Abstract
Over the last two decades, adeno-associated viruses (AAVs) have been widely used as viral vectors in gene therapy due to their ability to infect both dividing and nondividing cells, broad tissue specificity, and favorable safety profile. Recombinant AAV (rAAV) production requires a helper virus, typically adenovirus (AdV), which provides essential genes for AAV replication. However, the increasing demand for safer and more efficient rAAV production methods led to the need to develop helper plasmids with minimal AdV components. In this study, we evaluate the impact of AdV E2 and E4 in the productivity and genome packaging of rAAV serotypes 2, 5, 8, and 9, produced by transient transfection. We designed and tested eight novel helper plasmids with different deletions in E2 and E4 genes. Results indicated that deletions in these genes significantly affected rAAV productivity and packaging, particularly for serotypes 8 and 9. Helper plasmids containing minimal essential genes—E2-DBP, E4orf6, and VA RNA—showed near to 10-fold reduction in viral genome packaging compared to the control. However, including E2 L4-22/33K and E4orf3 regions significantly improved viral production, particularly for serotypes 8, and 9. In this study, we also demonstrated that the full E4 gene is crucial to achieving high full-empty ratios, minimizing the production of empty capsids, and enhancing viral release into the culture medium of rAAV8. Accordingly, we created a smaller plasmid, without adenoviral structural proteins that allows a similar rAAV production across all tested serotypes. Overall, these findings provide insights into the genetic requirements for efficient rAAV production and highlight the importance of the E2 and E4 regions for optimizing viral yield and quality. This approach could lead to the development of improved strategies for large-scale rAAV vector production by enabling safer and more cost-effective systems.
INTRODUCTION
Adeno-associated viruses (AAVs) are nonenveloped viruses composed of an icosahedral protein capsid and a single-stranded DNA (ssDNA) genome of 4.7 kilobases (kb) flanked by two inverted terminal repeats (ITRs). 1 The AAV genome contains two genes, rep and cap, controlled by three viral promoters. The promoter p5 codes for Rep78 and Rep68, which are involved in DNA replication and regulation of viral promoters. 2 –4 The p19 promoter activates the transcription of Rep52 and Rep40 while p40 codes for structural proteins (VP1, VP2 and VP3) and accessory proteins (MAAP and AAP). 5
AAV presents several advantages as a gene therapy vector, including the ability to infect both dividing and non-dividing cells, multiple serotypes that provide broad tissue specificity, and a favorable safety profile. 6 –10 Classified as Dependovirus, AAVs are non-replicative and require co-infection with a helper virus, such as an adenovirus (AdV), to replicate. 11 The presence of wild-type AdV critically modifies the cellular environment, enabling AAV replication and the formation of new virions. AdV serotypes 2 and 5 are the most used for AAV production. 12
The AdV genome is a linear double-stranded DNA (dsDNA) molecule that includes five early genes (E1A, E1B, E2, E3, and E4) and one late transcription unit (L1 to L5). Typically, the genes described as the main helper genes in AAV replication are E1A, E1B-55k, E2-DBP, E4orf6, and viral-associated RNA (VA RNA). 11
The E1A gene is the first to be expressed during AdV infection, leading to the activation of the E2A helper gene and AAV p5 promoter. 13,14 The activation of E1B follows, blocking E1A-induced apoptosis and forming a complex with E4orf6. This interaction promotes the synthesis of the viral ssDNA genome and AAV mRNA export. 15 –17 E2A codes for a ssDNA binding protein, the DBP, that recruits the AAV genome into the nucleus and stimulates DNA replication in vitro. 18 –20 E2A gene products are also responsible for AAV promoter regulation, mainly of p19 and p40, that controls the expression of minor Rep proteins and structural proteins, respectively. 21 Recently, a new promoter inside the 100K region of the E2A gene, named L4P, was discovered. 14 This promoter is activated by E1A, Iva2 and E4orf3 proteins leading to the expression of L4-22K and L4-33K proteins. 14 These proteins seem to be required for AdV and AAV viral DNA encapsidation, regulation of mRNA production, and stability. 22,23 The E4 gene is composed of seven open reading frames (ORFs), however, the interplay between some ORFs of the E4 gene and AAV replication is not fully understood. For example, although the E4orf3 protein interacts with other helper genes such as E2A and E1A, 24 it is not considered essential for recombinant AAV (rAAV) production, as it has been described as having redundant functions with the E4orf6 product. 25 The E4orf6 protein, apart from its interaction with E1B55K, is also responsible for the inhibition of the Mre11-Rad50-Nbs1 (MRN) complex involved in DNA repair mechanisms. 26 Nevertheless, it is also reported that these protein complexes can favor AAV capsids and Rep52 degradation, decreasing the production of rAAV serotype 5. 26 This negative effect can be compensated by VA RNA, which acts as a complex mediator. 27
The most used methods to produce rAAV are triple transient transfection and stable producer cell lines. In transient transfection, cells that constitutively express AdV E1A/E1B genes—typically 293/293T-based mammalian cells—are transfected with plasmid vectors containing the gene of interest (GOI), AAV rep and cap genes, and a helper plasmid containing AdV genes. 28 This technique offers significant advantages, including the absence of infectious AdV in viral preparations and considerable flexibility for serotype and transgene exchange. 29,30 However, the requirement for large quantities of plasmid DNA (pDNA) and the limitation of producing only one batch per transfection not only increase costs but are also the primary sources of batch-to-batch variability. 31 –33
To address these challenges, rAAV production can alternatively be performed using stable producer cell lines, in which the AAV components and/or the transgene, are stably integrated into the host cell genome. 34 Several strategies have been implemented in this regard, and viral production is typically initiated either by infection with AdV (e.g., in HeLa or A549 cells) or by induction of helper factor expression in cell lines where these components are also integrated (e.g., 293 cells) (reviewed in Fernandes et al. 34 ). The latter cell lines offer an increased safety profile and the potential for lower production costs due to the absence of the AdV helper virus in the production process. However, these platforms have yet to achieve titers comparable to those obtained through transient transfection-based production methods. 35,36 This discrepancy could be attributed to (1) inadequate suppression of toxic AAV and helper component expression, leading to cell instability and reduced rAAV production over time, 37 and/or (2) insufficient understanding of the minimal components required for efficient rAAV production.
Advancing our knowledge of the biological mechanisms underlying intracellular rAAV production is crucial to overcoming these challenges and developing more efficient production systems. Therefore, our study aims to elucidate the interactions between AAV and helper virus components by evaluating the impact of E2 and E4 AdV genes within the context of rAAV production by transient transfection. Four rAAV serotypes, 2, 5, 8, and 9 were tested.
