Abstract
DddA-derived cytosine base editors (DdCBEs) enable the targeted introduction of C•G-to-T•A conversions in mitochondrial DNA (mtDNA). DdCBEs work in pairs, with each arm composed of a transcription activator-like effector (TALE), a split double-stranded DNA deaminase half, and a uracil glycosylase inhibitor. This pioneering technology has helped improve our understanding of cellular processes involving mtDNA and has paved the way for the development of models and therapies for genetic disorders caused by pathogenic mtDNA variants. Nonetheless, given the intrinsic properties of TALE proteins, several target sites in human mtDNA are predicted to remain out of reach to DdCBEs and other TALE-based technologies. Specifically, due to the conventional requirement for a thymine immediately upstream of the TALE target sequences (i.e., the 5′-T constraint), over 150 loci in the human mitochondrial genome are presumed to be inaccessible to DdCBEs. Previous attempts at circumventing this requirement, either by developing monomeric DdCBEs or utilizing DNA-binding domains alternative to TALEs, have resulted in suboptimal specificity profiles with reduced therapeutic potential. Here, aiming to challenge and elucidate the relevance of the 5′-T constraint in the context of DdCBE-mediated mtDNA editing, and to expand the range of motifs that are editable by this technology, we generated DdCBEs containing TALE proteins engineered to recognize all 5′ bases. These modified DdCBEs are herein referred to as αDdCBEs. Notably, 5′-T-noncompliant canonical DdCBEs efficiently edited mtDNA at diverse loci. However, they were frequently outperformed by αDdCBEs, which exhibited significant improvements in activity and specificity, regardless of the most 5′ bases of their TALE binding sites. Furthermore, we showed that αDdCBEs are compatible with the enhanced DddAtox variants DddA6 and DddA11, and we validated TALE shifting with αDdCBEs as an effective approach to optimize base editing outcomes. Overall, αDdCBEs enable efficient, specific, and unconstrained mitochondrial base editing.
INTRODUCTION
Mitochondria are semiautonomous organelles with a central role in energy metabolism that contain a circular and multicopy genome, mitochondrial DNA (mtDNA), which in humans encodes 37 genes critical for oxidative phosphorylation. 1 –7 Pathogenic mtDNA variants are prevalent in ∼1 in 5,000 people and are causal in currently incurable metabolic disorders. 8 –12 Mitochondrial base editing has recently emerged as a potential therapeutic approach for these mtDNA-based diseases. 13 –16 Notably, given the challenging nature of exogenous RNA import into mitochondria, the effective manipulation of mtDNA is enabled by all-protein systems. 16 –20 In light of their rapid and accessible engineering, transcription activator-like effectors (TALEs) are the most commonly used DNA-binding domains in most current mitochondrial base editing technologies. 16,21 –36
DddA-derived cytosine base editors (DdCBEs), consisting of pairs of mitochondrially targeted fusion proteins composed of a TALE, one half of a split dsDNA deaminase toxin (DddAtox), and a uracil glycosylase inhibitor, represent the most extensively developed tools for mtDNA editing. 16,30 –34 Based on prior work on TALE-DNA interactions in native contexts and in gene targeting applications in the nuclear compartment, 23 –27,37 –47 the target flexibility of DdCBEs on mtDNA is presumed to be constrained by the requirement of a 5′ thymine (5′-T) in their TALE binding sites, restricting over 150 loci in the human mitochondrial genome. 13,48 This design guideline stems from the specific interaction between the highly conserved N-terminal domain (NTD) of wild-type TALEs and the 5′-T of their target sequences. 37 –40 Despite being generally followed, the significance of this constraint for the design of effective DdCBEs is yet to be characterized. 13,16,30 –34
Here, we sought to elucidate the relevance of the TALE 5′-T rule for DdCBE-mediated mitochondrial base editing, as well as to expand the targeting scope and design flexibility of this technology. To this end, building on our recently established system for the assembly of TALE-guided deaminases, 49 –51 we generated DdCBE variants that contain a previously developed TALE NTD engineered to accommodate any 5′ base. 52 We designated these unconstrained DdCBEs as αDdCBEs. We conducted direct comparisons between DdCBEs and αDdCBEs in six mitochondrial genes, ND4, ND2, ATP6, CO1, TC, and TL1. Remarkably, we noted that breaking the 5′-T rule did not obligately preclude mtDNA editing with DdCBEs. However, αDdCBEs consistently outperformed canonical DdCBEs, thereby supporting unconstrained mtDNA editing as a potential strategy for disease modeling and gene therapy applications.
MATERIALS AND METHODS
Construction of FusX-compatible DdCBE backbone plasmids
All backbone plasmids were made via restriction cloning. A list including the source material for each construct is provided in Supplementary Table S1. In general, insert and vector bands were separated by agarose gel electrophoresis and purified with the Monarch DNA Gel Extraction Kit obtained from New England Biolabs (NEB). Ligations were done with the Quick LigationTM Kit (NEB). NEB® Stable Competent E. coli (C3040H) were used for propagation, following the high efficiency transformation protocol specified by the manufacturer, and incubating plates and liquid cultures at 30°C. Plasmids were purified with the QIAprep Spin Miniprep Kit (Qiagen) and sequence-verified via whole-plasmid sequencing (Primordium Labs).
Assembly of DdCBE-encoding plasmids
All DdCBE-encoding plasmids used in this study were assembled via the FusX TALE Base Editor (FusXTBE) platform. 49 –51 Briefly, following standard design rules, 13,53 DdCBEs were designed in silico with TALE Writer 50,51 and SnapGene. Specifically, TALE repeat arrays were designed to target between 15 and 17 bp, separated by spacers ranging from 11 to 18 bp long. 13,50,53 A list of all TALE binding sites is provided in Supplementary Table S2. DdCBE-encoding plasmids were assembled via Golden Gate cloning. 49 –51 Primers used for colony PCR (see Supplementary Table S3) were synthesized as standard DNA oligos by Integrated DNA Technologies (IDT). NEB® stable competent E. coli (C3040H) were used for propagation. Plasmids were purified with the QIAprep Spin Miniprep Kit (Qiagen) and sequence-verified via whole-plasmid sequencing (Primordium Labs).
Generation of TALE-free constructs
Plasmids encoding TALE-free MTS–G1397-split DddAtox/DddA6/DddA11–UGI were generated with the Q5® Site-Directed Mutagenesis (SDM) Kit (NEB), using the corresponding herein developed backbone plasmids as templates for each final construct containing the DddAtox, DddA6, or DddA11 C- or N-terminal halves. 13,53 Cloning was carried out following the manufacturer’s instructions with the provided NEB® 5-alpha competent E. coli cells. Primers for SDM (see Supplementary Table S3) were designed using NEBaseChanger (NEB) and synthesized as standard DNA oligos (IDT). Plasmids were purified with the QIAprep Spin Miniprep Kit (Qiagen) and sequence-verified via whole-plasmid sequencing (Primordium Labs).
Mammalian cell culture and lipofection
HEK293T cells (CRL-3216TM, ATTC) were maintained at 37°C and 5% CO2. The cells were cultured in high-glucose DMEM (Thermo Fisher Scientific) supplemented with 10% (v/v) fetal bovine serum (Thermo Fisher Scientific) and 100 U ml−1 penicillin–streptomycin (Thermo Fisher Scientific). LipofectamineTM 3000 Transfection Reagent (Thermo Fisher Scientific) was used for lipofections. In brief, 24 h before lipofection, 0.3 × 106 cells/well were seeded in 6-well plates. Then, lipofections proceeded with 500 ng per monomer for DdCBEs and TALE-free constructs to make up 1,000 ng of total plasmid DNA, 13,53 and 1,000 ng of plasmid DNA for monomeric DdCBEs (mDdCBEs). 48 Cells were collected for genotyping at 72 h post-transfection.
