Abstract
Purpose:
Oxidative stress contributes to the pathogenesis of vision-impairing diseases. In the retina, retinal pigment epithelium (RPE) and Müller cells support neuronal homeostasis, but also contribute to pathological development under stressed conditions. Recent studies found that the investigational drug risuteganib (RSG) has a good safety profile, provided protection in experimental models, and improved visual acuity in patients. The present in vitro study evaluated the effects of RSG in RPE and Müller cell lines stressed with the oxidant hydrogen peroxide (H2O2).
Methods:
Human RPE (ARPE-19) and Müller (MIO-M1) cell lines were treated with various combinations of RSG and H2O2. Trypan blue assay was used to investigate the effect of compounds on cell viability. Gene expression was measured using RNA sequencing to identify regulated genes and the biological processes and pathways involved.
Results:
Trypan blue assay found RSG pre-treatment significantly protected against H2O2-induced cell death in ARPE-19 and MIO-M1 cells. Transcriptome analysis found H2O2 regulated genes in several disease-relevant biological processes, including cell adhesion, migration, death, and proliferation; ECM organization; angiogenesis; metabolism; and immune system processes. RSG pre-treatment modulated these gene expression profiles in the opposite direction of H2O2. Pathway analysis found genes in integrin, AP-1, and syndecan signaling pathways were regulated. Expression of selected RSG-regulated genes was validated using qRT-PCR.
Conclusions:
RSG protected cultured human RPE and Müller cell lines against H2O2-induced cell death and mitigated the associated transcriptome changes in biological processes and pathways relevant to the pathogenesis of retinal diseases. These results demonstrate RSG reduced oxidative stress-induced toxicity in two retinal cell lines with potential relevance to the treatment of human diseases.
Background
Retinal diseases represent the leading causes of blindness in developed countries. 1 Age-related macular degeneration (AMD) and diabetic retinopathy (DR) account for a majority of these cases and are characterized by a mix of pathological neovascularization, inflammation, metabolic dysregulation, and tissue degeneration.2,3 It has become increasingly clear that mitochondrial dysfunction and oxidative stress play key roles in the disease pathogenesis.4–6 Elevated reactive oxygen species (ROS) levels and evidence of oxidative damage, such as lipid peroxidation and oxidation of proteins and DNA, have been observed in AMD7,8 and DR9,10 patients. The retina is particularly vulnerable as ROS can be readily generated due to high levels of local oxygen tension, photoirradiation, metabolic activity, and reactive molecules (eg, polyunsaturated fatty acids, lipofuscin).11,12 When dysregulated, prolonged oxidative stress contributes to reduced cellular functionality and cell death.
The retina is an architecturally complex tissue composed of diverse cell types with distinct physiological functions. Of those, Müller glia and retinal pigment epithelial (RPE) cells maintain the neural retina homeostasis by facilitating the transport of nutrients and waste between the retina and the vascular system, recycling of light-sensing chromophores, and secretion of growth factors and antioxidants.13–16 Müller glia span the entire neural retina and support both neurons and cone photoreceptors, while RPE cells form a monolayer subjacent to the photoreceptors and primarily support both cone and rod photoreceptors. 13 RPE cells also provide light absorption and filtration functionality and have high levels of antioxidants to expedite removal of resultant ROS.15,17 Importantly, RPE cells have enhanced capability to withstand oxidative stress and serve to protect the surrounding retinal cells from oxidative damage, functions that has been found to be diminished at old age.18,19
Under pathological conditions, these cells are activated to provide an initial neuroprotective role while their primary functions become dysregulated.13,14,20,21 Prolonged Müller and RPE dysfunction is thought to contribute to the pathological development of neovascularization, inflammation, and degeneration of the retina.
Current treatment primarily focuses on reducing neovascularization and edema with anti-VEGF drugs, but none is approved for targeting retinal degeneration. Risuteganib (RSG) is an RGD-derived (arginine–glycine–aspartate) oligopeptide first designed for binding and inhibiting integrin activity and later found to reduce integrin expression and have antioxidative and cytoprotective properties. RSG is under clinical investigation for treatment of AMD, diabetic macular edema (DME), and dry eye disease. Several clinical trials showed that RSG has a good safety profile and improved visual acuity in dry AMD and DME patients.22,23 In a preclinical model of human RPE cells stressed with cigarette smoke toxin, RSG treatment reduced cell death and ROS level, upregulated cytoprotective genes, and improved mitochondrial bioenergetics and metabolic activity. 24 In RPE cells constructed to contain AMD donor mitochondria, RSG exposure reduced the expression of apoptosis and angiogenesis genes. 25
Altogether, previous studies found that RSG can improve visual acuity, possibly through reduction of oxidative damage and reestablishment of retinal homeostasis.
Although RSG showed promising therapeutic potential, its effect on retinal cells requires further investigation. This study aimed to identify the effect of RSG and hydrogen peroxide (H2O2), a major source of ROS primarily generated during aerobic respiration, 26 in human Müller (MIO-M1) and RPE (ARPE-19) cell lines. Cells were treated with RSG and H2O2, followed by trypan blue assay for cell viability, RNA sequencing (RNA-seq) for whole transcriptome analysis, and quantitative real-time PCR (qRT-PCR) for validation of gene expression. The combined exposure to both RSG and H2O2 reveals the drug's therapeutic effects in the context of H2O2-induced oxidative stress. A better understanding of the drug mechanism could lead to the development of novel therapies for these debilitating diseases.
Methods
Cell culture reagents and conditions
The human Müller cell line MIO-M1 27 was kindly provided by Professor Astrid Limb (University College London). Cells were cultured in Dulbecco's modified Eagle's medium (DMEM) with 4.5 g/L glucose, glutaGRO (Corning Cellgro, Manassas, VA), 10% fetal bovine serum (FBS; Sigma, St. Louis, MO), penicillin 100 U/mL, and streptomycin sulfate 0.1% mg/mL (Omega Scientific, Inc., Tarzana, CA). ARPE-19 cells 28 were obtained from ATCC (CRL-2302; Manassas, VA) and cultured in DMEM mixture 1:1 Ham's F-12 medium (Corning, Manassas, VA), with 10% FBS, penicillin 100 U/mL, streptomycin sulfate 0.1% mg/mL, gentamicin 10 mg/mL, and amphotericin B 2.5 mg/mL (Omega Scientific, Inc.). Allegro Ophthalmics (San Juan Capistrano, CA) provided RSG; H2O2 was purchased from Sigma. Upon receipt, the cells were split 5 times to generate frozen cell stocks.
