Abstract
Penicillin-binding protein 2x (PBP2x) mutations that occur during the selection with beta-lactams are located within the central penicillin-binding/transpeptidase (TP) domain, and are believed to mediate resistance by interfering with the formation of a covalent complex of the active site serine with the antibiotic. We now investigated the effect of two point mutations found in two independently obtained laboratory mutants that are located at the surface of the TP domain with their side chains facing outside (G422D respectively R426C). They have no significant effect on resistance to cefotaxime in vivo or on binding to Bocillin™FL to the active site in vitro using purified PBP2x derivatives, thus apparently do not affect the active site directly. In contrast, in silico modeling revealed that they affect van der Waal's interactions with the PASTA1 (PBP and serine/threonine kinase associated) domain of the C-terminal extension and a noncovalent cefuroxime molecule found in the X-ray structure of an acylated PBP2x, suggesting some effect of the mutations on the interaction of the TP domain with PASTA1 and/or with the antibiotic associated with PASTA1. The effect of the PASTA domains on covalent binding of PBP2x to Bocillin FL was then investigated using a series of soluble truncated PBP2x derivatives. Deletion of 127 C-terminal residues, that is, of both PASTA domains, decreased binding dramatically by ∼90%. Surprisingly, deletion of only 40 amino acids resulted in the same phenotype, whereas the absence of 30 amino acids affected binding marginally by 10%, documenting a crucial role of the C-terminal domain for beta-lactam binding.
Introduction
In Streptococcus pneumoniae, resistance to beta-lactam antibiotics is mediated by mutations in PBPs. Such mutated PBPs exhibit reduced affinity to beta-lactams and thus higher concentrations of the antibiotics are required for inhibition. Three of the six pneumococcal PBPs are main players in the development of penicillin resistance. The two essential hmm class B PBPs 2x and 2b each confer low-level resistance, whereby PBP2x mediates resistance to both penicillins and cephalosporins whereas PBP2b contributes mainly to penicillin resistance since it does not interact with cefotaxime and other compounds with similar side chains. 7 The hmm class A PBP1a is required for high resistance level in the presence of a mutated PBP2x and/or PBP2b. PBP2x is among the best investigated PBPs; multiple mutations have been characterized in clinical isolates and in laboratory mutants as well, 6 and it was the first PBP whose structure was resolved by X-ray crystallography. 17
Among the many mutations that occur in PBP2x of resistant clinical isolates and laboratory mutants, at least some have been shown to mediate resistance to beta-lactams by altering the overall structure of the active site, thus interfering with the antibiotic binding to the active site serine (for review, see Hakenbeck et al. 6 ). Curiously, the X-ray structure of an acylated PBP2x revealed two cefuroxime molecules in the protein; in addition to the one covalently bound to the active site serine, another antibiotic molecule was noncovalently associated and located in the cleft between the TP domain and the C-terminal extension. 4 The C-terminal extension is attached to the TP domain via a long flexible linker region and consists of two repeated structural motifs named PASTA domains, “Penicillin-binding protein And Ser/Thr kinase Associated.” 22 The PASTA domain occurs in some hmm PBPs and in eukaryotic-like serine/threonine kinases of a variety of bacteria. Since beta-lactams are structural analogues of muropeptides, 21 it has been hypothesized that the C-terminal PASTA domains of PBP2x as well as in the other proteins interact with muropeptides, the substrate molecules of PBPs. 22 This has been verified for several bacterial serine-threonine kinases, including StkP of S. pneumoniae.13,16,20 However, no experimental evidence has been obtained so far to confirm this conclusion for PBP2x.
Among the point mutations identified in resistant strains, the two changes G422D and R426C occur in two independently isolated cefotaxime-resistant laboratory mutants. Curiously, they are located on the surface of the TP domain at a great distance from the active site with their side chains facing the cleft between the TP domain and the C-terminal extension. We therefore analyzed the effect of these two mutations in more detail and investigated the impact of the PASTA domains of PBP2x on beta-lactam binding.
Materials and Methods
Bacterial strains, plasmids, and growth conditions
S. pneumoniae R6 and cefotaxime-resistant mutants have been described. 11 S. pneumoniae strains were grown in liquid culture without shaking in a casein-based semisynthetic medium supplemented with 0.2% yeast extract. 10 Minimal inhibitory concentrations (MICs) of beta-lactam antibiotics were tested by agar dilution on blood agar plates (3% sheep blood). A narrow range of antibiotic concentrations was used in order to detect subtle differences in susceptibility between different transformants.