Genomic DNA isolation from mammalian cell culture
At 72 h post-transfection, the cell medium was aspirated, the cells were washed with 500 µL 1× DPBS without calcium or magnesium (Thermo Fisher Scientific), trypsinized with 500 µL 1× Trypsin-EDTA (0.5%) without phenol red (Thermo Fisher Scientific) for 5 min at 37°C and collected in microcentrifuge. Total genomic DNA (including mitochondrial DNA) was purified using the DNeasy Blood & Tissue Kit (Qiagen) following the manufacturer’s instructions and stored at −20°C until further downstream processing.
High-throughput sequencing of genomic DNA samples
Genotyping primers were designed using Primer-BLAST, 54 querying the Homo sapiens genome assembly hg38 for primer pair specificity. In detail, to increase PCR specificity for the intended mitochondrial target sequences, PCR was biased against the amplification of nuclear mitochondrial pseudogenes (NUMTs) 55 by aligning the 3′ ends of candidate primers with specific single-nucleotide mismatches between intended mitochondrial targets and potential unintended nuclear templates, and accordingly adding 3′-terminal phosphorothioate (PS) bonds to the primers. This strategy avoids 3′-terminal editing of the mismatched primers by the 3′−5′ exonuclease activity of Q5® High-Fidelity DNA Polymerase (NEB), increasing PCR specificity. 56
Primers including the partial Illumina® forward and reverse adapter sequences, in addition to barcodes for sample multiplexing, were synthesized as UltramerTM DNA oligos (IDT). Afterward, genomic sites of interest were amplified with the Q5® High-Fidelity 2X Master Mix (NEB) using conventional thermocycling conditions. Then, PCR products corresponding to the same experimental condition but with different barcodes were combined after agarose gel electrophoresis, purified using the Monarch DNA Gel Extraction Kit (NEB), confirmed via Sanger sequencing (Genewiz), and submitted to next-generation sequencing (NGS, Amplicon-EZ with partial adapters, Genewiz). Alternatively, if the samples were not multiplexed, the PCR products were individually purified with the QIAquick PCR Purification Kit (QIAGEN), confirmed via Sanger sequencing (Genewiz), and submitted to NGS (Amplicon-EZ without partial adapters, Genewiz). A list of all genotyping primers is provided in Supplementary Table S4.
Analysis of high-throughput sequencing data
In multiplexed samples, the paired-end read FASTQ files generated by NGS were demultiplexed and analyzed utilizing the CRISPRessoPooled tool within CRISPResso2. 57,58 Similarly, if the samples were not multiplexed, the paired-end read FASTQ files were analyzed with the CRISPRessoBatch tool within CRISPResso2. 57,58 In general, DdCBE spacer sequences were used as the guide sequence input. Besides, for each replicate in each experimental condition, the sequence of the amplicon corresponding to the target site, plus the respective barcode if demultiplexing, was used as the amplicon sequence input. Unless otherwise stated, the quantification window size was set to 8 or 10, and the quantification window center was set to −8 or −10. All optional parameters were set to NA. 13,57
The output allele frequency table was used to determine the overall on-target editing in each sample, calculated as the percentage of aligned reads with C•G-to-T•A conversions within a spacer. 48 Likewise, the output nucleotide percentage table was used to calculate the editing activity at each cytosine within each spacer, as well as the proximal off-target editing within each amplicon. 29,48 In detail, similar to the methodology followed in the development of zinc-finger DdCBEs (ZF-DdCBEs), average amplicon-wide off-target editing was quantified as the sum of all C•G-to-T•A conversions within an amplicon, excluding its corresponding DdCBE spacer, over the total number of C•G base pairs within that amplicon. 29 Calculations were done in Microsoft Excel.
Targeted amplicon sequencing for nuclear DNA off-target analyses
Based on previous reports, nested PCR was performed to amplify a TALE-dependent off-target site within the NUMT MTND4P12, and conventional PCR to amplify a frequent TALE-independent off-target site at chr8:37153384C (hg38). 13,59,60 Primers for the first PCR (PCR1) in the nested PCR strategy were synthesized as standard DNA oligos (IDT). Primers for the generation of amplicons for NGS were synthesized as UltramerTM DNA oligos (IDT), including barcodes and Illumina® adapters. PCR was done with the Q5® High-Fidelity 2X Master Mix (NEB). After PCR1 in the nested PCR strategy, amplicons were purified with the QIAquick PCR Purification Kit (Qiagen), and 10 ng were used as template DNA for the second PCR. Amplicons for targeted deep sequencing were purified as detailed in the section “High-Throughput Sequencing of Genomic DNA Samples”” and submitted to NGS (Amplicon-EZ with partial adapters, Genewiz).
The methodology for data analysis was similar to the approach described in the section “Analysis of High-Throughput Sequencing Data.” In detail, to determine the overall nuclear DNA off-target editing at MTND4P12, the CRISPResso2 output allele frequency table was used to calculate the percentage of aligned reads with C•G-to-T•A conversions within the pseudospacer (i.e., the nuclear DNA region analogous to the genuine target spacer in mtDNA). In addition, the output nucleotide percentage table was used to calculate the editing activity at each cytosine within the pseudospacer. Similarly, the nucleotide percentage table was used to quantify the nuclear DNA off-target editing at the abovementioned TALE-independent off-target locus.
Sanger sequencing of genomic DNA samples and data analysis
For the 5′ nucleotide precedence analyses at the ATP6 locus, genomic DNA from ATP6-edited cells and the corresponding controls was purified as described in the section “Genomic DNA Isolation from Mammalian Cell Culture.” Afterward, PCR was conducted with the Q5® 2X Master Mix (NEB) and ATP6 Sanger sequencing primers (listed in Supplementary Table S4), which were designed as detailed in the section “High-Throughput Sequencing of Genomic DNA Samples” (without adapters or barcodes) and synthesized as standard DNA oligos with 3′ PS bonds (IDT). Then, PCR products were visualized by electrophoresis in a 1% agarose gel, purified with the QIAquick PCR Purification Kit (QIAGEN), and submitted to Sanger sequencing (Genewiz). The resulting trace (.ab1) files were analyzed in the EditR server. 61 Briefly, DdCBE spacer sequences were used as the guide sequence input. 50 In addition, the 5′ starts and 3′ ends of the trace files were trimmed to exclude bases with quality scores lower than 40, and the p value cutoff for calling base editing was set to 0.01. Besides, to exclude noise from low-confidence measurements in the calculation of the average editing efficiencies, these were computed per replicate as the mean of the predicted editing at cytosines within the spacer that corresponded to highly significant (p ≤ 0.01) editing events, compared with untreated controls. Statistical analyses were conducted using two-tailed unpaired t tests in GraphPad Prism 10.
RESULTS
Mitochondrial base editing with αDdCBEs
In general, most TALEs require a thymine base immediately upstream of their target sequences for efficient TALE-DNA binding. 37 –40 Thus, canonical DdCBEs, which contain standard TALE proteins, are predicted to induce efficient mtDNA editing only in 5′-T-compliant formats, that is, when the TALE target sequences are preceded by a thymine. 13,48 Consequently, given the ability of the modified TALE NTD to recognize all 5′ bases, 52 we hypothesized that αDdCBEs can edit mtDNA as efficiently as standard, 5′-T-compliant DdCBEs, regardless of the TALE 5′-T rule. Moreover, we expected 5′-T-noncompliant DdCBEs to induce poor editing efficiencies relative to 5′-T-compliant DdCBEs. To test these hypotheses, we compared pairs of DdCBEs and αDdCBEs in both 5′-T-compliant and 5′-T-noncompliant formats at four mitochondrial loci in HEK293T cells (Fig. 1). To avoid variations in base editing outcomes as a result of spacer variability, we maintained fixed spacer sequences at each target locus by lengthening or shortening the TALE binding sites from the 5′ ends by a maximum of 2 bp each.
