell wall of Streptococcus pneumoniae contains an unusually complex wall teichoic acid (WTA), which has identical repeating units as the membrane-anchored lipoteichoic acid (LTA). Both polymers share a common cytoplasmic pathway of precursor synthesis, but several TA enzymes have remained elusive. Bioinformatic analysis of the genome of various pneumococcal strains, including choline-independent mutant strains, has allowed us to identify the missing TA genes. We present here the deduced complete pathways of WTA and LTA synthesis in S. pneumoniae and point to the variations occurring in different pneumococcal strains and in closely related species such as Streptococcus oralis and Streptococcus mitis.
Introduction
In most species, the structure of WTA is different from that of LTA, and both polymers are synthesized by different pathways.9,75,89 The majority of TAs are chains of multiple glycerol phosphate or ribitol phosphate (Rib-P) repeating units that may carry D-alanine and/or sugar residues.2,61 The former increases the cell's resistance to antimicrobial compounds, but the role of the latter is less clear, although some bacteriophages require TA-bound sugars for infecting the bacterial cell. The TAs of S. pneumoniae are unusual in several aspects: first, in this species, the WTA and LTA chains have identical repeating unit structures and length distribution, indicating that they are produced in the same biosynthetic pathway. Second, the repeating units have an unusually complex structure containing the rare amino sugar 2-acetamido-4-amino-2,4,6-trideoxygalactose (AATGal), glucose (Glc), Rib-P, and two N-acetylgalactosamine (GalNAc) residues, each of which carries a phosphorylcholine (P-Cho) moiety (Fig. 1). 27
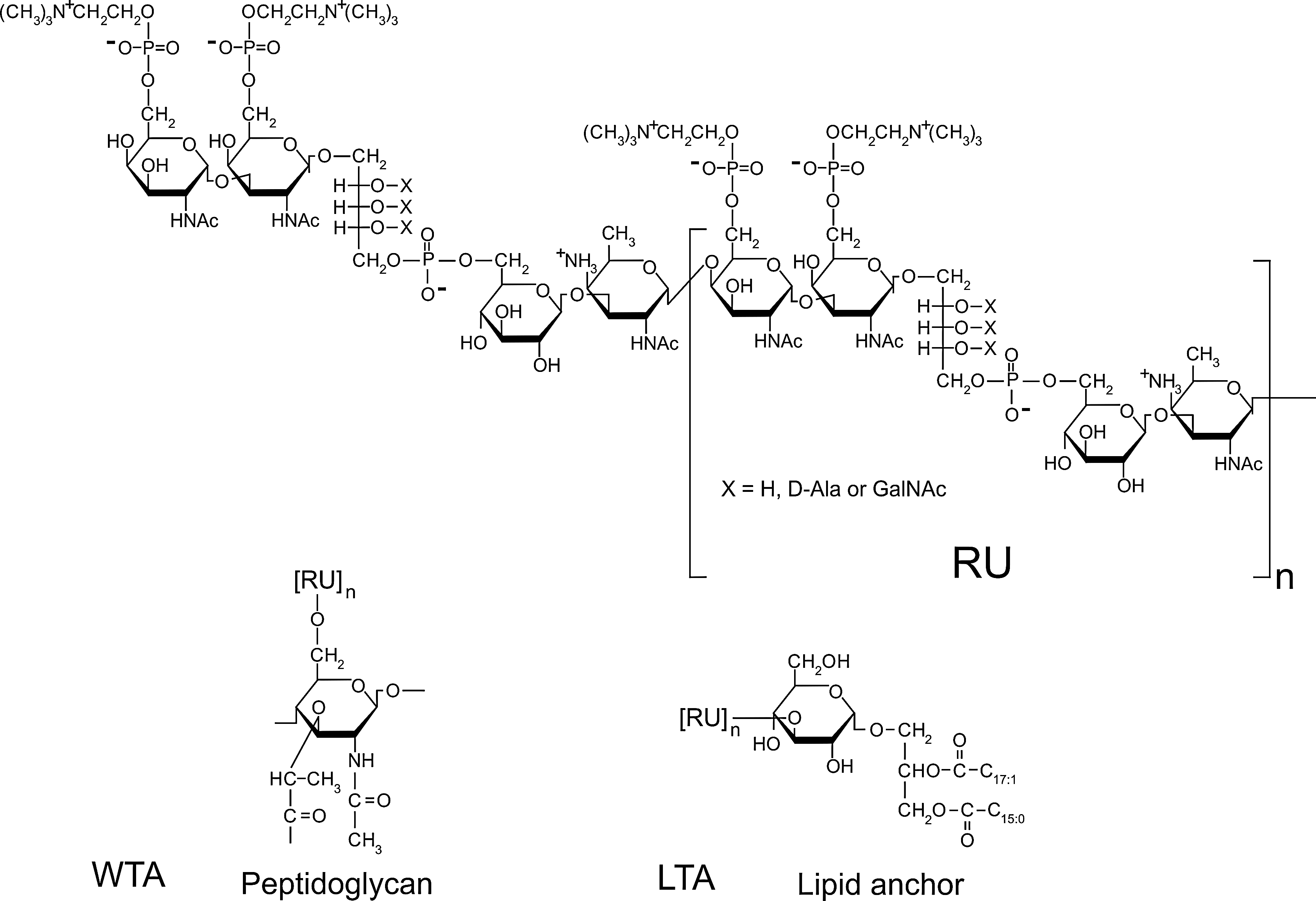
Structure of teichoic acid (TA) in S. pneumoniae. The revised structure of the TA repeating units (RU) according to Seo et al. 71 is shown. The RU contains 2-acetamido-4-amino-2,4,6-trideoxygalactose (AATGal), glucose (Glc), ribitol phosphate (Rib-P), and two N-acetylgalactosamine (GalNAc) residues, each of which carries a phosphorylcholine (P-Cho) moiety. The hydroxyl groups of Rib may be substituted by D-Alanine (D-Ala) or GalNac. The anchors for TA are also shown. RUs are attached to the glycolipid glucose-diacylglycerol (Glc-DAG) to yield lipoteichoic acid (LTA) or to peptidoglycan via N-acetylmuramic acid (MurNAc) to produce wall teichoic acid (WTA). n, number of repeating units.