All experiments used cells at passage 5; cells were cultured for 48 h, then counted and plated at a density of 0.5 × 106 cells per well, following by another 24-h culture before treatment. Cells were at subconfluent density at the onset of treatment as confluent cells have been found to be significantly more resistant to oxidative stress. 29
Trypan blue assay
ARPE-19 (n = 3) and MIO-M1 (n = 8–9) cells in 6-well plates were treated according to the following 5 regimens: (1) untreated control, (2) RSG for 36 h, (3) untreated for 24 h, then H2O2 for 12 h, (4) RSG pretreatment for 24 h, then H2O2 for 12 h, and (5) RSG for 24 h, then RSG and H2O2 cotreatment for 12 h; conditions labeled as control, RSG, H2O2, H2O2 + RSG pretreat, and H2O2 + RSG cotreat, respectively.
Four hundred micromolar of RSG was used because it is the cell culture equivalence of the clinical dose (1.0 mg injection into 4 mL of vitreous volume). 22 One hundred micromolar of H2O2 was used as this concentration was found to induce moderate cytotoxicity in ARPE-19 30 and MIO-M1 31 cells, which simulates the elevated oxidative stress that leads to progressive cell death in the relevant retinal diseases where RSG is currently being investigated. After exposure, cells were incubated for 48 h in fresh media before cell harvest, and cell viability measurement by trypan blue assay with an automated ViCell analyzer (Beckman Coulter, Inc., Fullerton, CA). The density of viable cells was measured and normalized to mean of control as 100%.
RNA-seq sample preparation and analysis
ARPE-19 (n = 6) and MIO-M1 (n = 6) cells in 6-well plates were treated according to the following 3 regimens: (1) untreated control, (2) untreated for 24 h, then H2O2 for 12 h, and (3) RSG pretreatment for 24 h, then H2O2 for 12 h; conditions labeled as control, H2O2, and H2O2 + RSG, respectively. Four hundred micromolar of RSG and 100 μM of H2O2 were used. After exposure, cells were incubated for 48 h in fresh media before cell collection. Total RNA was extracted using the RNeasy Mini Kit (Qiagen, Hilden, Germany), and DNA was removed with TURBO DNA-free (Thermo Fisher Scientific, Waltham, MA). RNA quality was measured with Bioanalyzer (Agilent Genomics, Santa Clara, CA).
RNA-seq libraries were prepared using the NEBNext Ultra RNA Library Prep Kit (New England Biolabs, Ipswich, MA) and sequenced on HiSeq 2500 (Illumina, San Diego, CA) to generate 10–15 million single-end, 100 base-pair reads per sample. RNA-seq reads were quality tested by FASTQC, 32 and then aligned by STAR 33 to human genome (GRCh38.p12) and transcriptome (GENECODE v30) references, followed by read quantification using featureCounts. 34
Principal component analysis (PCA) was used to visualize the RNA-seq data set along the top 3 dimensions that captured the most variance, using the top 15,000 expressed genes calculated by the DESeq2's VST method. 35 Differential expression analysis was performed using edgeR 36 with analysis limited to genes with count per million of 4 or greater in at least 6 samples (referred to as expressed genes). Differentially expressed (DE) genes have a false discovery rate (FDR) <0.05 and are either upregulated or downregulated based on signs of log2-fold change (log2FC).
DE genes were submitted to goseq 37 for enrichment of gene ontology biological processes38,39 and NCI Nature biological pathways 40 that are overrepresented in the gene list. Biological processes or pathways with FDR <0.05 were considered to be enriched, that is, significantly overrepresented in the list of DE genes. Enriched biological processes were condensed and visualized with REVIGO 41 with similarity metric set to small; selected representative processes were labeled. Biological processes were grouped into categories by first summarizing those with an FDR <0.001 using REVIGO, and then a manual review of the results to form categories. Heatmaps were used to visualize gene expression levels, which were averaged by condition and then normalized to mean expression across all samples; genes were clustered by Euclidean distance. 42 ERSSA evaluates whether a sufficient sample size was used; analysis was performed with filter cutoff = 4, log2FC cutoff = 0.25, and 50 subsamples per replicate level. 43
Quantitative real-time PCR
ARPE-19 (n = 6) and MIO-M1 (n = 6) cells in 6-well plates were used for qRT-PCR. Three hundred nanograms of RNA was reverse transcribed into cDNA with SuperScript IV VILO Master Mix (Thermo Fisher, Waltham, MA) on a ProFlex PCR system (Thermo Fisher). qRT-PCR was performed with PowerUp SYBR Green Master Mix on a QuantStudio 5 Real-Time PCR System. Predesigned SYBR Green primers were used (KiCqStart SYBR Green Primers [Sigma, Burlington, MA] and QuantiTect Primer Assays [Qiagen], Table 1). HPRT1 was the reference gene used.
Description of Genes Analyzed by Quantitative Real-Time Polymerase Chain Reaction
Data availability
The RNA-seq raw data are available at NCBI SRA database (accession PRJNA723610). RNA-seq analysis scripts are available at https://github.com/zshao1/RSG_H2O2. All other relevant data are within the article and its supplementary figures and tables.
Statistical analysis
Trypan blue data are subjected to 1-way ANOVA test with multiple comparison correction by the Benjamini, Krieger, and Yekutieli method using GraphPad Prism (Version 9.0, San Diego, CA). The q value is defined to be the adjusted P value from the ANOVA test (ie, P value after multiple comparison correction). A q value ≤0.05 is considered statistically significant. qRT-PCR fold values were calculated using the 2−ΔΔCt formula. 44
Results
RSG pretreatment reduced cell viability loss
ARPE-19 and MIO-M1 cells used in this study were confirmed to express RPE cell markers (RLBP1/CRALBP, OTX2, and PMEL/PMEL17) 45 and Müller cell markers (GFAP, GLUL, and SLC1A3), 46 respectively (Supplementary Fig. S8 and Supplementary Table S1). Trypan blue assay was used to investigate the effect of H2O2 and RSG on cell viability (Fig. 1A).