MATERIALS AND METHODS
Plasmid construction
The Helper plasmid used in this study contained AdV helper factors E2A, E4, and VA RNA and was obtained from AAV Helper Free System (#240071, Agilent Technologies, Santa Clara, CA). This plasmid, herein named Ctr (control), was used as the backbone for eight additional helper constructs (Fig. 1, Supplementary Table S1). The generation of recombinant helper plasmids was performed using gene synthesis services by IDT (Integrated DNA Technologies, Coralville, IA) and/or by PCR DNA amplification (Phusion High-Fidelity DNA Polymerase, Thermo Fisher Scientific, Waltham, MA). PCR products and gel extraction were purified using Illustra GFX PCR DNA and Gel Band Purification kit (GE Healthcare, Chicago, IL). Restriction enzymes were from NEB. Inserts were assembled by an In-Fusion cloning strategy (Takara Bio, Otsu, Shiga, Japan). The resulting plasmids were transformed into Stellar™ Competent Cells (Takara Bio) and selected in the presence of 100 μg/mL ampicillin. The plasmid sequence was screened by restriction enzyme digestion and DNA sequence analysis (STAB VIDA, Lisbon, Portugal).
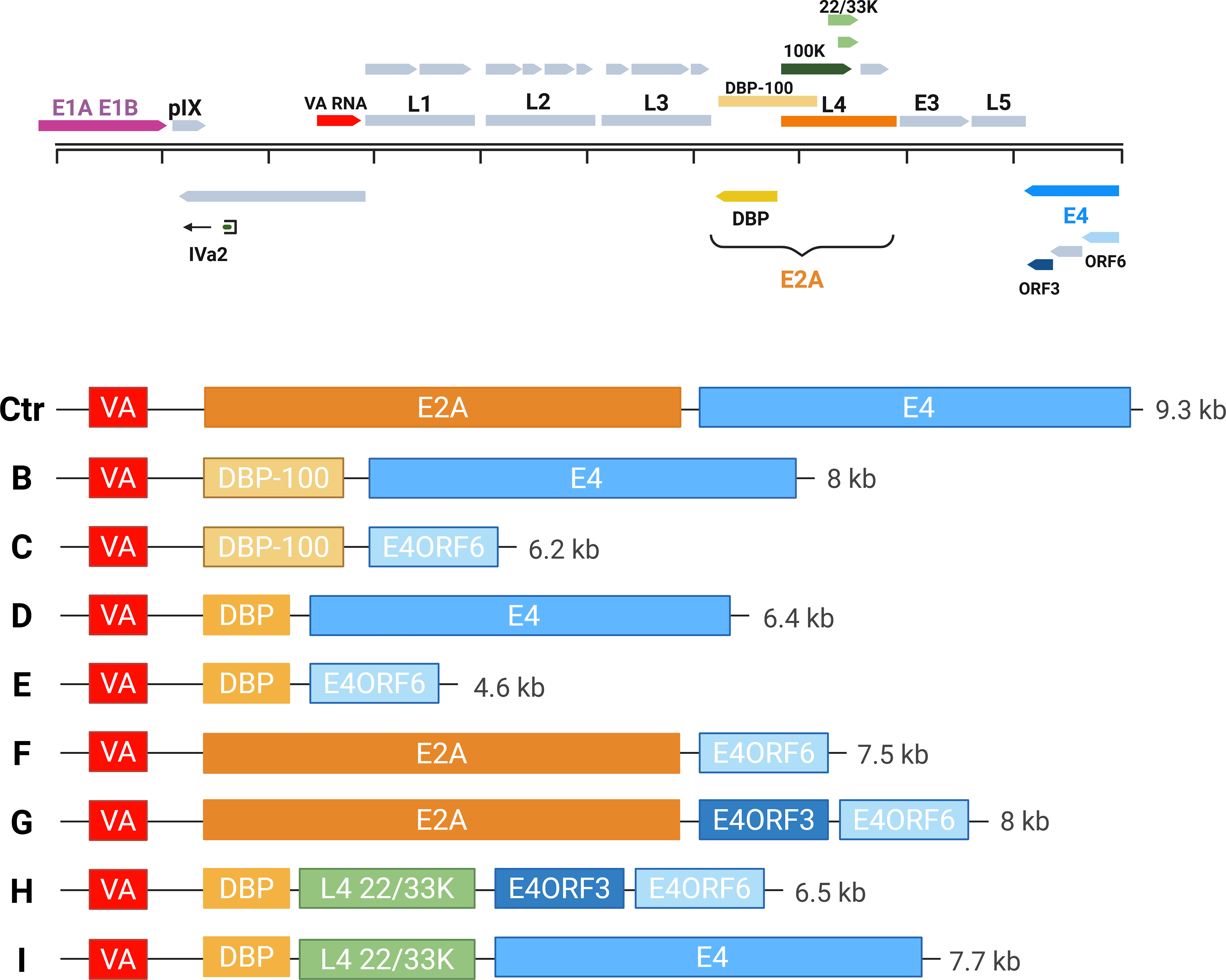
Design of new helper plasmids. Schematic representation of the adenovirus serotype 2 genome and the different combinations of E2 and E4 helper genes from AdV serotype 2 present in each of the tested helper constructs, along with the size of each helper cassette. All helper plasmids tested contain the same VARNA. Ctr corresponds to the control backbone of pHelper. pE only contains the minimal essential helper genes described in the literature, the E2A-DBP and E4orf6. The AdV genes involved in AAV replication are marked in color. The E1A/E1B gene is present in the HEK293 cells. E2A-DBP corresponds to the DNA binding protein region within the E2A gene. DBP-100 is a combination between a portion of L4-100K and the previously mentioned E2A-DBP. L4-22/33K comprises the L4P promoter and the regions 22K and 33K of L4, within the E2A gene.
