Abstract
Increasing antibiotic resistance of bacterial pathogens has drawn the attention to the potential use of bacteriophage endolysins as alternative antibacterial agents. Here we have identified, characterized, and studied the lytic potential of two endolysins, Lys168 and Lys170, from phages infecting Enterococcus faecalis. Lys168 and Lys170 belong to the cysteine, histidine-dependent amidohydrolases/peptidases (CHAP) and amidase-2 protein families, respectively. Lys168 is quite a unique enterococcal phage endolysin. It shares 95% amino acidic identity with the endolysin of Staphylococcus aureus phage SAP6, which in turn is distantly related to all known CHAP endolysins of S. aureus phages. Lys170 seems to be a natural chimera assembling catalytic and cell-wall-binding domains of different origin. Both endolysins showed a clear preference to act against E. faecalis and they were able to lyse a high proportion of clinical isolates of this species. Specifically, Lys168 and Lys170 lysed more than 70% and 90% of the tested isolates, respectively, which included a panel of diverse and typed strains representative of highly prevalent clonal complexes. Lys170 was active against all tested E. faecalis VRE strains. The quasi specificity toward E. faecalis is discussed considering the nature of the enzymes' functional domains and the structure of the cell wall peptidoglycan.
Introduction
Enterococci belong to the normal bacterial flora of the intestinal tract of humans and several animals and can be found in environmental soil, water, plants, and food. Although they are considered commensal bacteria, at least Enterococcus faecalis and Enterococcus faecium species are regarded as relevant opportunistic pathogens, being associated with nosocomial, and, to a lesser extent, community-acquired infections. Typical enterococcal infections occur in hospitalized patients with underlying conditions. Both species have been described as the second most common cause of wound and urinary tract infections and the third most common cause of bacteraemia, 43 and can also be involved in neonatal sepsis, 32 peritonitis, device-related infections, and endocarditis.9,14,42 The massive use of antibiotics in human healthcare systems and animal production has increased the incidence of antibiotic-resistant enterococci, 37 some of which are already intrinsically resistant to a broad range of antibiotics, including cephalosporins, sulphonamides, and low concentrations of aminoglycosides. 15 In the last decades, there has been a dramatic increase of E. faecalis and E. faecium infections due to resistant strains to vancomycin (VRE), for long considered the last resource when all other classes of antibiotics failed; therefore, the search for alternative antibacterials to combat these pathogens has become an immediate need.
Enzybiotics are an example of new potential antibacterials; among these, bacteriophage endolysins have been one of the most intensively explored.10,13,30 Endolysins are enzymes encoded by double-stranded DNA bacteriophages that cleave the bacterial cell wall peptidoglycan. This activity is essential to promote bacterial host cell lysis at the end of the phage life cycle, thus allowing an efficient escape of the viral progeny from infected cells. 40 The vast majority of known endolysins from phages infecting Gram-positive bacteria feature a well-conserved domain architecture, in which the N-terminal region carries one or two enzymatically active catalytic domains (CD) and the C-terminus motifs responsible for cell wall binding (CWBD). 12 These enzymes are designed to attack one or two of five major bonds in the peptidoglycan network. 23 The rationale behind utilization of endolysins as antibacterial agents is that, in principle, they should retain their lytic potential when added exogenously as recombinant enzymes.
Three different E. faecalis phage endolysins, belonging to two amidase families, have been reported before and their killing efficacy toward Enterococcus studied in vitro. These are PlyV12, encoded by phage Φ1, 51 EFAL-1 produced by phage EFAP-1, 46 and ORF9 from phage φEF24C. 48 In addition to the capacity to lyse their natural target, E. faecalis, the enzymes were also reported to act on the related species E. faecium. Moreover, EFAL-1 could also lyse some streptococcal isolates, whereas PlyV12 showed the broadest lytic spectrum by also acting against several streptococcal and staphylococcal strains. 51
In this study we have identified, produced, and purified two phage endolysins, Lys168 and Lys170, encoded in the genome of two E. faecalis phages, F168/08 and F170/08, respectively. Lys168 represents a novel endolysin among enterococcal phages as it carries a CD from the CHAP family. We have studied the lytic action of both endolysins against different Gram-positive pathogenic bacteria, which included a panel with representatives of the most prevalent VRE clonal complexes in nosocomial infections. The results obtained with Lys170 call for a reappraisal of those obtained with ORF9, since these two endolysins are virtually identical.
Materials and Methods
Bacteria, phages, culture media, and growth conditions
The Escherichia coli cloning strain XL1-Blue MRF′ and its derivatives were grown at 37°C with aeration in a Luria-Bertani (LB) medium. 39 The E. coli expression strain CG61 41 and its derivatives were grown in LB in the same conditions, except that the incubation temperature was 28°C before the induction of protein production and 37°C afterward. When appropriate, the LB medium was supplemented with kanamycin (30 μg/ml) and/or ampicillin (100 μg/ml) for plasmid selection.
The lytic action of enterococcal phage endolysins was assayed in 193 bacterial clinical isolates (Table 1 and Supplementary Tables S1, S3, and S5; Supplementary Data are available online at www.liebertonline.com/mdr). Table 1 lists a panel of 28 E. faecalis and 21 E. faecium typed strains recovered from patients of a Portuguese hospital between 2004 and 2006 27 (see Supplementary Table S3 for a detailed description of these strains). Table 1 also includes the two-model E. faecalis VRE strains V583 and MMH594. Supplementary Table S1 corresponds to 99 clinical isolates from the Technophage's collection, 73 E. faecalis and 26 E. faecium, which were obtained from different Portuguese community and hospital settings between 2005 and 2007. The lytic action of recombinant enzymes was also tested in clinical isolates of other bacterial species from the Technophage's collection, namely, against Streptococcus pneumoniae (n=10), Streptococcus pyogenes (n=8), Streptococcus agalactiae (n=8), Staphylococcus aureus (n=9), Staphylococcus haemolyticus (n=4), and Staphylococcus epidermidis (n=4) (Supplementary Table S5).
PFGE, pulsed field gel electrophoresis; NA, not applied.
The growth media for these bacteria were purchased from Biokar Diagnostics. Enterococcal and staphylococcal strains were cultured in either Brain Heart Infusion or Tryptic Soy Broth, whereas streptococci were propagated in the Todd Hewitt Yeast broth. Liquid cultures of Enterococcus and Streptococcus species were grown at 30°C and/or 37°C, without aeration, while those of Staphylococcus were incubated at 37°C with aeration.
When necessary, culture media were supplemented with 1.5% or 0.7% agar to obtain solid or soft-agar plates, respectively. E. faecalis phages were isolated, purified, and propagated by standard methods,6,18 in either soft-agar media or liquid broth supplemented with CaCl2 and MgCl2 (5 mM each). Phage F168/08 and F170/08 propagation hosts were E. faecalis clinical isolates 1518/05 and 926/05, respectively. (Supplementary Table S1).