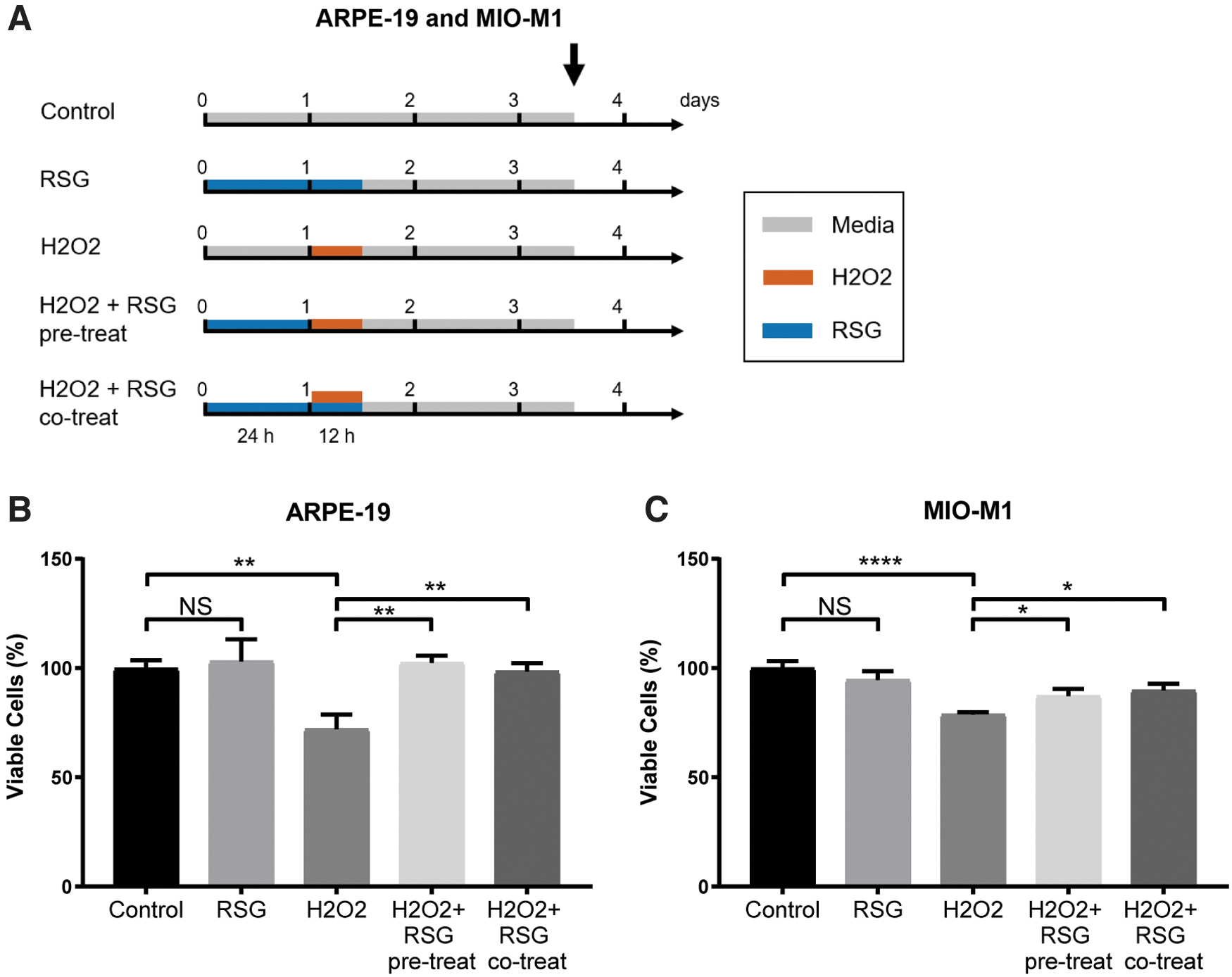
RSG protected against H2O2-induced cell viability reduction.
In both cell lines, RSG treatment had no effect, while H2O2 significantly reduced cell viability to 0.72 × (q = 0.004) and 0.79 × of control (q < 0.0001) in ARPE-19 and MIO-M1 cells, respectively (Fig. 1B, C). In cells pretreated with RSG for 24 h before H2O2 exposure, cell viability was significantly improved to 1.03 × (q = 0.0044) and 0.87 × of control (q = 0.043) in ARPE-19 and MIO-M1 cells, respectively. Similarly, in cells pretreated with RSG followed by cotreatment with both RSG and H2O2, cell viability was significantly improved to 0.99 × (q = 0.004) and 0.90 × of control (q = 0.020) in ARPE-19 and MIO-M1 cells, respectively.
RNA-seq revealed transcriptome changes by H2O2 and RSG
RNA-seq was used to probe the associated global transcriptome profile in control, H2O2, and RSG pretreatment + H2O2 conditions (Fig. 2A). Treatment with RSG alone was not evaluated because of the following: (1) it had a nonsignificant effect on cell viability and (2) previous study found it had a minimal effect on the transcriptome of human RPE cells. 24 RSG pretreatment followed by RSG and H2O2 cotreatment was also not evaluated since both RSG pretreatment regimens showed comparable protection. All samples had excellent RNA quality and sequencing quality (Supplementary Table S1). PCA-based visualization of the RNA-seq data set showed distinct separation by condition in both ARPE-19 (Fig. 2B) and MIO-M1 (Fig. 2C) cells, indicative of changes in the transcriptome after H2O2 exposure and RSG pretreatment.