Cell lines and culture conditions
293 viral production cells (VPC) (Gibco, Thermo Fisher Scientific) were adapted in-house to serum-free media BalanCD (FUJIFILM Irvine Scientific, Tokyo, Japan) with 6 mM GlutaMAX (Gibco, Thermo Fischer Scientific), by sequential approach following a similar strategy as described here. 38 293 VPC cells grew under suspension conditions using vented shake-flasks (Corning Life Sciences, Corning, NY), at 37°C in a humidified atmosphere with 8% (v/v) CO2 and an agitation rate of 140 rpm. HeLaRC32 (ATCC-CRL-2972) and HT1080 human epithelial adherent cells (ATCC-CCL-121), were obtained from the American Type Culture Collection (Manassas, VA), and cultured in adherent conditions and expanded using Dulbecco’s modified Eagle’s medium (DMEM) (# 10–013-CVR Corning, Berlin, Germany) with 10% (v/v) fetal bovine serum (FBS, Gibco, Thermo Fischer Scientific) and maintained at 37°C in a humidified atmosphere containing 8% CO2. Cell concentration and viability were evaluated by the trypan blue exclusion method and counted using CytoSMART live-cell imager (CytoSMART, Eindhoven, The Netherlands).
rAAV production by triple transient transfection of 293 VPC cells
For rAAV production, 24 h prior to transfection 293 VPC cells were seeded at 1.4 × 106 cells/mL. On transfection day, cells were diluted to a final density of 2 × 106 cells/mL and transfected with 1 μg of plasmid DNA per 106 viable cells, at a 1:1.5 (w/w) DNA: PEIpro (Polyplus, Strasbourg, France) ratio. The plasmid ratio for each helper plasmid, each pRC, and each transgene was 1:1:1 by mass. pDNA and PEIpro were diluted in BalanCD medium supplemented with 6 mM of GlutaMax (1/10 volume of the cells) and incubated at room temperature for 12 min before addition into cell culture. Cells were incubated at 37° C, 8% CO2, shaking at 140 rpm. Then, 72 h post-transfection (hpt) cells were harvested to measure the concentration of viable cells and the percentage of GFP-positive cells. The remaining culture was centrifuged for 10 min, 250 × g. The supernatant (extracellular fraction) was filtered at 0.45 μm and stored for analysis. The viruses from the cellular pellet (intracellular fraction) were retrieved using a lysis buffer and methodology described in. 39 For rAAV2 only the intracellular fraction was recovered. Three independent replicates were carried out for each rAAV study, always including the original helper plasmid as a Ctr. The virus present in each fraction was quantified by ELISA for total particles (TP) and qPCR for viral genomes (VG).
The AAV transgene plasmid pITReGFP is flanked by viral ITRs and contains an eGFP fluorescent reporter gene, under the control of a CBA (CMV-Chicken-β-actin) promoter. The pRC8 (Addgene No. 112864) and pRC9 (Addgene No. 112865) were a kind gift from Dr. James M. Wilson, University of Pennsylvania; pRC5 (Addgene No. 104964) was a gift from Melina Fan; pRC2 was commercially available from AAV helper-free system from Agilent Cat. No. 240071).
Flow cytometry data acquisition and analysis
Flow cytometry was conducted in a Gallios Flow Cytometer (Beckman Coulter, Brea, CA) to quantify the percentage and fluorescence intensity of GFP-positive cells. Single cells were isolated from debris and doublets based on their side scatter (SSC) versus forward scatter (FSC) characteristics. Data analysis was performed with FlowJo Software versus 10.9.0 (BD Biosciences; Franklin Lakes, NJ).
rAAV characterization
The quantification of TP of each rAAV production was performed by ELISA using specific Xpress ELISA kits (PROGEN Biotechnik GMBH, Heidelberg, Germany). Absorbance was quantified on an Infinite PRO NanoQuant (Tecan, Männedorf, Switzerland).
The amount of VGs was quantified by qPCR as described here. 40 Briefly, a DNAse step was performed to eliminate free DNA, followed by Proteinase K treatment. The treated sample was then diluted in TE buffer (10 mM TRIS, 1 mM EDTA—pH 8). For qPCR, a calibration curve was performed using a linearized transgene vector (pITReGFP), with a primer/probe set for eGFP in LightCycler 480 Instrument II (Roche Applied Science, Penzberg, Germany).
To assess infectivity of rAAV2 productions, HT1080 cells were seeded at 3.5 × 104 cells/cm2, in 24-well plates. To titrate serotypes 5, 8, and 9, HeLaRC32 cell lines were seeded at 4.0 × 104 cells/cm2, in 24-well plates. After 24 h, both cell lines were transduced with 200 μL of viral supernatant at different dilutions (1:100 to 1:96 000), in DMEM with 2% (v/v) FBS. For HeLaRC32, AdV5 at a multiplicity of infection (MOI) of 50 was also added to viral dilutions. 4 h post-infection, fresh DMEM with 10% (v/v) FBS medium was added to all wells. Transduction results were evaluated at 24 h (AAV5, 8, and 9) or 48 h (rAAV2) by flow cytometry. Transduction titers were calculated based on the percentage of GFP-positive cells, the cell count at the time of infection, and the dilution factor of the viral supernatant. For reliable titers an average of 2 dilutions was used, ensuring >2% of positive cells.
Infectious viral titers were calculated using the following equation:
Statistical analysis
The data were expressed as mean ± SD and analyzed with GraphPad Prism versus 8.0.1 software (San Diego, CA). Statistical significance was determined using multiple t-tests followed by the Holm-Sidak multiple comparisons test, with α = 0.05. Each row was analyzed individually, without assuming a consistent SD. Correlations were determined using Pearson’s correlation coefficient (r) with a 95% confidence interval. A nearly two-fold variation in viral titer was observed across two production runs (1st rAAV study and 2nd rAAV study). In each run, was always used helper Ctr. Therefore, we considered a helper construct to have a significant impact on rAAV production only if it exceeded this variation threshold.
RESULTS
Design and development of new helper plasmids
To produce rAAV by triple transient transfection, several plasmids containing AdV helper components are available, such as Agilent pHelper (#240071) Aldevron pALD-X80 (#5017–10), pALD-Help (#5082–10), or pAdDeltaF6 from Addgene (#112867). These plasmids have the full AdV E2A gene, all seven ORFs of the E4 gene, and the VA RNA. Some, such as pALD-X80 and pAdDeltaF6, also carry structural AdV components such as fiber and hexon proteins. Given that only E2A-DBP, E4orf6, and VA RNA are typically mentioned as essential for rAAV production, we aimed to explore whether rAAV production could be achieved and/or optimized using only these essential helper genes. Additionally, we sought to determine which additional components, if any, may have the greatest impact on viral production. For that, we used the Agilent pHelper plasmid as the Ctr as well as the backbone to generate eight new helper plasmids (pB to pI). All used helper plasmids contain the same VA RNA, and several combinations of AdV helper E2 and E4 genes while maintaining their internal promoters (Fig. 1). Helper plasmids pB, pD, and pI contain only deletions in E2 gene; the pF and pG only in E4 gene and plasmids pC, pE, and pH, deletions in both E2 and E4 gene. The main AdV components evaluated were the E2-DBP, E2-100K, E2 L4-22/33k, E4orf6 and E4orf3. The helper pE contains only the minimal essential AdV genes (VARNA, E2-DBP, E4orf6). This systematic approach enabled us to evaluate the minimal genetic requirements for a successful rAAV production. To increase the safety of the production system, no structural AdV genes are present in the helper plasmids.
