Abstract
Adeno-associated viral (AAV) vectors are increasingly used for preclinical and clinical cardiac gene therapy approaches. However, gene transfer to cardiomyocytes poses a challenge due to differences between AAV serotypes in terms of expression efficiency in vitro and in vivo. For example, AAV9 vectors work well in rodent heart muscle cells in vivo but not in cultivated neonatal rat ventricular cardiomyocytes (NRVCMs), necessitating the use of AAV6 vectors for in vitro studies. Therefore, we aimed to develop an AAV that could efficiently express genes in NRVCMs, human engineered heart tissue (hEHT), and mammalian hearts. The production of AAV6 vectors results in lower yields compared with AAV9. Hence, we used random AAV9 peptide libraries and selected variants on NRVCMs at the vector genome and RNA levels in parallel. The enriched library variants were characterized using high-throughput analysis of barcoded variants, followed by individual validation of the most promising candidates. Interestingly, we found striking differences in NRVCM transduction and gene expression patterns of the AAV capsid variants depending on the selection strategy. AAV variants selected based on the vector genome level enabled the highest transduction but were outperformed by AAVs selected on the RNA level in terms of expression efficiency. In addition, we identified a new AAV9 capsid variant that not only allowed significantly higher gene expression in NRVCMs compared with AAV6 but also enabled similar gene expression in murine hearts as AAV9 wild-type vectors after being intravenously injected into mice. Moreover, the novel variant facilitated significantly higher gene expression in hEHT compared with AAV9. Therefore, this AAV variant could streamline preclinical gene therapy studies of myocardial diseases by eliminating the need for using different AAVs for NRVCMs, hEHT, and mice.
INTRODUCTION
Adeno-associated viral (AAV) vectors are well-established tools for gene delivery, and so far, seven AAV-based gene therapies were clinically approved by the Food and Drug Administration or European Medicines Agency to treat monogenic disorders. 1,2 However, the broad application of AAVs is still limited by known challenges such as immunity due to high levels of neutralizing antibodies, species-dependent differences in transduction and/or expression efficiency, and low organ specificity potentially resulting in dose-dependent toxicity when applied in vivo. 3
Cardiac gene transfer in small and large animal models is efficient with AAV serotype 9 (AAV9) vectors in particular for gene transfer into cardiomyocytes. However, limitations of AAV9 vectors are low transduction efficiencies in neonatal rat ventricular cardiomyocytes (NRVCMs) and human-induced pluripotent stem cell-derived cardiomyocytes. 4,5 Thus, preclinical in vitro studies frequently rely on the application of AAV6 vectors, 6,7 posing the risk that results may not translate well to in vivo studies with AAV9.
The issue of efficiency and specificity has been addressed by many groups in recent years using different methods to generate recombinant AAV capsids with novel features. These methods include rational design (such as domain swapping, generation of capsid chimeras, peptide or protein insertions, and others; further reviewed in Pupo et al.) 8 as well as directed evolution approaches. To accomplish the latter, large libraries of randomized capsids are created, for example, by using error-prone polymerase chain reactions (PCRs), 9 DNA family shuffling of various AAV wild-type capsid domains, 10,11 or by displaying random peptide sequences on the AAV capsid surface. 12,13 These libraries are then selected through multiple rounds based on their ability to target distinct cell types or tissues, or their ability to evade neutralizing antibodies.
Such directed evolution approaches have resulted in various distinguished AAV capsid variants with selected benefits. For instance, AAV2-ESGHGYF specifically expresses in the murine lung and was selected using a peptide library in vivo. 14 Moreover, the shuffled capsid AAV-DJ that was selected on human hepatocytes exhibits strong liver tropism in mice. 10 AAV-PHP.B is capable of penetrating the blood–brain barrier and transduces astrocytes and neurons in the murine central nervous system. 15 Further novel capsid variants have been reported and reviewed elsewhere. 16,17
In the early stages of AAV capsid library screening, selections were performed using helper virus co-infection. Later, the focus shifted to selecting AAV capsid variants based on their ability to transduce cells using PCR amplification of vector genomes. 17 However, the efficacy of the AAV capsid depends not only on high cellular uptake through targeting specific receptors but also on efficient intracellular trafficking to the nucleus, capsid uncoating, and transgene transcription and translation. 16,18,19 The development of RNA-based selection systems such as TRACER 20 and DELIVER 21 has allowed for a greater emphasis on the expression of transgenes in the selection of AAV capsids.
In this study, we aimed to systematically compare the directed evolution on the transduction and gene expression levels by performing separated DNA and RNA selections. We generated an AAV9 random peptide library and selected it on NRVCMs as the most frequently used preclinical cardiac cell culture model. AAV9 has been shown to mediate the strongest transgene expression in the murine heart after intravenous delivery compared with the other naturally occurring serotypes. 22 –24 Furthermore, AAV9 vector productions usually yield higher titers than the prevalent AAV6. However, its efficiency in vitro is low. 5 Therefore, we chose AAV9 as a library scaffold to obtain capsid variants with improved in vitro gene transfer capability. One of the capsids we identified from the AAV library selections, AAV9-AERYTKY, exhibits higher transgene expression in cardiomyocytes in vitro and has the potential to be utilized as a single AAV in both cultivated cardiomyocytes and mouse models.
MATERIALS AND METHODS
Primary NRVCM isolation
NRVCMs were isolated from 1- to 3-day-old Wistar rats (Charles River) as previously described. 25 The cells were kept in Dulbecco's modified Eagle medium (DMEM) with 2 mM L-glutamine (Gibco), 100 U/mL penicillin, and 100 µg/mL streptomycin (Pen Strep, Gibco) +10% (v/v) fetal bovine serum (FBS, Capricorn Scientific) at 37°C and 5% CO2. After seeding for 24 h, the cells were transduced with AAVs.
Animal study
Animal experiments were carried out under the guidelines from directive 2010/63/EU of the European Parliament on the protection of animals used for scientific purposes with the approval of the regional council (permission no. V242-56596/2022). Animals were housed under standard conditions with a 12-h-light/12-h-night cycle. Water and food were offered ad libitum. Barcoded AAV vector libraries were injected systemically via tail vein injection into 8-week-old male C57BL/6N mice (Charles River). Each mouse was injected with 5 × 1011 viral genomes in a total volume of 100 µL. Mice were sacrificed 2 weeks after injection by cervical dislocation and harvested organs were snap-frozen in liquid nitrogen.
Single AAV vector injections for comparison of AAV9-AERYTKY and AAV9 wild type were performed at the Amsterdam University Medical Centre and conducted according to the local legislation (permission no. AVD11800202114455). AAV vectors were injected systemically via retro-orbital injections into 8-week-old female FVB mice (Janvier). Each mouse was injected with 2.5 × 1011 viral genomes in a total volume of 100 µL. Mice were sacrificed after 3 weeks by inhaling carbon dioxide followed by cervical dislocation. The apex of the heart and the other organs were snap-frozen. The base of the heart and a part of the liver were used for histological analysis.
AAV9 random peptide insertion library and selection process
The AAV9 random heptapeptide insertion library was generated and used for in vitro biopanning as described earlier 26 with the exception that the plasmid library was directly used for production of the AAV library.