Phosphorylcholine is very rare in bacteria. S. pneumoniae is the only species whose growth is known to depend on exogenous choline, which is exclusively metabolized to decorate the TA chains. 77 The amino alcohol choline serves as an anchor for the class of choline-binding proteins, among which are peptidoglycan hydrolases and adhesions. 35 Choline can be replaced in the growth medium by ethanolamine (or putrescine 86 ), which then decorates TA in lieu of choline. 78 However, cells grown with ethanolamine in the absence of choline grow in chains and fail to lyse in the stationary phase due to the lack of activity of the septum-cleaving LytB glucosaminidase and the autolysin LytA, both of which are choline-binding proteins. 79 A similar phenotype is observed in several choline-independent mutant strains that carry unsubstituted TAs when grown in the absence of choline.15,31,72,92
Earlier studies by Fischer suggested the following repeating units present in both WTA and LTA: Glc-AATGal-GalNAc(P-Cho)-GalNAc(P-Cho)-Rib-P. 28 Consistent with this structure, hydrofluoric acid, which hydrolyses the TA chains at phosphomono and phosphodiester bonds, released the Glc-AATGal-GalNAc-GalNAc-Rib moiety. Complete LTA of this type, containing the glycolipid anchor, has been chemically synthesized.59,60,70 However, recent mass spectrometry analysis of whole LTA molecules isolated from pneumococci suggested a revised structural model in which the biosynthetic repeating unit begins with AATGal and has the following structure: GalNAc(P-Cho)-GalNAc(P-Cho)-Rib-P-Glc-AATGal (Fig. 1). 71 While the different models predict only minor differences in the composition of the components, they have important implications on the biosynthetic pathway: The previous model predicts that the synthesis of the repeating unit begins with the attachment of ribitol-P to the upr-P membrane anchor, whereas the new model predicts that the first sugar to be attached to the membrane anchor is AATGal. In this communication, we assume the revised model of LTA to suggest the pathway of TA synthesis.
The biosynthetic pathway of TA synthesis has been well studied in B. subtilis and S. aureus, in which most of the enzymes have been identified and characterized. However, most of the pneumococcal TA enzymes have remained elusive, except those encoded by genes in the lic1/lic2 genomic region. In this work, we propose novel TA genes, which, together with the known ones, account for the complete pathways for the synthesis of LTA and WTA in S. pneumoniae. Our assignments of TA genes are based on bioinformatic searches, known mutations in choline-independent mutant strains, and the revised structural model of the LTA molecule. We also present a comparative analysis of TA genes in closely related streptococcal species.
Methods
The genomic sequences of S. pneumoniae R6 (accession number AE007317 42 ), Streptococcus mitis B6 (FN568063 16 ), and Streptococcus oralis Uo5 (FR720602 65 ) were used for the identification of genes involved in TA biosynthesis.
The predicted protein sequences were searched against the nonredundant protein sequence database provided by the National Center for Biotechnology Information using BLAST software (www.ncbi.nlm.nih.gov/BLAST/). 54 To access protein family data, the Pfam database was used (http://pfam.sanger.ac.uk/). 24
Transmembrane helices in membrane proteins were predicted with online tools for topology prediction TMHMM 2.0 51 and SOSUI. 39 Putative signal peptides were identified by using SignalP. 4
Results and Discussion
Biosynthesis of pneumcoccal TAs
TA biosynthesis in S. pneumoniae requires multiple cytoplasmic and membrane-associated steps. Based on several known components, bioinformatic analysis, and the revised structure of LTA, we suggest here the complete pathway of the formation of the precursors common for WTA and LTA, its polymerization to the TA chains, and the transport of these chains across the cytoplasmic membrane (Fig. 2). The TA chains are then attached to either the peptidoglycan to form the WTA or the glycolipid anchor to form LTA. In the next sections, we present the various known enzymes of the pathway and suggest the hypothetical, missing components, beginning with the cytoplasmic membrane steps for the synthesis of lipid-linked AATGal, the completion of the core repeating unit, and the attachment of phosphorylcholine and additional substituents. The synthesis of the TA repeating unit requires the products of at least 16 known and hypothetical genes, present in four regions: the 8-gene lic1/lic2 region (with the licA, licB, licC, licD1, licD2, tarI, tarJ, and tacF genes); the five genes in the lic3 region (with the licD3, spr1221, spr1222, spr1223, and spr1224 genes); and two 2-gene regions, spr0091-spr0092 and spr1645-spr1655. The genes and enzymes known and proposed to be involved in TA biosynthesis are depicted in Fig. 2 and Table 1.
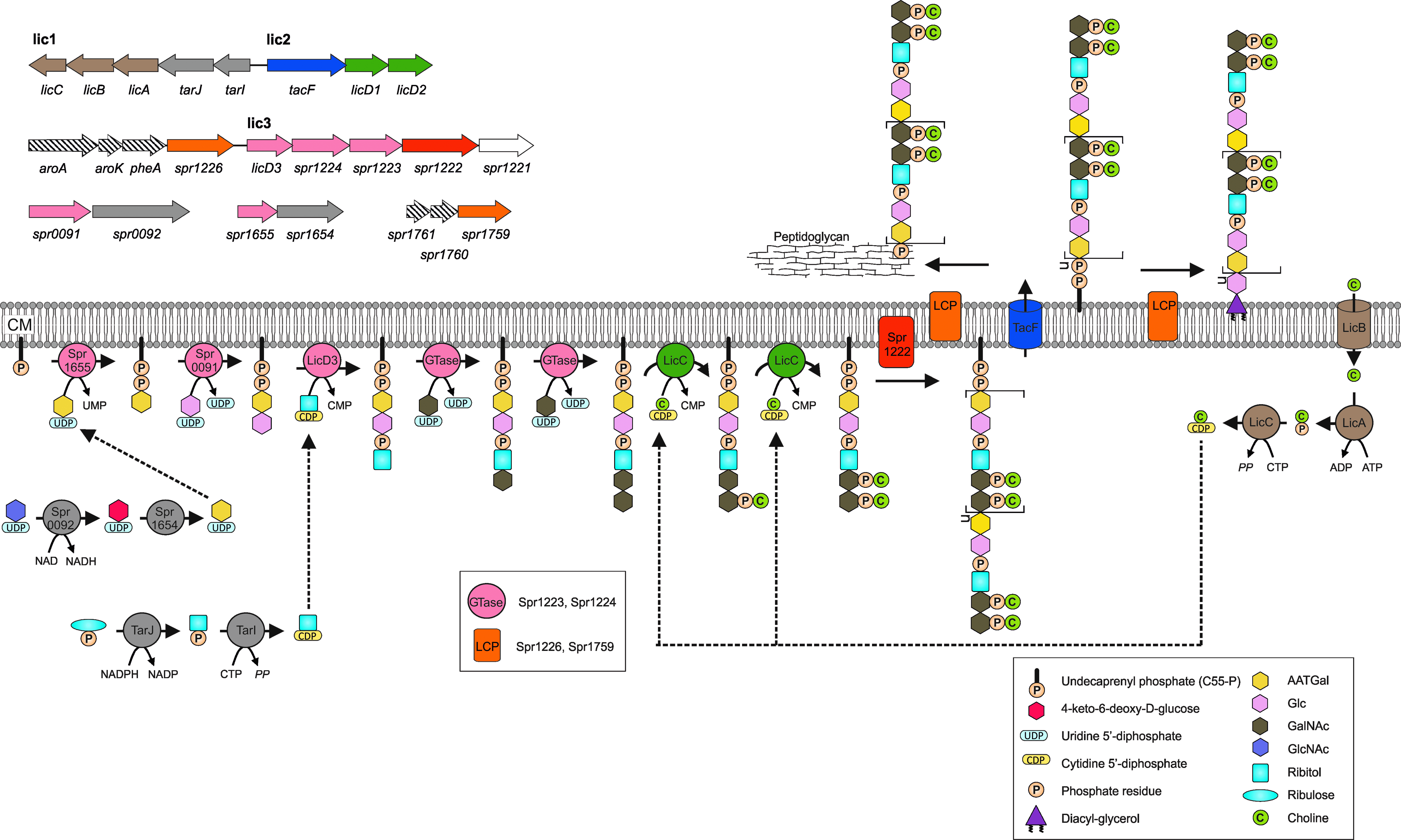
Proposed pathway for the biosynthesis of TAs in S. pneumoniae and the genetic organization of the corresponding genes. The pathway generally applies to S. pneumoniae. The gene/protein names and numbers are from S. pneumoniae R6. The different colors of genes and the corresponding proteins represent distinct steps of TA biosynthesis: gray, formation of the TA precursors of RU; dusky pink, biosynthesis of RU; brown, choline uptake and activation; green, decoration of the RU with choline; red, polymerization of the precursor; blue, transport across the membrane; orange, linking TA chains to peptidoglycan and the glycolipid anchor. The spr1221 has no predicted role in the pathway and is depicted by an open arrow. The dashed arrows indicate the genes encoding proteins that are not involved in TA biosynthesis. Enzymes predicted to be involved in repeating unit RU biosynthesis were drawn close to the membrane although not all of them were predicted to be membrane proteins (see Table 1 for details). CM, cytoplasmic membrane; AATGal, 2-acetamido-4-amino-2,4,6-trideoxygalactose; Glc, glucose; GalNAc, N-acetylgalactosamine; GlcNAc, N-acetylglucosamine; UMP, uridine-monophosphate; CMP, cytidine-monophosphate; GTase, glycosyltransferase; LCP, LytR-Cps2A-Psr family protein.