The Rep/Cap plasmids (pRC) used in this study contained the rep gene from serotype 2 and the cap gene from each serotype, 2, 5, 8, or 9, without any modification. The same eGFP-transgene plasmid was used in all viral productions.
Assessing rAAV productivity using different helper constructs
Two independent studies were carried out, always including the original helper plasmid as a Ctr. Three independent replicates were carried out for each rAAV study. In the first study, we assessed the rAAV production of 3 serotypes (AAV5, 8, and 9) with helper plasmids pB to pF; in the second study, we used pG to pI, and the AAV2 serotype was included. Viral vectors were collected from both intracellular and extracellular fractions. rAAV2 production was evaluated only the intracellular fraction, as its strong binding to cellular HSPG (heparin sulfate proteoglycans) severely impacts viral release into the supernatant. 41 The virus present in each fraction was quantified separately using ELISA for TP and qPCR for VG (Figs. 2 and 3).
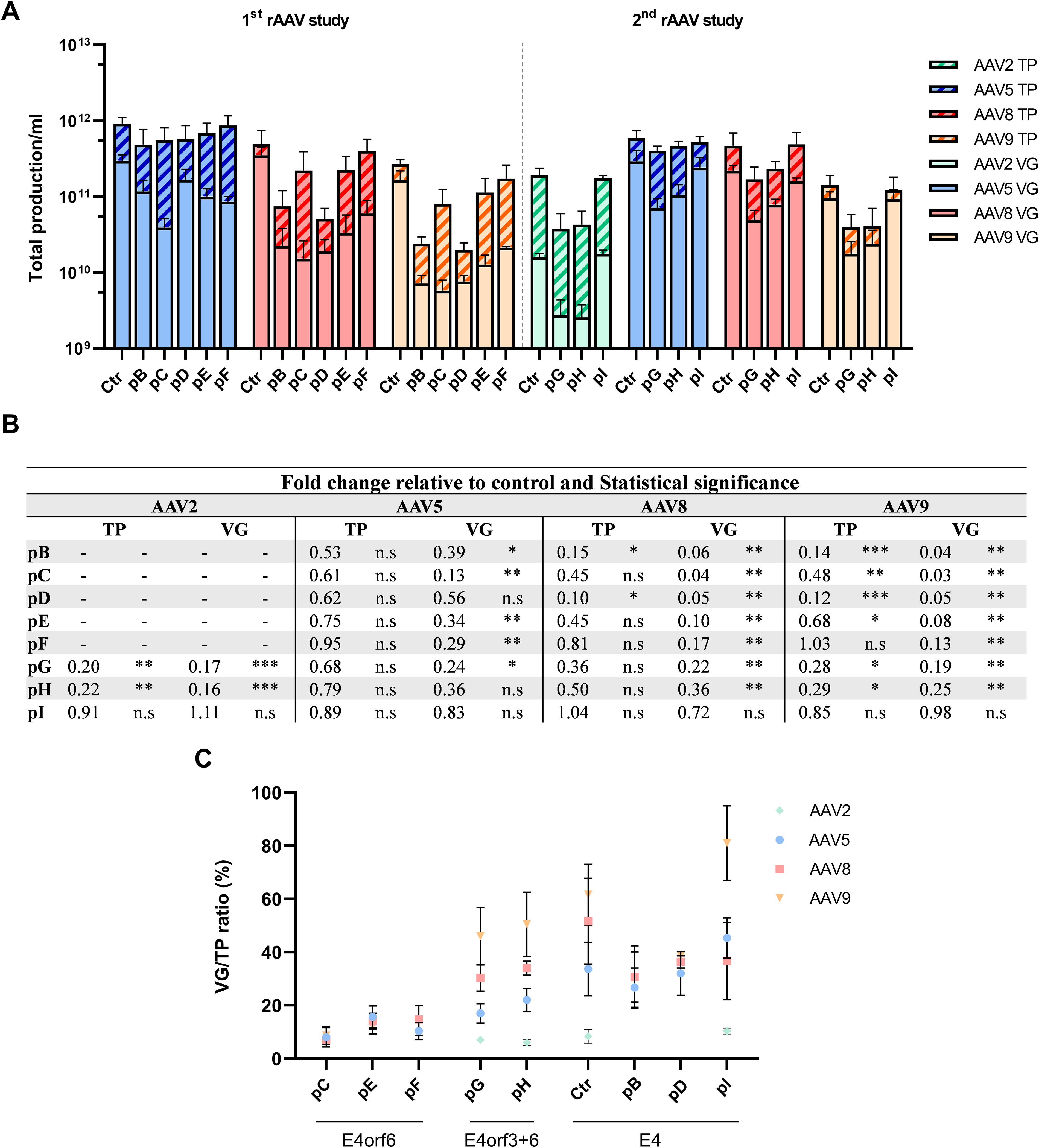
Characterization of rAAV production from four serotypes with different helper plasmids. In the first study, rAAV from serotypes 5, 8, and 9 were produced using pB - pF helper plasmids. For the second study, rAAV from serotypes 2, 5, 8, and 9 were produced using helper plasmids pG - pI. All rAAV were harvested 72 hpt from both intra and extracellular fractions, except for rAAV2 which were only harvested from intracellular fraction. The productivity was characterized by
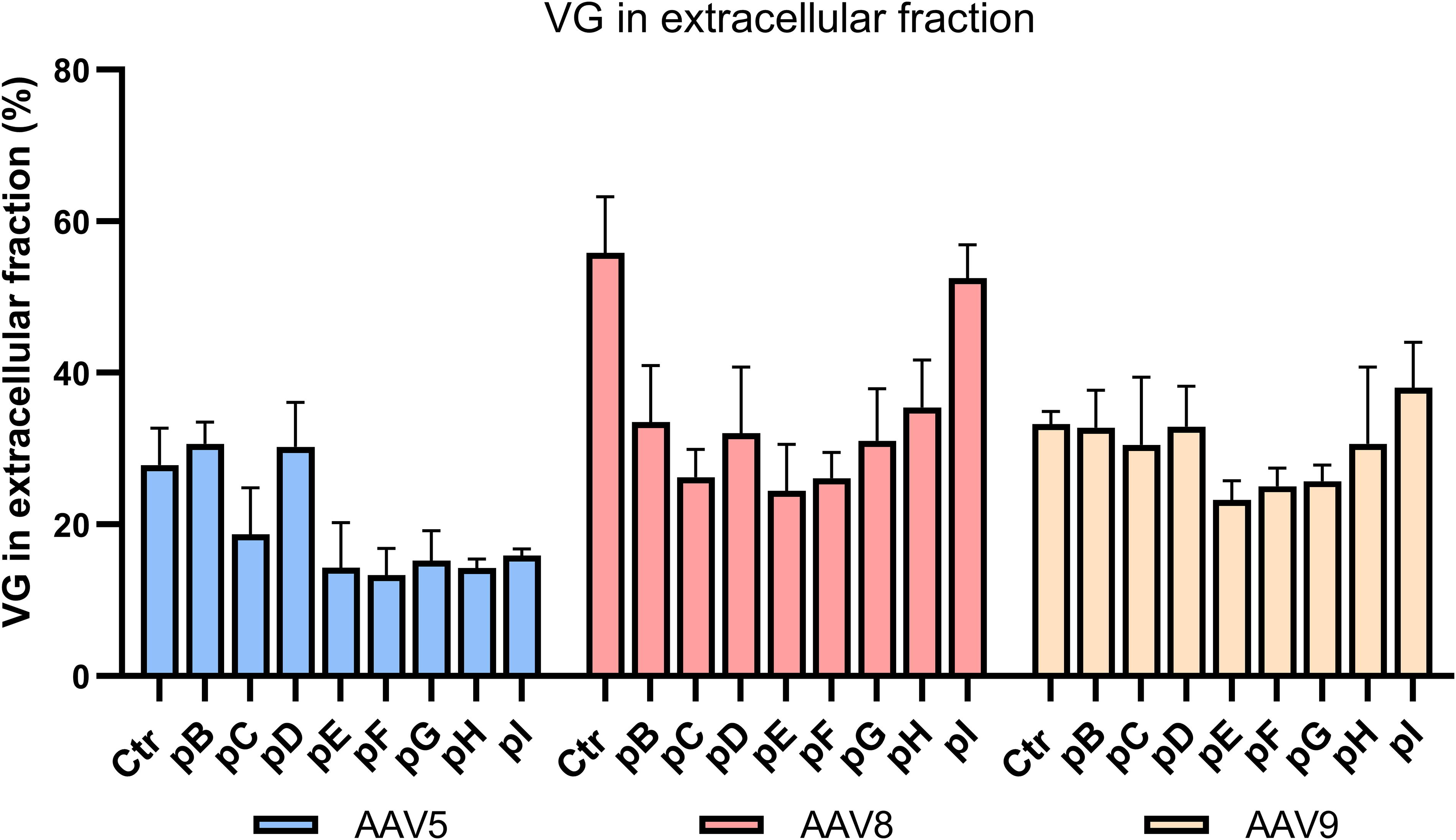
rAAV distribution using different helper plasmids. Distribution of VG full capsids in the cellular fractions according to each helper plasmid used, for serotypes 5, 8, and 9. Values presented correspond to the percentage of VG in the extracellular fraction. AAV2 capsid localization values were not included since only the intracellular fraction was analyzed. Data are presented as the mean of three independent replicates for each study.
Deletions in the AdV E2 and E4 can severely impact rAAV packaging ability
The quantification of all produced virus (TP) showed that serotype 5 was not significantly affected by the different helper plasmid combinations, showing less than a 2-fold reduction in TP across all used helper plasmids (Fig. 2A and B). In contrast, for serotypes 8 and 9, the majority of generated helper plasmids led to a significant decrease in TP (Fig. 2A and B). Helper plasmids containing only parts of the E2A gene combined with the full E4 gene (pB and pD) showed the highest reduction of rAAV TP compared to control, up to 10-fold for rAAV8 and up to 13-fold for rAAV9. Conversely, helper plasmids which included only E4orf6 (pC, pE, and pF) or E4orf3 + orf6, had a lower or no impact on the production of TP (Fig. 2A and B), regardless of E2 regions present.
