Abstract
The use of a helper plasmid to replace adenovirus infection for adeno-associated virus (AAV) manufacturing has been common practice for decades. Adenovirus E4, E2a, and VA RNA genes are sufficient to support efficient AAV replication. In an effort to ensure that all transfected DNA has a functional role in AAV production, deletions were introduced to the E4 and E2a genes to determine if any portions were dispensable. Although a 900 bp deletion in the E2a intron did not have an impact, the removal of open reading frames (orf) 1–4 from the E4 gene resulted in a doubling of AAV productivity. The E4Δorf1-4 deletion was associated with a reduction in E4orf6 transcripts, along with an increase in Rep and Cap transcripts and protein levels, which corresponded to increased AAV productivity in crude lysate. The final product of these studies was a helper plasmid, termed OXB-Helper_3, that is >3.4 kb smaller than the original control plasmid and resulted in ∼2× improvement in vector genome productivity across multiple capsid serotypes, genome designs, and transfection platforms.
INTRODUCTION
The wild-type AAV genome is ∼4.7 kb, encoding only two genes, Rep and Cap, each of which produces a number of proteins. 1,2 However, AAV requires additional support from a helper virus in order to complete its replication cycle. 3 Various large DNA viruses, including herpes simplex virus and baculovirus, can serve as a helper; however, adenovirus is most commonly used in recombinant AAV manufacture. 4 The use of replication-competent adenovirus has generated some concern for the safety of AAV therapies as it was difficult to ensure total removal of the helper virus from the final product. 5 The vast majority of the field has replaced live helper virus with a helper plasmid containing the necessary adenovirus genes to enable AAV replication. A plasmid expressing E4, E2a, and VA RNA genes was determined to be sufficient to replace adenovirus infection. 6,7 E1a and E1b are also required helper genes for AAV production 8 ; however, they are usually supplied through the use of HEK293 cells, which have been transformed using these adenoviral sequences, and as such, E1a/b are not present on helper plasmids. 9
The E4 gene encodes multiple distinct polypeptides expressed early during adenovirus infection. E4 open reading frame (orf) 1, E4orf1, is involved in adenovirus oncogenesis, whereas E4orf2 has been detected in infected cells, but its role is unknown. 10,11 The functions of E4orf3 and E4orf6 are redundant, and E4orf4 plays a noncritical role in regulating protein phosphorylation. 12,13 E4orf6/7 binds the cellular transcription factor E2F and dimerizes at a pair of E2F binding sites in the E2a early promoter causing transactivation. 14,15 The roles of each E4 orf have been reviewed in detail; however, the only one with clear requirement for AAV production is E4orf6. 16,17 E4orf6 generates a 34 kDa protein that forms a complex with E1b-55K to perform a variety of functions. The E4orf6/E1b-55K complex enhances viral DNA replication by preventing concatemer formation during synthesis through the disruption of the MRN complex and the DNA-damage response. 18,19 This complex is responsible for the accumulation of viral mRNA in the cytoplasm as well as the shutdown of host cell transcript export across the nucleus. 20,21 In addition, E4orf6 binds p53 and prevents the cell from completing apoptosis. 22 The E2a gene contains two promoters, an early and a late, that temporally regulate expression of the 72 kDa DNA-binding protein (DBP). DBP binds to single stranded DNA in a sequence independent manner and functions to increase the processivity of AAV DNA replication by stimulating elongation and stabilizing the displaced ssDNA from nuclease degradation. 23 –25 In addition, DBP has regulatory roles in enhancing AAV p5 promoter activity as well as suppression of adenovirus E4 promoter during later stages of the infection. 26 DBP is involved in the establishment and maturation of viral replication compartments (RC) in the nucleus and has been shown to interact with Rep proteins within these RCs. 27,28 Adenovirus expresses two highly structured, noncoding virus-associated RNAs (VA RNAs) that play an important role in regulating viral protein synthesis. Protein kinase R (PKR) normally inhibits global protein translation as an anti-viral defense; however, VA RNA binds PKR and prevents this translation shutdown, allowing expression of viral proteins to continue. 29,30
There are a number of commercially available adenovirus-based helper plasmids that vary in size and specific sequence, but all minimally contain E4, E2a, and VA RNA genes. The entire E2a gene spans ∼5.3 kb, yet the E2a promoter and DBP coding sequence account for a significantly smaller portion of this. It remains unclear whether the presence of large introns is essential for appropriate DBP expression. 31 In addition, it has previously been suggested that E4orf6 is sufficient for AAV production, 32 potentially allowing for removal of approximately half of the E4 gene. A minimized helper plasmid has the potential to improve plasmid manufacturability while ensuring all transfected DNA has a functional role in AAV production. In this study, we sought to determine if any of the current helper plasmid sequence is “junk” in the context of AAV manufacturing.