-, no information available; licA, licB, licC, licD1I, and licD2 are not essential in choline-independent strains.15,21,46
Transmembrane helices were predicted with online tools (TMHMM and SOSUI) for topology prediction; C, cytoplasmic; M, membrane; TMH, transmembrane helices.
Reference for gene essentiality.
Verified experimentally.
Shown to be membrane associated.
spr1759 deletion strain may contain secondary mutations. 43
TA, teichoic acid.
Synthesis of Glc-AATGal-PP-upr
The new model of the LTA structure derived from mass spectrometry analysis shows that the repeating unit begins with the sugar AATGal. 71 Assuming this model, the first membrane step in the synthesis of the repeating unit should be the transfer of AATGal 1-phosphate to the undecaprenyl phosphate (upr-P) lipid anchor. UDP-activated AATGal, which is also found in some capsule types, has been predicted to be synthesized in two steps by the products of the spr1654 and spr0092 genes based on their sequence similarity to the genes in Shigella sonnei. 1 Accordingly, the predicted dehydrase Spr0092 converts UDP-GlcNAc to UDP-4-keto-6-deoxy-GlcNAc, which is the substrate for the aminotransferase Spr1654 to form UDP-AATGal. 1 In Bacteroides fragilis, the synthesis of the repeating unit of an exo-polysaccharide is initiated by the addition of AATGal from UDP-AATGal to the upr-PP lipid anchor, forming AATGal-PP-upr, and this reaction is catalyzed by the gene product of wcfS. 13 The product of the S. pneumoniae spr1655 gene has 44% sequence identity to WcfS, making Spr1655 a likely candidate for this function. 71
In the next step, a glucose residue is attached to AATGal-PP-upr, requiring a glycosyltransferase. Interestingly, the uncharacterized Spr0091 is a predicted glycosyltransferase. Should Spr0091 be the enzyme for adding a glucose to AATGal-PP-upr, then all four genes required for the synthesis of Glc-AATGal-PP-upr would locate to two chromosomal two-gene regions, spr0091-spr0092 and spr1654-spr1655. Consistent with this hypothesis, the spr0091 gene appears to be essential 74 as would be expected for a role in the synthesis of the TA repeating unit.
Although the Glc residue could be added by one of the two predicted glycosyltransferases, Spr1223 or Spr1224, encoded by genes in the lic3 region (see below), we predict that Spr0091 adds glucose based on the unusual TA found in a serotype 5 strain, which contains galactose instead of glucose in the repeating unit. 81 At the genomic position equivalent to spr0091, the serotype 5 strain 70585 18 contains a gene encoding a glycosyltransferase of the same family but with very limited sequence identity to Spr0091. By contrast, the glycosyltransferases corresponding to Spr1223 or Spr1224 are virtually identical in both strains. Therefore, the replacement of Spr0091 by another glycosyltransferase (SP70585_0164) strongly suggests that these enzymes incorporate glucose and galactose, respectively. Homologs of the sp70585_0164 gene are also present in strains GA17570 and GA07463, suggesting that they may contain galactose in their TA.
Synthesis of activated ribitol and the attachment of ribitol to Glc-AATGal-PP-upr
The lic1 region contains the two genes, tarI and tarJ, for the synthesis of activated ribitol (CDP-ribitol). The corresponding enzymes have been isolated, and their enzymatic activities were demonstrated. 3 TarJ is an NADPH-dependent alcohol dehydrogenase catalyzing the synthesis of ribitol 5-phosphate from ribulose 5-phosphate. TarI is the cytidylyl transferase for the synthesis of CDP-ribitol from ribitol 5-phosphate and CTP. The crystal structure of TarI with and without bound CDP has been solved by providing the rationale for the observed substrate specificity of the enzyme. 3 Both tarI and tarJ were shown to be essential genes under laboratory conditions.3,93
We hypothesize that CDP-ribitol is the substrate for a phosphotransferase reaction transferring ribitol-phosphate onto Glc-AATGal-PP-upr to form Rib-P-Glc-AATGal-PP-upr (P, phosphate; Fig. 2). A good candidate for the enzyme catalyzing this reaction is the hypothetical phosphotransferase Spr1125 (LicD3). This proposed role of Spr1125 in the transfer of ribitol-phosphate is backed up by genetic data: In the choline-independent mutant strain R6Cho−, the whole lic3 gene region (spr1221-spr1225) has been replaced by foreign S. oralis DNA containing TA genes, 46 directly linking the lic3 region to TA synthesis. Indeed, our bioinformatic analysis indicates that the spr1221-spr1225 genes likely represent a novel pneumococcal TA gene cluster based on the alteration of this region in the choline-independent strain, the hypothetical function of these genes from sequence similarity searches, and the structure of the TA repeating unit (Table 1). 71
Completion of the repeating unit core structure
To complete the repeating unit core structure, two GalNAc residues need to be attached to Rib-P-Glc-AATGal-PP-upr. The lic3 (spr1221-spr1225) region contains two genes, spr1123 and spr1124, encoding hypothetical glycosyltransferases, which we consider the prime candidates for the attachment of the two GalNAc residues from the UDP-GalNAc precursor. So far, none of these hypothetical glycosyltransferases has been characterized.