Identification and bioinformatics analysis of phage endolysins
Genomes from E. faecalis phages F168/08 and F170/08 were extracted from CsCl-purified lysates 49 and their complete nucleotide sequence was determined (service purchased to Macrogen). DNA homology searches were carried out with BLASTN, 52 using the NCBI's nonredundant nucleotide sequences database. Phage putative genes were recognized by integrating the results obtained with GeneMark.hmm and MetaGeneAnnotator web software.3,29 Identification of F168/08 and F170/08 endolysin genes was based on BLASTP homology searches 1 with deduced gene products, against the NCBI's nonredundant protein sequence database, and on prediction of protein functional domains using NCBI's CDD 26 and Pfam (http://pfam.sanger.ac.uk/). Putative linkers connecting protein functional domains were assigned with SVM, 8 using the SVM-joint output. Multiple protein sequence alignments were performed with ClustalW2. 20
Cloning of Lys168 and Lys170 endolysin genes
The coding sequence of endolysins Lys168 and Lys170 was polymerase chain reaction (PCR)-amplified from phage DNA using a high-fidelity Pfu DNA Polymerase (Fermentas Molecular Biology Tools, Thermo Scientific). The forward and reverse primers used to amplify lys168 carried at their 5′ end, the restriction sites NcoI and XmaI, respectively, whereas the corresponding primers for lys170 amplification carried BspI and XmaI sites. Both products were purified using the High Pure PCR Product Purification Kit (Roche Applied Science), double-digested with the appropriate restriction enzymes, and ligated to the pIVEX2.3d expression vector (Roche Applied Science), which had been previously restricted with NcoI and XmaI. This vector is designed to drive the expression of cloned genes under the control of the phage T7 φ10 promoter and to allow the production of the corresponding proteins C-terminally fused to a hexahistidine tag. Ligations were used to transform the E. coli strain XL1-Blue MRF′ as previously described. 5 Transformants were selected in the presence of 100 (μg/ml ampicillin and screened for the presence of the desired recombinant plasmids by PCR using insert and vector complementary primers. Plasmid DNA from positive clones was extracted (Pure Link Quick Plasmid Miniprep Kit; Invitrogen), and the correct DNA structure was confirmed by endonuclease restriction and DNA sequencing (Macrogen). The constructs pDP1 and pDP2 are pIVEX2.3d derivatives carrying lys168 and lys170, respectively.
Production and purification of the endolysins Lys168 and Lys170
E. coli strain CG61, which overproduces the phage T7 RNA polymerase upon temperate upshift, 41 was transformed with plasmids pDP1 and pDP2, and the transformants were selected at 28°C in the presence of 100 (μg/ml ampicillin and 30 (μg/ml kanamycin. The ability of CG61 derivatives to produce soluble and active Lys168 and Lys170 was first checked by their culturing over a dense lawn of autoclavated enterococcal cells, incorporated in a soft-agar LB medium, and confirming the presence of lysis halos around E. coli colonies (Supplementary Fig. S2).
Selected clones of each endolysin were grown at 28°C until an optical density at 600 nm (OD600) of 0.3–0.5, after which protein production was induced by moving the cultures to a shaking water bath set to 42°C. After 45-min induction, the cultures were transferred to an incubator at 37°C and agitated for an additional 3 h. The cells from induced cultures were pelleted by centrifugation (8,000 g, 30 min, 4°C) and resuspended in 1/50 volume of a lysis buffer (20 mM Hepes-Na, 500 mM NaCl, 20 mM imidazole, 1% glycerol, and 1 mM dithiothreitol [DTT], pH 8.0) supplemented with 1× Complete Mini EDTA-free Protease Inhibitor Cocktail (Roche Applied Science). The cells were kept on ice and disrupted by sonication (Vibra Cell MS2T; Sonic Materials) by performing about 10 bursts of 1 min (amplitude 5, pulse 3, 30–40 W) intercalated with pauses of 1 min. Insoluble material was sedimented by centrifugation (10,000 g, 30 min, 4°C). The supernatant corresponding to the total soluble protein extract was filtered through a 0.22 μm filter, and endolysins were purified by affinity chromatography using HisTrap™ HP columns (GE Healthcare) coupled to an AKTA-Prime system (GE Healthcare). The column and elution buffers had the same composition of the lysis buffer, except that the imidazole concentration in the elution buffer was 500 mM. The eluted fractions were analyzed by sodium dodecyl sulfate–polyacrylamide gel electrophoresis and Coomassie blue staining. 19 Endolysins from pure fractions were pooled, concentrated, and changed to an imidazole-free, phosphate-based endolysin buffer (50 mM phosphate-Na, 500 mM NaCl, 25% glycerol, and 1 mM DTT, pH 8.0) using HiTrap Desalting columns (GE Healthcare). Protein concentrations were determined by the Bradford method (Bio-Rad Laboratories) using bovine serum albumin as the standard. The enzymes were divided in small aliquots and kept at −80°C.
Evaluation of endolysin lytic action against bacterial pathogens
The capability of endolysins Lys168 and Lys170 to induce lysis of clinical strains from different bacterial species was evaluated by two different assays. The endolysins were tested against a large number of bacterial isolates by spotting different enzyme quantities in dense lawns of viable target cells, which were prepared as follows. Enterococcal and streptococcal strains were cultured overnight at 30°C without aeration. Typically, these cultures reached an OD600 of about 0.8–1.0. Staphylococcal cultures at this OD600 were prepared after 1:200 dilution of overnight cultures and growth at 37°C with aeration. Cells from these cultures were recovered by centrifugation and resuspended in 1/100 volumes of the correspondent growth medium. A 300-μl sample of these cell suspensions was incorporated in a lysis assay buffer (25 mM phosphate-Na, 250 mM NaCl, 1% glycerol, and 1 mM DTT, pH 8.0), supplemented with 0.7% agar and poured in a Petri dish. Four protein quantities of each endolysin (5, 1, 0.2, and 0.04 μg, in 10 (μl final volume) were spotted on each strain lawn and, after an overnight incubation at 37°C checked for the presence of lysis halos. These were evaluated and scored (− to +++) according to their relative diameter and transparency (Supplementary Fig. S3).
Bacterial cell lysis was also studied in liquid medium. Selected strains were grown until an OD600 of 0.3–0.4, centrifuged, and the cells were recovered in 1/2 volume of lysis assay buffer. Cell suspensions were challenged with the indicated endolysin concentrations and the OD600 variation followed over time. At the end of each assay the surviving colony forming units (CFU)/milliliter was determined. Negative controls were equally prepared, except that endolysin buffer was added instead of endolysin.