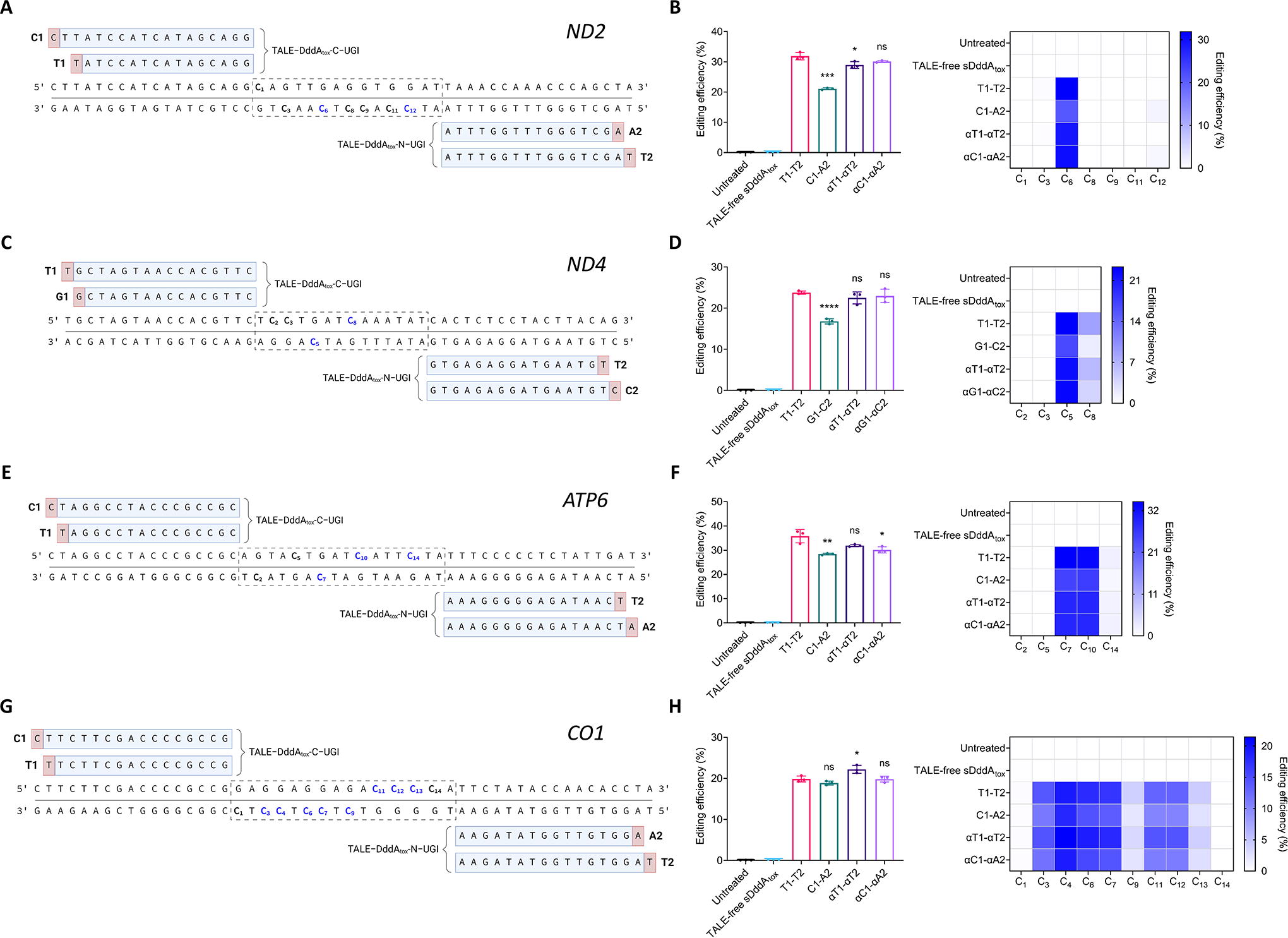
Mitochondrial base editing with 5′-T-compliant and 5′-T-noncompliant DdCBEs and αDdCBEs.
In addition to the untreated condition, TALE-free MTS–split DddAtox–UGI (labeled as TALE-free sDddAtox) was used as a negative control. Additionally, DdCBE pairs are generally referred to as N1-N2 (N = A, C, G, or T), denoting the most 5′ base of either the left (1) or the right (2) TALE. Accordingly, αDdCBEs are designated as αN1-αN2. Hence, T1-T2 denotes a 5′-T-compliant DdCBE pair, which corresponds to a positive control. Besides, unless otherwise noted, all base editors were designed with G1397-split DddAtox in the C-to-N configuration, that is, left TALE–DddAtox-C–UGI + right TALE–DddAtox-N–UGI. 13
Based on previous work on the development of DdCBEs and the FusXTBE platform, the mitochondrial genes ND2, ND4, and CO1 were chosen for these experiments. 13,50 In addition, the target site within the ATP6 locus was selected due to its sequence structure, which enabled the testing of several combinations of base editors with TALEs preceded by any 5′ base (explored in detail in Fig. 3).
At the ND2 locus, we observed that C1-A2 was the least active base editor, reaching overall editing efficiencies of ∼21%, whereas T1-T2, αT1-αT2, and αC1-αA2 displayed editing frequencies ranging from ∼29% to ∼32%. In addition, all ND2 base editors resulted in similar mutation patterns (Fig. 1A, B). Similarly, at the ND4 site, we found that G1-C2 induced editing efficiencies of ∼17%, the lowest compared with T1-T2, αT1-αT2, and αG1-αC2, which installed edits at frequencies between ∼22% and ∼24%. Interestingly, both G1-C2 and αG1-αC2 resulted in more specific mutation patterns within the spacer than their 5′-T-compliant counterparts (Fig. 1C, D). Comparably, at the ATP6 locus, C1-A2 was less efficient than T1-T2, with editing frequencies of ∼28% compared with ∼36%, respectively. Moreover, both αT1-αT2 and αC1-αA2, which displayed editing frequencies of up to ∼32%, were nearly as effective as T1-T2. Notably, all ATP6 base editors displayed similar editing patterns (Fig. 1E, F). Unexpectedly, at the CO1 site, all base editors displayed similar levels of activity and mutation patterns, with overall efficiencies ranging from ∼19 to ∼22%. Besides, despite the preference of DddAtox for cytosines in TC motifs, 13 several non-TC motifs within the CO1 spacer were efficiently edited (Fig. 1G, H).
Collectively, these results suggest that canonical DdCBEs can effectively edit mtDNA even if their respective TALEs break the 5′-T rule. However, αDdCBEs tend to perform similarly to 5′-T-compliant DdCBEs and outperform 5′-T-noncompliant DdCBEs, thereby surpassing canonical DdCBEs in regard to design flexibility.
Characterizing off-target editing by αDdCBEs
Seeking to characterize the specificity profiles of αDdCBEs relative to DdCBEs, based on an approach reported by Willis et al., 29 we calculated the normalized ratios between the on-target (i.e., within the spacer) and average amplicon-wide off-target editing efficiencies for each base editor. These quantities enabled us to conduct direct comparisons between the overall performance of αDdCBEs in contrast to DdCBEs, both in terms of their on-target editing activities and proximal off-target effects.