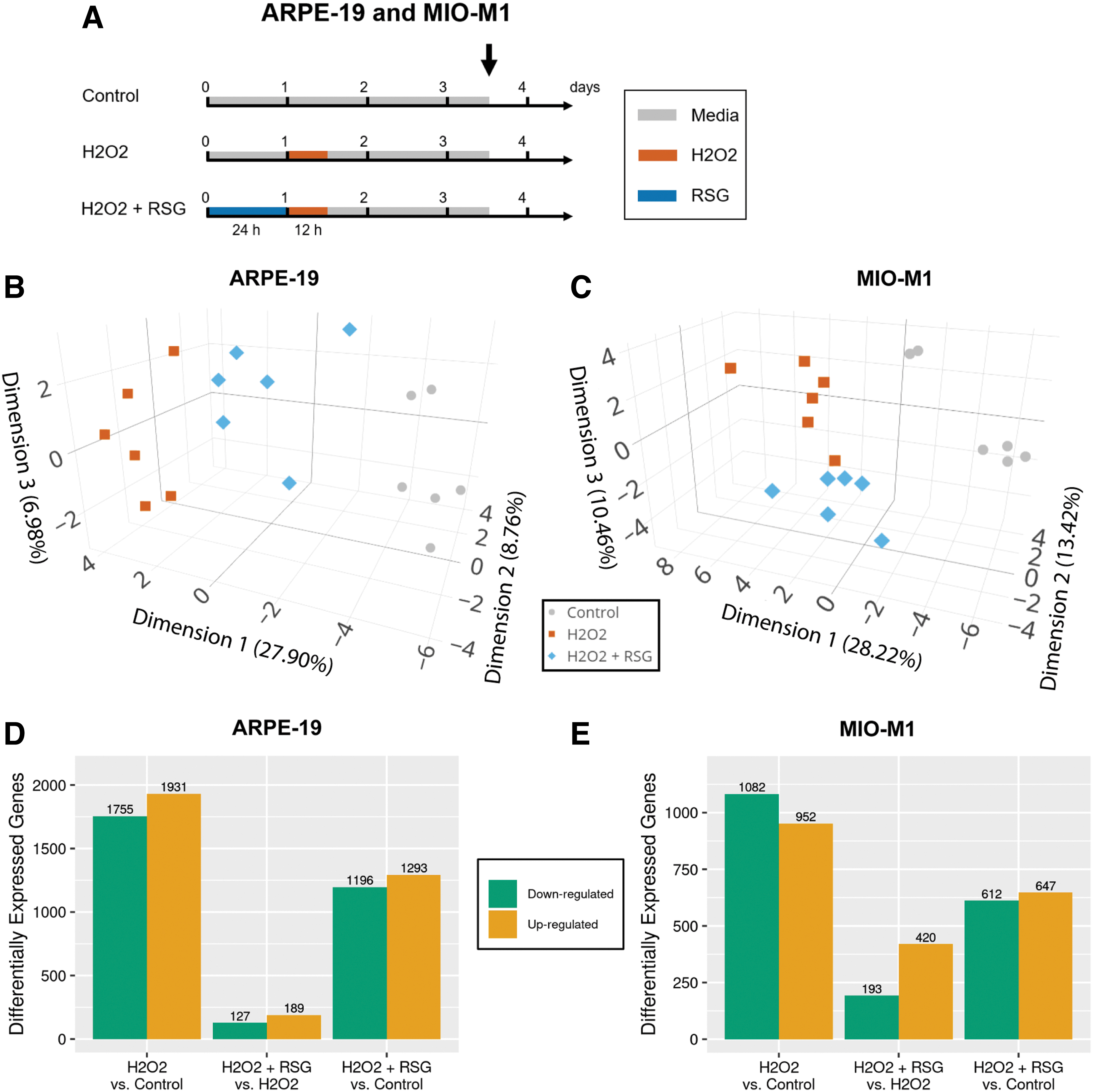
Transcriptome analysis of cells after H2O2 exposure and RSG pretreatment.
Differential expression analysis was performed to identify the DE genes that can be contributed to H2O2 exposure and RSG pretreatment. In both cell models, H2O2 exposure is associated with more DE genes (H2O2 vs. control; 3,686 in APRE-19 and 2,034 in MIO-M1) than RSG pretreatment (H2O2 + RSG vs. H2O2; 316 in ARPE-19 and 613 in MIO-M1), Fig. 2D, E, and Supplementary Fig. S1. In addition, cells pretreated with RSG had fewer DE genes (H2O2 + RSG vs. control; 2,489 in ARPE-19 and 1,259 in MIO-M1) compared with those with no treatment before H2O2 exposure (H2O2 vs. control).
Next, ERSSA confirmed that a sufficient sample size was used in all comparisons to produce a meaningful number of DE genes. As sample size approached n = 6, the number of biological replicates in this study, the average discovery trend plateaued in all comparisons (Supplementary Fig. S2). Additional samples are unlikely to significantly improve the current differential expression discovery.
RSG suppressed transcriptome changes by H2O2
Biological processes that are overrepresented (ie, statistically enriched) in the DE genes were identified. In ARPE-19 cells, 590 and 236 biological processes were enriched with H2O2- and RSG pretreatment-regulated genes, respectively (condensed and visualized in Fig. 3A, B). Of the 236 processes enriched with RSG pretreatment-regulated genes, a majority (61%, 145/236) were also enriched with H2O2-regulated genes (Fig. 3C). The enriched processes were further grouped into major categories, which found that H2O2 uniquely modulated processes related to cell communication, cytoskeleton organization, exocytosis, the immune system, and response to cytokines. Significantly, both H2O2 and RSG pretreatment regulated genes involved in cell adhesion, migration, death, and proliferation; angiogenesis; development; extracellular matrix (ECM) organization; metabolism; and response to growth factor (GF) and stimulus.
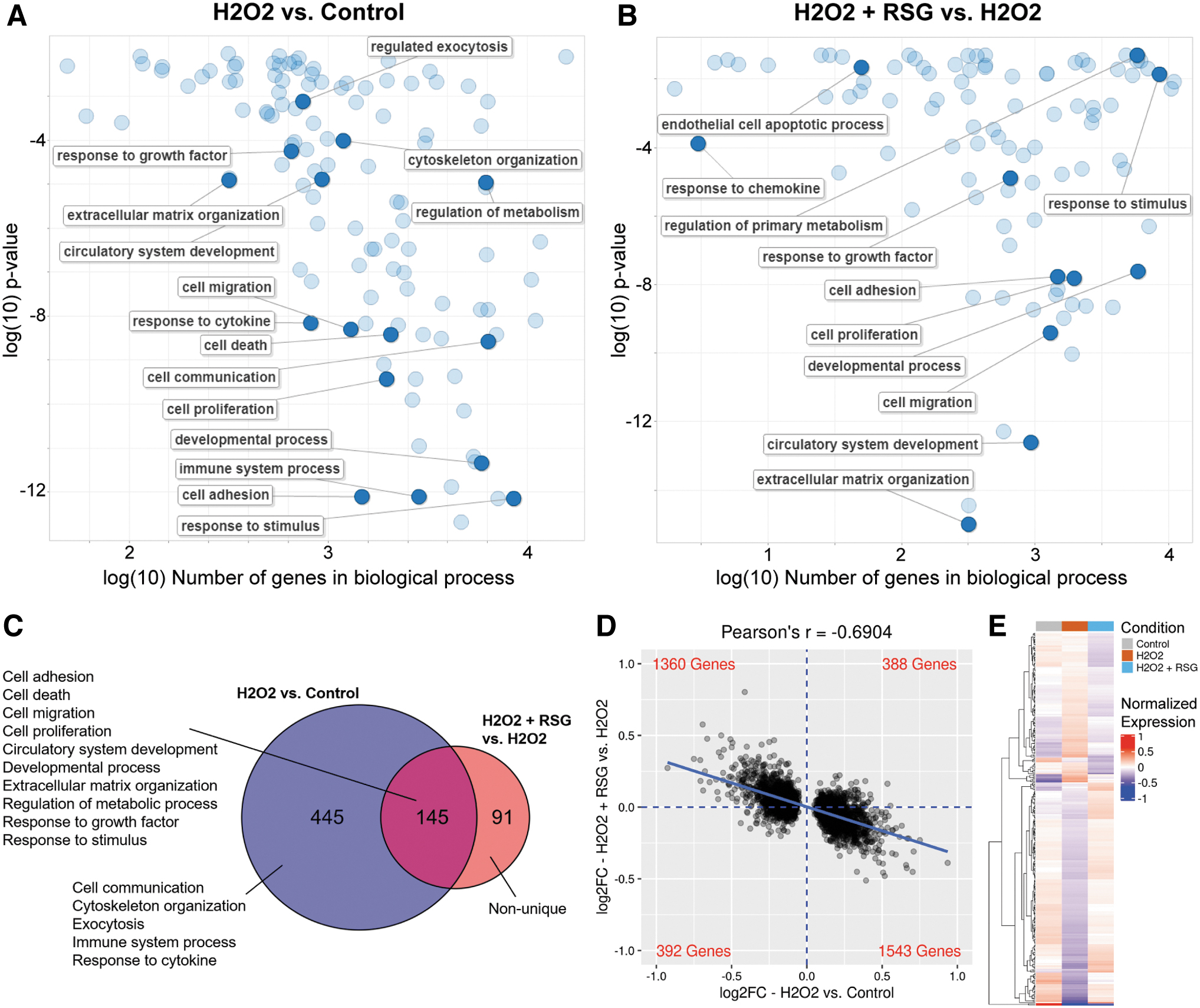
RSG pretreatment suppressed H2O2-induced expression changes in ARPE-19 cells.