Identification of bacterial species
When necessary, discrimination between E. faecalis and E. faecium was performed by a PCR-based approach using species-specific primers targeting the ddl gene. Primers for E. faecalis were CACCTGAAGAAACAGGC (forward) and ATGGCTACTTCAATTTCACG (reverse), with an amplicon size of 475 bp. 7 For E. faecium, the amplicon size was 1091 bp using primers GAGTAAATCACTGAACGA (forward) and CGCTGATGGTATCGATTCAT (reverse). 17 For identification purposes, Enterococcus type strains obtained from the Deutsch Sammlung von Mikroorganismen and Zellkulturen collection (DSMZ; Braunschweig, Germany) were used as references, namely, Enterococcus faecalis DSM 20478 and Enterococcus faecium DSM 20477.
Results
Bioinformatics of enterococcal phage endolysins Lys168 and Lys170
We have recently determined the nucleotide sequence of the genome of two E. faecalis phages from the Technophage's collection, F168/08 and F170/08. Sequence analysis by bioinformatics tools identified an open reading frame in each phage genome, whose deduced amino acid sequences had a high sequence identity with known or putative phage endolysins, and which featured conserved domains involved in the hydrolysis of the bacterial cell wall peptidoglycan. Therefore, these proteins were assigned as the endolysins of phages F168/08 and F170/08 and were designated as Lys168 and Lys170, respectively.
Lys170 is basically identical to the previously described endolysin ORF9 of E. faecalis phage φEF24C,47,48 showing a single amino acid substitution over its 289 amino acid sequence. Both the enzymes carry in their amino terminal region a CD of the amidase-2 family (Fig. 1A and Supplementary Fig. S1), whose members include zinc amidases that have N-acetylmuramoyl-L-alanine amidase activity. 4 This type of activity was confirmed experimentally for ORF9. 48 Lys170 (as well as ORF9) appears to be a natural chimera of intergeneric origin since its N-terminal CD is highly similar to that of lactobacilli amidases, whereas its C-terminal region, probably containing the CWBD, reveals a high sequence identity to that of enterococcal amidases (Fig. 1B).
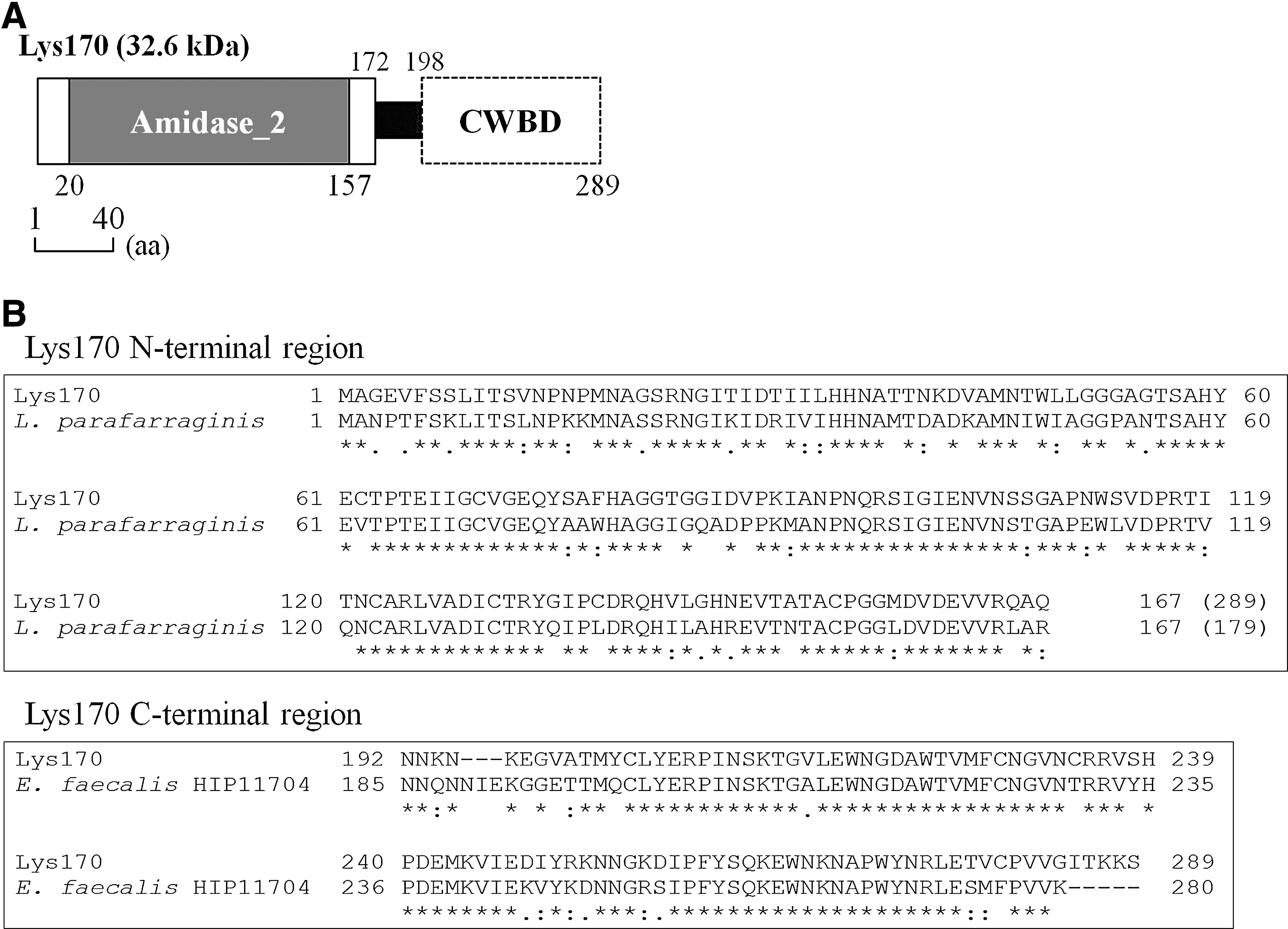
Domain architecture and sequence relatedness of Lys170.
In silico analysis of Lys168 identified a conserved domain of the CHAP family2,21,35 (Fig. 2A and Supplementary Fig. S1) in the first half of the protein. This protein family includes enzymes that cleave different amide bonds in the peptidoglycan network. Unexpectedly, Lys168 shared 95% identity with a protein assigned as “amidase” from Staphylococcus aureus phage SAP6 (GenBank AEM24735.1). In addition, the F168/08 genome shared between 80% and 94% sequence identity over 68% of the SAP6 genome (BLASTN analysis), which translated into a high sequence similarity between the products encoded by the homologous portions of both the genomes. In addition to its close relationship to the SAP6 endolysin, the Lys168 CD shared a significant identity with the CHAP domain of a single E. faecalis protein (strain TX0104) and with that of other S. aureus phage proteins (Fig. 2B). The latter, however, are ∼600aa multifunctional proteins associated with the virion structure and are thought to assist DNA entry into the host cells at the initial steps of infection.34,36 Lys168 C-terminal region had no equivalent homologs besides that of the already-mentioned endolysin from phage SAP6.