At the ND2 locus, C1-A2, αT1-αT2, and αC1-αA2 resulted in an ∼1.8-fold reduction in average amplicon-wide off-target editing compared with T1-T2 (Fig. 2A). Accordingly, given their high on-target editing activities (Fig. 1B) and relatively low off-target effects (Fig. 2B, left), αDdCBEs considerably outperformed their canonical counterparts at the ND2 site (Fig. 2B, right). In contrast, all ND4 base editors introduced off-target editing at frequencies below 0.2% throughout the amplicon (Fig. 2C). However, given the relatively low on-target activity displayed by G1-C2 (Fig. 1D), and the moderately higher off-target effects caused by T1-T2 compared with the other pairs (Fig. 2D, left), at the ND4 locus, αDdCBEs outperformed DdCBEs (Fig. 2D, right). Similarly, all ATP6 base editors introduced off-target cytosine conversions at rates below 0.5% (Fig. 2E). However, both T1-T2 and C1-A2 resulted in somewhat higher average amplicon-wide off-target editing than αT1-αT2 and αC1-αA2 (Fig. 2F, left). Consequently, at the ATP6 site, αDdCBEs performed better than DdCBEs (Fig. 2F, right). In contrast, at CO1, the 5′-T-compliant pairs resulted in higher average amplicon-wide off-target editing efficiencies compared with the 5′-T-noncompliant pairs (Fig. 2G and Fig. 2H, left). Therefore, both 5′-T-noncompliant CO1 base editors outperformed the 5′-T-compliant DdCBE pair, T1-T2, with αC1-αA2 showing the highest overall performance (Fig. 2H, right).
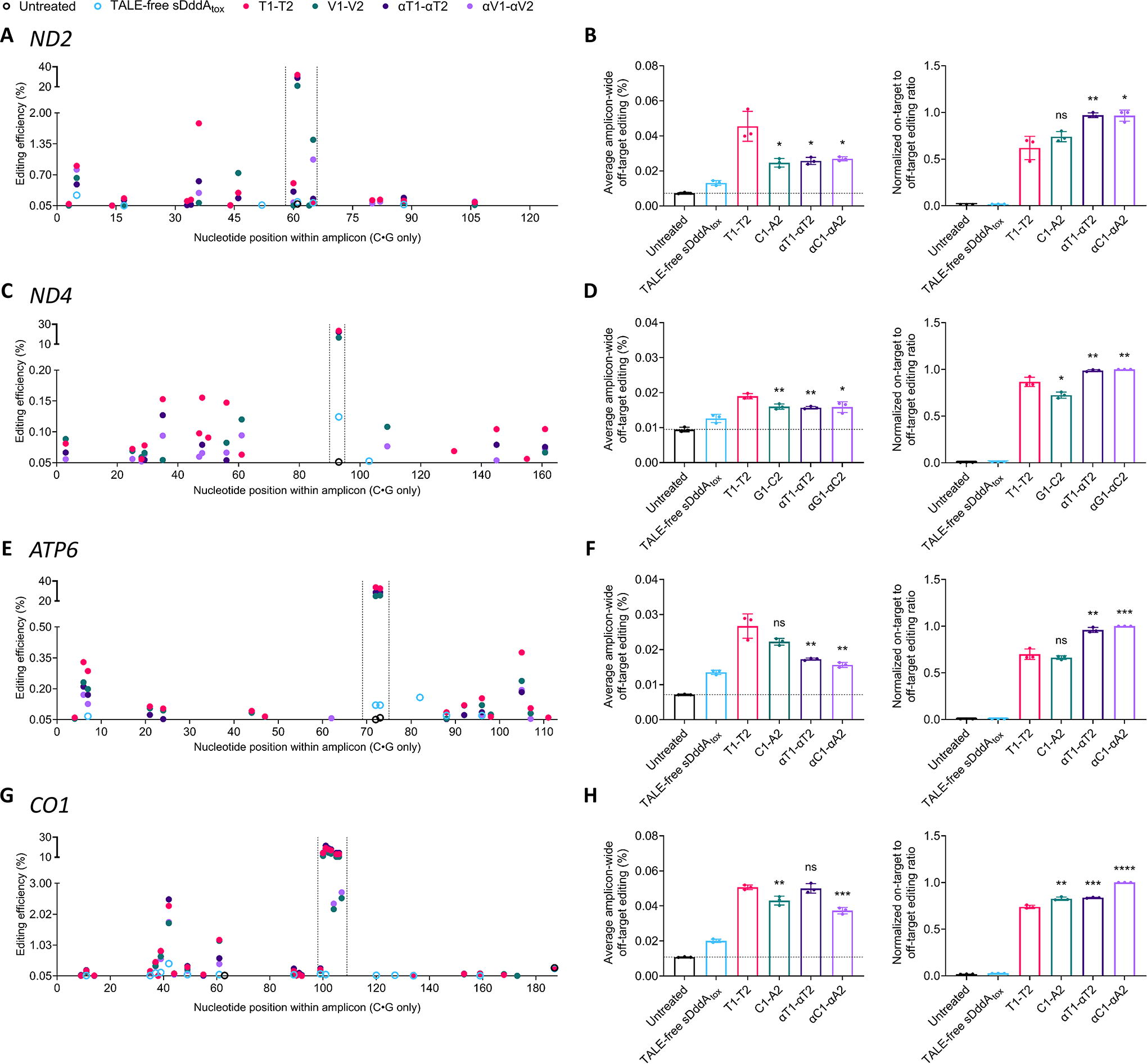
Proximal off-target effects by 5′-T-compliant and 5′-T-noncompliant DdCBEs and αDdCBEs.
Subsequently, to further characterize the specificity profiles of αDdCBEs relative to DdCBEs, we investigated their nuclear off-target effects at a TALE-dependent site (MTND4P12) and a TALE-independent site (chr8:37153384C, hg38) in ND4-edited cells (Supplementary Fig. S1). 13,60 Notably, the MTND4P12 off-target and the ND4 on-target sequences differ by a single G/A mismatch (Supplementary Fig. S1A). Remarkably, at MTND4P12, T1-T2 resulted in off-target editing efficiencies of ∼16%. In contrast, all other pairs achieved frequencies of ∼0.2% (G1-C2), ∼7.5% (αT1-αT2), and ∼1% (αG1-αC2) (Supplementary Fig. S1B,C). On the contrary, at the herein examined TALE-independent off-target region, which was previously identified by Lei et al. as a frequently observed nuclear off-target across DdCBEs, and shares no sequence homology with the ND4 on-target sequence, the editing efficiencies remained substantially similar among base editors. Nonetheless, αG1-αC2 resulted in moderately higher cytosine conversion rates relative to all other editors, although at efficiencies below 0.2%. Besides, TALE-free sDddAtox led to off-target editing with frequencies of ∼1% (Supplementary Fig. S1D).
As a whole, these results suggest that, in the scope of proximal off-target effects in mtDNA and, potentially, nuclear editing at TALE-dependent off-target sites, αDdCBEs tend to be more specific than standard, 5′-T-compliant DdCBEs, thereby outperforming them in terms of specificity.
Comparative analyses of the on-target activities of DdCBEs and αDdCBEs preceded by all 5′ bases
We then explored whether αDdCBEs consistently led to on-target (within a spacer) base editing enhancements relative to DdCBEs, regardless of the 5′ bases of their TALE binding sites. To this end, we identified ATP6 as a locus accessible by base editors containing TALEs targeting sequences preceded by A, C, G, or T, with moderate variability across the resulting spacers. We designed TALE proteins containing between 15 and 17 repeats, and delimiting spacers ranging from 13 to 18 bp long. Of note, all spacers contained a bottom- and a top-strand cytosine approximately halfway through, and an additional top-strand cytosine toward the 3′ end (Fig. 3A).