Uptake and activation of choline and attachment of phosphorylcholine
S. pneumoniae cannot synthesize choline and strictly depends on the presence of exogenous choline for growth. 77 Choline uptake and activation involves the products of the essential licA, licB, and licC genes, which are homologs of the choline uptake genes of Haemophilus influenzae, a Gram-negative bacterium that decorates its lipopolysaccharide with phosphocholine residues. 23 The lic genes can be deleted in different choline-independent S. pneumoniae mutant strains abrogating the incorporation of choline into the cell wall, even in the presence of exogenous choline.29,30,46 A choline-independent capsule type 2 strain unable to decorate its cell wall with choline showed drastically reduced virulence in different models of infection. 47 LicB is predicted to be an integral membrane protein with 10 transmembrane helixes 23 consistent with the membrane localization of GFP-LicB fusion proteins. 21 LicB is similar to the BetT betain permease of Escherichia coli, which, together with the just mentioned genetic evidence, suggests that LicB is the transporter for the uptake of choline. Once taken up into the cytoplasm, choline becomes phosphorylated, an activity that has been detected in pneumococcal cell fractions and which is most likely catalyzed by the presumed choline kinase LicA. 90 Phosphorylcholine is then activated to CDP-choline by the cytidylyl transferase LicC, which belongs to the nucleoside triphosphate transferase family.66,91 LicC is purified and kinetically characterized. 11 The crystal structure of LicC explains the specificity for CTP and phosphocholine and suggests a direct role for Mg2+ in the positioning of the substrates. 52 GFP-fusions of both LicA and LicC localize in the cytoplasm consistent with the absence of predicted membrane domains. 21
The CTP-activated choline is the substrate for the attachment of a phosphoryl choline moiety to each of the two GalNAc residues to the repeating unit. These reactions occur after the completion of the core structure and before the transport of the subunits across the membrane, either at the repeating unit monomers or in the polymerized chains or both. The phosphotransferases responsible for the attachment of phosphocholine residues are most likely encoded by the licD1 and licD2 genes. 94 LicD1 but not licD2 is essential, and an licD2 mutant has a reduced content of cell wall choline residues. It is impaired in the adherence to human alveolar cells and shows reduced virulence in the intraperitoneal mouse model. 94 Although LicD1 and LicD2 are predicted to be soluble, both the cytoplasmic GFP-fusion proteins are localized to the cell membrane, which is consistent with the substrate, the undecaprenyl phosphate-bound TA repeating unit or chain, being attached to the cell membrane. 21 Either the LicD proteins have a yet unidentified membrane attachment site or a binding partner or the substrate localizes them to the membrane.
Polymerization of the precursor and transport across the membrane
Polymerizing the TA chains and flipping them across the cytoplasmic membrane most likely involves integral membrane proteins. Although it is not known which step occurs first in B. subtilis and S. aureus, the TA chains are polymerized before or during the time they are transported across the membrane.75,89 Each of the lic1/lic2 and lic3 regions contains one gene predicted to encode for a highly hydrophobic membrane protein, which we assign to the two missing membrane steps in TA synthesis: Spr1222 has 10 or 11 predicted transmembrane helices and might catalyze the polymerization of the TA precursors under the release of undecaprenyl pyrophosphate (upr-PP). TacF (Spr1150) belongs to the family of transmembrane transporters with 14 predicted transmembrane helices and is most likely responsible for the transport of upr-PP-linked TA chains across the membrane. TacF is an essential gene 74 and, interestingly, a single point mutation in tacF renders strain R6Chi independent for exogenous choline 15 (see TA Pathway in Choline-Independent S. pneumoniae Strains).
Structural variation and TA modifications
The TA core structure is common for most pneumococcal isolates tested with one exception: The WTA of a serotype 5 strain contains a galactose residue instead of the usual glucose in its repeating unit. 81
There are several reports on structural variations in pneumococcal TA and additional modifications. In S. aureus and B. subtilis, TA can be modified by D-alanylation, which is catalyzed by the products of the dltXABCD genes. 61 This modification increases the number of positive charges in the cell wall, thereby contributing to the cellular resistance against cationic antimicrobial peptides. S. pneumoniae has not been known to have the D-alanine modification in its WTA or LTA.27,28 However, homologs of the dltXABCD genes have been recently identified in the pneumococcal genome. Indeed, the standard laboratory strain R6 contains a mutation in the dltA gene, resulting in a stop codon shortly after the translation start, explaining the absence of D-alanine modification in this strain. 50 Repair of the dltA mutation resulted in a strain that contained D-alanine in its cell wall. 50 D-alanine has also been shown to be bound to the ribitol OH groups in the LTA of Fp23, a nonencapsulated mutant of the capsule type 4 strain TIGR4. 19 Similar to other pathogens, the inactivation of dltA leads to increased sensitivity to cationic antimicrobial peptides. These findings suggest that pneumococci have a functional pathway for the D-alanine modification of TAs (Fig. 3). 50