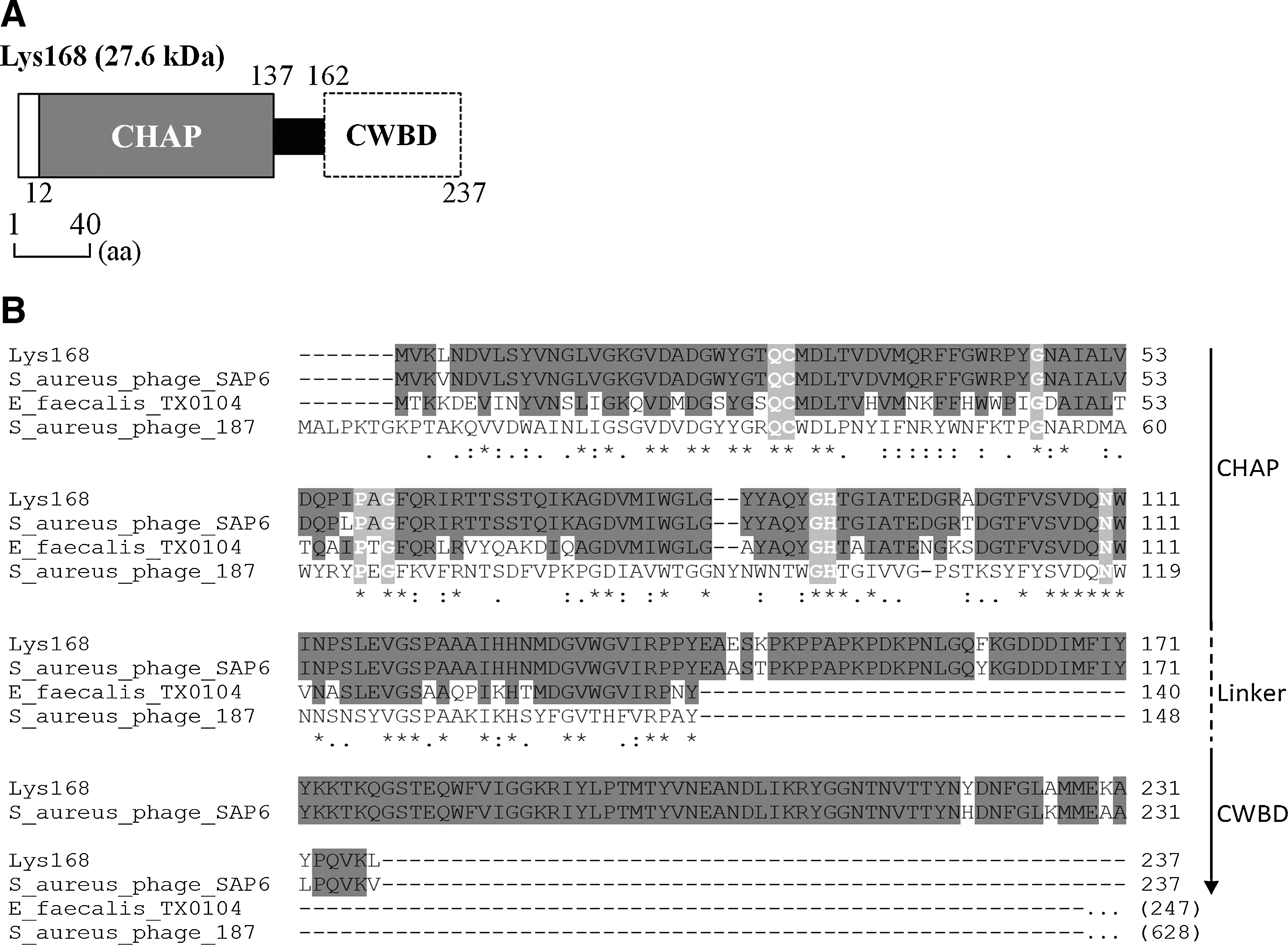
Domain architecture and sequence relatedness of Lys168.
Endolysins from phages infecting Gram-positive bacteria display a typical domain architecture in which the N-terminal CD and C-terminal CWBD are connected by a linker sequence. 12 Although the CDs of Lys168 and Lys170 could be delimited in their N-terminal portions using the bioinformatics tools (see above), these failed to recognize any known CWBD in their C-terminal regions. We could, however, predict the location of the central linker domain in each endolysin; based on this, we inferred the probable position of CWBD (Figs. 1A, 2A and Supplementary Fig. S1).
Heterologous production and purification of endolysins Lys168 and Lys170
The genes encoding Lys168 and Lys170 were PCR-amplified and cloned in the E. coli expression vector pIVEX2.3d, which allowed the production of the endolysins C-terminally fused with a hexahistidine tail (see Materials and Methods). E. coli clones producing the enzymes in their active form were initially selected by growing transformants on a dense lawn of autoclavated E. faecalis target cells and checking for the presence of lysis halos around the E. coli colonies (Supplementary Fig. S2). Medium-scale protein production from the selected clones allowed us to obtain substantial amounts of soluble Lys168 and Lys170 with the expected molecular weight, which were subsequently purified by affinity chromatography using nickel columns. Endolysins of pure fractions from the affinity chromatography were changed to an imidazole-free, sodium phosphate-based buffer by performing a desalting step (Fig. 3).