Escherichia coli JM101 was used as host strain for the plasmids derived from the expression vector pGEX-6P-1tet, 1 a derivative of pGEX-6P-1 (GE Amersham Biosciences) in which ampR is replaced by tetR. E. coli JM101 containing pGEX-6P-1tet derivatives were grown at 37°C with shaking in Luria-Bertani broth supplemented with 20 μg/ml of tetracycline. Transformation of E. coli JM101 was performed following published procedures. 18
Construction of PBP2x derivatives
Plasmids encoding PBP2x glutathione-S-transferase (GST) fusions containing the desired PBP2x mutations were constructed by overlap extension polymerase chain reaction (PCR)8,9 using the oligonucleotide pairs PM11/PM48 and PM12/PM51 (G422D), PM11/PM49 and PM12/PM52 (R426C), or PM11/PM50 and PM12/PM53 (G422D/R426C) (Table 1). For construction of truncated PBP2x derivatives, DNA fragments covering various regions of the PBP2x gene of S. pneumoniae R6 were obtained by PCR amplification of S. pneumoniae R6 pbp2x using the oligonucleotides listed in Table 1. For all PCRs, the high fidelity enzyme iProof Polymerase (BioRad) was used. DNA fragments were subsequently cloned into the expression vector pGEX-6P-1 tet 1 and transformed into the E. coli strain JM101. The presence of the desired mutation was verified by DNA sequencing of the plasmids. Only in plasmid pPM19 expressing the N-terminal domain of PBP2x, a mutation occurred in codon 53 (G to A) that resulted in an Arg53His change, but this was not considered to be relevant for beta-lactam-binding studies of the peptide since this peptide was used as negative control.
Restriction sites are underlined.
Altered nucleotides for site-directed mutagenesis are marked by small letters.
Amino acids (aa) covered by the cloned pbp2x fragment.
Primers used for the construction of the plasmids.
Overexpression and purification of PBP2x derivatives
The GST-PBP2x proteins overexpressed in E. coli strain JM101 from the plasmids (Table 1) were purified from the cytoplasmic fraction as follows. To exponentially growing cultures 1 mM isopropyl-β-D-thiogalactopyranoside was added at OD600=1.2, followed by incubation at 15°C with shaking overnight. All purification steps were performed on ice. After centrifugation at 8,000 g for 10 min, cells were resuspended in 35 ml PBSKMT (25 mM sodium phosphate, 150 mM NaCl, 3 mM KCl, 1 mM MgCl2, and 0.1% Tween 20, pH 7.3) and lysed by sonication. Soluble proteins were obtained after centrifugation at 21,000 g in J2-21 centrifuge (Beckman) for 60 min. GST fusion proteins were purified from supernatants using a glutathione-Sepharose column (GSH-Sepharose High performance; Amersham Biosciences) equilibrated in PBSKMT buffer. The column was washed with 20 ml PBSKMT and 20 ml PBSKM (PBSKMT buffer minus Tween 20) and GST-PBP2x derivatives and finally eluted with Elution buffer (10 mM GSH and 50 mM Tris-HCl [pH 8.0]). The Elution buffer was exchanged with PBSKMT by gel filtration (PD10; Amersham Biosciences). Purified PBP2x derivatives were stored at −80°C; no loss of activity in terms of beta-lactam binding was noted during a several month period.
Labeling and detection of penicillin-binding proteins
PBP2x derivatives were overexpressed in E. coli described and purified as previously. For labeling, 10 μl of purified PBP2x proteins corresponding to 500 ng in PBSKMT buffer was incubated with 10 μl Bocillin™FL 2 (5 μM final concentration if not otherwise stated) for 10 min at 37°C as described. 11 Proteins were separated on sodium dodecyl sulfate-polyacrylamide gels by gel electrophoresis using 11% acrylamide (ratio of acrylamide:bisacrylamide=37.5:1). PBP-Bocillin FL complexes were visualized with a FluorImager at 488 nm (GE-Healthcare), and analyzed using the ImageQuant Software.
In silico modeling of PBP2x
The X-ray structure of 1QMF 4 was used to model the effect of mutations with the program Swiss Protein Viewer (www.expasy.org/spdbv/). 5
Results
Structural impact of the PBP2x mutations Gly422Asp and Arg426Cys by in silico modeling
Three respectively four mutations in PBP2x occur in two independently obtained cefotaxime-resistant laboratory mutants C503 and C606 isolated during successive selection steps with increasing concentrations of cefotaxime, in addition to mutations in other genes12,23 (Fig. 1). The fourth respectively third mutation in PBP2x, Gly422Asp in C606 and Arg426Cys in C503, are located at a great distance from the active site serine (>16 Å) on a loop between helix 6 and helix 6a at the surface of the TP domain (Fig. 2). Using the structure of the acylated form of PBP2x obtained with cefuroxime, 4 a network of hydrogen bonds ranging between 2.24 and 3.18 Å becomes visible in the PBP2x structure between the TP domain, the PASTA domains of the C-terminal extension, and the cefuroxime molecule located in the cleft between the two domains (Fig. 3). In the wild-type protein, Arg426 is involved in multiple hydrogen bonds: the side chain is connected to Asp648 and Glu651, and to both oxygen of the COO-group of the cefuroxime beta-lactam ring; Arg654 of PASTA1 connects to the backbone O of Arg426 and to Thr425, and the cefuroxime molecule is hydrogen bonded to the backbone O of Pro424 (Fig. 3, top). The mutation Cys426 allows two new hydrogen bonds (arrows in Fig. 3) that link the Cys side chain with Thr425 and Arg654, but the interaction with Asp648, Glu651, and the cefuroxime molecule is lost, and in case of Asp422, a hydrogen bond of 2.25 Å to the side chain of cefuroxime is established (Fig. 3, bottom). This suggests some effect of the mutations on the interaction of the TP domain with PASTA1 and/or with a putative substrate molecule located in this region.