Remarkably, the highest impact on rAAV productivity was observed in full particles (VG). For serotype 5, the helper plasmids with only E4orf6 (pC, pE, and pF), resulted in a significant reduction of packaged genomes (Fig. 2A and B). For serotypes 8 and 9, the expression of just E2-DBP and/or E2-DPB plus 100k region negatively impacted rAAV production at VG levels (pB to pE) (Fig. 2A and B). The highest VG reduction, up to 30-fold, was obtained with helper plasmid pC, which contains E2-DPB plus 100k region combined with E4orf6.
The results also show that having only the E2-DBP, E4orf6, and VARNA resulted in a decreased VG viral production that can be 10-fold lower for rAAV8 and rAAV9 (Fig. 2).
Adding L4-22/33K region in combination with full E4 gene rescued rAAV8 and rAAV9 titers
To determine whether rAAV packaging during transient transfection is influenced by the L4-22/33K region—as previously observed with AdV and in rAAV production using HeLaRC32 cell lines infected with helper AdV 23,42 —we incorporated additional components such as the E2-L4-22/33K and E4orf3, generating helper plasmids pG - pI. Having just the E2-DBP + E2-L4-22/33K regions or the addition of E4orf3, allowed an improvement in VG titers, compared to other helpers (Fig. 2A and B). The inclusion of E2-DBP + E2-L4-22/33K combined with the entire E4 gene (as in pI) allowed the production of TPs and VGs at similar (rAAV2, 5, 8, and 9) levels of the helper Ctr (Fig. 2A and B).
rAAV quality seems mostly related to the E4 gene
Having the full E4 gene in the generated helper plasmids resulted in a full-empty particle ratio (VG/TP %) that was comparable to that obtained with the helper Ctr (Fig. 2C). The helper plasmids with E4 gene modifications resulted in lower VG/TP ratios across all serotypes. For instance, in serotype 8, the VG/TP ratio dropped from 50%, obtained with plasmid helper control, to below 15% with the pB - F plasmids (Fig. 2C). However, the presence of the E4orf3 region partially rescued rAAV quality, as observed with later helper plasmids generated, pG and pH (Fig. 2C). These results highlight the importance of having the full E4 gene, or at least both E4orf3 and E4orf6, to minimize the number of empty particles.