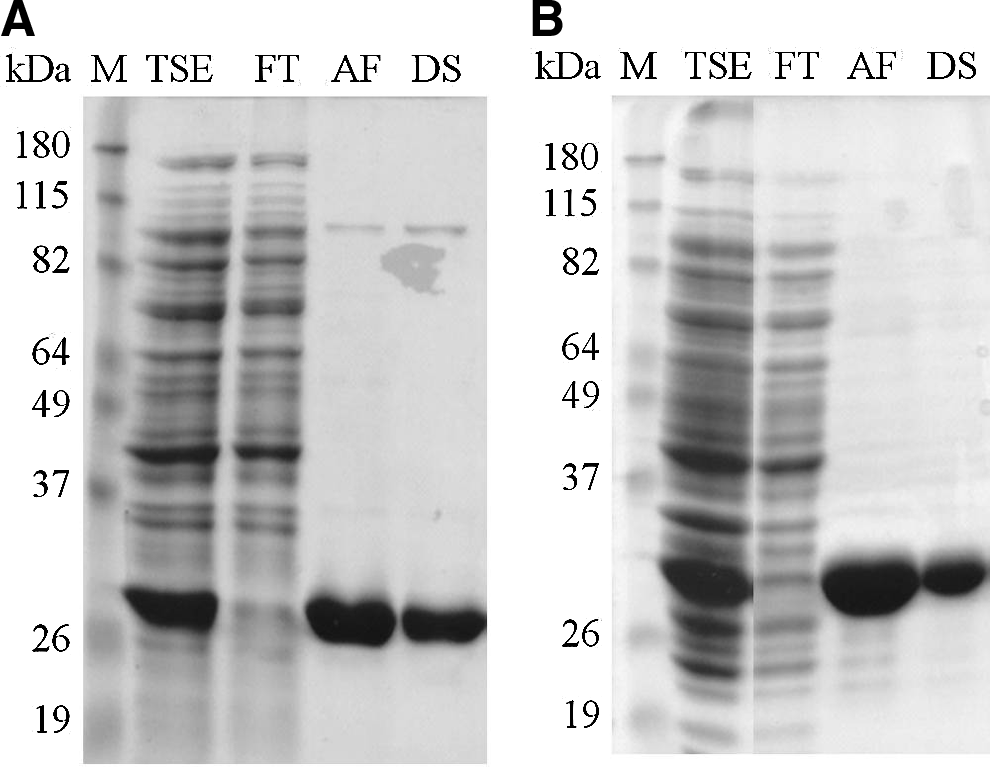
Analysis of endolysins Lys168
Lytic action of Lys168 and Lys170 against enterococcal clinical strains
In a preliminary assay we assessed the lytic action of purified Lys168 and Lys170 against a panel of enterococcal clinical isolates from the Technophage's collection, which were provided by different Portuguese clinical settings and isolated from different infection contexts. This panel was composed of 73 E. faecalis and 26 E. faecium isolates (Supplementary Table S1). Four different amounts of each endolysin (5, 1, 0.2, and 0.04 μg) were spotted on a dense lawn of viable cells from each isolate, which was produced by incorporating the cells from exponentially growing cultures in a soft-agar, phosphate-buffered medium (see Material and Methods). Lytic activity was qualitatively evaluated by scoring the relative diameter and turbidity of the lysis halos produced after overnight incubation at 37°C (Supplementary Fig. S3).
When applied in its highest quantity (5 μg), Lys170 produced a discernible lysis halo in 97% and 54% of E. faecalis and E. faecium isolates, respectively, whereas Lys168 lysed 81% and 42% of these. When we scored the percentage of susceptible isolates for the lower amounts of each endolysin, it became clear that Lys170 had a higher lytic action than Lys168 (Supplementary Fig. S4). In addition, for each tested enzyme quantity, Lys170 almost always produced clearer and larger lysis halos than Lys168 (Supplementary Table S2). The results from this preliminary study indicated that Lys170 had a better lytic performance than Lys168 and suggested that both endolysins preferentially lysed E. faecalis strains.
The isolates from the panel referred to above were not typed, and thus the diversity within each Enterococcus species was unknown. To gain more insight on the lytic potential of each endolysin against these enterococcal species, the enzymes were equally assayed in a panel of 51 multi-resistant typed strains, 30 E. faecalis, and 21 E. faecium (Table 1 and Supplementary Table S3), 49 of which were the cause of infections in a Portuguese hospital, over a 3-year period. These strains displayed a high-level resistance to gentamicin and included VREs of clonal complexes E. faecalis-CC2 and E. faecium-CC17, which are highly prevalent in nosocomial settings and disseminated worldwide. 27
We observed that 5 μg of Lys168 and Lys170 was still able to induce lysis of more than 70% and 90% of the E. faecalis strains, respectively, but only up to 10% of the E. faecium strains were susceptible to the endolysins. The percentage of lysed strains decreased just slightly when the quantity of the applied Lys170 was lowered to 0.04 μg. In contrast, this percentage was significantly diminished when Lys168 quantity dropped to 0.2 and 0.04 μg (Fig. 4). As described above, Lys170 produced clearer and larger lysis halos than Lys168 (Supplementary Table S4). These results confirmed the highest lytic capacity of Lys170 and the clear preference of both endolysins toward E. faecalis when compared to E. faecium.
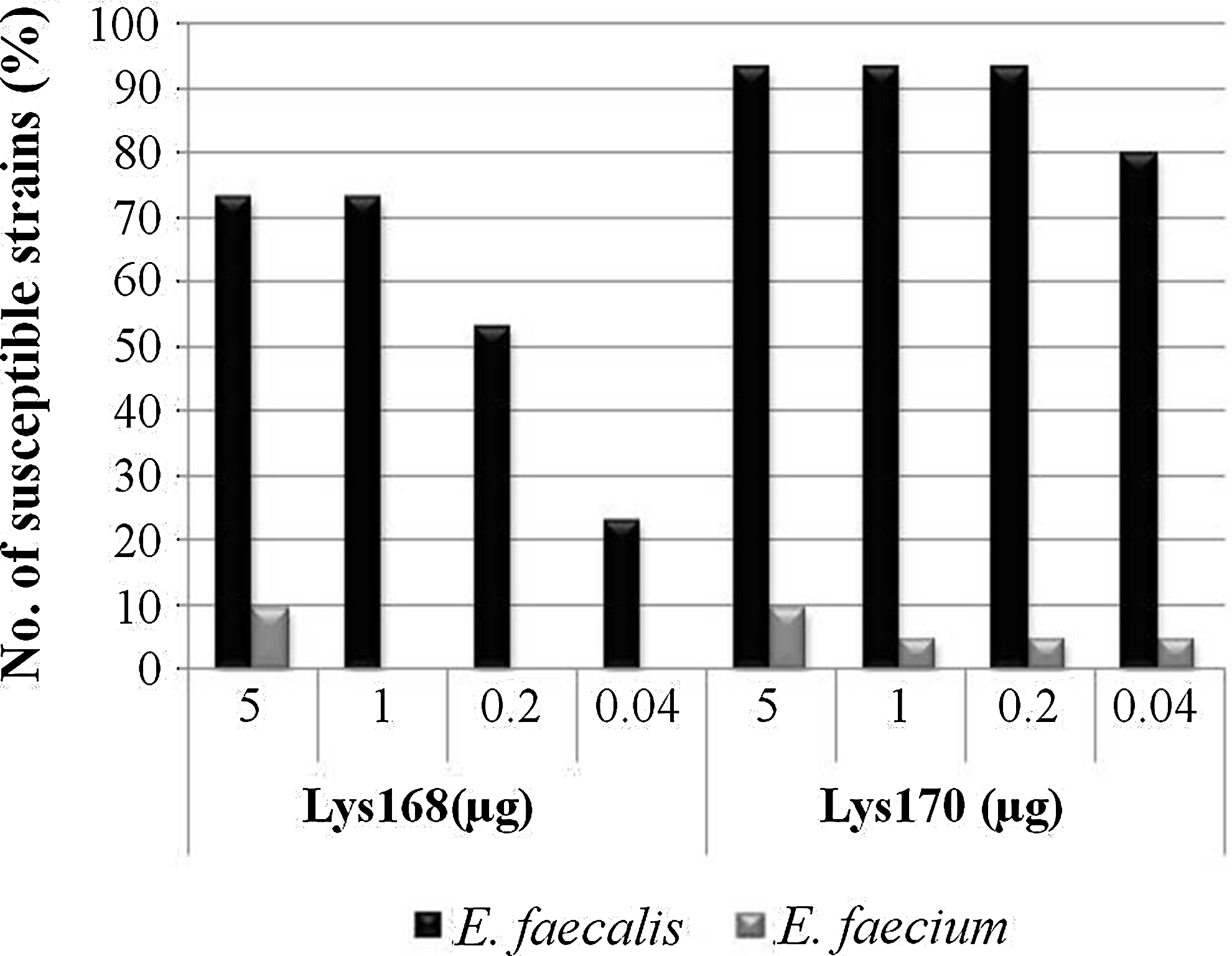
Lytic action of Lys168 and Lys170 against a panel of diverse, typed clinical strains of E. faecalis (n=30) and Enterococcus faecium (n=21). The percentage of strains that presented lysis halos is plotted as a function of each endolysin quantity.