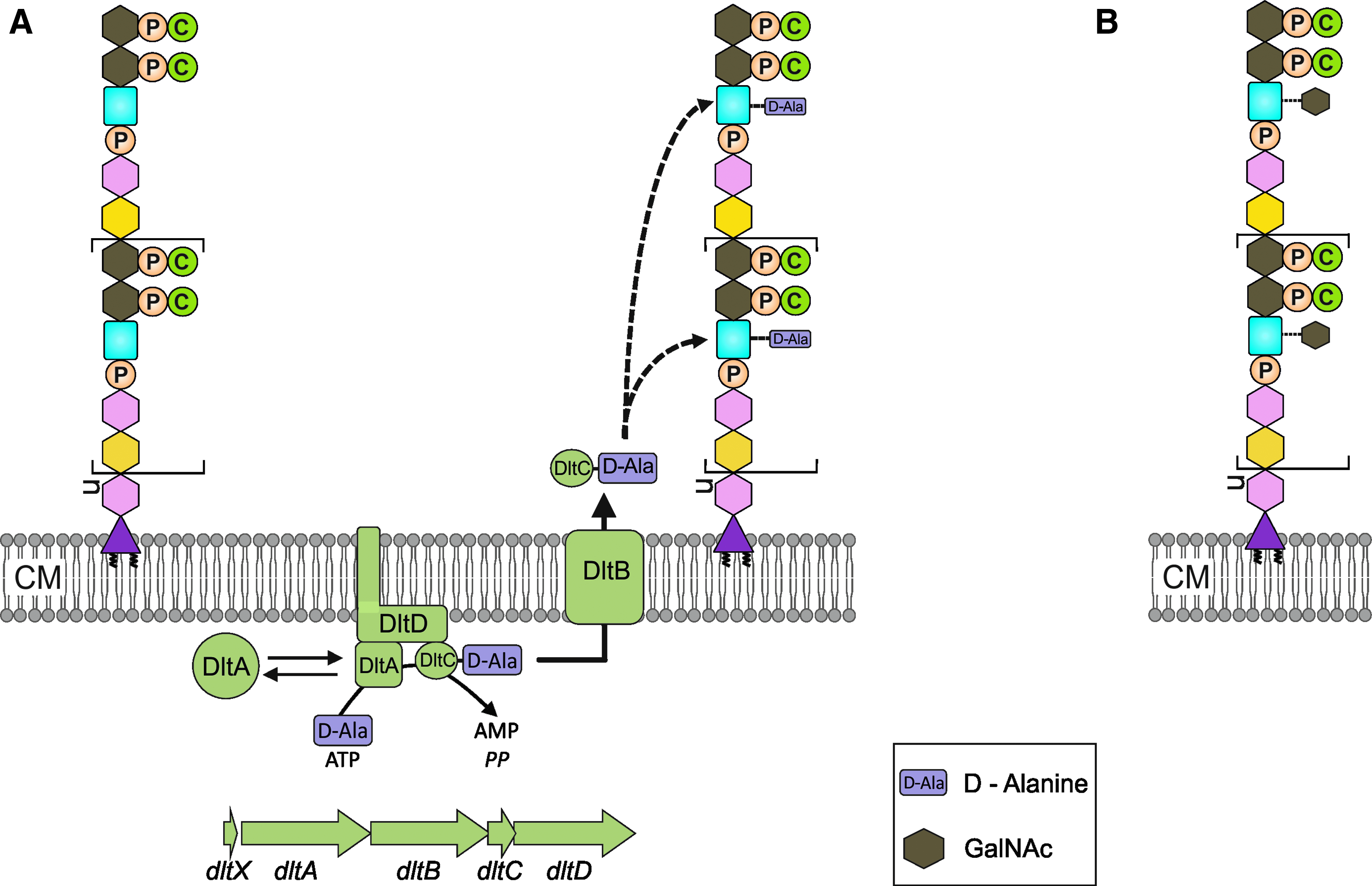
Decoration of TA with D-alanine or N-acetylgalactosamine.
Various species also attach monosaccharides to glycerol or ribitol residues in their TA. While the role of these modifications is not known, the sugar-loaded TAs can serve as specific recognition sites of certain bacteriophages and, consequently, mutants lacking these sugar residues are resistant to these phages. 2 LTA from strains R6 and Fp23 isolated with a mild butanol extraction method carried ribitol-bound GalNAc residues 19 ; obviously, this modification has previously escaped detection when chloroform-methanol was used to purify LTA. Whether or not the GalNAc residues are utilized by pneumococcus phages for cell wall attachment is not known. In addition, the enzyme responsible for this modification, the stage in TA synthesis in which GalNAc is added, and the cellular role of the GalNAc residues have remained elusive.
Synthesis of the lipid anchor of LTA
According to the revised LTA structure, 71 the lipid anchor is glucosyl-diacylglycerol (Glc-DAG), which is one of the two glycolipids present in S. pneumoniae. The second glycolipid is GalGlc-DAG. 10 A glucose residue is attached to DAG by the glycosyltransferase Spr0982, 5 and GalGlc-DAG is produced by a second glycosyltransferase CpoA, 22 the gene of which had been originally isolated as a β-lactam resistance determinant. 32
Linking TA chains to peptidoglycan and the glycolipid anchor
Unlike in many other bacteria, pneumococci have the same unusually complex structure in WTA and LTA, and we propose that the biosynthetic pathway for the synthesis of the upr-PP-linked TA chains is used for both polymers. Consistent with this view, the TA chains of WTA and LTA are chemically identical 28 and LTA is not used as a precursor of WTA, 8 indicating that the pathway branches before the last steps after the synthesis of the upr-PP-linked TA chains: In this scenario, the transfer of the TA chain to peptidoglycan (or the peptidoglycan precursor) yields WTA, and the transfer of the TA chain on the glycolipid anchor yields LTA.
A recent publication has identified members of the widespread LCP family of phosphotransferases, named after the representative members LytR, Cps2A, and Psr, as being responsible for the attachment of anionic polymers to peptidoglycan. 44 Many species, including S. pneumoniae, have several LCP proteins with sometimes redundant or semi-redundant functions. All these are anchored to the cell membrane near their N-terminus, with the typical LCP domain located outside the cytoplasm. Interestingly, the crystal structure of Cps2A shows a polyprenol (pyro)phosphate lipid tightly bound inside a hydrophobic cavity of the protein; these lipids resemble substrate or product analogs of the phosphotransfer reaction. 44 A paper in the same issue of MDR reports on the role of Cps2A and LytR in the attachment of capsular polysaccharide to peptidoglycan. 20 It is likely that the pneumococcal LCP proteins are collectively responsible for the attachment of capsular polysaccharides and of TA chains to peptidoglycan, the latter producing WTA, and of TA chains to the glycolipid anchor to form LTA. However, more work is required to decipher the role of each of the LCP proteins in these processes.
Genetic Organization and Regulation of TA Biosynthesis Genes
Figure 2 depicts the six genomic loci in which the TA biosynthetic genes are clustered. We have searched the promoter regions of these loci (Table 2) for sequence motifs in order to define putative coregulation. Inspecting 200 bp up- and 100 bp downstream of the transcriptional start sites did not help in the identification of any common regulatory motif. Therefore, it is unlikely that one common mechanism controls the expression of all TA biosynthetic genes. Instead, these appear to be organized in eight independent transcriptional units. In this section, we will review the available data on the regulation of these units.
-10, −35 regions and the TG motif are shown in gray. Binding sites for CiaR are shown in bold and underlined.
Distances are calculated from the transcriptional start site.
Transcriptional start site experimentally determined.
Transcriptional start site predicted.
Assuming a later start preceded by a Shine–Dalgarno sequence. 73
The repeat sequence for CiaR binding is on the opposite strand.
The lic1 genes are cotranscribed as demonstrated by northern blot analysis. 47 Transcription starts at two promoters P1tarI and P2tarI (Table 2) previously designated P1/2spr1149, 36 which have a similar strength as judged from promoter probe studies (Halfmann and Brückner, unpublished results). Transcription from P1tarI is strongly dependent on the response regulator CiaR and is also controlled by an as yet unknown regulator. 37 P2tarI appears to be constitutively active and alone ensures efficient lic1 operon expression regardless of the activity of P1tarI. Hence, the physiological significance of the CiaR regulation of P1tarI is not clear. Transcription from P1tarI produces an lic1 mRNA with a rather long untranslated region (199 nt) at the 5′-end, which could be an indication for post-transcriptional control. 87
The genetic organization of the lic2 locus strongly suggests that the three genes constitute an operon, and a promoter has been identified in front of tacF (Table 2). The expression of licD2 was found to be stimulated by choline starvation, 17 but the underlying regulatory mechanism has not been characterized. The tacF promoter is not regulated by CiaR, although it is located relatively close to the P1 tarI promoter of the lic1 operon.