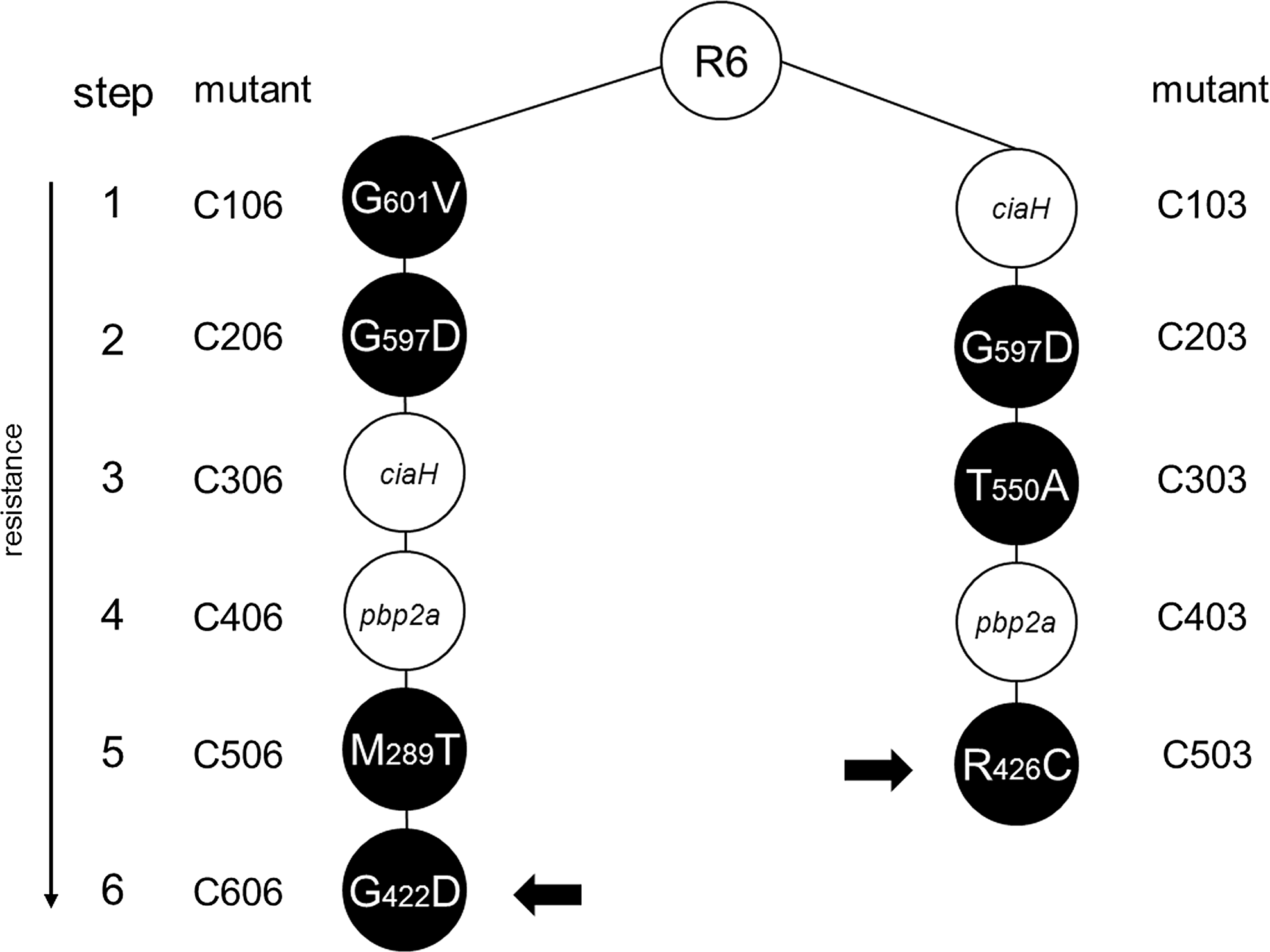
Cefotaxime-resistant laboratory mutants of the two lineages C003 and C006. Mutants were obtained from the parental R6 strain by successive selection on increasing cefotaxime concentrations. During each step, mutations in pbp2x (black circles), pbp2a, or ciaH were selected; the amino acid changes in penicillin-binding protein 2x (PBP2x) are indicated. The small black arrows mark the mutations investigated here.