For the selections, 0.5 to 1 × 106 cells were seeded into six-well plates (Greiner Bio-One). After 24 h, the cells were washed with 1× dulbecco's phosphate-buffered saline (DPBS) and then transduced with the AAV9 peptide library at an multiplicity of infection (MOI) of 1 × 105 in serum-free DMEM. Three hours post-transduction, the cells were washed three times with 1× DPBS, and fresh DMEM +10% FBS was added. After 72 h, cells were harvested, DNA and RNA were isolated, and used as templates for PCR amplification of the region containing the random heptapeptide with the following primers: forward: 5′-AGTCATGATAACCAACGAAG-3′, reverse: 5′-GAAAGTTGCCGTCCGTGT-3′. The resulting PCR product was then purified (NucleoSpin Plasmid Kit, Macherey-Nagel), digested with BglI, and cloned into the AAV9 backbone plasmid pKV-AAV9Lib/BB 26 using the SfiI restriction sites. The resulting plasmid libraries were then used to generate the preselected AAV9 libraries. In the first round of selection, only DNA was processed to produce the primary library, but after the second round of selection, DNA and RNA were employed separately to generate secondary libraries. After the third rounds of the selection process, PCR products containing the insertion region of the gDNA and cDNA were analyzed by Illumina-based amplicon sequencing (GENEWIZ, Azenta Life Sciences).
Selected peptide motifs were cloned into plasmid p5E18-VD2/9-SfiI1759 to produce AAV9 capsid variants using the helper plasmid pDGΔVP 27 in a three-plasmid system. 26
AAV production and titration
AAV production was carried out as described elsewhere. 28 In brief, human embryonic kidney (HEK)293T cells (ATCC) were seeded into a 5-layer CellStack Culture Chamber (Corning) or an appropriate number of 15 cm petri dishes 24 h prior to transfection. The transfection of cells was carried out using polyethylenimine (Polysciences) and a two- or three-plasmid system for wild-type AAVs and AAV capsid variants, respectively. Following a 72 h incubation period, the cells were harvested and lysed to extract the AAVs, which were subsequently purified through a discontinuous iodixanol (OptiprepTM, Progen) gradient ultracentrifugation. The AAV containing fraction was extracted and the iodixanol exchanged to 1× DPBS using Zeba Spin Desalting columns (7K MWCO, Thermo Scientific). If necessary, AAVs were concentrated in Vivaspin centrifugal concentrators (10 kDa MWCO, Sartorius).
For viral genome quantification, alkaline lysis was performed, followed by quantitative PCR (CFX96 Real-Time PCR System, Bio-Rad) utilizing the SensiFAST SYBR No-ROX Kit (Meridian Bioscience) as described previously. 28 The AAV9 peptide libraries were titrated via primers for REP2 (forward: 5′-AAGTCCTCGGCCCAGATAGAC-3′, reverse: 5′-CAATCACGGCGCACATGT-3′) and a respective plasmid standard. In addition, barcoded AAV variants were quantified targeting the cytomegalovirus (CMV) promoter (forward: 5′-CATTATGCCCAGTACATGACC-3′, reverse: 5′-GAAATCCCCGTGAGTCAAACC-3′) and a viral standard.
Barcoded next-generation sequencing analysis
To validate multiple AAV capsid variants simultaneously, we followed the method previously described by Weinmann et al. 29 Each capsid was packed with a CMV promoter-driven enhanced yellow fluorescent protein (eYFP) reporter genome containing a unique barcode in the 3′-untranslated region. The individual AAVs were mixed equimolar and concentrated if necessary. For in vitro validation, cells were transduced with an MOI of 1 × 105. For in vivo validation, mice were injected intravenously with 5 × 1011 vg. The barcoded library for in vivo validation contained a total of 52 AAV variants. After DNA and RNA isolation, the barcode-containing region of the gDNA and cDNA was amplified via PCR using the following primers: forward: 5′-CGACAACCACTACCTGAGCTAC-3′ and reverse: 5′-GGCTGGCAACTAGAAGGCACA-3′. The amplified DNA was then purified and analyzed through Illumina-based amplicon sequencing. Reads were normalized to the read populations of the mixed input library before transduction or injection. We applied a Python script previously described by Weinmann et al. to identify barcode reads and normalize the data. 29
YFP quantification by flow cytometry
To quantify in vitro transduction, 2–4 × 105 cells per well were seeded in a 12-well plate (Greiner Bio-One). After 24 h, the cells were washed with 1× DPBS, and serum-free DMEM was added for the transduction with distinct AAVs with an MOI of 1 × 104. After 24 h, cells were washed again with 1× DPBS, and fresh DMEM +10% FBS was added. Three days post-transduction, the medium was replaced with 1× DPBS, and the mean fluorescent intensity was determined using a BioTek Citation 5 Imaging Reader (excitation 488 nm, emission 530 nm) from Agilent. The cells were detached with trypsin-ethylenediaminetetraacetic acid solution (Gibco), washed with 1× DPBS, and a single-cell suspension was generated in Falcon Round Bottom Polystyrene Tubes (Corning). Positive eYFP-expressing cells were detected with the LSRFortessaTM (BD Biosciences) using the FACSDivaTM software. Vital cells were separated using FSC-A against SSC-A, and doublets were extracted using FSC-H against FSC-A. The eYFP-positive gate for FITC-A was set using control cells treated with PBS.
Human engineered heart tissue transduction
The generation of human engineered heart tissues (hEHTs) was performed as described previously. 30,31 Each hEHT contained 1 × 106 human-induced pluripotent stem cell-derived cardiomyocyte (hiPSC-CM) cells (in-house cell line UKEi001-A) and was cultured in 24-well plates containing DMEM, 1% penicillin/streptomycin, 10% horse serum (Gibco), 10 µg/mL insulin (Sigma-Aldrich), 33 µg/mL aprotinin from bovine lung (Serva Electrophoresis), and 2 mM tranexamic acid (Sigma-Aldrich) for 14 days prior transduction. The AAVs were added to the culture medium with an MOI of 2.5 × 104 (AAV9 wild type and capsid variants) or 2.5 × 103 (AAV6 wild type), as established in previous experiments. hEHTs were incubated for 24 h before transferring them to fresh medium. Medium was changed in 2–3-day intervals. After 7 days, images were captured (90% intensity, 30 ms, and 4× magnification) with an EVOS FL microscope (Peqlab), and fluorescence intensities were quantified using ImageJ. Furthermore, contractile force was measured directly before the transduction and 7 days after (EHT Technologies GmbH) as previously described (2–4 h after medium change). 30
IMMUNOSTAINING
For immunocytochemistry staining, NRVCMs were seeded on collagen-coated coverslips in 12-well plates. Cells were fixed using 4% paraformaldehyde (PFA, Sigma-Aldrich) for 10 min, washed with 1× DPBS, and treated with 0.1% (v/v) Triton X-100 (Serva Electrophoresis), 2.5% (w/v) BSA (Carl Roth) for at least 1 h at room temperature. Cells were incubated with anti-α-actinin antibody (Dilution 1:400, monoclonal, mouse; Sigma-Aldrich #A7811) and anti-green fluorescent protein (GFP) AF488 conjugate (Dilution 1:400, polyclonal, rabbit; Invitrogen, Thermo Scientific #A21311) overnight at 4°C. Then, cells were washed with 1× DPBS and incubated with antimouse AF647 (dilution 1:400, polyclonal, chicken; Invitrogen, Thermo Scientific #A21463) and 4′,6-diamidino-2-phenylindole (DAPI) (Dilution 1:1000, Sigma-Aldrich) for 1–2 h at room temperature. The coverslips were mounted on microscope slides using FluorPreserveTM Reagent (Calbiochem) and examined with a Keyence BZ-X810 fluorescence microscope (Keyence) at 20× magnification.
Histological analysis was performed by fixing hearts and livers in 4% PFA overnight and embedding in paraplast (Leica Biosystems). Sections of 3–4 µm thickness were prepared using the HistoCore Multicut (Leica Biosystems), then blocked with 0.1% (v/v) Triton X-100, 2.5% (w/v) BSA for 1 hour, and stained with anti-GFP AF555 conjugate (Dilution 1:400, polyclonal, rabbit; Invitrogen, Thermo Scientific #A31851) overnight at 4°C followed by DAPI for 1 h at room temperature. Images were taken using a laser scanning microscope 800 (Carl Zeiss) at 20× magnification.