The strong overlap prompted a closer evaluation of the gene expression changes. When all 3,686 genes regulated by H2O2 were evaluated, there was a notable inverse relationship (r = −0.6904) between the expression changes associated with H2O2 exposure and RSG pretreatment (Fig. 3D). The same inverse expression profile was observed in each of the biological process categories regulated by both the H2O2 and RSG pretreatments (Supplementary Fig. S3). In addition, the expression level of RSG pretreatment-regulated genes matched more closely with the control condition than the H2O2 condition (Fig. 3E).
Lastly, an analysis of the overlap in the DE genes showed that almost all (95%, 208/219) of the genes regulated by both the H2O2 and RSG pretreatments had expression changes in opposite directions, that is, genes upregulated by H2O2 exposure were downregulated by RSG pretreatment and vice versa (Table 2). Among them, genes with the largest expression changes are involved in cell proliferation and death (FOS, FOSB, EGR1, CTGF, PDGFA, and FAIM), stress response (SGK1), and cell adhesion and migration (CYR61, THBS1, COL4A1, COL5A1, POSTN, and ADAMTS9). RSG pretreatment also upregulated 24/27 (88.9%) of “wound healing” genes, which are involved in restoring integrity to a damaged tissue following an injury (Supplementary Fig. S7A).
Differentially Expressed Genes in ARPE-19 Cells
Genes upregulated, downregulated, or not DE are labeled as up, down, or no change, respectively.
A comparable expression profile was observed in MIO-M1 cells. In this study, 402 biological processes were enriched with H2O2-regulated genes and 347 with RSG pretreatment-regulated genes, of which 55% (192/347) were enriched by both (Fig. 4A–C). When biological processes were grouped into categories, 13 major categories were found to be regulated by both treatments, including cell adhesion, migration, communication, death, and proliferation; angiogenesis; development; ECM organization; immune system process; metabolism; and response to GF, cytokine, and stimulus. Evaluation of the 2,034 genes regulated by H2O2 showed an inverse relationship (r = −0.6472) between the expression changes associated with the 2 treatments (Fig. 4D).
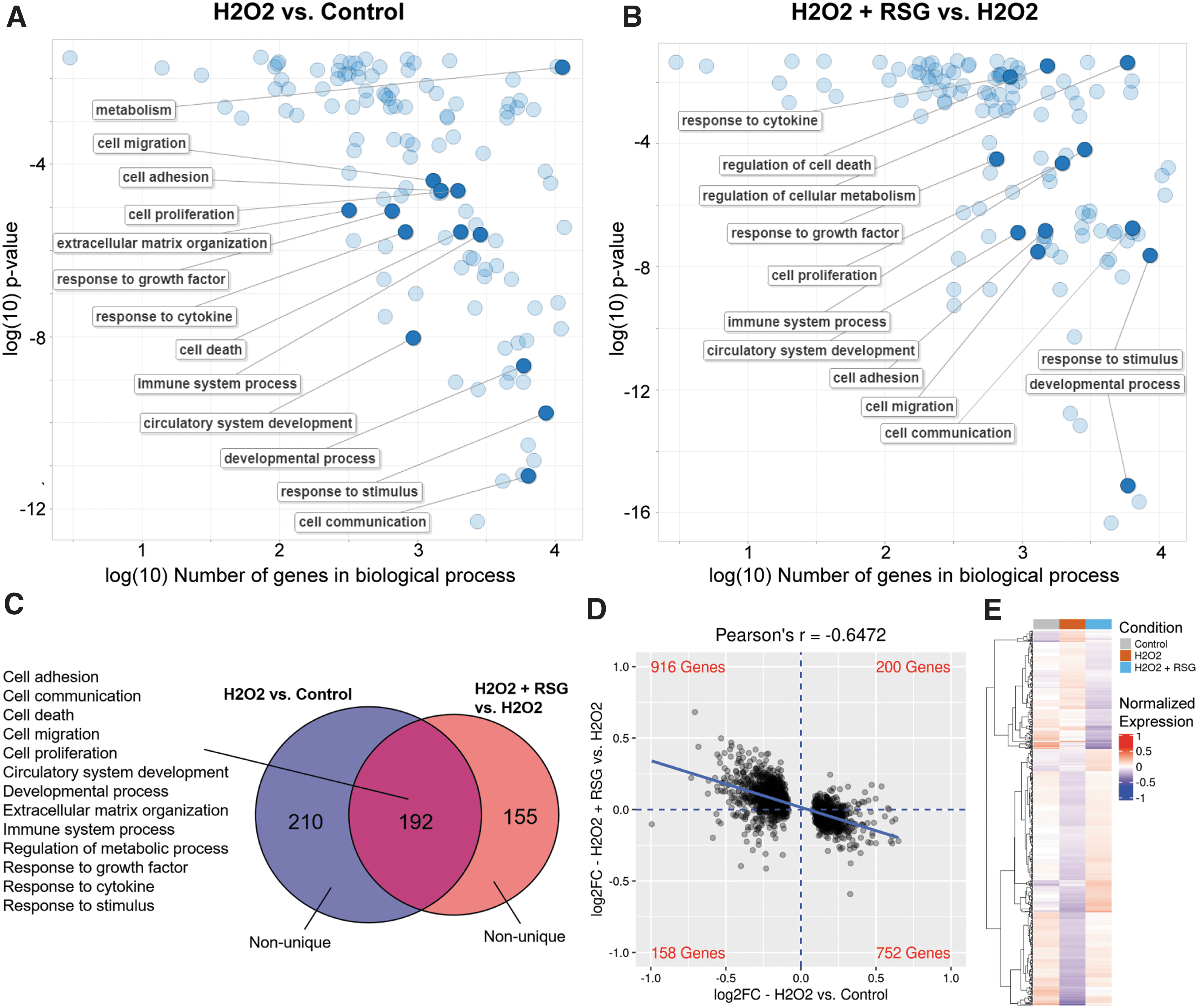
RSG pretreatment suppressed H2O2-induced expression changes in MIO-M1 cells.