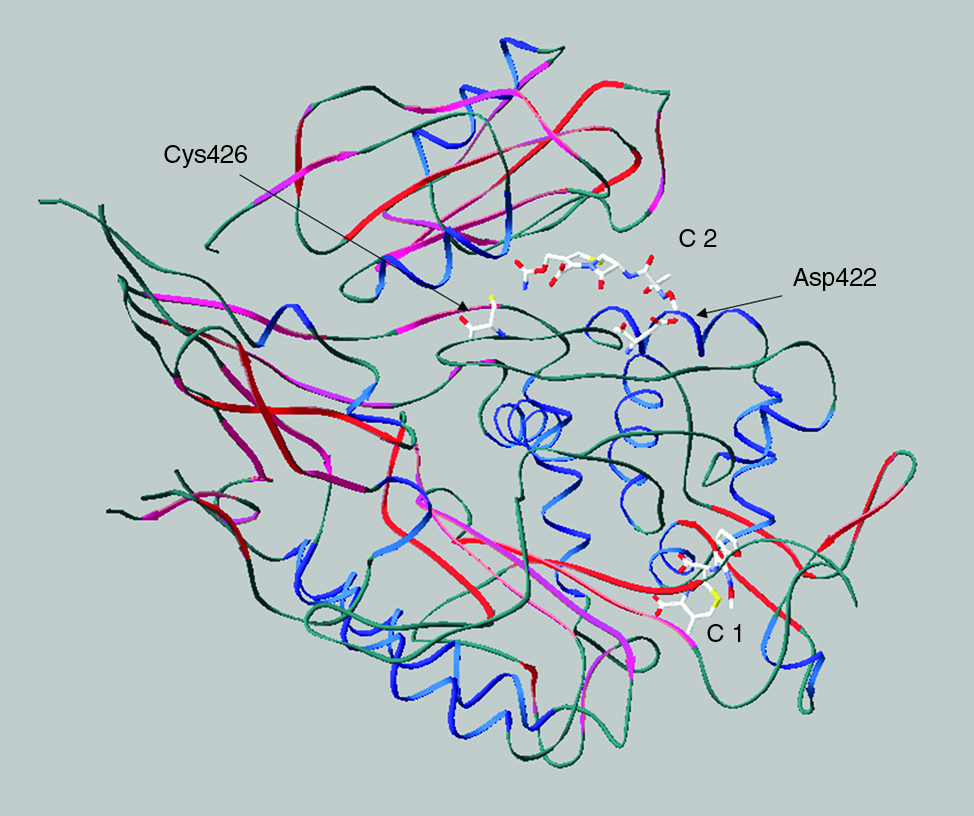
Position of the mutations Arg422 and Cys426 in PBP2x. The structure of acylated PBP2x with cefuroxime molecule is shown (1QMF4). C1 and C2 indicate the covalent respectively noncovalent cefuroxime molecules in the X-ray structure. The mutations are marked. α-Helixes are shown in blue, and beta-sheets in red.
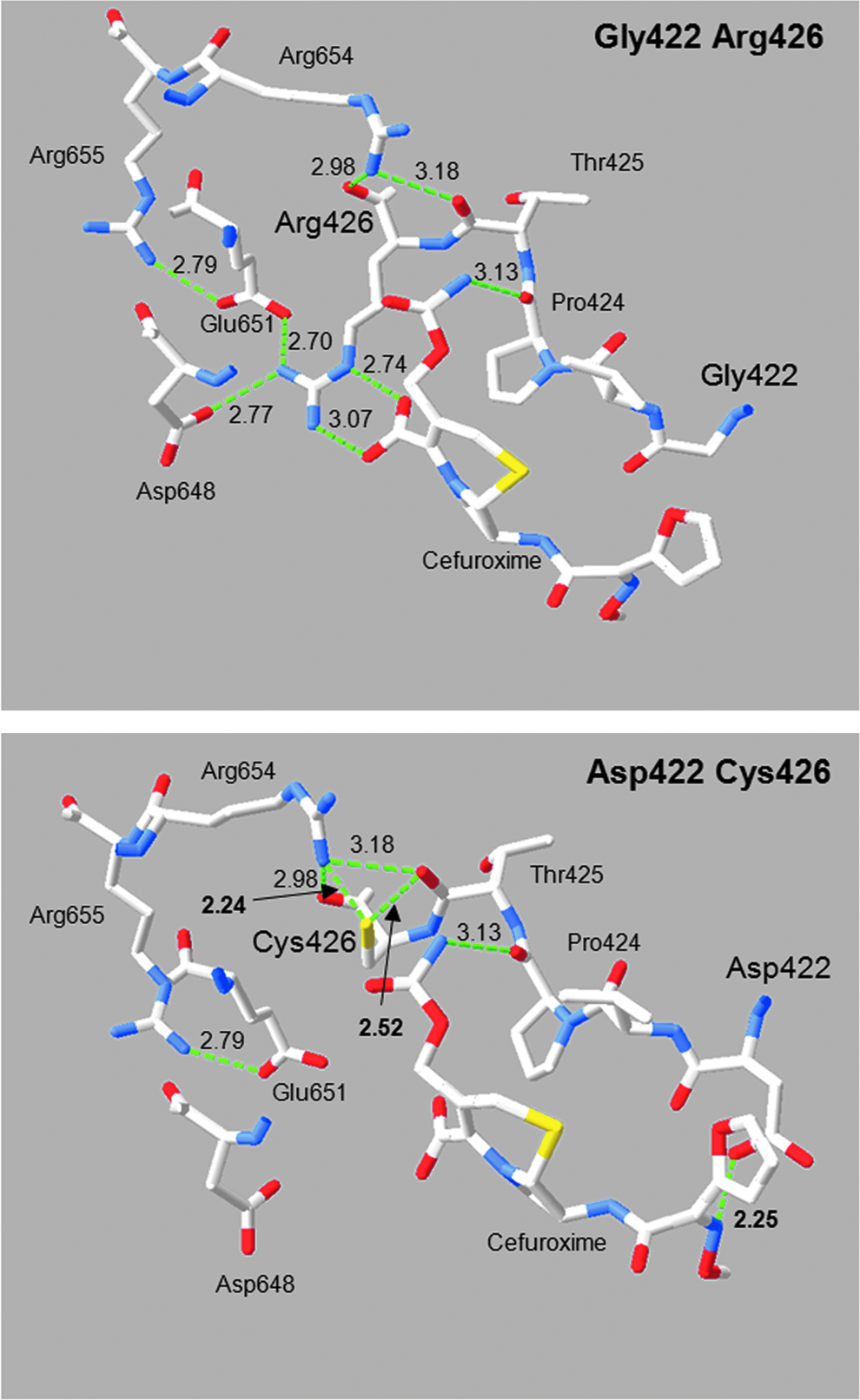
Impact of the hydrogen bonding network by the mutations Gly422Asp and Arg426Cys. The structure of PBP2x 1QMF4 was modeled with the program Swiss Protein Viewer with calculated H-bonds (green dotted lines). Alterations in the hydrogen bonding network between wild type (top: R6 [Gly422/Arg426]) and mutants (Asp422 [C606] and Cys426 [C503]) are shown. The numbers refer to the hydrogen bond distances in Å. Important amino acid residues and the cefuroxime molecule are indicated. Blue, red, and yellow mark nitrogen, oxygen, and sulfur molecules.