The large lic3 TA gene cluster came into view, because it is replaced by exogenous S. oralis DNA in the choline-independent S. pneumoniae R6Cho− strain (see next) 46 The cluster of five genes is apparently cotranscribed starting at a promoter upstream of licD3 (Fig. 2 and Table 2) and ending at a transcriptional terminator behind spr1221. The last gene, spr1221, is highly conserved in S. pneumoniae and S. mitis. Its possible function in the TA pathway is not known. Spr1221 lacks sequence similarity to the proteins of known function. It has two predicted transmembrane helices flanking a large cytoplasmic loop and a C-terminal extracytoplasmic tail. The large cytoplasmic part of Spr1221 may aid the assembly of other members of the TA biosynthetic machinery to the inner side of the membrane by protein-protein interaction.
The spr1226 gene, also called psr, encodes one of the three LCP family proteins presumably involved in the attachment of TA to Glc-DAG or peptidoglycan and/or of capsular polysaccharides to peptidoglycan.20,44 Psr locates adjacent to licD3 and is the last gene of a separate 4-gene transcriptional unit expressed from a promoter in front of aroA containing no other TA gene (Fig. 2 and Table 2). Instead, the aroA, aroK, and pheA genes are required for the biosynthesis of aromatic amino acids. 58 The expression of the second lcp gene, lytR (spr1759), has been analyzed in some detail. 43 The gene is constitutively transcribed during exponential growth from a promoter that is located two genes further upstream of lytR (Fig. 2). In addition, lytR expression is enhanced during the development for genetic competence development 38 when transcription starts at the promoter of comM encoding a protein required to protect S. pneumoniae from the peptidoglycan hydrolase CbpD. 12 Thus, lytR is the last gene of the transcriptional units of three or four genes, respectively (Fig. 2). The function of the proteins encoded by spr1760 and spr1761 is not known. Inactivation of lytR could only be achieved at the expense of secondary and yet unknown genomic mutations,38,43 and the resulting lytR mutant strain showed aberrant cell division and premature lysis. 43 The third LCP gene, cps2A, is coregulated with the other capsule genes. Cps2A and LytR are required for the full expression of the cell wall anchored capsule in the capsular type 2 strain D39. 20
The spr0091-spr0092 and spr1655-spr1654 genes constitute two operons. One canonical promoter sequence is present in front of spr1655 (Table 2). Several putative promoters could be predicted upstream of spr0091 (Table 2). A degenerate CiaR binding site is present in the second promoter region, but it is not known whether it is functional. Three of the genes in these loci, spr0091, spr1654, and spr1655, are essential, and there is no information about the essentiality of spr0092.74,76
The proteins encoded in the dltXABCD gene cluster modify TA by D-alanylation. A canonical promoter structure with −35 and extended −10 regions has been mapped upstream of dltX (Table 2), 50 consistent with the recent notion that dltX is a part of the dlt operon. 49 The dltX gene is transcribed and translated as determined by translational fusions (Halfmann and Brückner, unpublished results), but no function is assigned to this small, 43 amino-acids-containing protein. The dlt operon is activated by CiaR as shown in microarray studies,14,53 although the typical features of CiaR-activated promoters, a hexamer repeat 10 bp upstream of the −10 region, are missing. 36 A CiaR-related repeat (TTTCAG-N5-TTTAAG) is found 49 bp upstream of the −10 region on the opposite DNA strand and weakly bound by CiaR in gel shift experiments. However, promoter probe experiments yielded no significant alterations of promoter activity in the absence of CiaR (Halfmann and Brückner, unpublished results). Thus, under normal conditions, the expression of dlt is independent of CiaR, and only the hyperactivation of CiaR by mutations in ciaH 55 increases the dltX promoter activity 1.4-fold. Since such a hyperactivation of CiaR has never been obtained by altered growth conditions or by imposing stresses to S. pneumoniae34,37,67 the physiological relevance of this subtle up-regulation remains unknown.
TA Pathway in Choline-Independent S. pneumoniae Strains
A unique property of S. pneumoniae is the nutritional requirement for choline, 63 which is metabolized exclusively to decorate the TA chains. Choline can be replaced by ethanolamine in the growth medium, allowing pneumococcal growth. However, ethanolamine-grown pneumococci are strongly impaired in daughter cell separation, autolysis, competence for genetic transformation, and virulence. 78 Several mutant strains of S. pneumoniae have been described that can grow in the absence of choline and ethanolamine.15,31,72,92 One set of mutants was isolated after several rounds of growth in a medium, in which choline was replaced by decreasing the concentrations of ethanolamine in each passage, finally omitting ethanolamine. The other mutants were obtained by transforming S. pneumoniae strains with DNA from S. oralis or S. mitis and selection on choline-free growth medium. The S. oralis donor strain ATCC35037 contains choline in its TA but has no auxotrophic requirement for it. 72 In contrast, the other donor, S. mitis SK598, incorporates ethanolamine in its TA even when choline is available. 7
In recent years, genome modifications present in different choline-independent pneumococcal mutants have been discovered. R6Chi, a mutant of S. pneumoniae R6 obtained by ethanolamine-depletion experiments, contains a point mutation in the putative TA flippase gene tacF, which leads to choline independency. 15 The tacF gene was also affected in another mutant isolated by serial passage, JY2190, and in the transformant P072 that has taken up S. mitis DNA. 31 In the latter study, it was suggested that two mutations in tacF improve the fitness of the mutant strain when grown in the absence of choline.
The genomic changes in the choline-independent transformant R6ho− obtained with S. oralis DNA were more dramatic. 46 The whole lic3 region including a number of genes in the neighborhood, almost 17 kb in total, was replaced by 20 kb of S. oralis DNA. A large part of this incoming DNA contained genes predicted to be involved in TA biosynthesis (see below for details). The S. oralis genes could apparently complement functions from the lost S. pneumoniae lic3 locus. Moreover, the entire lic2 operon containing the essential tacF and licD1 genes could be deleted in R6ho. 46 Therefore, the new lic region from S. oralis combines genes for functions in TA biosynthesis that are encoded in separate loci in S. pneumoniae, including the export of the TA precursor by a variant of the flippase TacF. Thus, so far, all known choline-independent S. pneumoniae strains have an altered tacF gene.