Analysis of each of the 13 biological process categories showed the same inverse correlation in expression changes (Supplementary Fig. S4). Similarly, the expression level of RSG pretreatment-regulated genes matched more closely with the untreated control than the H2O2 condition (Fig. 4E). Lastly, an overlap in DE genes showed that a majority (93%, 264/284) of them had expression changes in opposite directions (Table 3). Among them, genes with largest expression changes are involved in cell proliferation and death (BMF, MEGF10, BACH2, and TEAD4), angiogenesis (THSD7A), and cell adhesion and migration (FMN1, MYO1, SYNE1, SYNE2, HMCN, MACF1, COL12A1, COL6A3, COL1A1, COL13A1, SPP1, and MMP3). RSG pretreatment also upregulated 21/40 (52.5%) “would healing” genes (Supplementary Fig. S7B).
Differentially Expressed Genes in MIO-M1 Cells
Genes upregulated, downregulated, or not DE are labeled as up, down, or no change, respectively.
Biological pathway enrichment analysis
Next, an analysis was done to identify the biological pathways that were overrepresented in the DE genes. In both cell models, 3 main groups of pathways were significantly enriched: integrin, AP-1, and syndecan signaling pathways (Table 4). In ARPE-19 cells, the beta1 integrin pathway was enriched with H2O2-regulated genes, while 3 integrin pathways (beta1, beta3, and integrins in angiogenesis), AP-1 transcription factor pathway, and 2 syndecan signaling pathways (syndecan 1 and 2) were enriched with RSG pretreatment-regulated genes. The beta1 integrin pathway was enriched by both analyses and showed the strongest statistical significance in the data set. In addition, a majority of pathway genes were downregulated by H2O2 and upregulated by RSG (Supplementary Table S1). The genes with the largest expression changes by RSG included FOS, EGR1, CCL2, and CXCL8 in the AP-1 and syndecan pathways, and COL4A1, COL5A1, THBS1, ITGA7, and MMP2 in the integrin and syndecan pathways.
Biological Pathways Enriched with Differentially Expressed Genes in ARPE-19 and MIO-M1 Cells
The number of DE genes found in an enriched pathway.
FDR, false discovery rate.
In MIO-M1 cells, 2 AP-1 pathways (AP-1 transcription factor, and targets of AP1 family members Fra1 and Fra2) and 2 integrin pathways (beta1 and integrin in angiogenesis) were enriched with H2O2-regulated genes, while 3 integrin pathways (beta1, beta3, and integrins in angiogenesis) and syndecan-1 signaling pathway were enriched with RSG pretreatment-regulated genes. Integrins in angiogenesis and beta1 integrin pathways were enriched by both analyses. Similar to ARPE-19, a majority of regulated pathway genes were downregulated by H2O2 and upregulated by RSG (Supplementary Table S1). The genes with the largest expression changes by RSG included COL1A1, COL6A3, COL4A4, COL12A1, COL9A3, and COL13A1 in the integrin and syndecan pathways, and CSPG4, LAMA1, LAMA2, and SPP1 in the integrin pathways.
Expression validation with qRT-PCR
RNA-seq revealed that H2O2 and RSG pretreatment regulated many genes in biological processes relevant to disease pathogenesis. To confirm RNA-seq findings, 12 RSG pretreatment-regulated genes each from ARPE-19 and MIO-M1 were analyzed by qRT-PCR with mRNA collected from cells processed for RNA-seq. The 12 genes included 6 genes involved in cell proliferation and death and 6 genes involved in ECM, cell adhesion, and cell migration (Table 1). For a majority of the genes (CCN1, CCN2, EGR1, FAIM, FOS, HBEGF, NEDD9, SGK1, and THBS1 in ARPE-19 cells; BACH2, BMF, COL1A1, COL6A3, EGR1, FMN1, MEGF10, MMP3, PGF, SGK1, and THSD7A in MIO-M1 cells), there was high consistency in gene expression profiles between RNA-seq and qRT-PCR (Supplementary Figs. S5 and S6).
Several genes (ADAMTS9, PDGFA, and POSTN in ARPE-19 cells; COL13A1 in MIO-M1 cells) showed less consistent expression profiles between the measurement methods, but their direction of expression changes appears to be uniform. Indeed, in both ARPE-19 and MIO-M1 cells, strong correlations (r > 0.93) were observed between RNA-seq and qRT-PCR data (Fig. 5).