G422D and R426C and beta-lactam resistance
The mutations Asp422 in C606 and Cys426 in C503 did not lead to a significant increase in resistance when the MIC values of the mutants were compared with their predecessors without them (C506 respectively C403). 15 Also, R6 transformants obtained with pbp2xC606 containing all four PBP2x mutations, or only three without the Asp422 mutation, did not differ in their cefotaxime MIC values (not shown). To test the effect of the individual mutation in the wild-type background, plasmids carrying pbp2x with either one of these mutations or both were constructed by site-directed mutagenesis to transform the parental R6 strain followed by beta-lactam selection. Narrow concentration ranges of cefotaxime or oxacillin (0.01 μg/ml intervals) were used to select transformants at or slightly above the MIC levels of the R6 strain. However, all attempts to obtain resistant colonies were unsuccessful, indicating that the mutations have no impact on beta-lactam susceptibility in the wild-type background. Next, soluble derivatives of PBP2x with a deleted membrane anchor carrying either one of the mutations or both were overexpressed in E. coli and Bocillin FL binding was tested with the purified proteins. No significant difference was observed in the binding kinetics for PBP2xD422 or PBP2xC426; only with the double-mutant PBP2xD422-C426 an ∼20% reduction in binding was observed (Fig. 4A–C). This suggests that the effect on beta-lactam binding to the active site serine is only marginally affected if at all by the single mutation D422 or C426.