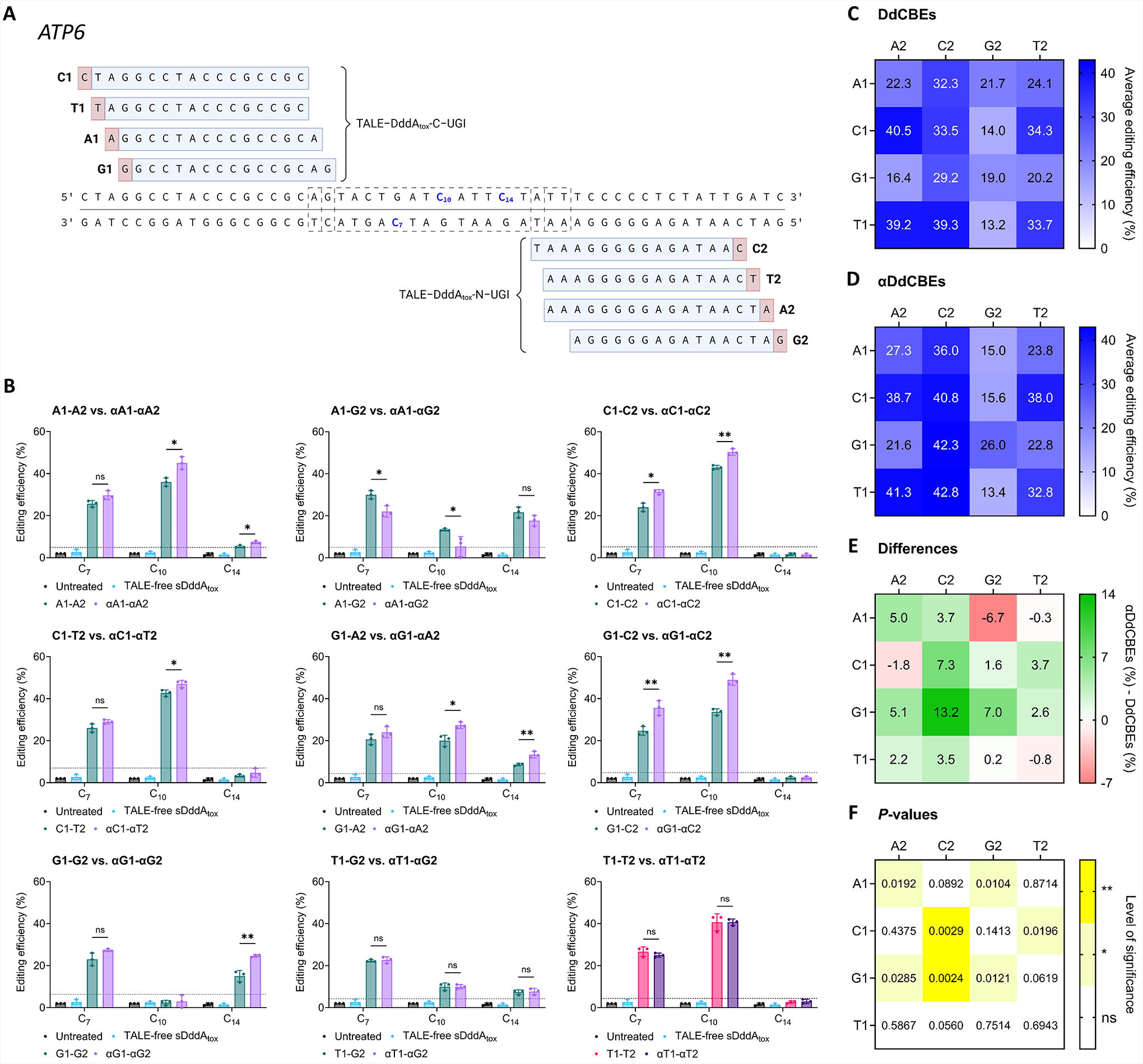
On-target editing efficiencies at ATP6 with DdCBEs and αDdCBEs preceded by all 5′ bases.
We observed that equivalent base editors (i.e., A1-A2 and αA1-αA2, A1-C2 and αA1-αC2, and so on) led to similar mutation patterns, although frequently with moderately different efficiencies. Representative comparisons between equivalent pairs are shown in Figure 3B, and the complete set is displayed in Supplementary Figure S2. Afterward, we calculated the average activities of each construct to elucidate the general differences between αDdCBE- and DdCBE-induced base editing frequencies at ATP6 (Fig. 3C, D). Then, we determined the differences between these quantities for each pair of equivalent base editors, along with their respective significance levels, facilitating the visualization of the overall αDdCBE-induced activity enhancements across comparisons (Fig. 3E, F). Notably, broad improvements in the efficiency of base editing with αDdCBEs relative to DdCBEs were observed in 6 out of the 16 total comparisons (Fig. 3C–F).
It is worth noting that ATP6 base editors containing G2/αG2 arms were generally the least effective across conditions (Fig. 3C, D); moreover, only αA1-αG2 led to a statistically significant overall activity reduction relative to its canonical counterpart (Fig. 3E, F). Furthermore, only G2/αG2-containing pairs, except for G1-G2 and αG1-αG2, targeted spacer sequences with lengths of 17 or 18 bp (Fig. 3A). DddA-derived base editors targeting spacers of such lengths often install lower editing efficiencies compared with pairs with spacers up to 16 bp long. 13,53 Thus, we hypothesized that reducing the spacers of G2/αG2-containing pairs to 16 bp or less would improve their editing efficiencies.
To test this hypothesis and characterize the effects of decreasing spacer length on the activities of A1-G2 versus αA1-αG2, we evaluated two additional sets of arms: G2.16/αG2.16 and G2.17/αG2.17 (Supplementary Fig. S3A). Interestingly, G2.16/αG2.16-containing pairs led to lower editing efficiencies relative to the initial G2/αG2-containing pairs (designated as G2.15/αG2.15 in Supplementary Fig. S3), as well as minimal αDdCBE-induced enhancements. Strikingly, G2.17/αG2.17-containing pairs, with shorter spacers, led to improvements in base editing efficiencies relative to the original pairs, as well as greater αDdCBE-induced base editing reductions (Supplementary Fig. S3B).
Overall, these results further indicate that DdCBEs and αDdCBEs can effectively edit mtDNA in 5′-T-noncompliant formats. However, αDdCBEs can lead to greater mtDNA editing efficiencies than their canonical counterparts, although in particular contexts the opposite can be observed.
αDdCBEs outperform DdCBEs at mtDNA sites with stretches without 5′-T nucleotides
Subsequently, we evaluated the effectiveness of αDdCBEs at target sites that, based on standard design principles, 13 cannot be accessed without breaking the TALE 5′-T rule. To this purpose, we first assessed an array of base editors at the tRNA-Cys-encoding gene TC. This locus is reportedly editable by mDdCBEs but not by dimeric DdCBEs. 48 In detail, we tested a standard pair (A1-T2), two partially modified pairs (αA1-T2 and A1-αT2), and a fully modified pair (αA1-αT2). In addition, we included two monomeric controls: mA1 and mT2, analogous to the left and right arms of the dimeric constructs (Fig. 4A). Unexpectedly, the dimeric base editors induced overall editing efficiencies ranging from ∼49% to ∼56%, well above mA1 (∼24%) and mT2 (∼14%) (Fig. 4B). Furthermore, A1-T2, αA1-T2, A1-αT2, and αA1-αT2 were considerably more specific than mA1 and mT2, which installed edits outside of the intended target sequence at frequencies of up to ∼6% and ∼3%, in that order (Fig. 4C and Fig. 4D, left). Therefore, the dimeric base editors far outperformed their monomeric counterparts, with αA1-αT2 exhibiting the greatest overall performance (Fig. 4D, right).