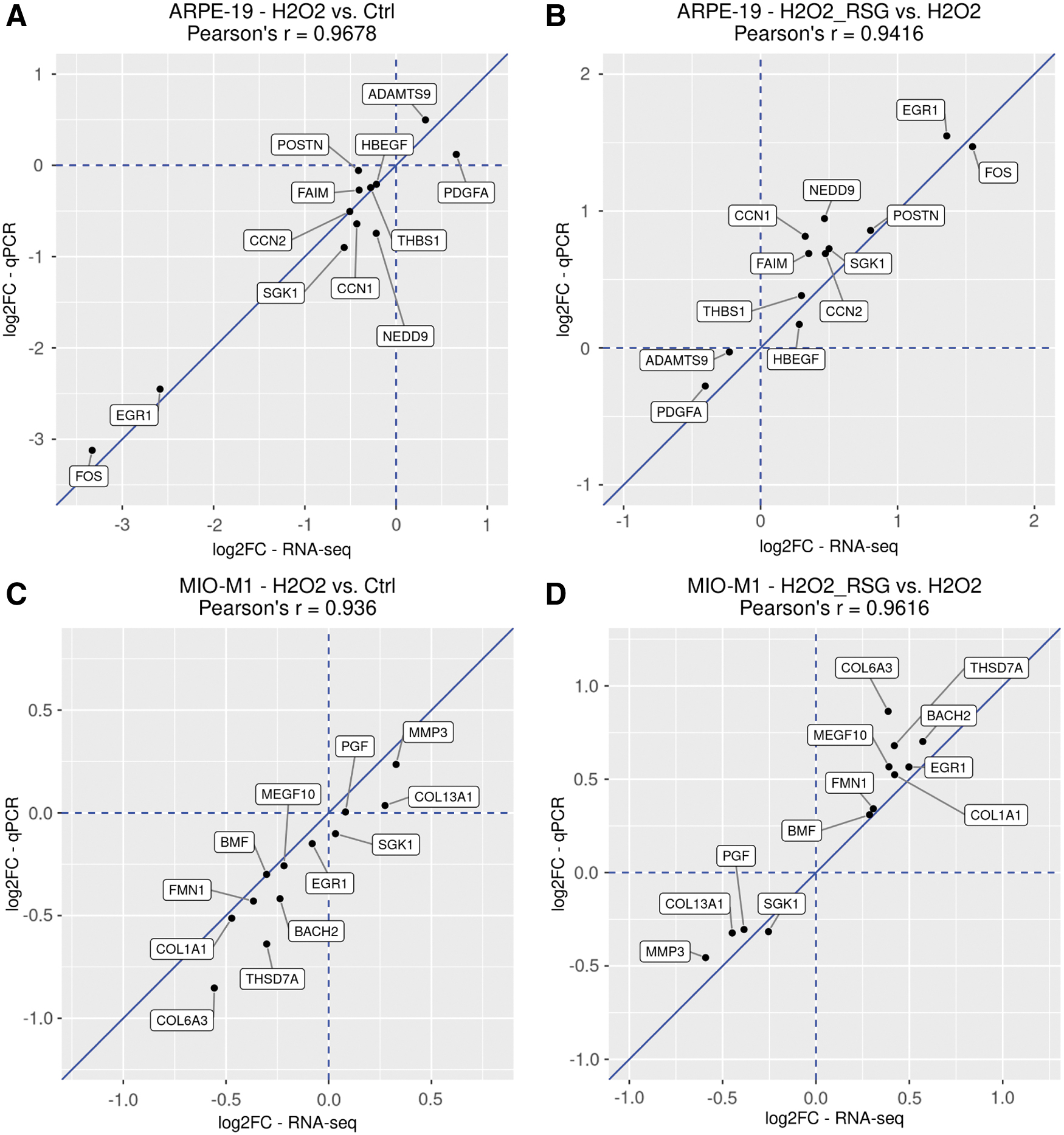
Expression fold change of selected genes analyzed by RNA-seq and qRT-PCR. RSG pretreatment-regulated genes identified through RNA-seq were selected for qRT-PCR validation. Fold change and Pearson's correlation values are visualized. ARPE-19 results are shown in
Discussion
AMD and DR are major causes of blindness in developed nations and oxidative stress contributes to the disease development. In the affected retina, RPE and Müller cells are activated to provide an initial neuroprotective role, but eventually contribute to the development of pathological neovascularization, inflammation, and tissue degeneration.14,20 While anti-VEGF therapies are routinely used to treat neovascularization in a subset of patients, no therapeutics are approved for reduction of oxidative stress and retinal degeneration. 47 This study used cell culture and transcriptomic approaches to study the effect of the oxidant H2O2 and the investigational drug RSG in human RPE (ARPE-19) and Müller (MIO-M1) cell lines.
H2O2 is a common by-product of cellular metabolism and is essential for cell growth at low levels. 48 However, excess H2O2 is cytotoxic and is routinely used experimentally to study oxidative stress and cell death. 49 H2O2 has shown to induce retinal cell apoptosis in both in vitro 50 and in vivo 49 settings. In this study, 12 h of H2O2 treatment significantly reduced cell viability in both ARPE-19 and MIO-M1 cells. In contrast, RSG had no detectable effect on viability of unstressed cells, while RSG pretreatment significantly reduced cell viability loss from H2O2. These findings corroborate earlier studies that found RSG protected human donor RPE cells damaged by H2O2 and a different toxin, hydroquinone.24,51
In addition, earlier work found that RSG pretreatment was much more effective in preventing cell death than post-treatment, perhaps because pretreatment prevents the cells from reaching a certain threshold the point at which they are committed to cell death and cannot be rescued. 52 This also connects to the clinical setting where RSG improved vision in dry AMD patients with early disease stage, before significant and irreversible retinal damage has occurred. 22 In the advanced stage of dry AMD (geographic atrophy), where retinal damage is extensive, 2 late-stage investigational drugs (Zimura and APL-2) can only slow down but not stop lesion growth. 53 Thus, we combined drug pretreatment and RNA-seq to further investigate the regulatory dynamics involved in H2O2 cytotoxicity and protection by RSG.
Using a sufficiently large set of biological replicates, the RNA-seq data showed H2O2 exposure altered the expression of 3,686 genes in ARPE-19 and 2,034 genes in MIO-M1 cells. In contrast, the effect of RSG pretreatment was relatively small, involving 316 genes in ARPE-19 and 613 genes in MIO-M1 cells. When evaluated against untreated control, both cell models pretreated with RSG before H2O2 exposure had fewer DE genes than cells treated with H2O2 alone, suggesting RSG suppressed the effects of H2O2. This is supported by an analysis of H2O2-regulated genes that found RSG pretreatment effectively mitigated expression changes associated with H2O2 exposure.
A functional analysis of the regulated genes identified the biological processes and pathways affected by H2O2 exposure and RSG pretreatment. Intriguingly, many disease-relevant biological processes were regulated by both treatments, including cell adhesion, migration, death, and proliferation; ECM organization; angiogenesis; metabolism; immune system process; and others. Regulation of cell death and proliferation genes is consistent with the observed changes in the cell viability experiment. Elevated oxidative stress has also been associated with the development of retinal neovascularization, 54 inflammation, 55 and metabolic dysfunction.56,57 In addition, reorganization of ECM and dynamic changes in cell adhesion and migration are key cellular processes during retinal disease progression, such as during neovascularization and inflammation.58,59 A more detailed analysis found that while many of the same biological processes were coregulated, they have opposite gene expression profiles, further indicating that RSG treatment protected cells by suppressing the effects of oxidative stress.