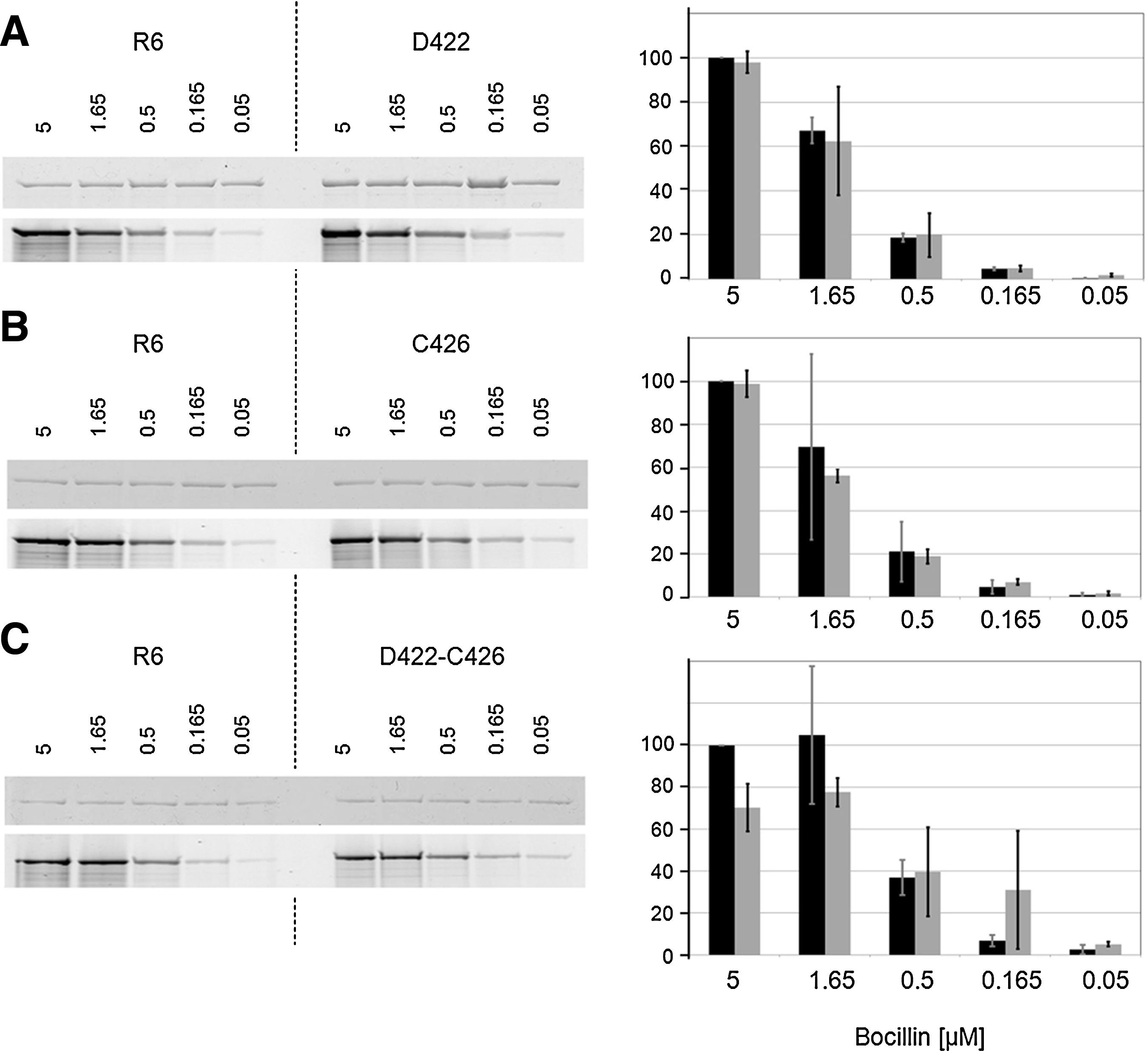
Binding of PBP2x mutant derivatives to Bocillin™FL. Glutathione-S-transferase (GST) fusion proteins were purified from Escherichia coli extracts, and incubated with Bocillin FL; per sample 500 ng protein in a total volume of 20 μl was used. After sodium dodecyl sulfate-polyacrylamide electrophoresis, PBP2x-Bocillin FL was visualized with a FluorImager and binding intensity was recorded as described in the Materials and Methods section. Left: fluorograms; the Bocillin FL concentrations (μM) are shown above the lanes. Right: mean values of two experiments; black: control (R6); gray: mutant.
Effect of C-terminal deletions on beta-lactam binding
Since some interactions between the TP domain and PASTA1 were implicated by the structural analysis as described previously, the effect of C-terminal deletions on Bocillin FL binding was investigated using a series of truncated soluble PBP2x derivatives (Fig. 5). Each of the PASTA domains consists of three beta-sheets and one α-helix, with PASTA1 ranging from amino acid residues 632–691, and PASTA2 from 692–750. 