DNA/RNA isolation and cDNA synthesis
Cultured cells were homogenized by transferring them onto QIAshredder (Qiagen) columns and centrifugation at 17,000 r.c.f. for 2 min. Tissue was disrupted with ceramic beads in a Precellys24 homogenizer (Peqlab) in a prior step and, subsequently, homogenized as described above. Isolation of total DNA and RNA both from cultured cells and tissues was performed using the AllPrep DNA/RNA/miRNA Universal Kit (Qiagen) according to the manufacturer’s instructions.
Reverse transcription of RNA into cDNA was carried out using LunaScript RT SuperMix Kit (New England Biolabs) adding 1 µg of RNA, if applicable.
Viral genome quantification and mRNA level determination
Ratios of viral genomes (vg) per diploid genomes (dg) were determined using multiplexed real-time quantitative reverse transcription PCR of isolated genomic DNA with iQ Multiplex Powermix (Bio-Rad). eYFP was quantified to determine the vg copy number (forward: 5′-GAGCGCACCATCTTCTTCAAG-3′ and reverse: 5′-TGTCGCCCTCGAACTTCAC-3′, probe: 5′-FAM-ACGACGGCAACTACAAGACC-3′; adapted from Weinmann et al.), 29 which was then normalized to the RPP30 copy number (forward: 5′-AGCCTGGGCTACTTGGT-3′ and reverse: 5′-CTGATGGCCGTGGTTTCTT-3′, probe: 5′-HEX-ACTTGTTTCTGTTTCTTGAGACAGGATCTC-3′; kindly provided by the Grimm lab in personal communication with K. Rapti) and divided by two to obtain the number of dg. For each well, 100–200 ng of genomic DNA was mixed with 62.5 nM per primer and 125 nM per probe in a 20 µL volume. A standard curve was generated by serial dilution of a plasmid comprising both amplicons.
mRNA levels were quantified by determination of eYFP levels (forward: 5′-GCATCAAGGTGAACTTCAAGATCC-3′ and reverse: 5′-ATGTGATCGCGCTTCTCGTTG-3′) and normalizing them to the housekeeping gene RPL32 (forward: 5′-CTGCTGATGTGCAACAAATCT-3′ and reverse: 5′-GCTGTGCTGCTCTTTCTACAAT-3′) using SensiFAST SYBR No-ROX Kit. In 20 µL volumes, 10 ng of cDNA were mixed with 200 nM per primer.
Statistical analysis
All results are presented as mean values ± standard deviation (SD) unless otherwise specified. Statistical analyses were conducted using a one-way analysis of variance (ANOVA) followed by Dunnett’s multiple comparison test. For the comparison of two groups, an unpaired two-tailed t-test was applied. p-Values <0.05 were considered statistically significant. Statistical analysis was carried out using GraphPad Prism 10.
RESULTS
Separated AAV9 random peptide library selections on the DNA and RNA levels result in enrichments of distinct AAV capsid variants
To obtain capsid variants with improved properties for cardiac gene transfer, we utilized a directed evolution approach with an AAV9 random peptide library on NRVCMs. Previous studies have reported the capability of AAV9 to display peptide variants and enrich distinct motifs. 15,21,26,32 However, most AAV library selections were based on amplification of the randomized region on the level of vector genomes. As approaches typically aim to improve gene expression, selections on the transduction level may not fully exploit the potential. 33 Thus, we aimed to characterize the effect of the selection process on the resulting AAV capsid variants by performing selections on either the DNA or RNA level. This could provide valuable insights into improving gene expression and transduction levels.
First, NRVCMs were transduced with the initial AAV9 peptide library. Then, the DNA was isolated and used to generate the first preselected library. To avoid the accumulation of falsely genotype–phenotype coupled or mosaic capsids with high gene expression in the high-diverse initial library, 34 the RNA level selection was omitted here. Previous studies demonstrated that about 95% of the capsids were eliminated after the first round of selection. 20
After the second round of transduction, both DNA and RNA were isolated from the cells and separately processed, resulting in DNA- or RNA-selected secondary libraries. These secondary libraries were further used in third selection rounds (Fig. 1A). Subsequently, gDNA and cDNA were analyzed via next generation sequencing (NGS), and enrichments of peptide motifs were clearly visible on all levels. However, the intensity of enrichment varied. Compared with the DNA-based selections, the RNA-based selections seemed to lead to single peptides with a higher level of enrichment, such as AERYTKY with 25.71% and PDRFGRP with 25.35% (Fig. 1B, 1C). Also, the cDNA revealed higher enriched peptides compared with the corresponding gDNA in both selection strategies (Supplementary Table S1). This was expected, as single peptides leading to strong transgene expression might directly occupy a large proportion of the total reads, as demonstrated earlier. 20
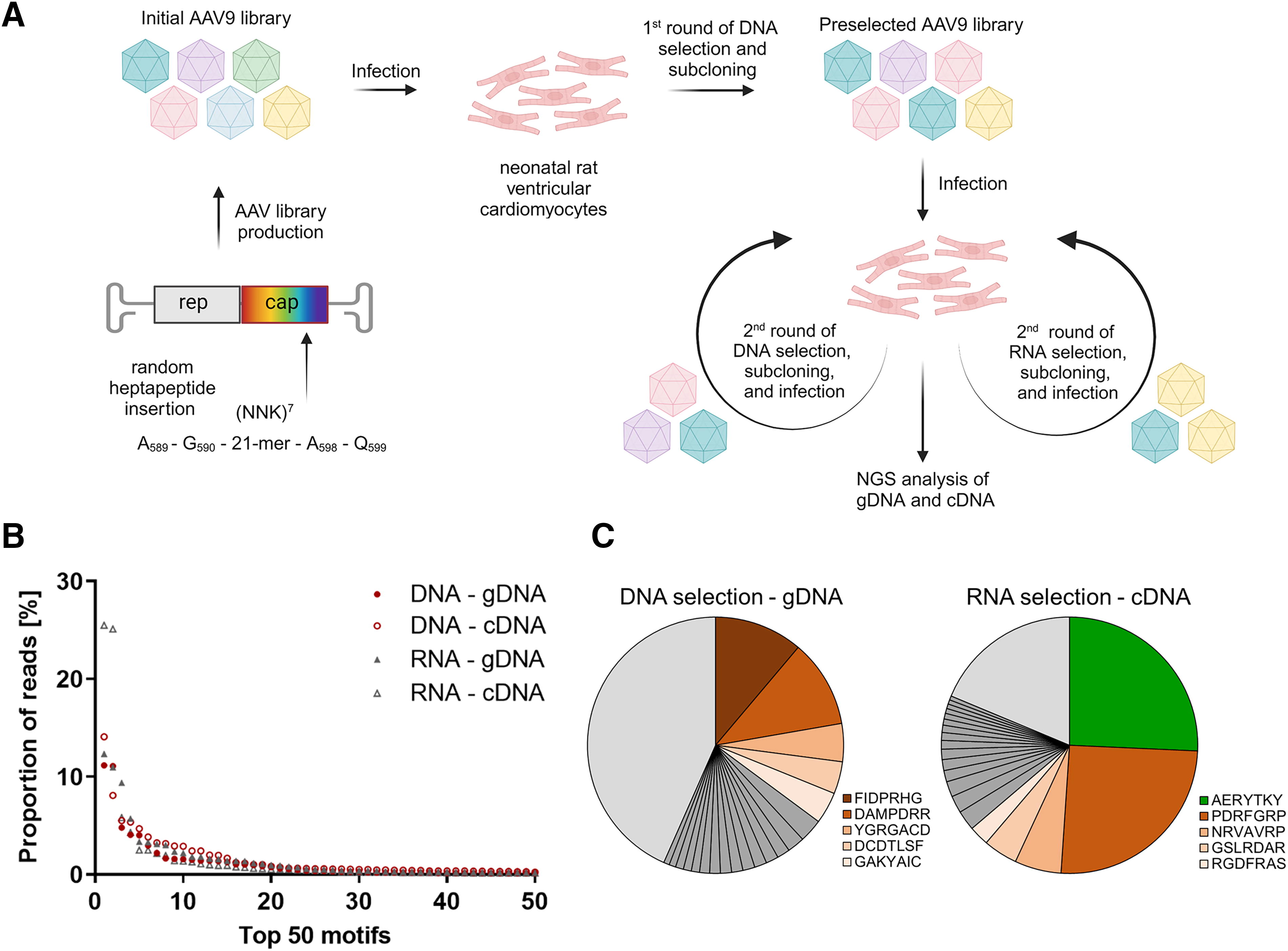
Selection of AAV9 peptide libraries on the DNA and RNA levels results in enrichment of distinct variants.