Lytic action of Lys168 and Lys170 against E. faecalis in liquid medium
The enterococcal endolysins also induced lysis of dense suspensions of viable E. faecalis cells prepared from exponentially growing cultures. The examples of Figure 5 show the lytic action of both endolysins against two target strains, one that was only susceptible to Lys170 in the spot assay (see above), the E. faecalis VRE strain V583 (Fig. 5A), and the other that was similarly lysed by both endolysins, the E. faecalis strain 1915/05 (Fig. 5B). The VRE strain V583 was challenged with 5 μg/ml of each endolysin or with a mixture of both enzymes, each at a concentration of 2.5 μg/ml (Fig. 5A). As expected, Lys170 induced fast and extensive cell lysis, with the OD600 of the suspensions decreasing to about 10% of the initial value within 30 min. At the end of the assay (t=90 min), the CFU/ml dropped to 1% of the initial value. Interestingly, although V583 seemed to be resistant to Lys168 in the spot assays, in the liquid medium this endolysin could still produce a rather gradual cell lysis, leading to a 60% reduction of the initial OD600 and to ∼80% cell killing during the time course of the assay. No significant synergistic effect was observed when the cell suspensions were treated with a mixture of both the enzymes, as the lysis profile and loss of cell viability were very similar to those observed with Lys170 alone.
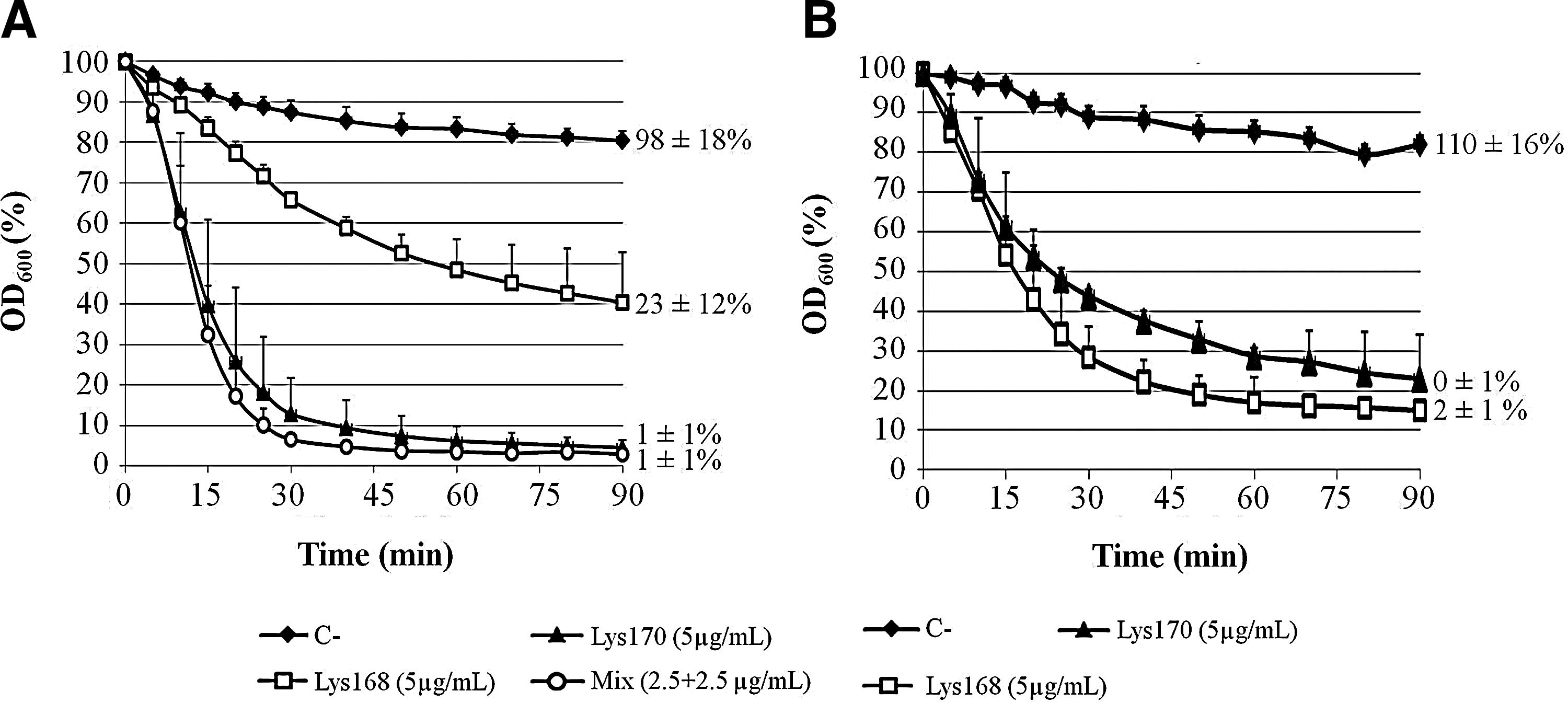
Lytic action of Lys168 and Lys170 in a turbidity assay using E. faecalis strains V583
The apparent similar efficacy of Lys168 and Lys170 in lysing strain 1915/05 in a soft-agar medium basically correlated with lysis induced by each endolysin in the liquid medium (Fig. 5B). Both the enzymes produced similar lysis curves, with the OD600 decreasing to about 20% of the initial after 90 min, although in this case Lys168 seemed to induce slightly faster and more extensive lysis than Lys170. Both endolysins were capable of killing ≥98% of the initial CFU/ml by the end of the assay.
Overall, we observed that the lysis profile of a particular E. faecalis strain when challenged in the liquid medium with the enterococcal endolysins, essentially correlated with the lysis efficiencies observed in the spot assay.