22 Deletion of the beta-sheets of PASTA2 had only little effect on beta-lactam binding (∼90% remained), whereas deletion of both PASTA domains, that is, of 127 C-terminal amino acids (aa), reduced binding to below 10% (Fig. 6A, B). In fact, the presence of PASTA1 plus the α-helix in PASTA2 appears to be crucial for antibiotic binding, since deletion of 40 aa that results in disruption of this α-helix already reduced binding to a similar extent (Fig. 6A, B). Deletion of 268 N-terminal aa abolished binding completely, probably since the N-terminal domain is closely interlinked with the TP domain and thus the deletion might affect the overall structure of the TP domain considerably. This is probably also the reason why no binding was observed with the TP domain alone. As negative controls, the N- and the C-terminal domains were used that do not covalently interact with the antibiotic (Fig. 6A, B). The results clearly establish a role of the PASTA domains in beta-lactam binding in PBP2x in context with the TP domain.
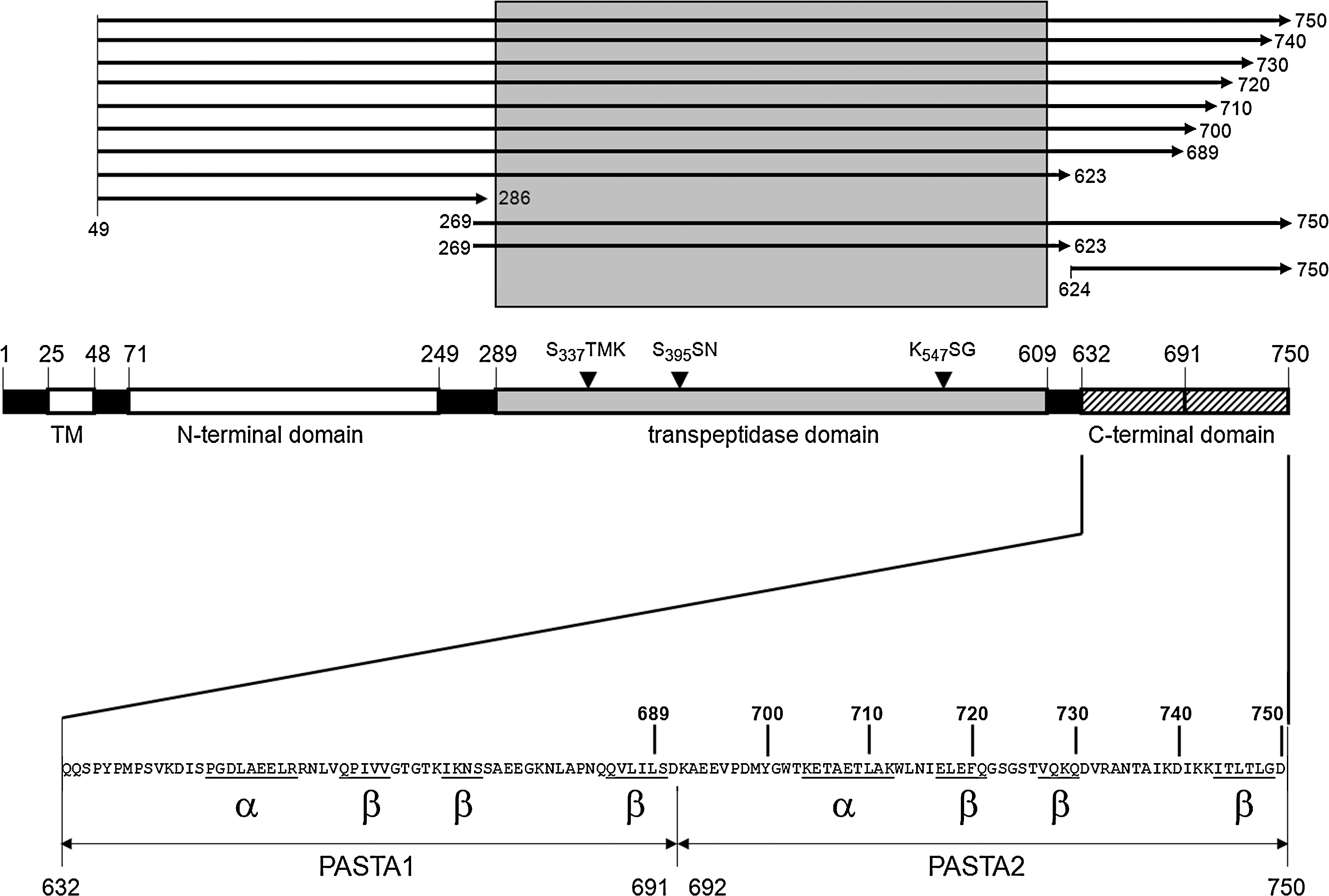
Truncated PBP2x derivatives. Top: the length of the various PBP2x derivatives is shown by arrows; first and last amino acids are indicated. Bottom: shown is the overall domain structure of PBP2x with the active site motifs, and the C-terminal extension with structural features of the two PBP and serine/threonine kinase associated (PASTA) domains is depicted in detail below.
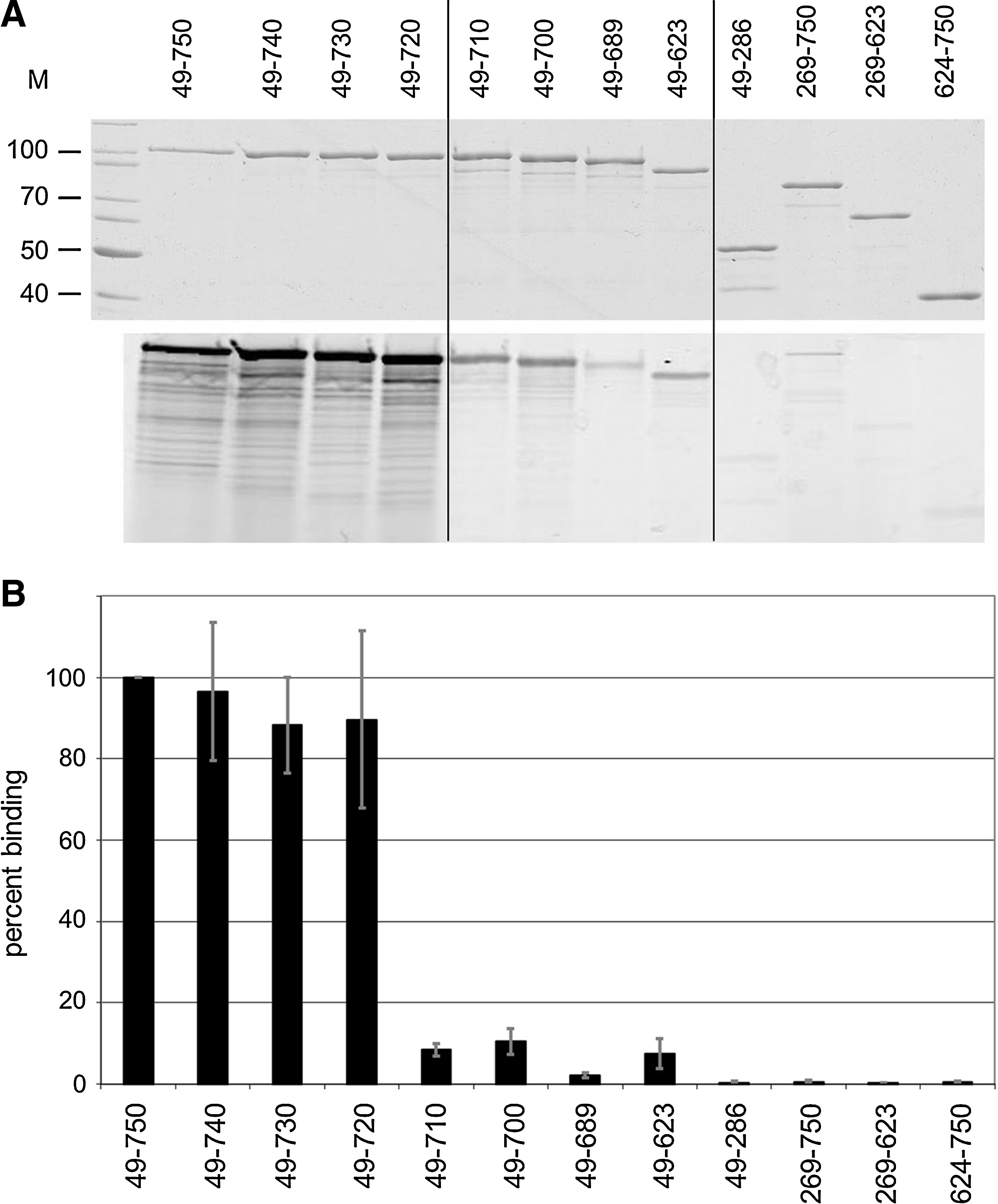
Binding of truncated PBP2x derivatives to Bocillin FL. Purified PBP2x derivatives were labeled with Bocillin FL as described in the Materials and Methods section.