A model for the choline dependency of pneumococcal growth has been proposed. Accordingly, TacF specifically transports phosphocholine-loaded TA chains across the membrane, ensuring that the cell wall is loaded with phosphocholine residues by not transporting choline-unloaded chains. 15 Hence, choline depletion results in a block in TA synthesis, which is lethal. Presumably, the tacF point mutation(s) in R6Chi, JY2190, and P072 and the exogenous tacF variant inR6Cho− allow the transport of choline-loaded and choline-unloaded TA chains, explaining why these mutants grow normally in the presence of choline and, unlike wild type, can grow in the absence of choline. 15
TA Biosynthetic Genes in S. oralis and S. mitis
The molecular analysis of the choline-independent transformant R6Cho− allowed an identification of the TA biosynthesis genes in S. oralis. Here, we have used the recently completed genome sequences of S. oralis Uo5 64 and S. mitis B6 16 to search for TA biosynthetic genes in these species closely related to S. pneumoniae. The proteins predicted to be involved in TA biosynthesis in these organisms are listed in Table 3, and the genetic organization of the corresponding genes is shown in Fig. 4.
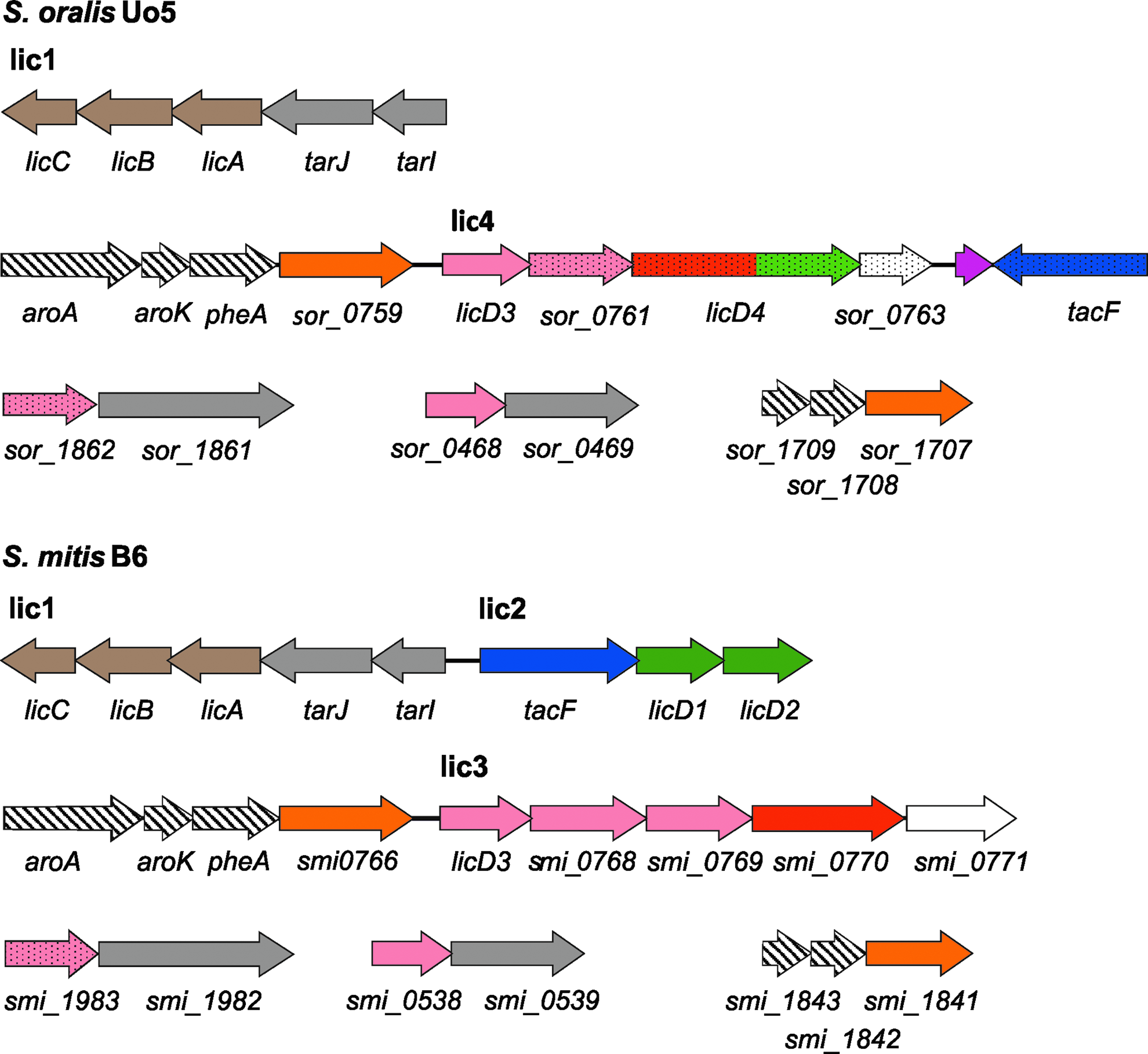
Genetic organization of TA biosynthetic genes in Streptococcus oralis Uo5 and Streptococcus mitis B6. The order of the TA loci and color codes are the same as for the Streptococcus pneumoniae loci shown in Fig. 2. The genes denoted by dotted arrows are different from S. pneumoniae.
Spr numbers represent S. pneumoniae R6.
S. pneumoniae 70585.
Homologs of the three genes proposed to encode proteins for the synthesis of AATGal-PP-upr are present in S. oralis Uo5. The gene for the glycosyltransferase Spr0091 performing the next step in TA biosynthesis, the addition of glucose to AATGal-PP-upr, is not detected. Instead, S. oralis Uo5 contains sor_1862, a glycosyl transferase gene with a high similarity to sp70585_0164 of S. pneumoniae 70585. Sp70585_0164 presumably adds galactose instead of glucose to AAT-Gal-PP-upr and, hence, we propose that the resulting TA intermediate in S. oralis Uo5 is Gal-AATGal-PP-upr. Similar to S. pneumoniae, the genes required for the initial steps in TA biosynthesis, sor_0468-sor_0469 and sor_1862-sor_1863, are clustered in two 2-gene operons.
According to our hypothesis for S. pneumoniae, a phosphotransferase is required to attach ribitol-phosphate to the TA precursor. The substrate CDP-ribitol is provided by TarI and TarJ, which are encoded in a genomic region in S. oralis Uo5 virtually identical to the lic1 operon of S. pneumoniae (Figs. 2 and 4). The predicted phosphotransferase gene licD3 begins the large lic4 TA biosynthetic operon, which was originally identified in the choline-independent transformant R6Cho− (see above). The predicted LicD3-mediated ribitol-phosphate transfer in S. oralis Uo5 results in the precursor Rib-P-Gal-AATGal-PP-upr.