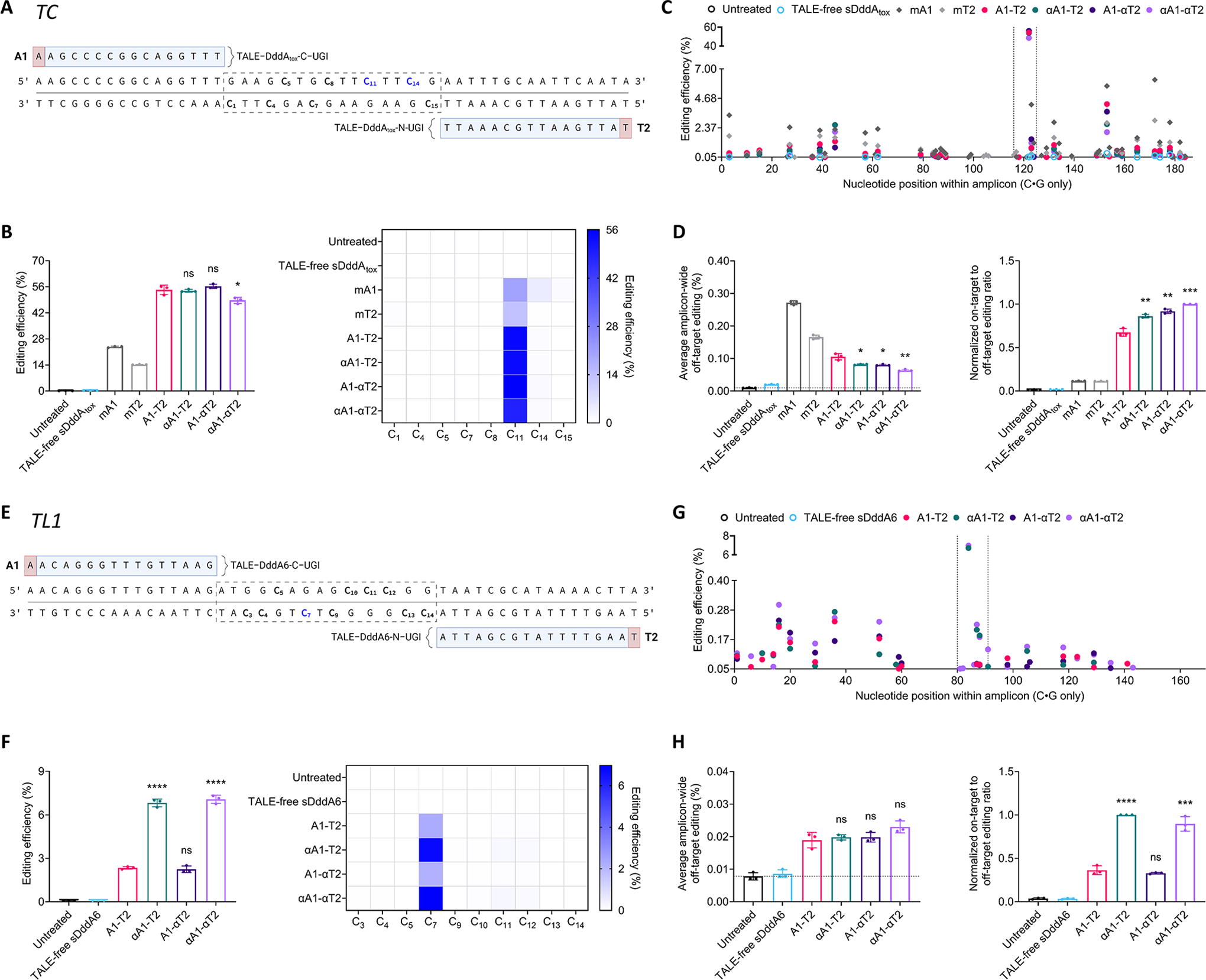
αDdCBEs effectively edit mtDNA at sites with stretches without 5′-T nucleotides.
To further evaluate the effectiveness of αDdCBEs at target sites lacking accessible 5′-Ts, we assessed a set of base editors at the tRNA-Leu-encoding gene TL1. Pathogenic variants in this gene, such as the broadly prevalent m.3243A>G, are linked to impaired oxidative phosphorylation and a wide range of complex disease outcomes. 62 –64 We initially observed that DddAtox was poorly active at TL1; hence, to obtain an ample range of activities for comparison purposes, we used DddA6, an enhanced variant of DddAtox, which showed improved editing efficiencies at this site (Supplementary Fig. S4). Of note, the denominations of the TL1 base editors are similar to those of the TC pairs (Fig. 4E). Besides, given that monomeric variants for DddA6 are yet to developed, 48,53 TL1-specific monomeric controls were not included. Remarkably, the αA1-containing pairs led to an approximately threefold increase in activity relative to A1-T2 and A1-αT2 (Fig. 4F). Moreover, all base editors resulted in similar specificity profiles; thus, the αA1-containing pairs significantly outperformed their A1-containing counterparts (Fig. 4H).
These results collectively suggest that dimeric DddA-derived base editors containing either canonical or unconstrained TALE NTDs, or both, can effectively access loci with stretches without 5′-Ts. Nevertheless, in some contexts, utilizing unconstrained TALEs to target sequences preceded by non-T nucleotides can facilitate the introduction of targeted modifications in mtDNA.
TALE shifting with αDdCBEs as a strategy to fine-tune mitochondrial base editing outcomes
The outcomes of DddA-derived base editors are partly determined by spacer length and the positions of the target cytosines within the spacer. 13,28,29,53 Given that these determinants are contributed by the DNA-binding domains, targeting a particular locus with different pairs of TALEs can lead to diverse editing outcomes. 32,65 Indeed, depending on their TALE proteins, the ATP6 base editors developed in this study resulted in distinct mutation patterns (Fig. 3B). Thus, focusing on the disease-relevant gene TL1 and based on its optimized TALE formats (Fig. 4E–H), we aimed to explore TALE shifting with αDdCBEs as a strategy to fine-tune mitochondrial base editing outcomes.
Seeking to further increase the editing efficiencies at TL1, and considering that DddA6-containing base editors can result in modest levels of activity (Fig. 4F), we switched to DddA11, a deaminase variant with higher relative editing efficiencies compared with both DddAtox and DddA6. 53 Importantly, the enhanced activity of DddA11 can lead to decreased target selectivity, since it can process cytosines in non-TC motifs, which tend to be poorly processed by DddAtox or DddA6. 53 Nonetheless, for precision genome editing applications, we reasoned that the unconstrained TALEs of αDdCBEs could be shifted around a target site to mitigate unintended editing events within a spacer.
To investigate this premise, we focused on the m.3242G>A variant at the TL1 locus, which is associated with various disease phenotypes, including mitochondrial myopathy. 66 –73 This mutation has been reported in patient tissues in both heteroplasmic (where wild-type and mutant mtDNA coexist) and homoplasmic states (where all mtDNA molecules contain the mutation). As a proof-of-concept, we attempted to install this point mutation at heteroplasmic levels in vitro. Notably, only a homoplasmic cellular model of the m.3242G>A variant has been developed to date, limiting the investigation of the heteroplasmic condition primarily to clinical observations and the analysis of patient tissues. 66 –70
In detail, we examined the performance of eight partially or fully modified base editors in installing the disease-associated variant m.3242G>A at the TL1 locus in HEK293T cells. This base transition is equivalent to C-to-T editing at C7 in the TL1 spacer region (Fig. 4E and Fig. 5A). For clarity, the partially modified pairs are denoted as αLDdCBEs, as only the left (L) arms contain unconstrained TALEs, and the fully modified pairs are referred to as αDdCBEs. Of note, TL1 αDdCBE 3NC corresponds to a pair with G1397-split DddA11 in the N-to-C configuration. All other pairs, including TL1 αDdCBE 3CN, contain G1397-split DddA11 in the C-to-N orientation.