Changes in E2 and E4 also affect the release of full particles to the culture medium
By separating intracellular and extracellular fractions, we were able to evaluate the impact of several AdV helper components in the release of rAAV5, 8, and 9. For rAAV5, except with plasmids containing the E2-DBP or E2-DBP + 100k region combined with E4 (pB and pD), we observed an increase of full particles remaining intracellularly compared with the control. For rAAV8, only when E2-DBP + 22/33k region was combined with E4 (pI) a similar proportion of VG to Ctr in the supernatant were observed. No major differences were observed for serotype 9 (Fig. 3).
Evaluation of rAAV transduction efficiency
The impact of different helper plasmids on viral transduction efficiency was evaluated in HT1080 for serotype 2 and in HeLaRC32 cell lines for the other serotypes. HT1080, a human epithelial cell line derived from a fibrosarcoma patient’s connective tissue, is widely used in rAAV transduction assays due to its susceptibility to AAV2 infection. 39 HeLaRC32, a stable rAAV packaging cell line with integrated rep and cap genes. HeLaRC32 cells are commonly used for rAAV titration to distinguish between infectious and noninfectious rAAV serotype 8 particles. 43 These results showed that for all serotypes, the pI helper plasmid yielded similar levels of transducing units (TU) compared to the control condition and even performed significantly better than the control in serotype 2 (Fig. 4A and B). Regarding pG and pH, we observed a significant decrease in TU, particularly in serotypes 2 and 5 (Fig. 4A). However, when normalized by the number of VG, these differences became non-significant (Fig. 4B). This shows that, as expected, the viral transduction was positively affected by the VG titer, correlating with the VG/TP ratio (Fig. 2C). 39
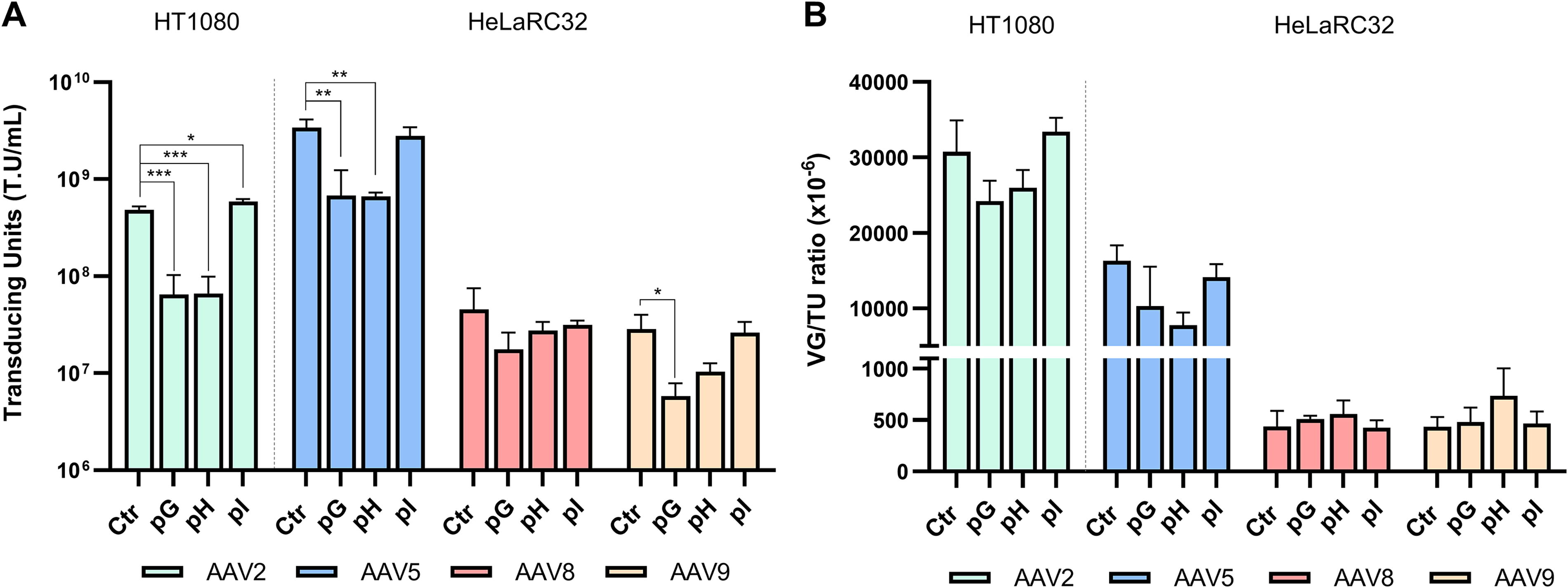
Characterization of rAAV transducing capacity with different helper plasmids.
Helper plasmids with E4 gene deletions resulted in increased eGFP intensity
In addition to viral productivity, we also evaluated cell number, viability, transfection efficiency, and fluorescence intensity in transfected cells at 72 hpt. The results indicate that the generated helper plasmids did not impact cell number, viability, or transfection efficiency (Fig. 5A, Supplementary Fig. S1). The transfected cells doubled their initial concentration, maintaining viability above 80% (Fig. 5A), with a transfection efficiency of 99% across all helper constructs and serotypes (Supplementary Fig. S1). Interestingly, in cells transfected with helper plasmids containing only the E4orf6 (helpers pC, pE, pF, pG, and pH), we observed a 1.5–3-fold increase in eGFP mean fluorescence intensity (MFI) across all tested serotypes (Fig. 5B). This increase was not observed in cells transfected with helper plasmids with full E4 (pB, pD and pI). Curiously, when eGFP MFI levels were compared with VG/TP ratios of the produced rAAV (Supplementary Fig. S2), a strong correlation was observed, indicating that higher eGFP MFIs correspond to lower full-empty ratios in serotypes 5, 8, and 9 (Supplementary Fig. S2A, B, and C).