The sor_0761 gene following licD3 encodes a glycosyltransferase of the pfam00535 family sharing 42% residues with Spr1223, presumably transferring one of the GalNac residues to the repeating unit. No other glycosyl transferase is present in the S. oralis lic4 locus in contrast to S. pneumoniae lic3, where a second glycosyl transferase gene (spr1124) is present. In S. oralis, either only one glycosyl residue is attached to Rib-P-Gal-AATGal-PP-upr, or Sor_0761 acts processive to add several glycosyl residues. The low-sequence similarity between Sor_0761 and Spr1223 and the absence of Spr1224 could indicate that a glycosyl residue other than GalNac is attached in S. oralis Uo5 at this step of the repeating unit biosynthesis. This hypothesis is consistent with the observation that monoclonal antibodies directed against the repeating unit of pneumococcal TA do not recognize the TA of S. oralis.6,48
The next gene, licD4, has no homolog in its entity in S. pneumoniae. It encodes a large membrane protein of 717 amino acids with two clearly defined regions. The N-terminal region is homologous to O-antigen ligase proteins from the pfam04932 family, whereas the C-terminal region has 35% to 32% sequence identity with the LicD1 and LicD2 proteins of S. pneumoniae. Consequently, this protein should serve two functions. The N-terminal region predicted to have 12 transmembrane helices might catalyze the polymerization of TA precursors, and the C-terminal LicD-type region could add phosphorylcholine. The substrate for this latter reaction, CTP-activated choline, is provided by proteins in the lic1 locus. Thus, S. oralis Uo5 harbors all the genes required for choline incorporation, consistent with the presence of phosphorylcholine in its TA.6,41 A second protein with an LicD domain is encoded downstream of licD4 and may add another phosphorylcholine residue to the repeating unit. The last gene in this operon is a truncated version of the phosphocholine esterase gene pce. 84
The tacF gene encoding the TA flippase is located opposite to the licD3-sor_0761-licD4 operon (Fig. 4). The S. oralis TacF protein shares 47% of the residues with TacF of S. pneumoniae. This divergence may be a consequence of the different TA repeating unit structures in both organisms.
The S. oralis Uo5 lic4 TA locus combines functions encoded in lic2 and lic3 in S. pneumoniae. Lic4 has replaced lic3 in the choline-independent S. pneumoniae R6Cho− introducing the S. oralis allelles for TA flippase (TacF) and functions for choline decoration, which explains that the lic2 locus harboring these functions becomes dispensable in R6Cho−. In this transformation experiment, DNA from S. oralis strain ATCC35037 was used, in which the essential TA genes are virtually identical to those of S. oralis Uo5. In addition to the TA genes, the lic4 region of strain ATCC35037 encodes a phosphorylcholine esterase (Pce) and a small protein of unknown function. 46 If S. pneumoniae and S. oralis indeed produce structurally different TA repeating units, the TA of R6Cho− should have a rather unique, “mixed” structure in which the repeating unit starts with Rib-P-Glc-AATGal, similar to S. pneumoniae, but the next residues should be specific for S. oralis.
The attachment of exported TA precursors to peptidoglycan or glycolipid anchor is predicted to rely on LCP family proteins, especially on Spr1226 and Spr1759, which are present in all strains of S. pneumoniae. Both proteins are encoded in S. oralis Uo5, the deduced proteins showing an identity of about 80%.
The TA gene content and their genetic organization are nearly identical in S. mitis B6 and S. pneumoniae R6, except for one gene. The smi1983 gene located in the operon equivalent to spr0091-spr0092 encodes a glycosyltransferase very similar to S. pneumoniae Sp70585_0164 70585 and to S. oralis Sor_1862. Therefore, S. mitis may contain galactose instead of glucose in its TA repeating unit. The structure of the repeating unit of another S. mitis strain, SK137, has been determined and is identical to that of S. pneumoniae. 6 No genomic information is available for this strain, but it could well be that the incorporation of glucose or galactose is a variable trait in S. mitis. The vast majority of S. mitis strains were recognized with monoclonal antibodies similar to the repeating unit of S. pneumoniae and to phosphorylcholine,6,48 corroborating the genetic analysis of S. mitis B6. Some S. mitis strains contained choline but had apparently another TA repeating unit structure, while others did not react with any of the antibodies. 6 Searching unfinished genomes available at NCBI revealed one S. mitis strain harboring TA biosynthetic genes just described for S. oralis Uo5. Therefore, S. mitis may contain S. pneumoniae-type or S. oralis-type TA or perhaps completely different molecules.
A Polyglycerolphosphate-Containing LTA in S. mitis?
Many Gram-positive bacteria have a much simpler LTA structure than the S. pneumoniae consisting of a polyglycerolphosphate (PGP) chain attached to a glycolipid for membrane anchoring. 25 S. aureus uses a single LTA synthase, designated LtaS, to polymerize PGP chains. LtaS and its homologs are membrane proteins with a large extracellular enzymatic domain, consistent with LTA synthesis occurring outside of the cell.62,64
Surprisingly, BlastP searches revealed a homolog of S. aureus LtaS in the S. mitis B6 genome. The similarity between LtaS (SAV0719) and Smi0753 covers the whole protein, showing 38% identity and 57% similarity. Smi0753 is predicted to be a membrane protein with a C-terminal extracytoplasmic domain and belongs to the PF00884 sulfatase family. Searching databases with Smi0753 yielded highly similar proteins in streptococcal species known to possess a PGP-type LTA. In contrast, LtaS homologs are not present in S. pneumoniae and S. oralis Uo5. The presence of an LtaS homolog in S. mitis B6 is consistent with the previous detection of a PGP-type LTA in culture supernatants of some S. mitis strains, 57 an observation that, however, was questioned in a later study. 40
The identification of an LtaS homolog in S. mitis B6 raises a number of questions. Does this species have two different LTA molecules, a conventional PGP-type LTA produced by the LtaS-like protein Smi0753 and a more complex LTA produced by the homologs of the pneumococcus-type TA pathway (Fig. 4 and Table 3)? If the answer is yes, then are the different LTA chains anchored to the same glycolipid and are they present concomitantly, and is one or the other synthesized under certain conditions? These questions should be experimentally addressed using strains with known genomic sequences.
Concluding Remarks
High-throughput sequencing has produced a wealth of genome sequences of S. pneumoniae and closely related species that contain WTA and LTA of an unusually complex chemical structure. Our bioinformatic analysis together with the phenotypic analysis of different choline-independent pneumococcal mutant strains has allowed us, for the first time, to deduce the complete pathway of TA synthesis in S. pneumoniae. Our analysis predicts so far unknown genes to be involved in the synthesis of the TA subunit, and we predict deviations form the pathway in certain pneumococcal strains and in S. oralis and S. mitis. These predictions will further guide biochemical work on the formation of a complex TA.
Footnotes
Acknowledgments
The authors would like to thank Anette Schedler for help in transcriptional mapping and Alexander Halfmann for CiaRH regulatory work. Work in Kaiserslautern was funded by the Deutsche Forschungsgemeinschaft (DFG; HA 1011/13-1). WV was funded by the Biotechnology and Biological Sciences Research Council (BBSRC).
Disclosure Statement
No competing financial interests exist.