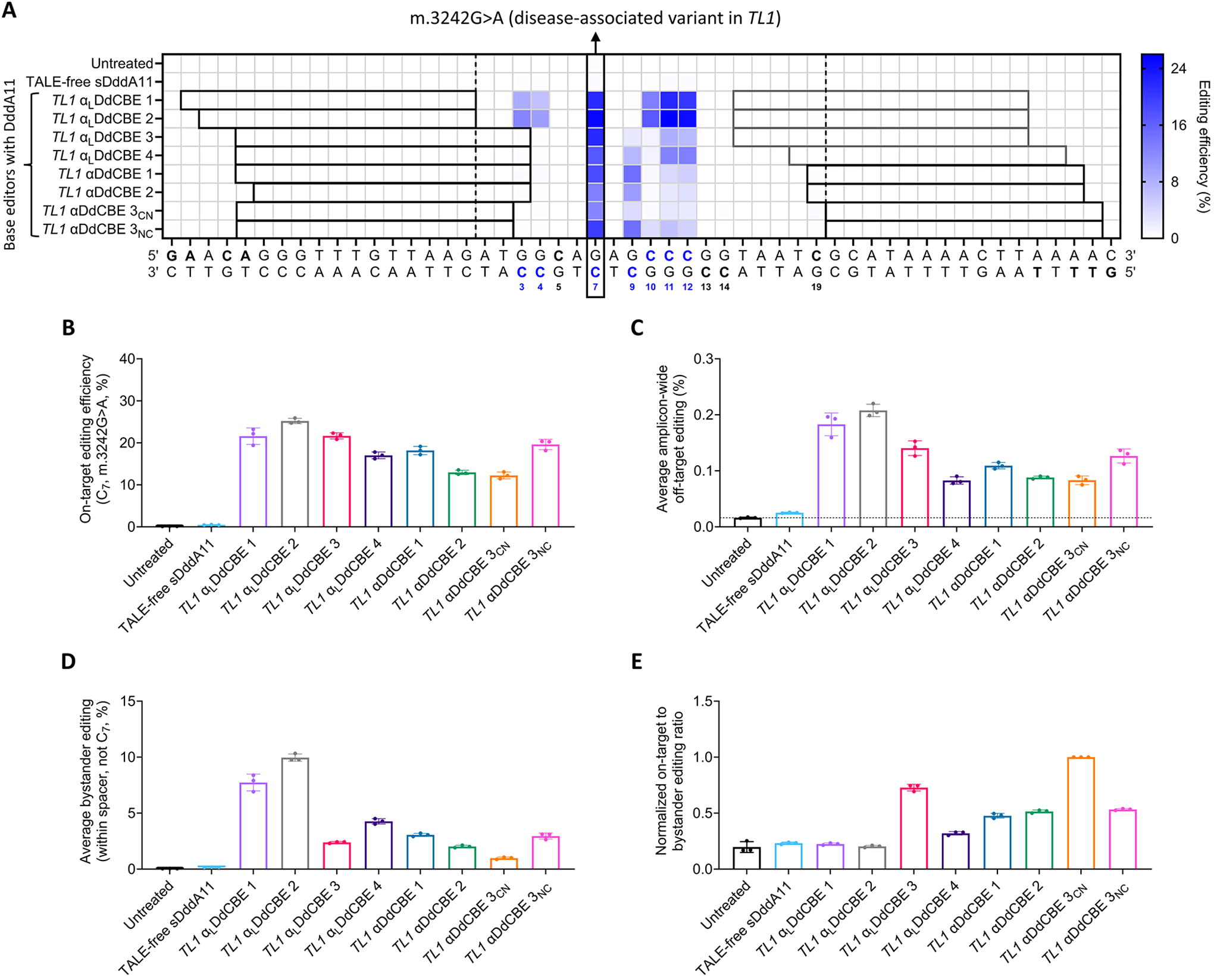
Fine-tuning mitochondrial base editing outcomes at the TL1 locus via TALE shifting.
It is important to emphasize that, in this context, only C-to-T editing at C7 within the overall spacer region (i.e., the installment of the m.3242G>A mutation) corresponds to on-target activity, while the conversion of other cytosines within a spacer is, by definition, bystander editing. 53,74 Likewise, as in previous analyses, editing events outside of a spacer are considered off-target effects. 29,74
In contrast to the low editing efficiencies observed at C7 with the DddA6-containing TL1 base editor αA1-T2 (∼7%, Fig. 4F), its DddA11-containing counterpart, TL1 αLDdCBE 2, edited C7 with substantially higher efficiencies (∼25%, Fig. 5A, B). However, while αA1-T2 resulted in amplicon-wide off-target editing at an average frequency of ∼0.02% (Fig. 4H, left), TL1 αLDdCBE 2 installed off-target edits at an average frequency of ∼0.21% throughout the amplicon (Fig. 5C). Besides, both base editors led to distinct editing patterns (Fig. 4F, right, and Fig. 5A). Furthermore, TALE shifting enabled the screening of additional DddA11-containing pairs with varying performance metrics, as summarized in Figure 5.
Overall, TL1-specific, DddA11-containing αLDdCBEs and αDdCBEs led to on-target editing efficiencies ranging from ∼12% to ∼25% (Fig. 5B). In addition, TL1 αDdCBEs can display improved levels of specificity compared with some TL1 αLDdCBEs (Fig. 5C and Supplementary Fig. S5). In detail, TL1 αDdCBEs 1, 2, and 3CN resulted in significantly less average amplicon-wide off-target editing frequencies compared with TL1 αLDdCBEs 1, 2, and 3, but greater or similar levels of off-target effects relative to TL1 αLDdCBE 4. Similarly, TL1 αDdCBE 3NC was more specific than TL1 αLDdCBEs 1 and 2, but not more specific than TL1 αLDdCBEs 3 and 4.
Regarding mutation patterns, C7 was efficiently edited by all pairs. In particular, TL1 αLDdCBEs 1 and 2, which displayed promiscuous editing profiles, were the only pairs to edit C3 and C4. In contrast, more than half the activity of TL1 αLDdCBE 3 and αDdCBE 3CN corresponded to editing at C7 (Fig. 5A). Furthermore, despite differing in spacer length by just 1 bp relative to TL1 αDdCBEs 1 and 2, TL1 αLDdCBE 4 displayed increased activity at C11 and C12 (p < 0.0001 for all comparisons) and decreased editing at C9 (p < 0.001 for both comparisons). Moreover, likely due to their opposite orientations of split DddA11, 53 TL1 αDdCBEs 3CN and 3NC resulted in distinct mutation patterns.
In addition to assessing on- and off-target editing activities (Fig. 5B, C), we measured each base editor’s average bystander editing (Fig. 5D). Notably, TL1 αDdCBE 3CN displayed the lowest mean bystander editing efficiencies (p < 0.001 for all comparisons), while TL1 αLDdCBE 2 showed the highest (p < 0.0001 for all comparisons). Subsequently, to quantify the performance of each base editor in efficiently and precisely installing the m.3242G>A variant, we calculated normalized on-target-to-bystander editing ratios (Fig. 5E), which are further informed by the off-target activities of each pair (Fig. 5C). Notably, TL1 αDdCBE 3CN outperformed all other pairs (p < 0.0001 for all comparisons), followed by TL1 αLDdCBE 3 (p < 0.001 for all comparisons).
In addition, we calculated normalized on-target-to-off-target editing ratios (Supplementary Fig. S6A). However, as bystander editing events are disregarded using this specific metric, these analyses do not fully reflect the performance of each base editor in precisely installing the m.3242G>A variant. Alternatively, we calculated normalized on-target-to-unintended (i.e., bystander and off-target) editing ratios (Supplementary Fig. S6B). However, when compared with the normalized on-target-to-bystander editing ratios (Fig. 5E), the ratios in Supplementary Figure S6B overstate base editor performance. Thus, for applications in which a specific point mutation is desired, on-target-to-bystander editing ratios along with amplicon-wide specificity data can facilitate base editor selection.
In the context of DddA-derived base editors, these results collectively demonstrate that TALE shifting, that is, utilizing different sets of TALEs to access a single target site, can result in enhanced editing outcomes, both in terms of activity and cytosine selectivity within the spacer regions.
DISCUSSION
In this study, we formally developed αDdCBEs for unconstrained mitochondrial base editing, which displayed improved performance compared with canonical DdCBEs across different target sites in human mtDNA in vitro. In addition, we demonstrated that αDdCBEs are compatible with DddAtox and its engineered variants, DddA6 and DddA11. Furthermore, we validated TALE shifting with αDdCBEs as an approach to fine-tune base editing outcomes. This method enabled the definition of diverse spacers at a single target site, regardless of the most 5′ nucleotides available to the TALEs. In terms of practical applications, TALE shifting can be leveraged to modulate bystander editing.