RNA-selected capsid variants display improved expression efficiencies
The top 10 enriched peptide motifs of both the DNA and the RNA results of both selection strategies were produced individually as AAV9 capsid variants (a comprehensive list of all variants can be found in Supplementary Table S1 and Supplementary Fig. S1). To facilitate individual and parallel validation, each AAV was equipped with an eYFP reporter gene driven by the constitutively-expressing CMV promoter and a unique 15 nucleotide barcode in the 3′-untranslated region (Fig. 2A). This linking of genomic information to the respective AAV capsid variant allowed for parallel quantification of the transduction and expression efficiencies, as previously published. 29,35
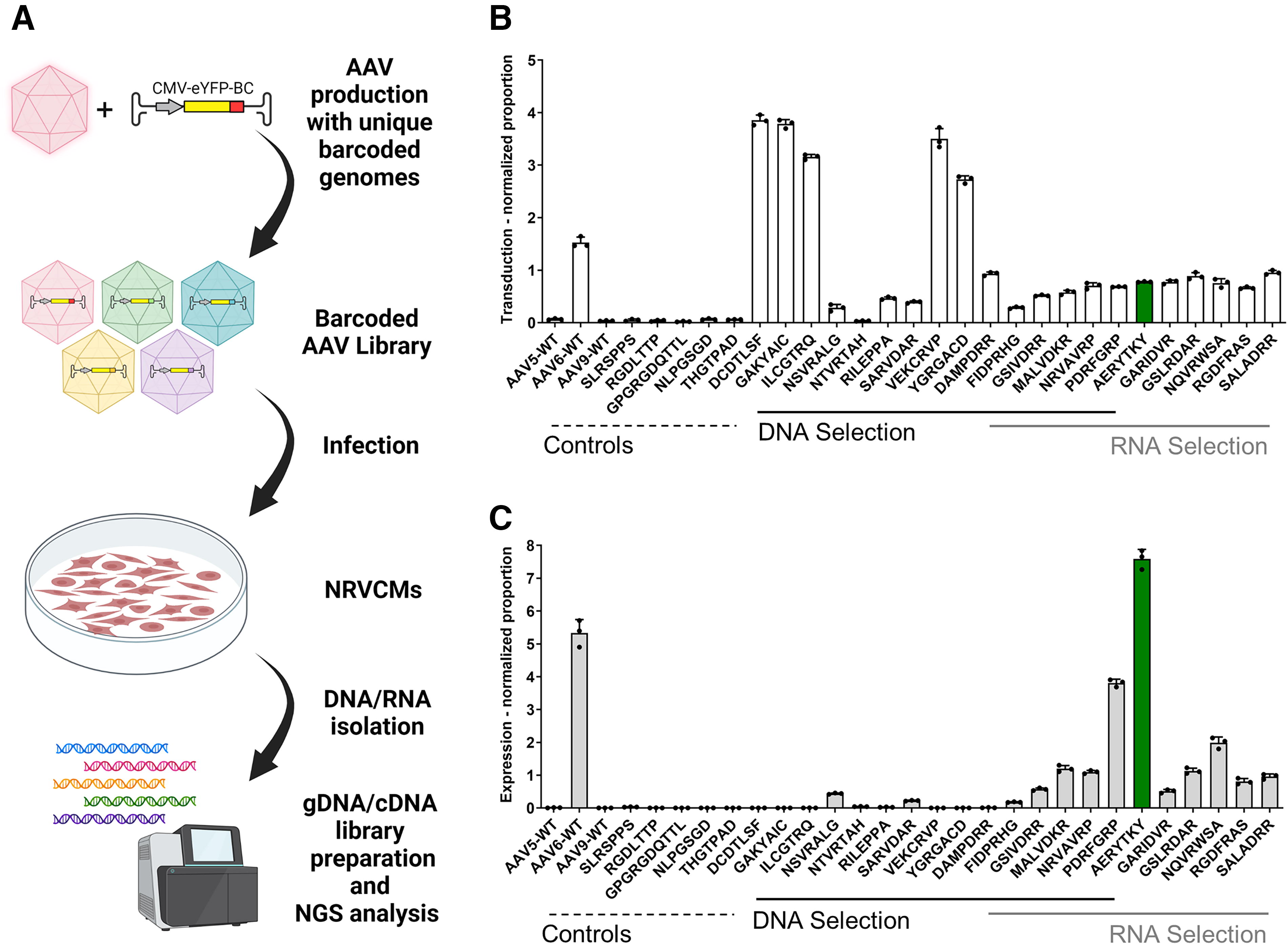
Parallel characterization of newly identified capsid variants at the levels of transduction and gene expression using NGS of barcoded libraries.
All variants were capable of being produced in a similar efficiency as AAV9 wild type, with yields ranging from 2.4 × 1011 vg to 2.4 × 1012 vg per production (Supplementary Fig. S2). The AAV capsid variants were equimolarly pooled to obtain a barcoded input library. In addition, certain peptide motifs known to target cardiac (NLPGSGD and THGTPAD) 36 or muscle tissue (RGDLTTP and GPGRGDQTTL) 21 as well as our previously identified endothelial targeting variant (SLRSPPS) 26 were cloned into the AAV9 capsid backbone, produced, and added to the library as benchmarks. Differences in abundance were monitored via NGS and considered for data normalization (Supplementary Fig. S3).
As expected, AAV6 was the only wild type showing reasonable transduction and expression in NRVCMs (Fig. 2B and 2C). Neither of the benchmark motifs seemed to efficiently transduce or express in NRVCMs, likely due to their inefficiency to target NRVCMs or their incapability to exhibit previously demonstrated effects when incorporated in our AAV9 backbone.
The AAV9 capsid variants derived from the independent selections clearly showed the anticipated differences in transduction and expression efficiency (Fig. 2B and 2C). The peptide motifs from the DNA selection demonstrated significantly higher transduction efficiencies but showed only very low to no transgene expression. Conversely, the RNA selection-based capsid variants demonstrated increased expression efficiencies compared to AAV9, with the peptide motifs PDRFGRP and AERYTKY exhibiting similar or even higher expression than AAV6, respectively. These results highlight the significant and selective impact of using either genomic DNA or RNA for selecting AAV peptide insertion libraries.