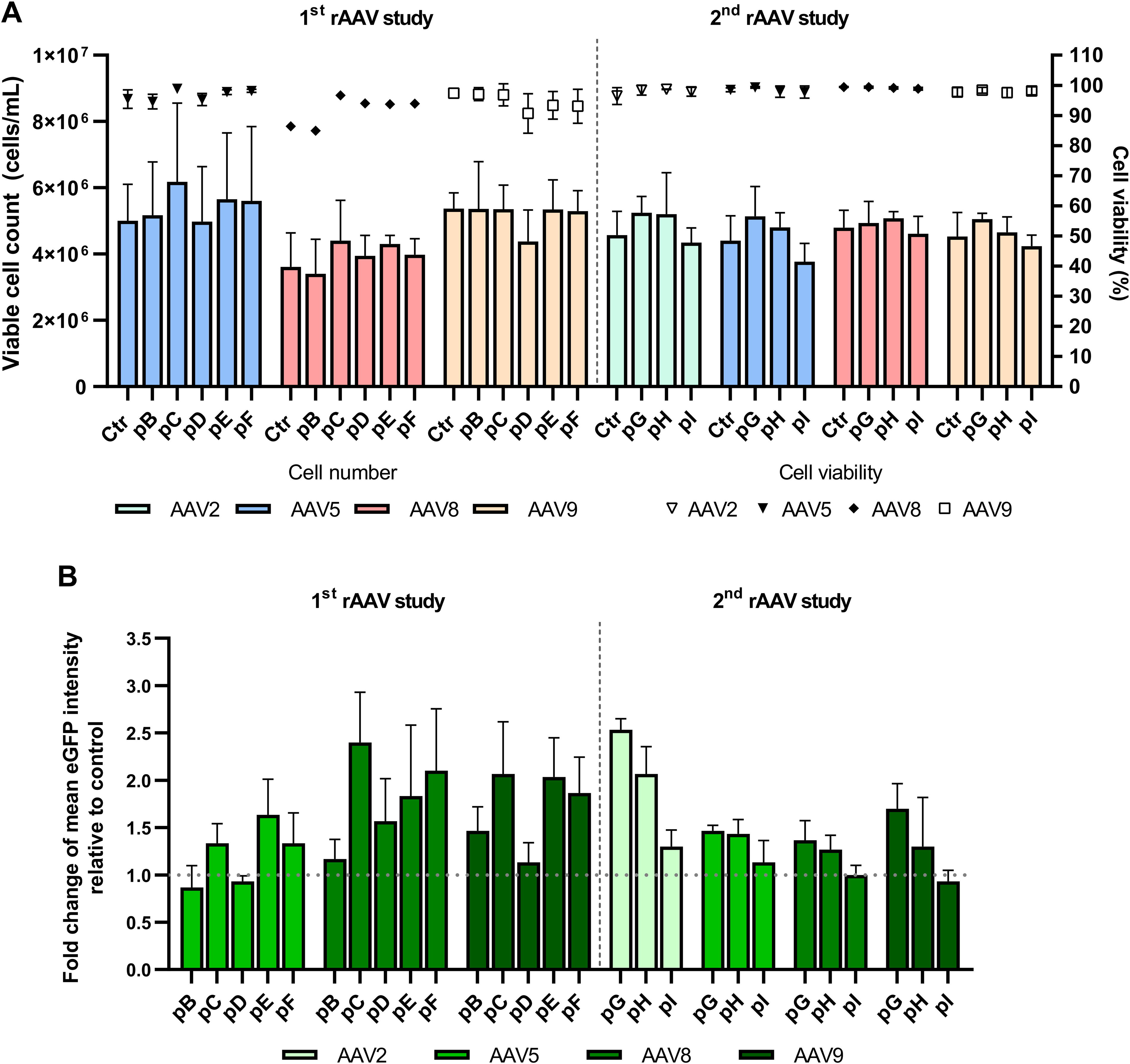
Post-transfection analysis of 293 cells.
DISCUSSION
The growing demand for safer and more efficient production methods for rAAV vectors has driven the evaluation of helper plasmids with minimal viral components. In this study, we generated eight novel helper plasmids containing the full VA RNA region and deletions in the E2 and E4 helper genes to assess: (i) total productivity, (ii) the amount of full particles (VG), (iii) viral release into the supernatant, and (iv) transducing ability. These findings offer key insights into the minimal genetic requirements necessary for the successful production of four rAAV serotypes.
Overall, our results suggest that deletions in the E2 and E4 genes have varying effects on total rAAV production and packaging efficiency, with some of these effects being serotype-dependent. Among the serotypes tested, rAAV5 was the least affected in terms of productivity and vector quality, as indicated by the VG/TP ratios (Fig. 2). In contrast, serotypes 2, 8, and 9 exhibited more similar responses to the deletions in the adenoviral helper genes. These differences may be related to the fact that rAAV8, rAAV9, and rAAV2 are phylogenetically closer to each other than rAAV5, which is the most distant. 44,45
The most pronounced effect was observed on the titer of packaged genomes (VG), while total production (TP) was only mildly reduced in comparison to the control (Fig. 2A, 2B). For rAAV5, no significant differences in TP titers were observed with any of the generated helper plasmids (Fig. 2A and B). This aligns with previous studies, which suggest that AAV5 is less reliant on helper viruses for productive infection, particularly for activating the p41 promoter that drives the expression of AAV5 capsid proteins. 46,47 In contrast, for rAAV8 and rAAV9, we observed a more significant decrease in TP titers, as the expression of capsid proteins in these serotypes is driven by the p40 promoter. Although the p40 promoter exhibits activity even in the absence of AAV Rep2 and AdV proteins, its activation is more dependent on adenoviral components compared to p41. 48 Interestingly, it appears that E2-DBP or E2-DBP/100K is unable to counteract the known negative effects of E4-derived proteins on rAAV8 and rAAV9 capsid production when the entire E4 gene is present. 26 This is evidenced by the fact that helper plasmids pB and pD resulted in the lowest TP titers (Fig. 2A and B). However, when only E4orf6 is present, the reduction in TP titer is either less pronounced or absent, suggesting that fewer E2 regions are required to mitigate the detrimental effects.
In this study, we demonstrated that the combination of the E2-DBP + E2-L4 22/33K regions, along with the full E4 gene (as in the helper plasmid pI), resulted in the highest viral production across all generated helper plasmids and tested AAV serotypes. With this helper plasmid, we achieved levels of productivity (TPs, VGs) and quality (VG/TP %) comparable to those of the control (Fig. 2). Previous studies have highlighted the importance of the L4-22/33K region in the late stages of AdV replication, specifically for efficient viral packaging, 14,22,49 –51 and in rAAV production using recombinant AdV helper virus. 23,42 In these reports, 23,42 the authors noted a 20-fold reduction in rAAV full particle production in the absence of the L4-22/33K region, a trend we also observed when this region was deleted from our helper plasmids, particularly for serotypes 8 and 9 (Fig. 2 and Supplementary Fig. S3A).