Activity of enterococcal endolysins against other Gram-positive pathogenic bacteria
The lytic activity of Lys168 and Lys170 was also evaluated in a few clinical isolates of other common Gram-positive pathogenic cocci (Supplementary Table S5) by performing the enzyme spot assay as described above. No obvious lysis halo could be discernible in any of the tested isolates even for the highest protein amount spotted (5 μg). The results suggest that Lys168 and Lys170 are evolutionarily designed to specifically act against Enterococcus species, particularly E. faecalis, if we consider the results described above.
Discussion
In this work we have characterized two endolysins, Lys168 and Lys170, from two phages infecting E. faecalis and have evaluated their bacterial cell lysis activity. As far as we know, only three additional E. faecalis phage endolysins have been described in the literature, PlyV12, EFAL-1, and ORF9.46,48,51 ORF9 is virtually identical to Lys170, and thus it will be omitted from this discussion, except in the part where we compare the lytic spectrum we obtained with Lys170 with that reported for ORF9 (see below).
Analysis of the primary sequence of these endolysins uncovered interesting features. The enzymes are clear examples of modular architecture by assembling different CDs and CWBDs of heterologous origin, thus generating endolysin diversity (Fig. 6). In fact, the four distinct endolysins referred to above have distantly related amino acid sequences, even when sharing CDs of the same family, as it is the case of Lys170 and EFAL-1 (Amidase-2 family). Remarkably, although designed to act on the same bacterial cell wall, each endolysin seems to carry a distinct CWBD, suggesting that several different ligands of that cell compartment might be targeted by the endolysins. These enzymes are completely unrelated to those identified in eight sequenced E. faecalis temperate phages, which encode endolysins with CD and CWBD of the Glyco-hydro-25 and LysM families, respectively. 50 Another striking feature is the lack of close similarity between the CD of Lys170, EFAl-1, and PlyV12 and that of E. faecalis peptidoglycan hydrolases of the same family. BLASTP analysis showed that the closest homologs of the endolysin CDs are those carried by enzymes from different bacterial species, some from different genera (Fig. 6). This is in clear contrast to other known phage/bacterial systems, where the CD of endolysins is closely related to that of the bacterial host autolysins.24,53 Lys168 CD was found to be closely related to a single E. faecalis peptidoglycan hydrolase encoded by the strains TX0104 and TX1341 (Fig. 6).
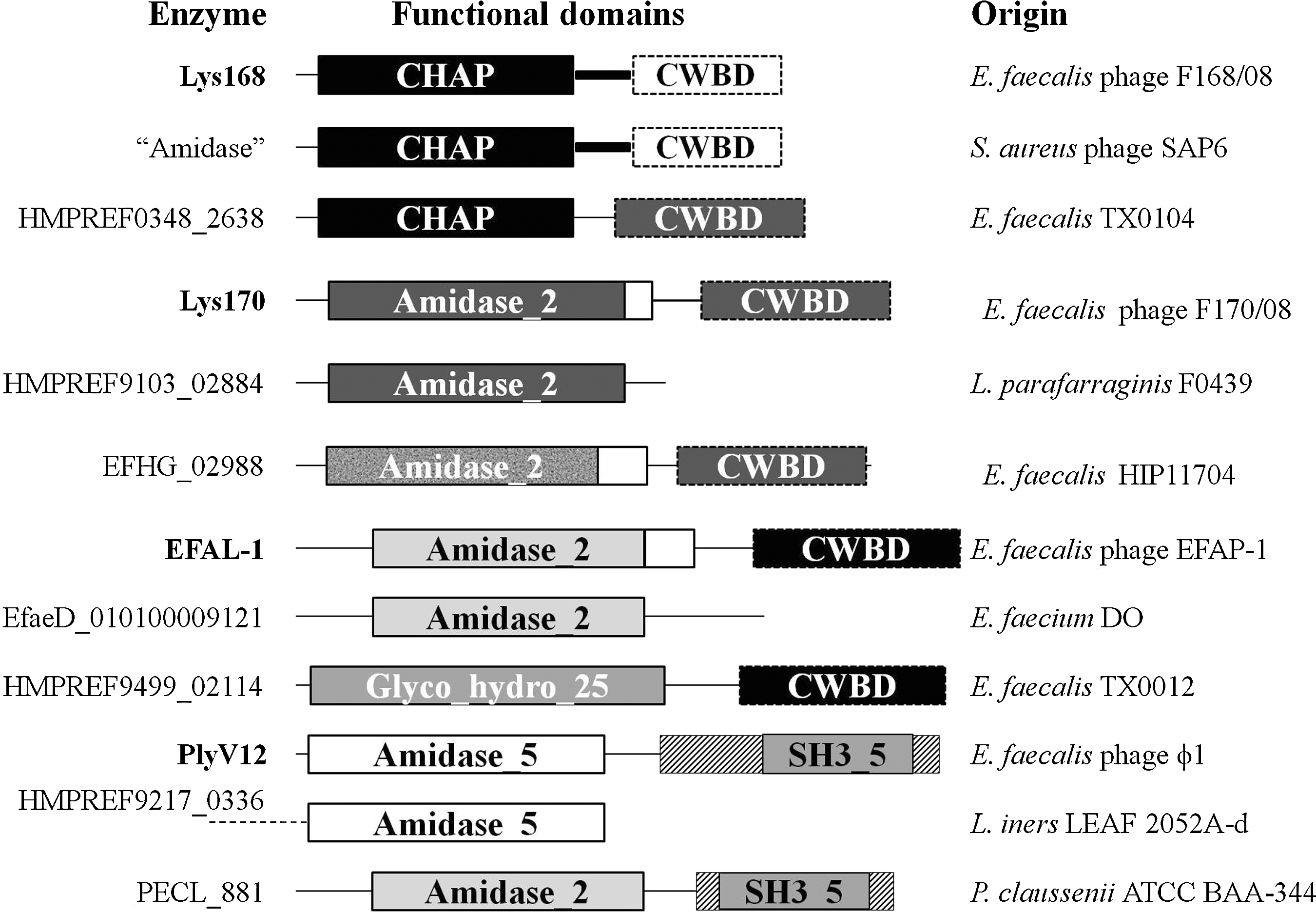
Nature, organization, and sequence relatedness of E. faecalis phage endolysin functional domains. The sequence similarity between functional domains is evidenced by using identical filling patterns. L., Lactobacillus; P., Pediococcus.
Uchiyama et al. 48 reported for ORF9 (identical to Lys170) a lytic spectrum of 97% and 60% against 35 and 10 nontyped E. faecalis and E. faecium isolates, respectively, which is very close to the results we obtained when Lys170 was tested in 73 and 26 clinical isolates of these species (Supplementary Fig. S4). However, when assayed in a panel of distinct and typed E. faecalis and E. faecium strains, Lys170 lytic range against E. faecium dropped to about 10% while maintaining that against E. faecalis (Fig. 4). The results show the importance of testing the lytic spectrum of endolysins on a reasonable number of strains with different known genetic backgrounds. We have thus concluded that Lys170 has a strong preference to act against E. faecalis. Both ORF9 and Lys170 were unable to lyse bacterial species outside the Enterococcus genus 48 (Supplementary Table S5).