Previous studies also found RSG beneficially regulated disease-relevant genes in RPE cells,24,25 but this is the first report of a similar behavior in a Müller cell line. To confirm the observed RNA-seq expression profiles, we used qRT-PCR to validate the expression of 12 genes each in ARPE-19 and MIO-M1 cells, selected for known function in ECM organization and cell adhesion, migration, death, and proliferation. Overall, strong positive correlations were detected between RNA-seq and qRT-PCR data, providing greater confidence in the whole transcriptome data set.
In the biological pathway analysis, 3 main groups of pathway were identified: integrin, AP-1, and syndecan signaling pathways. Similar to the biological process analysis, there was considerable overlap between the pathways affected by H2O2 and RSG pretreatment. Interaction between integrins and their ECM ligands influences most cellular functions, including cell adhesion, migration, and survival decisions. 60 Under oxidative stress conditions, integrin trafficking and expression changes have been reported to regulate cell adhesion and survival.60,61 In the context of RSG biology, regulation of integrin signaling pathways is not surprising as the drug has been shown to modulate expression of integrin genes and integrin-associated processes such as cell adhesion and migration.24,25
However, regulation of AP-1 and syndecan pathways by RSG under oxidative stress condition has not previously been reported. The AP-1 transcription factor is known to play a critical role in regulating a diverse array of cellular response to pro-oxidant conditions, including activation of antioxidative processes, inflammation, and cell survival decision.62,63 Regulation of AP-1 pathways by RSG may be significant, as AP-1 has been shown to be involved in both angiogenesis and inflammatory signaling in the retina. 64
Syndecans are a small family of membrane receptors known to interact with actin cytoskeleton, ECM, GF, intracellular kinases, and integrins to regulate cell survival, proliferation, adhesion, and angiogenesis. 65 It has been documented that oxidative stress induces syndecan activation and is associated with immune chemotaxis, wound healing, and fibrogenesis in diseases. 66 While our expression data suggest that RSG acts to minimize peroxide's effect on these pathways, the exact role it plays in regulating these pathways during elevated oxidative stress condition is unclear and warrants further investigation.
A vast amount of existing literature indicates that the pathophysiology of human retinal diseases is incredibly complex67–69 and cannot be captured by any single experimental model.70–72 The immortalized RPE and Müller cell lines used in the study are cultured in vitro, isolated from other relevant retinal cells, including photoreceptors, microglia cells, and endothelial cells, that play a role in disease pathophysiology. In addition, the ARPE-19 cells in this study were stressed at subconfluent density as it has previously been reported that high-density RPE cell culture, which better resembles a physiological RPE monolayer, is highly resistant to peroxide stress. 29 Nonetheless, cell culture models are highly reproducible and can provide insight into the underlying disease mechanisms and potential therapeutic interventions. In this study, we demonstrated that oxidative stress-induced cell death in in vitro RPE and Müller cell lines is accompanied by profound transcriptome changes in disease-relevant processes.
These observations suggest that oxidative stress not only activates cell death but also modulates other pathways that could further exacerbate retinal dysfunction and degeneration. Indeed, the current literature supports the existence of positive regulation between oxidative stress, inflammation, and angiogenesis in AMD, 73 DR, 74 and other disorders. 75 Given the important role oxidative stress has been recognized in the development of retinal diseases, multiple antioxidants have been tested in preclinical and clinical settings with varying levels of success.53,76 At present, the only accepted therapeutic intervention for dry AMD is a combination of antioxidant vitamins and minerals, but the beneficial effect is modest. 77 For RSG intravitreal injection, 3 Phase 2 clinical trials demonstrated significant visual acuity improvement in patients with dry AMD and DME.22,23
Our current findings suggest RSG's therapeutic effect may involve favorably regulation of cell death, neovascularization, inflammation, and metabolic dysfunction, which are involved in the pathophysiology of the clinically studied retinal diseases.24,25 These results expand our understanding of RSG's mechanism of action and support further clinical trials in the currently targeted indications and other pathologically relevant retinal diseases such as retinitis pigmentosa and glaucoma. More broadly, these pathological mechanisms are present in many human diseases, where RSG may potentially serve as a novel therapeutic option.
Conclusions
In summary, we found RSG has a cytoprotective effect in ARPE-19 and MIO-M1 cells against H2O2-induced cell death. Whole transcriptome analysis showed genes in many disease-relevant biological processes (including cell adhesion, migration, death, and proliferation; ECM organization; angiogenesis; metabolism; and immune system processes) were regulated by both H2O2 exposure and RSG pretreatment, but in opposite directions. Pathway analysis showed novel insight that under oxidative stress conditions, RSG regulated integrin, AP-1, and syndecan pathways. Altogether, our results suggest RSG exhibits cytoprotective properties in cells under oxidative stress, which could lead to novel therapy for retinal degenerative diseases.
Footnotes
Acknowledgment
The authors thank Jan Beiting for her helpful suggestions in revising the article.
Authors' Contributions
Conceptualization: Z.S., J.P., H.K., V.K., and M.C.K. Formal analysis: Z.S., M.C., and M.C.K. Funding acquisition: J.P., H.K., V.K., and M.C.K. Investigation: Z.S., M.C., and S.A.R. Methodology: Z.S. and M.C. Project administration: M.C.K. Resources: Z.S., M.C., S.A.R., J.P., H.K., and V.K. Software and visualization: Z.S. Writing—review and editing: Z.S. and M.C.K.
Author Disclosure Statement
Z.S., J.P., H.K., and V.K.: employees of Allegro Ophthalmics, LLC.
M.C.K.: research programs are supported by the Discovery Eye Foundation, a 501(c)3 nonprofit organization. She serves as a board member for DEF. The terms of this arrangement have been reviewed and approved by the University of California, Irvine, in accordance with its conflict-of-interest policies. M.C.K. collaborates with Allegro Ophthalmics, LLC.
M.C. and S.R.A.: None.
Funding Information
This work was supported by the Discovery Eye Foundation, National Eye Institute R01 EY027363, and, in part, by an unrestricted departmental grant from Research to Prevent Blindness. M.C.K. was the primary recipient of these awards. Allegro Ophthalmics, LLC, funded Z.S., J.P., H.K., and V.K. The Sponsors/Funders did not play any role in the study design, data collection and analysis, decision to publish, or preparation of the article.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