AAV9-AERYTKY leads to significantly increased reporter gene expression in NRVCMs
To verify the results from the barcode sequencing approach, individual AAVs harboring an eYFP reporter were used to monitor transgene expression through flow cytometry and fluorescence detection (Fig. 3). NRVCMs were transduced with individual AAV9 capsid variants. Capsids that have showed no expression in the parallel validation, such as AAV9 wild type and GAKYAIC, barely led to eYFP-positive cells (p < 0.0001). In contrast, all variants with high expression efficiency in the barcode sequencing approach resulted in significantly improved NRVCM targeting, with 65% to 83% eYFP-positive cells compared with AAV9 wild type, with only 1% eYFP-positive cells. Furthermore, taking into account the mean fluorescence intensity of the cells, AERYTKY and PDRFGRP noticeably outperformed the current benchmark AAV6 by 7.0- and 3.4-fold (p < 0.0001), respectively (Fig. 3A).
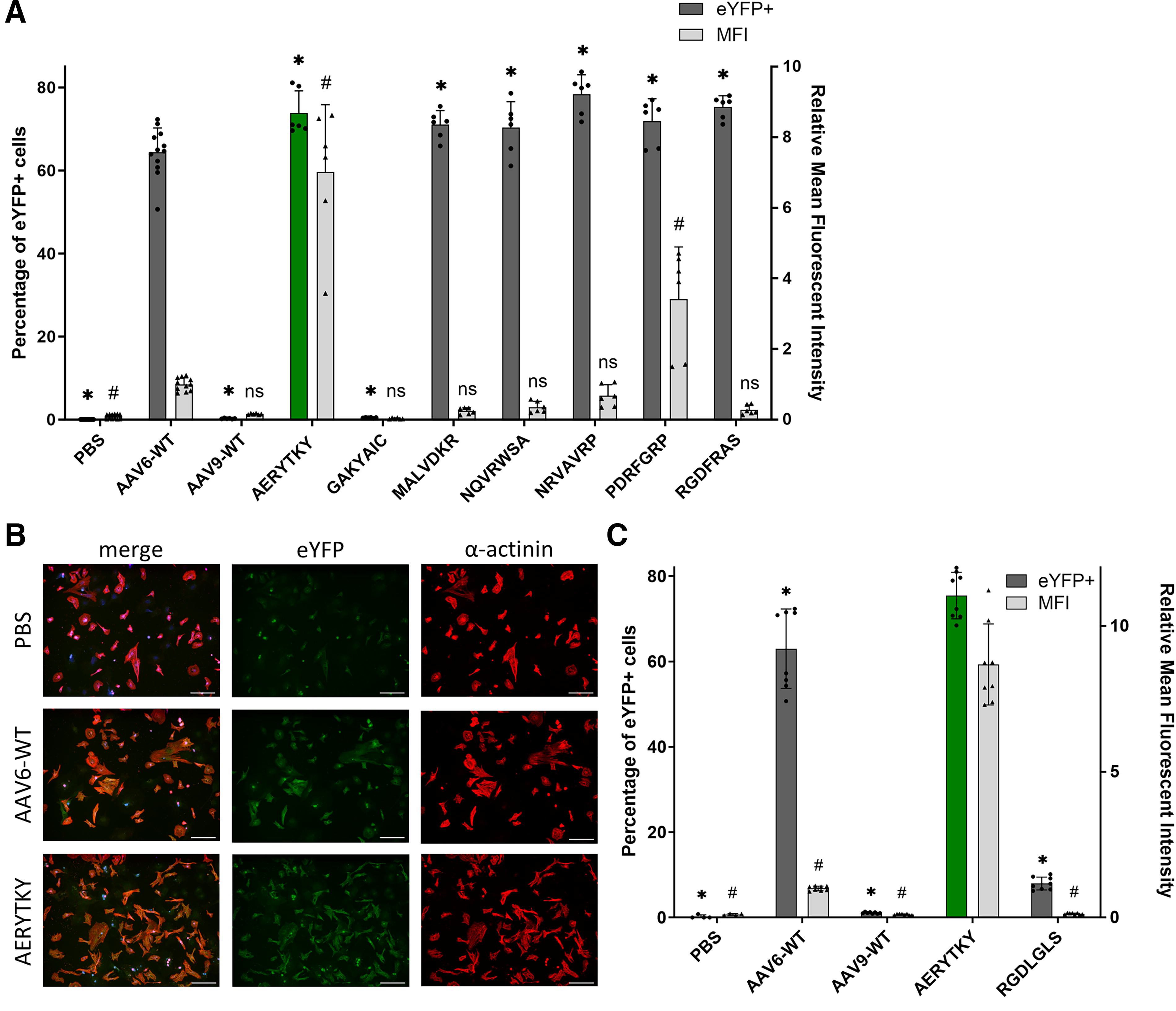
Characterization of individual AAV capsid variants in vitro.
To further demonstrate the significantly improved transgene expression of AERYTKY in NRVCMs, AAV vectors from two independent productions were validated again side-by-side by flow cytometry and visualized by immunocytochemistry (Fig. 3B and 3C, Supplementary Fig. S4). The capsid variant RGDLGLS, which is a benchmark capable of targeting cardiac tissue in vivo, was examined for its NRVCM transduction efficiency. 29 Interestingly, RGDLGLS barely enhanced the transduction to approximately 8% eYFP-positive cells (p < 0.0001), further emphasizing the potential of the capsid variant AERYTKY as new AAV gene therapy tool for in vitro cardiac research.
AAV9-AERYTKY enables cardiac gene transfer in vivo comparable with AAV9 wild type
The newly validated AAV9 capsid variants were then tested for their capability of in vivo cardiac gene transfer in the murine model employing the CMV-driven eYFP reporter. All selected capsids and some additional benchmarks known for targeting specific tissues in vivo were combined into a single barcoded library for simultaneous screening (Supplementary Fig. S5). The library was injected intravenously, and 2 weeks later, tissues were collected and gDNA and cDNA analyzed via NGS. To compare the results of multiple tissues, total viral genomes per diploid genomes were determined (Supplementary Fig. S5) and used for normalization. The normalized expression efficiencies were then utilized to generate a heat map (Fig. 4A). Notably, the transcriptional profile of the benchmarks is consistent with previous reports and demonstrates the high suitability of the simultaneous characterization. For instance, in our data, AAV2-ESGHGYF 14 and AAV-DJ 10 showed almost exclusive expression in lung and liver, respectively (Fig. 4B, Supplementary Fig. S5). The overall highest expression in the liver was enabled by AAV8 wild type, while the muscle-targeting motifs RGDLGLS and RGDLTTP exhibit highest expression in both skeletal muscle and heart (Fig. 4B).