Importantly, the relevance of the 5′-T constraint in the context of mitochondrially targeted, TALE-guided deaminases remained uncharacterized. 13,16,30 –36 By functionally exploring this design rule, we found that it often acts as a moderate limiting factor for efficient mtDNA editing with DdCBEs, rather than as an obligate requirement. In fact, several 5′-T-noncompliant DdCBEs efficiently edited their target sites, although occasionally at moderately decreased levels of on-target activity or increased frequencies of proximal off-target editing compared with αDdCBEs, which often outperformed their canonical counterparts (Supplementary Table S5).
In contrast to our findings, 5′-T compliance for optimal TALE activity in the nuclei of plant and animal cells is broadly supported by the frequent presence of thymine preceding the targets of natural TALE proteins, and by the standard guidelines for nuclear-targeted TALE-based technologies. 23 –27,37 –47 These platforms include TALE scaffolds specifically engineered to circumvent the 5′-T constraint through amino acid substitutions in the TALE NTD. 52,75 –80 However, TALE sensitivity to the 5′-T constraint is influenced not only by the interaction between the canonical TALE NTD and a 5′-T on the target DNA, 39,52,76,79 but also by the number and composition of repeats in the TALE protein. 44,77,81,82
In particular, in an in vitro reporter assay, TALEs containing between 9.5 and 13.5 repeats showed a strong dependence on the 5′-T rule, while TALEs with 17.5 repeats were, in comparison, less influenced by this constraint. 77 It should be noted that FusX-based technologies, including the DdCBEs in this study, are currently limited to TALEs with 14.5, 15.5, or 16.5 repeats, 49 –51 lengths associated with optimal TALE binding affinity in cell-free assays. 82 Thus, given the number of repeats in the TALEs within the DdCBEs tested here, we speculate that the observed flexibility in the 5′-T rule might be partially explained by the TALE-length-dependent stringency of this requirement. 77
Of note, the number of repeats in reported TALE-based technologies for mtDNA editing ranges from 8.5 to 19.5 repeats. 13,32 –36,48,50,53,59,65,83,84 Despite this variability in TALE length across systems, the effects of repeat number on the performance of TALE-based mitochondrial base editors remain unclear. Notably, our findings suggest that αDdCBEs containing TALEs with 14.5 or 15.5 repeats, particularly in 5′-T-noncompliant formats, can be moderately more active and specific than their canonical counterparts, suggesting an enhanced design flexibility under these conditions. In contrast, some αDdCBEs containing TALEs with 16.5 repeats resulted in less on-target activity than canonical DdCBEs.
Given these observations, it might be tempting to attempt to reduce the stringency of the 5′-T constraint in canonical DdCBEs by utilizing long TALEs (≥16.5 repeats). However, this potential strategy is yet to be thoroughly evaluated. Furthermore, each additional repeat in a single TALE protein introduces 34 amino acids (102 bp) to its sequence. 37,38 For gene therapy applications, these increments in transgene size could hinder packaging into some delivery vectors.
Overall, we recommend the following approach for the use of αDdCBEs. First, target sites where αDdCBEs can be more applicable than DdCBEs can be identified by verifying whether 5′-T-compliant DdCBEs can be designed, based on standard design guidelines. 13,53 Accordingly, we suggest using αDdCBEs if the target contains only one or no binding sites for 5′-T-compliant TALEs, as ignoring the 5′-T constraint can result in loss of performance. In detail, for αDdCBE design, we recommend spacers between 11 and 18 bp long, and TALEs with 14.5 or 15.5 repeats, irrespective of the most 5′ nucleotides of their targets. In addition, users may test αDdCBEs with short TALEs (8.5 to 13.5 repeats).
Of note, we suggest careful consideration to spacer composition, which can be modulated via TALE shifting with αDdCBEs. In general, narrow spacers can be defined to limit the number of bystander cytosines around a specific target nucleotide. Furthermore, αDdCBEs might facilitate testing of additional spacers at sites where only a limited set of 5′-T-compliant DdCBEs can be designed. Regarding deaminase domain selection, our work is consistent with the previous recommendations by Mok et al. 53 Finally, we hypothesize that other effector domains, such as DddA homologs or chimeric deaminases, 16,32 –36 will also work well with unconstrained TALEs.
Study limitations
In this study, αDdCBEs were developed using immortalized human cells in vitro. Further characterizations in clinically relevant cell types and in vivo models are needed to continue to validate our observations. Moreover, comparisons between DdCBEs and αDdCBEs were conducted at a limited number of sites, and although most measurements were obtained via NGS, some base editing activities were measured using Sanger sequencing, from which low-frequency variants cannot be detected. Likewise, although amplicon-wide analyses are highly informative, 29 genome-wide surveys will enhance our understanding of the overall specificity of αDdCBEs.
CONCLUSIONS
We determined that the TALE 5′-T constraint for mitochondrial base editing with DdCBEs can act as a moderate limiting factor, rather than a definitive requirement. For cases in which this design rule hampers canonical DdCBE performance, αDdCBEs can be more effective, particularly with 5′-T-noncompliant TALEs containing 14.5 or 15.5 repeats. In general, αDdCBEs provide an alternative for testing and can display increased on-target activity or reduced off-target editing relative to canonical DdCBEs.
Footnotes
ACKNOWLEDGMENTS
We thank Dr. David Liu and Dr. Beverly Mok (Broad Institute of MIT and Harvard) for kindly providing various DdCBE plasmids, as detailed in Supplementary Table S1.
AUTHORS’ CONTRIBUTIONS
S.R.C.: Conceptualization, methodology, software, validation, formal analysis, investigation, data curation, writing—original draft, writing—review and editing, visualization, project administration, and funding acquisition. B.W.S.: Methodology and writing—review and editing. K.J.C.: Writing—review and editing, and funding acquisition. P.D.: Resources, writing—review and editing, and supervision. S.C.E.: Resources, writing—review and editing, supervision, project administration, and funding acquisition.
DATA AVAILABILITY STATEMENT
DdCBE and αDdCBE plasmids used in this study will be made available through Addgene. The FusX system for TALE assembly is available on Addgene (Kit # 1000000063). The general architecture of the base editors used in this study is detailed in the Supplementary Sequences. Main source data are provided as Supplementary Material. Raw chromatogram files (.ab1) from Sanger sequencing have been deposited in a repository publicly available at ![]() Raw paired-end read FASTQ files from high-throughput sequencing have been deposited in the NCBI Sequence Read Archive under accession code PRJNA1111950.
Raw paired-end read FASTQ files from high-throughput sequencing have been deposited in the NCBI Sequence Read Archive under accession code PRJNA1111950.
PREPRINT DISCLOSURE
AUTHOR DISCLOSURE
The Mayo Foundation for Medical Education and Research is the current assignee for a patent on “Methods and Materials for Assembling Nucleic Acid Constructs” (US20180002707A1), which includes the FusX TALE assembly system used in this study.
FUNDING INFORMATION
This work was supported by an NIH grant 1U01AI142773-01 (S.C.E. and K.J.C.), NIH grant 1R01063904 (S.C.E.), the Mayo Foundation for Medical Education and Research (S.R.C.), the 2022 ASCB International Training Scholarship Program (S.R.C.), the 2021 and 2023 Gateway to Mitochondrial Medicine Grants from the United Mitochondrial Disease Foundation and the North American Mitochondrial Disease Consortium (S.R.C.), the 2020 Mayo Clinic Department of Molecular Medicine Small Grant (S.R.C), and the Harry C. and Debra A. Stonecipher Predoctoral Fellowship (S.R.C.).
SUPPLEMENTARY MATERIAL
Supplementary Figure S1
Supplementary Figure S2
Supplementary Figure S3
Supplementary Figure S4
Supplementary Figure S5
Supplementary Figure S6
Supplementary Table S1
Supplementary Table S2
Supplementary Table S3
Supplementary Table S4
Supplementary Table S5
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