The next most effective helper plasmid in our study was pH, which contains the same E2 region as pI but with only the orf3 and orf6 regions of E4. E4orf3 is known to enhance viral replication by interacting with the E1A 14 gene and the L4P promoter (within the E2A gene), thereby facilitating the transcription of E2A L4-22/33K. 14,52 Given its role in replication, E4orf3 was incorporated into our set of minimal regions, as demonstrated in helper pH, which presents a promising alternative, especially in the development of stable cell lines. This is particularly important as pH lacks the E4orf4 gene, which encodes a protein known for its cytotoxic effects on cells. 53
Another key observation from our study was the impact of the E4 region on the quality of rAAV preparations and the release of viral particles from cells into the supernatant. Helper plasmids containing the full E4 gene (pB, pD, and pI) produced rAAV preparations with higher VG/TP ratios for serotypes 5, 8, and 9 (Fig. 2C), and also improved the release of rAAV5 and rAAV9 VGs into the supernatant (Fig. 3). This enhancement does not appear to be due to decreased cell growth or viability, as these factors were similar across all tested conditions (Fig. 5A). It is important to highlight that, for pB and pD, the higher VG/TP ratio results from a concurrent reduction in both TP and VG (Fig. 2A and B), reinforcing the role of the L4-22/33K region in both packaging efficiency and capsid formation (present in helper pI). Interestingly, since E4orf3 forms a nuclear scaffold that sequesters cellular proteins and facilitates SUMOylation—a process crucial for rAAV export 54 —we expected a more significant increase in VG release into the supernatant with helper plasmids pG and pH. However, this increase was not as pronounced as anticipated, suggesting that rAAV export may require additional adenoviral E4 components beyond E4orf3 for optimal efficiency.
Recently, two additional reports were published that also highlight the importance of L4-22/33K for rAAV production. 41,55 In the work by Adsero et al., several AdV E2 components were placed under the control of heterologous promoters and overexpressed to enhance rAAV9 production. 55 Similar to our findings, they suggest that full E2 100K is dispensable, and only the E2 L4 region, together with the E2A core promoter, is necessary for efficient viral production 55 (Supplementary Fig. S3B). We observed comparable results for other rAAV serotypes: (i) having E2-DBP or E2-DBP-100K (as in pC and pE) delivers similar results; and (ii) the presence of L4-22/33K can restore the VG levels, as observed with plasmids pG and pI (Fig. 2). In our study, the VG titers of rAAV9 appeared to be generally higher, likely because we retained either the full E4 (as in pI) or both E4orf3 and E4orf6 (as in pG) in the same helper construct along with the tested E2 components (Fig. 2). This suggests a synergistic effect between these E4 regions and the E2 regions (Fig. 2). More recently, Johari et al. also confirmed that E2 L4-22/33K is essential for optimal rAAV8 production. 56 They reported a slight improvement in VG titers when combining this E2 region with E4orf3 and E4orf6/7 or just E4orf6/7. 56 While we did not test E4orf6/7 in our study, we evaluated full E4, E4orf6 alone, and E4orf6 in combination with E4orf3 (Fig. 1). Our findings showed a recovery in VG titers when both E4orf3 and E4orf6 were included, but the most significant improvement was seen when the E2 L4-22/33K region was combined with the full E4 (Fig. 2, Supplementary Fig. S3A and C). This trend was consistent across all four tested serotypes.
Curiously, Doshi et al. 57 observed that eliminating E4 can enhance rAAV production. However, their study differs from ours in several experimental conditions: (i) they used adherent cells with 10% serum, whereas we used suspension cultures under serum-free conditions; (ii) our helper factors were expressed under native promoters, maintaining conditions similar to standard triple transient transfection, while Doshi et al. utilized strong constitutive promoters (EF1α and/or CMV); and (iii) we used a different helper plasmid as a control, with our initial rAAV9 productivity being up to 6-fold higher (VG/mL) than that reported by Doshi et al. with their DF6 control plasmid. Another recent study by Lieshout et al. 58 also found that removing E4 from rAAV production led to reduced productivity, particularly in full particles. However, in contrast to our findings, they observed that retaining only E4orf6 alongside full E2 was sufficient to maintain viral productivity. Since Lieshout et al. employed a double transient transfection approach, while we used a triple transient transfection strategy with a different molecular ratio, these methodological differences may explain the discrepancies in rAAV productivity. Taken together, these findings highlight the intricate interplay between helper and AAV components, suggesting potential compensatory mechanisms that influence rAAV yield and quality.
An additional finding of this study was the observed correlation between eGFP MFI and the quality of produced rAAVs (Supplementary Fig. S2). Specifically, plasmids that produced lower VG/TP ratios (such as those containing only E4orf6) were associated with higher eGFP MFI in transfected cells. This indicates that eGFP expression might serve as a predictor of viral vector production quality. The underlying reasons for this could include: (i) the accumulation of unpackaged rAAV DNA transgenes within the cell, which remain accessible to the cellular transcription machinery and can be freely transcribed, in contrast to packaged DNA that is protected within the viral capsid; and (ii) a reduction in the levels of Rep proteins, which could relieve their known inhibitory effects on heterologous promoters, 59 thereby enhancing the expression of eGFP driven by the CBA promoter. Further studies are required to clarify this observation, such as the evaluation of free unpackaged DNA, and the quantification of Rep protein expression, after transfection with each generated helper plasmid.
Together, these findings emphasize the critical role of the E2 region, along with E4orf3 and E4orf6, in the production of several rAAV serotypes through transient transfection. Given that 293 cells contain stably integrated AdV pIX protein, its potential involvement in rAAV production cannot be completely ruled out.
In summary, we were able to identify a minimal set of adenoviral elements and develop a smaller helper plasmid, able to match the rAAV production obtained with the currently used pHelper in transient transfection. This smaller plasmid not only simplifies the production system but also reduces the genetic load, potentially leading to safer and more stable production cell lines. The insights gained from this study facilitate the development of a helper-free stable cell line, further enhancing the safety and efficiency of rAAV vector production.
Footnotes
ACKNOWLEDGMENTS
S.F. and J.G. contributed equally to this work. The authors acknowledge the financial support received from Fundação para a Ciência e Tecnologia/Ministério da Ciência, Tecnologia e Ensino Superior (FCT/MCTES, Portugal) through national funds to iNOVA4Health UIDB/04462/2020 and UIDP/04462/2020) and the Associate Laboratory LS4FUTURE (LA/P/0087/2020) and MVF acknowledge the financial support received from PhD fellowship UI/BD/151256/2021 within the scope of the PhD Program in Bioengineering—Cell Therapies and Regenerative Medicine.
AUTHOR DISCLOSURE
The authors declare no conflict of interest.
FUNDING INFORMATION
No funding was received for this article.
SUPPLEMENTARY MATERIAL
Supplementary Figure S1
Supplementary Figure S2
Supplementary Figure S3
Supplementary Table S1
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