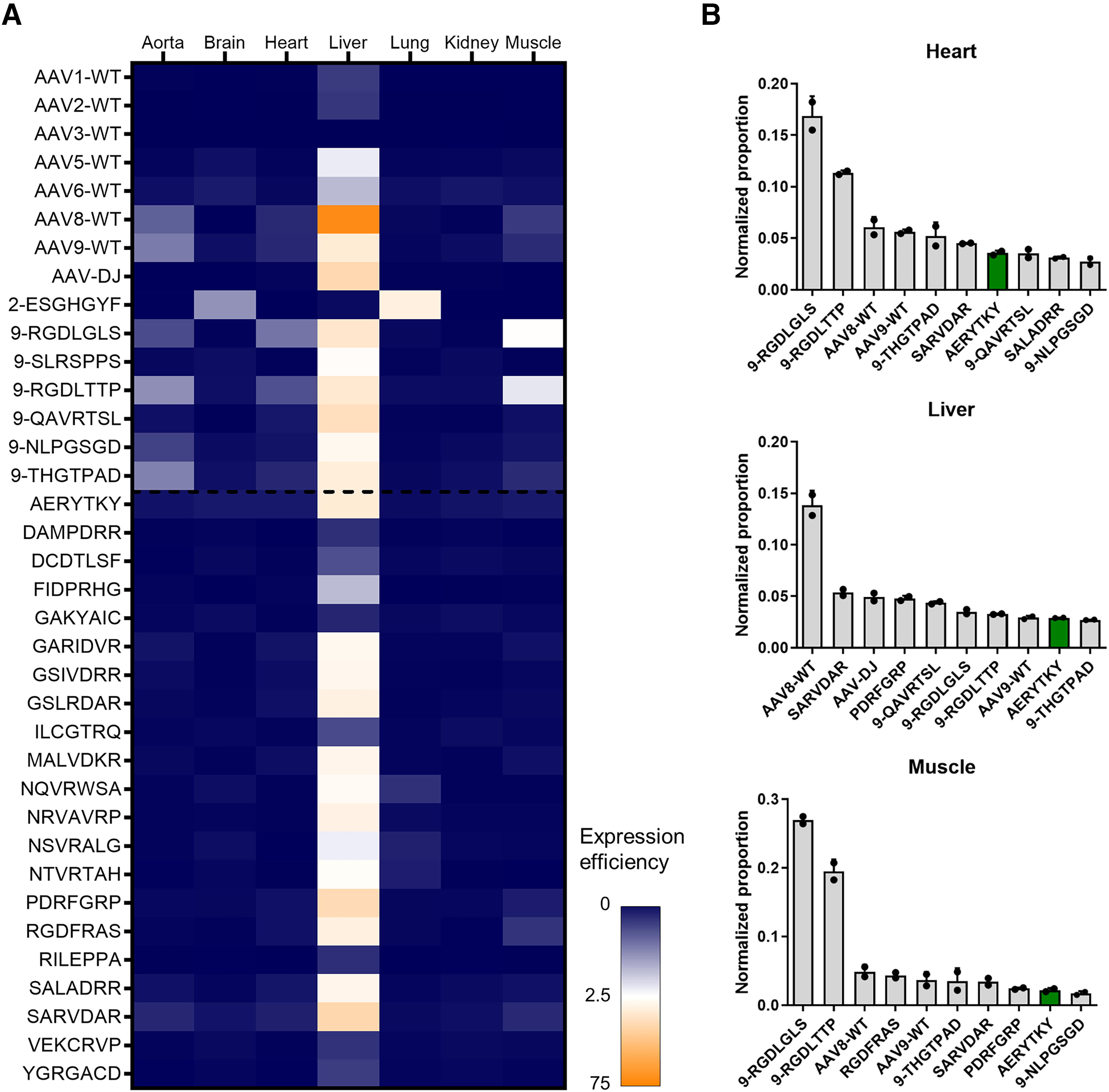
Parallel evaluation of the expression efficiencies of newly selected capsids in vivo. All selected and produced capsids were combined into a barcoded AAV library. Which also included various AAV wild types and known benchmark motifs from literature. This barcoded library was injected systemically in the tail vein of male C57BL/6N mice (5 × 1011 viral genomes per mouse), and tissues were harvested after 2 weeks. DNA and RNA were isolated and then analyzed via NGS to determine the number of reads of the barcoded region, which was normalized to the abundance in the input library and to the total viral genomes per diploid genomes (Supplementary Fig. S5).
Despite testing various AAV9 capsid variants in the mouse heart, none of them revealed an improvement of transgene expression in the heart in vivo when compared with AAV9 wild type or the respective benchmarks. This is not surprising because the selections were performed in vitro. Nevertheless, some variants showed detargeting from the liver either on the gene expression (Fig. 4) or transduction level (Supplementary Fig. S6).
We observed that the cardiac benchmark RGDLGLS was not able to efficiently transduce NRVCMs (Fig. 3C). Accordingly, we propose AERYTKY as an alternative for in vitro and in vivo studies in cardiomyocytes using a single vector. Our barcoded parallel comparison with AAV9 wild type in mice revealed that AERYTKY did not exhibit higher expression in the liver, unlike SARVDAR and PDRFGRP. Hence, we studied AERYTKY individually after intravenous injection into mice and compared it directly with AAV9 wild type for further in vivo validation (Fig. 5). Quantification of viral genomes per diploid genomes indicated a similar biodistribution of both vectors in a set of organs (Fig. 5B). In addition, expression levels quantified on the mRNA level were comparable across all tissues, with a notable difference in the brain (p = 0.012, Fig. 5C). These findings were also corroborated by representative sections of hearts and livers (Fig. 5A) which confirm similar cell-type tropism of AERYTKY.
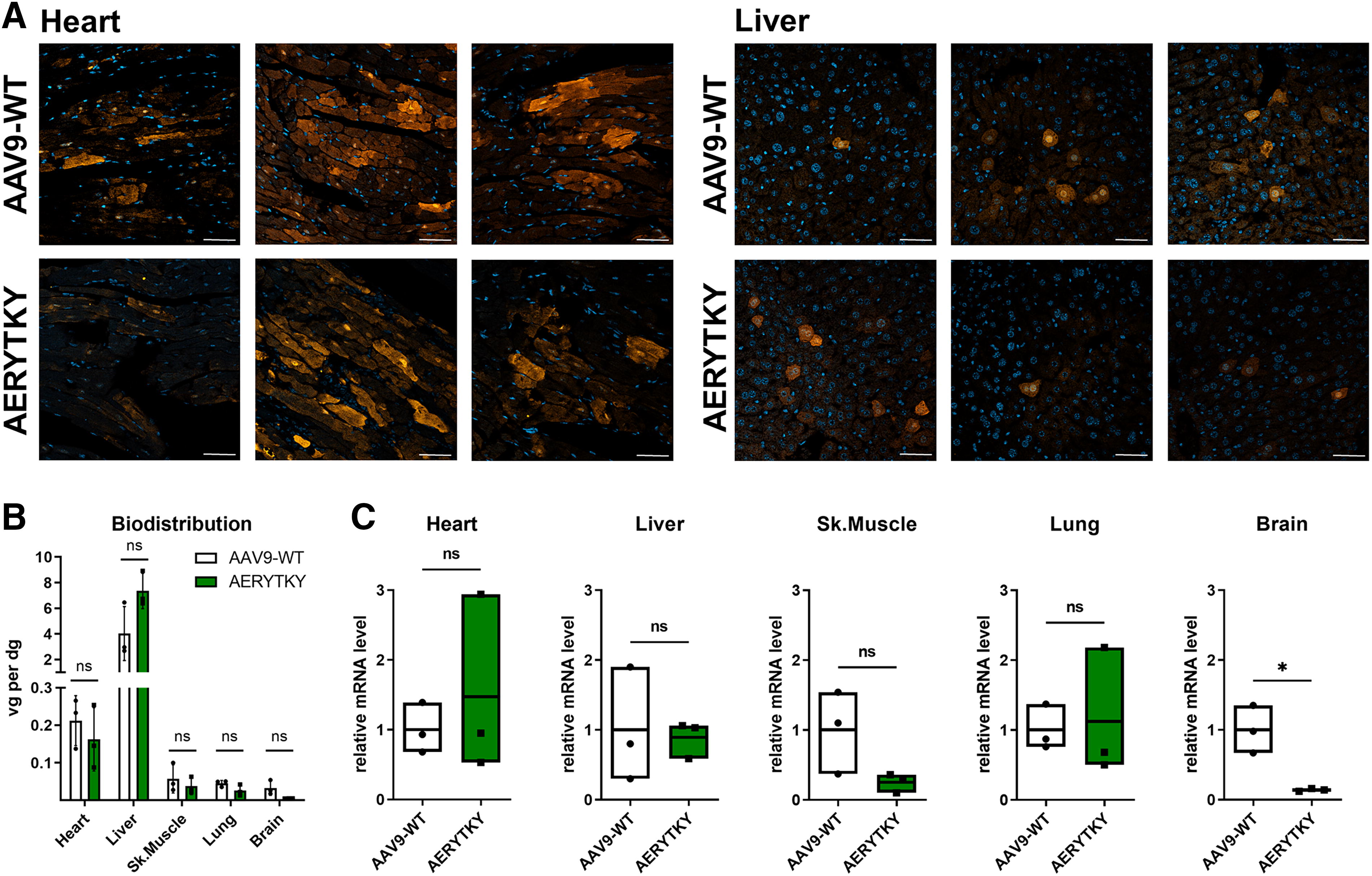
Comparison of AAV9-AERYTKY with AAV9 wild type in vivo. Expression efficiencies of AAV9-AERYTKY and AAV9 wild type were compared individually in mice in vivo. AAV vectors were systemically applied into female FVB mice (2.5 × 1011 viral genomes per mouse) using retro-orbital injections and tissues were harvested after 3 weeks.