Discussion
Despite the availability of a high-resolution structure of PBP2x, the impact of many point mutations identified in laboratory mutants and clinical isolates on beta-lactam binding and enzymatic function is still not clear (for review, see Hakenbeck et al. 6 ). It is remarkable that most mutations described in laboratory mutants do not map close to the active site, although all of them occur within the TP domain of the protein.12,15 The two mutations investigated here—Gly422Asp and Arg426Cys—are positioned at the surface of the TP domain with their side chain directed toward the C-terminal domain interacting with Glu651 and Arg654 (Cys426) or with no other amino acid (Asp422). Thus, they do not appear to affect the active site at all. In agreement with this notion, the contribution of these two mutations to beta-lactam resistance appears to be marginal if detectable at all, neither in the context of mutations in other genes present in the original mutants C503 and C606, nor if introduced into the genetic background of the parental R6 strain. In other words, there is no indication that they interfere with the covalent binding of the beta-lactam inhibitor. In contrast, they curiously appear to interact with the cefuroxime molecule noncovalently located between the two domains in the structure of an acylated PBP2x 4 (see Fig. 2).
Why do these mutations occur in the cefotaxime-resistant mutants? We believe it to be unlikely that they are spontaneous mutations not associated with the selection procedure for two reasons. Both are located in close vicinity on the same structural element that links the two α-helixes 6 and 6a with their side chains directed toward the C-terminal extension. Moreover, both occur after the introduction of other (two or three) mutations in pbp2x and in mutants that also contain mutations in ciaH encoding the histidine protein kinase CiaH (see Fig. 1). It has been shown that PBP2x mutations associated with the resistance phenotype are functionally not neutral, but that cellular growth and cell morphology of PBP2x mutants are dependent on a functional CiaRH system.14,24 It is possible that the two mutations Gly422Asp and Arg426Cys are directed to complement the functional deficiency of the mutated PBP2x. Since the part of the beta-lactam ring that is associated with the PBP2x PASTA1 domain through van der Waal's interactions is analogous to D-Ala-D-Ala, the C-terminal amino acids of muropeptides that are attacked during the transpeptidation reaction of PBPs, 21 it has been suggested that the PASTA1 domain binds uncrosslinked murein. 22 Indeed, binding to ampicillin and D-Ala-D-Ala as well has been shown for the PASTA domains of the S. pneumoniae serine/threonine kinase StkP, 13 and also the Bacillus subtilis PrkC protein binds to peptidoglycan fragments. 20 Thus, the effect of the two PBP2x mutations might be to modulate the interaction with uncrosslinked muropeptides associated with the PASTA domains, which would explain why they confer no resistance.
Another feature of the C-terminal extension was apparent when binding to bocillin was investigated in a series of C-terminal-truncated PBP2x derivatives. Whereas deletion of the last 40 residues reduced binding to bocillin only slightly to ∼90%, deletion of another 10 aa, that is, disruption of the α-helix of PASTA2, dramatically affected binding, resulting in only 10% residual binding (see Fig. 5). This suggests that the overall fold of the C-terminal extension is strongly disturbed in the 40-aa-truncated protein, and an indirect effect on the TP domain cannot be excluded. Further structural analysis of the mutant proteins—and a detailed kinetic analysis preferentially with cefotaxime, the selective antibiotic—and substrate molecules might help to shed more light on the role of the C-terminal PASTA domains of PBP2x and its interaction with substrate and inhibitor molecules.
Footnotes
Acknowledgments
The authors thank Linda Kunsch and Julia Bishop for help in the construction of plasmids, Barbara Koch for help during construction of pbp2x mutants, and Dieter Höltje and Paulus van Hooft for helpful discussions. This work was supported by the DFG (Ha 1011/11-1), the EU (BI04-CT98-0424), and the Stiftung Rheinland-Pfalz für Innovation (15202-386261/580).
Disclosure Statement
There are no commercial associations with P.M., J.S., K.T., or R.H. that might create a conflict of interest in connection with submitted article. No competing financial interests exist.