Lys170 is most likely an N-acetylmuramoyl-L-alanine amidase, as this was the activity experimentally determined for ORF9. 48 Amidases cleave the amide bond that links the N-acetyl muramic acid of glycan strands to the L-alanine residue of the peptide stems. This bond and the nature of the linked residues are common to the vast majority of bacterial cell wall peptidoglycans, including that of E. faecium. 44 Why is then the E. faecalis cell wall the preferred substrate of Lys170? The ability of a given endolysin to cleave the bacterial cell wall depends on the integration of four major factors: (i) binding of the CWBD to a specific ligand of the cell wall, (ii) dependence of the CD activity on CWBD binding, (iii) CD affinity to its substrate, and (iv) the presence of the peptidoglycan bond that is specifically cleaved by the CD. 25 BLASTP analysis of Lys170 CWBD showed that this domain shares only significant similarity with those from E. faecalis enzymes. This suggests that Lys170 CWBD binds to an epitope that is predominantly found in the E. faecalis cell wall and that this binding is important for the endolysin lytic action. This epitope eventually exists in a few strains of the related species E. faecium, explaining why some strains of this species are susceptible to Lys170.
Lys168 also displayed a preferred lytic action against E. faecalis cells, acting poorly and in a much reduced number of E. faecium typed strains (Fig. 4). As referred to above, Lys168 CWBD is unrelated to that of Lys170, and thus the CHAP endolysin must recognize an epitope different from that targeted by Lys170. The peptidoglycan hydrolases of the CHAP family cleave different bonds of the murein structure, although a recent survey of the literature suggests that when present in bacterial autolysins, the CHAP domain specifies amidase activity, whereas in phage endolysins it seems to confer an endopeptidase activity. 22 The latter activity typically cleaves the amino acidic bridges that cross-link the peptidoglycan stem peptides,28,33 which can be different among the bacterial species as it happens, for example, in E. faecalis, E. faecium, S. aureus, and S. agalactiae. 44 Assuming this type of activity for Lys168 it could be easily explained the specificity of the endolysin toward E. faecalis cell wall. However, a recently constructed chimera composed of the Lys168 CHAP domain and the CWBD of an S. aureus endolysin proved to be very efficient in lysing several bacterial species, including a large number of S. aureus clinical strains 11 (see below). It is therefore more likely that the Lys168 CHAP domain specifies the amidase activity and that the enzyme specificity toward E. faecalis cell wall is conferred by its CWBD.
The lytic spectrum of the other two putative amidases, EFAL-1 and PlyV12 (Fig. 6), has been also studied. In contrast to what we have observed with Lys168 and Lys170, PlyV12 was reported to have a broad lytic spectrum, displaying different degrees of activity against E. faecium and several streptococcal and staphylococcal strains. 51 The authors provided a possible explanation for the broad lytic spectrum of PlyV12, which relied on some sequence relatedness between the enzyme CD and that of endolysins from phages infecting the susceptible bacterial species, 51 although these endolysins are not the closest PlyV12 homologs, as mentioned above (Fig. 6). It was also suggested that PlyV12 CWBD might target a cell wall epitope that is common to the different bacteria. 51
The significant sequence relatedness observed between the PlyV12 CD and that of streptococcal and staphylococcal phage endolysins was not verified for Lys170. Lys168 though exhibited 95% sequence identity with the endolysin of S. aureus phage SAP6 and significant similarity with virion-associated lysins of staphylococcal phages (Figs. 2 and 6). Despite this fact, Lys168 failed completely to induce lysis of all tested staphylococcal isolates, including those of S. aureus (Supplementary Table S5). This suggests that the few differences observed between Lys168 and SAP6 endolysins (Fig. 2) are on key residues that determine the specificity of these enzymes and that these most likely reside in the CWBD (see above). In fact, and as referred before, when we exchanged the Lys168 CWBD by that of an S. aureus phage endolysin the resulting chimera could efficiently lyse S. aureus. 11
The endolysin EFAL-1 was also reported to display a broad lytic spectrum against E. faecalis and E. faecium. 46 Although this enzyme was tested in a reduced number of isolates (13 E. faecalis and 7 E. faecium) and no information was provided about their diversity, the fact is that the enzyme seems to be a natural chimera assembling a CD and a CWBD closely related to those from E. faecium and E. faecalis cell wall lytic enzymes, respectively 46 (Fig. 6). This may explain the ability of EFAL-1 in lysing these two bacterial species. No significant sequence similarity was observed between the CD of Lys170 and Lys168 and that of E. faecium lytic enzymes (Fig. 6).
In conclusion, the results here presented indicate that endolysins Lys168 and Lys170 are good candidates for the specific elimination of E. faecalis, including VRE strains, for either sanitation or therapeutic purposes. The efficacy of these endolysins in animal models of E. faecalis infections is currently under study.
Note Added in Proofs
In this article, the authors highlight the striking sequence similarity between their enterococcal phage F168/08 and a phage named SAP6, which was assigned in sequences databases as being from Staphylococcus aureus. The close relatedness between these two phages translated in almost identical endolysins both at the gene and protein levels. However, after the paper has been accepted for publication, the SAP6 genome and endolysin entries in sequences databases, GenBank JF731128 and AEM24735, respectively, were corrected and it turned out that SAP6 appears also to be an Enterococcus faecalis phage. Readers should take this into account when reading the article.
Footnotes
Acknowledgments
D.P.'s work has been supported through a Ph.D. fellowship (SFRH/BD/64177/2009) from Fundação para a Ciência e a Tecnologia (FCT, MCTES, Portugal). The typed enterococcal strains were characterized through the FCT grant POCI/SAU-ESP/58030/2004 (awarded by R. Mato).
Disclosure Statement
Technophage has proprietary rights over E. faecalis phages F168/08 and F170/08 and their endolysins Lys168 and Lys170. All authors reviewed and approved the present article before submission. All authors declare that there are no conflicts of interest. All authors contributed to the elaboration of the article. D.P. and S.F. were responsible for endolysin cloning, production, and purification; D.P. executed all experiments evaluating lytic action of endolysins; C.L. and F.A.S. were responsible for phage isolation, purification, and extraction of phage DNA; C.L. performed phage genomic analysis; S.S. was responsible for Enterococcus species identification by PCR; F.L., R.M., P.C.-S., M.P., and C.S.-J. were responsible for the experimental design and work supervising; C.S.-J. co-ordinated the collaborative work of the involved laboratories.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