AAV9 capsid variants increase transduction of hiPSC-derived engineered heart tissues
Finally, we investigated the most promising AAV9 capsid variants for their ability to transfer genes into human iPSCs that were differentiated into cardiac organoid-like tissue (hEHT). After transducing hEHTs with the variants, we took images after 7 days to examine eYFP fluorescence (Fig. 6A, 6B) and monitored the contractile force before and after transduction to evaluate the viability of the hEHTs (Fig. 6C). Considering previous experiments, AAV6 wild type was examined simultaneously at the established 10-fold lower dose as benchmark control to allow comparison with AAV9 capsid variants in terms of fluorescence intensity. Consistent with our in vitro findings, most of the examined peptide motif insertions significantly improved the transduction efficiency. In particular, AERYTKY, NRVAVRP, and RGDFRAS exhibited a 4.9-, 7.8-, and 11.2-fold increase in eYFP intensity compared with AAV9 wild type (p < 0.0001), respectively. Our results clearly demonstrate that these novel AAV capsid variants could be potential candidates to support preclinical cardiovascular research involving hiPSC-CMs and differentiated cardiac organoids.
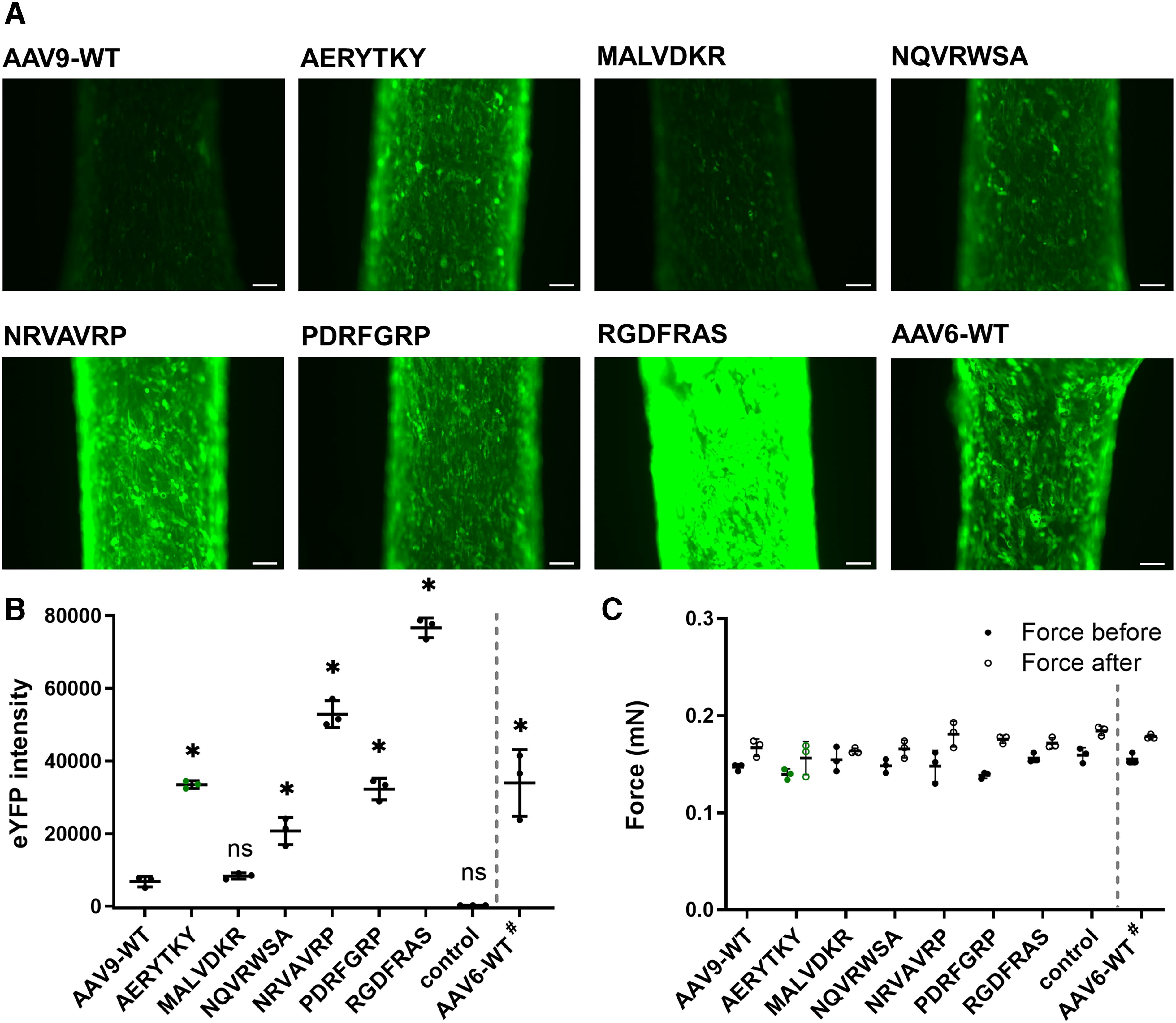
Evaluation of selected AAV capsid variants on human engineered heart tissue. The AAV capsids that showed the highest expression efficiency in vitro were tested for their ability to transduce hEHTs. The hEHTs were transduced with an MOI of 2.5 × 104 (AAV9 wild type and variants) or 2.5 × 103 (
DISCUSSION
The tailoring of AAV capsids toward specific tropisms and new beneficial properties is an ongoing and dynamic scientific pursuit. We aimed to use directed evolution approaches to facilitate transduction of cardiomyocytes. To improve the selection process of AAV random peptide library screenings, we compared selections on the DNA and RNA levels. As proof of concept, we found AAV capsid variants that have better transduction capability and/or transgene expression. Furthermore, we also identified AAV9-AERYTKY as a capsid variant that yields high titers in production and allows stable transgene expression in NRVCMs, which is even superior to the commonly applied AAV6 wild type. Finally, we could show that this variant allows an efficient gene expression in cardiomyocytes in mice and hEHTs.
Since the first publications on AAV peptide library selections in 2003, numerous groups have adapted and improved the process. For example, they have introduced PCR-based selections, 37 restricted the selection to specific cell types utilizing Cre recombinase or tissue-specific promoters, 15,21 and integrated machine learning into the AAV library context. 38 Performing AAV library selections based on the viral DNA or the expressed RNA in the target cells improved the selection toward capsid variants with high transduction or gene expression, respectively, which was discussed frequently. 16,17,19 During the course of this study, this was additionally evaluated by the Lisowski lab. 33
In this study, we used an initial AAV9 peptide library to enrich peptide motifs. This was done by processing the DNA or the RNA after AAV transduction. Through this process, we identified peptide motifs that either boosted cell entry, for example, by superior receptor targeting, or higher expression of the transgene, for example, by improved intracellular trafficking to the nucleus, uncoating of the capsid shell, or second-strand synthesis. Our data show clear enrichment of peptides at all levels, with significant differences between DNA- and RNA-based selections, consistent with previous findings. 33 While some peptide motifs can be found in both kind of selections, unique motifs are enriched when evaluating the top ten of each selection. The NRVCM selections show the expected impact of improved transduction by DNA-selected AAV variants and enhanced expression by RNA-selected variants. The majority of peptides displayed these benefits: DCDTLSF, GAKYAIC, and VEKCRVP exhibited increased transduction efficiency but unaffected expression, whereas moderately transducing capsid variants AERYTKY and PDRGFRP showed significantly higher transgene expression. Some motifs found in both selections, such as GSIVDRR, MALVDKR, and NRVAVRP, only slightly improved transduction and expression.
Interestingly, all peptide variants containing a cysteine in the introduced 7mer led to high levels of transduction in vitro but exhibited no gene expression. Similarly, these variants demonstrated detargeting from the liver in vivo and had higher transduction in other organs compared with AAV9 wild type but still enabled almost no transgene expression. AAV9 and AAV2 wild types contain only four conserved cysteine residues distal from the surface-exposed loop utilized for the heptamer insertion. In AAV2, it has been shown that the disulfide bonds between these cysteine residues are nonessential for the formation of the multi-subunit virion. 39 It is still unclear whether the improved transduction in vitro and liver detargeting in vivo is based, for example, on the hydrophobic effects of the cysteine residues. It is possible that the expression after cell entry might be impaired due to formation of disulfide bonds with nonviral components or interacting capsids, which could then inhibit capsid uncoating and viral genome release. However, further in-depth investigation is needed in the future.
In early AAV peptide library selections, cells latently transduced with the AAV library were co-infected with wild-type adenovirus to directly generate the preselected library progeny. 10,12,13,26,40 –46 This attempt resulted in AAV capsid variants that require both successful transduction and sufficient expression of the rep and cap genes. In 2009, the first PCR-based approaches were introduced after DNA isolation without the need of helper virus co-infection using AAV peptide display 37 and chimeric shuffled libraries 47 and were subsequently applied in further studies especially since it was easier-to-use for directed evolution in vivo. 14,15,48 –50 Some groups coupled the selection process (based on DNA enrichment) to the transgene expression by adding an expression marker such as a fluorophore 36,51 or Cre recombinase 32,52 to the vector genome. Recently, the selection was performed by isolating and processing RNA from specific tissues, 20,21 leading to the development of highly expressive capsid variants, forming the basis of this study. Westhaus et al. conducted a comparison study of adenovirus co-infection with PCR-based DNA and RNA selections on human hepatocytes in a xenograft mouse model. 33 Interestingly, one capsid variant, AAV-RC06, demonstrated high transgene expression while exhibiting rather inefficient transduction. Our findings support this notion that capsids recovered from the RNA selection were significantly more effective.
AAV9 vectors robustly transduce cardiomyocytes in mice, but achieve comparatively disappointing efficiencies in NRVCMs in vitro. NRVCMs are a crucial cell culture model for many cardiac diseases, including the induction of hypertrophy. However, studies in cultivated cardiomyocytes such as NRVCMs and cardiomyocytes differentiated from iPSCs require the generation of AAV6 vectors for efficient transduction. 4,5 Thus, it would be preferable to use one AAV vector for both in vitro and in vivo studies. Our discovered variant AAV9-AERYTKY might be versatile as one-vector solution for improved gene transfer in vitro while being directly applicable for studies in mice. In addition, we investigated the ability of our improved AAV variants to transduce engineered heart tissues differentiated from human iPSCs. The emergence of cardiac organoid technologies has significantly advanced cardiovascular research by allowing investigation of more accurate models of the human heart. This includes overcoming limitations of 2D monolayer cultures such as contractility, electrophysiology, and cell–cell and extracellular matrix interactions. 53,54 Using our new capsid variants for hEHT transduction, we achieved a significant increase in eYFP intensity and, hence, transgene expression. Among the efficient variants was also one containing an RGD motif that is known for strong interactions with RGD-binding integrin heterodimers facilitating targeting of cardiac and muscle tissue. 21,29
Previously published AAV variants have demonstrated notable differences in transduction efficiency across murine or human models. 1,55 Such species-specific effects are a known limitation of directed evolution approaches, as the mechanisms driving these improvements often remain elusive without additional investigation and may not be conserved between species. 8 A prominent example is the murine brain-targeting AAV vector PHP.B, which proved ineffective and toxic in nonhuman primates. 15,56 The encouraging initial results on human iPSCs suggest that the AAV9 capsid variants presented here could have potential translational value in large animal models, including nonhuman primates, and may ultimately support gene therapy approaches in the (pre)clinical context. Therefore, further in-depth investigations, particularly in mammalian models, are essential to elucidate the underlying mechanism enhanced by the introduced capsid modifications and to validate the efficacy, safety, and broader applicability of these novel vectors.
Furthermore, although AAV9 shows the strongest cardiac expression among AAV wild-type vectors after systemic application in mice, AAV6 has been shown to mediate high expression after indirect intracoronary injections 23 and, recently, strong gene transfer in organotypic heart slices. 57 In this context, our AAV9 capsid variants might facilitate gene transfer using alternative administration routes where barriers such as the immune response, overcoming the endothelial layer, or interaction with serum proteins are circumvented. 44
CONCLUSION
Taken together, we demonstrated the effects of AAV capsid variants derived from AAV9 peptide library selections based on the DNA and the RNA levels. Our RNA-selected capsids exhibit strong transgene expression in cardiomyocytes, outperforming the AAV6 wild type. Our lead candidate, AAV9-AERYTKY, demonstrated high expression in NRVCMs and could be directly employed for in vivo experiments. In addition, we present human iPSC data, showing significant transduction of engineered heart tissue using our capsid variants. In future studies, AAV9-AERYTKY could serve as cardiotropic AAV vector through alternative administration routes and should be further evaluated in large animal models.
Footnotes
ACKNOWLEDGMENTS
The authors gratefully thank Angela Lorenzen, Katrin Timm-Richert, and Arnie Boenders for invaluable technical assistance. The authors would also like to thank Sabine Michalewski and Claudia Kremling for assisting with paraffin sectioning and staining. The authors thank Thomas Schulze, Department of Experimental Pharmacology and Toxicology, University Medical Center Hamburg Eppendorf, for performing the EHT transduction and contractile analysis. The authors would like to acknowledge the support of the CYTO Kiel, Cytometry Facility of the UKSH. Graphic figures were created with BioRender.com.
DATA AVAILABILITY STATEMENT
The data can be obtained from the corresponding author upon reasonable request.
AUTHORS’ CONTRIBUTIONS
L.H.: conceptualization, methodology, investigation, formal analysis, visualization, and writing—original draft; L.C.S.: software, investigation, and formal analysis; P.M.V.S., A.M., A.R.B., and T.J.: methodology, investigation; S.S.H., A.K.R., D.G., D.F., G.J.J.B., T.E., and D.S.: conceptualization, formal analysis, project administration, and supervision; O.J.M.: conceptualization, methodology, investigation, formal analysis, project administration, supervision, visualization, and writing—original draft.
AUTHOR DISCLOSURE
L.H., L.C.S., T.J., G.J.J.B., T.E., and O.J.M. are inventors on a patent application covering AAV variants reported in this article. G.J.J.B. reports ownership interest in PacingCure B.V. A.R.B. is an employee of PacingCure B.V. The other authors declare no conflict of interest.
FUNDING INFORMATION
This work was supported by the
SUPPLEMENTARY MATERIAL
Supplementary Figure S1
Supplementary Figure S2
Supplementary Figure S3
Supplementary Figure S4
Supplementary Figure S5
Supplementary Figure S6
Supplementary Table S1
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