METHODS
Plasmids
All experiments utilized an Oxford Biomedica (OXB) dual plasmid, 33 containing a genome expressing GFP and luciferase through the use of an internal ribosomal entry site (IRES) (G1) and the AAV9 capsid sequence unless otherwise noted in the figure legend. Details about the four additional genomes (G2–G5) evaluated can be found in Supplementary Table S1. The helper plasmid sequences are based on adenovirus type 2 (National Center for Biotechnology Information accession J01917.1), and the specific coordinates of each gene, the deletions and plasmid sizes, can be found in Table 1 and Supplementary Table S2. The individual helper gene plasmids and all deletion plasmids were generated using site directed mutagenesis. All plasmids were produced at industry grade with GenScript Biotech.
Description of Individual Helper Gene Plasmids and the Evaluated E4 and E2a Deletions
VA RNA, virus-associated RNA.
AAV production
VPC 2.0 cells (Thermo Fisher) were cultured in 125 mL shake flasks with a 50 mL working volume in Expi293 media (Thermo Fisher). Cells were transfected at 2E6 viable cells/mL with 0.75 µg of total DNA per 1E6 cells. Polyethyleneimine (PEI, Polyscience) was used as the transfection reagent at a 1:1.5 ratio PEI: DNA. All transfections used equal molar plasmid ratios unless otherwise specified in the figure legend. Seventy-two hours post-transfections, cells were lysed using a chemical lysis buffer and crude lysate was centrifuged at 3,000 g for 10 min to pellet cell debris.
Crude lysate VG and capsid productivity
Clarified crude lysate was used to determine VG and capsid productivity. VG productivity was determined by standard droplet digital PCR (ddPCR) as previously outlined. 33 Capsid productivity was determined using a commercially available enzyme-linked immunosorbent assay (ELISA) kit for AAV2, AAV5, AAV6, AAV8, or AAV9 (PROGEN). The full capsid values from crude lysate were calculated by dividing the VG titer by the capsid ELISA titer.
Western blot analysis
Clarified crude lysate was mixed with a loading dye, and 10 µg of total protein was loaded onto a precast gel with a protein ladder. The gel was transferred onto a nitrocellulose membrane using the iBlot transfer system. The membrane was incubated in blocking buffer (LI-COR BioSciences) before incubation with primary antibody overnight at 4°C. The B1 anti-Cap primary antibody (American Research Product) was used at a 1:1,000 dilution, and 303.9 anti-Rep (American Research Product) was used at a 1:250 dilution in blocking buffer. A polyclonal antibody was raised against DBP by immunizing New Zealand rabbits with the full-length DBP recombinant protein, and the resulting anti-sera were purified by affinity resin (GenScript Biotech). The anti-DBP polyclonal was used at a 1:20,000 dilution. The membrane was incubated at a 1:10,000 in anti-mouse or anti-rabbit secondary antibody (LI-COR Biosciences), before being imaged on the Odyssey CLx. REVERT total protein stain (LI-COR Biosciences) was applied to the same gel to ensure equal loading.
Reverse transcription quantitative PCR analysis
Cell pellets were collected prior to lysis and RNA was extracted using RNeasy plus mini kit (QIAGEN). Samples were run on Qubit to ensure adequate RNA quality (RIN > 8.0). cDNA was generated using superscript IV first-strand synthesis system (Invitrogen). cDNA was quantified by real-time quantitative PCR using TaqMan Universal PCR master mix (Thermo Fisher) with primer and probe sets listed in Supplementary Data S1. Relative expression levels were determined with the ΔΔCT threshold cycle (CT) quantification method, 34 using GAPDH as a housekeeping reference gene.
RNA sequencing and data processing
RNA sequencing and data processing are described in Supplementary Data S1.
Flow cytometry
Flow cytometry is described in Supplementary Data S1.
Residual RepCap quantification
Residual RepCap quantification is described in Supplementary Data S1.
RESULTS
E4, E2a, and VA RNA genes are all required for optimal AAV production
To enhance the efficiency of manipulating the helper plasmid, each helper gene was isolated onto its own individual plasmid. Combinations of the individual E4, E2a, and VA RNA plasmids were compared to the full-sized control helper to assess productivity. VG and capsid productivity from the three individual plasmids were equivalent or slightly elevated compared to the control helper plasmid (Fig. 1). The individual helper plasmids also allowed for confirmation that all three genes were required for maximum productivity. Transfection with no E4 plasmid resulted in 50× decrease in VG productivity but only a 4× decrease in the capsids produced compared with the individual helper gene control (Fig. 1). Without E2a, productivity decreased by 144× and 31× for VGs and capsids respectively, demonstrating the largest impact of the three helper genes (Fig. 1). Without VA RNA, there was a relatively minor decrease in both VG and capsid productivity compared with removal of E4 or E2a; VGs decreased by 3× and capsids decreased by 6× (Fig. 1). Although VA RNA removal was less significant than the others, there is still an advantage to inclusion of all three helper genes for optimal productivity.
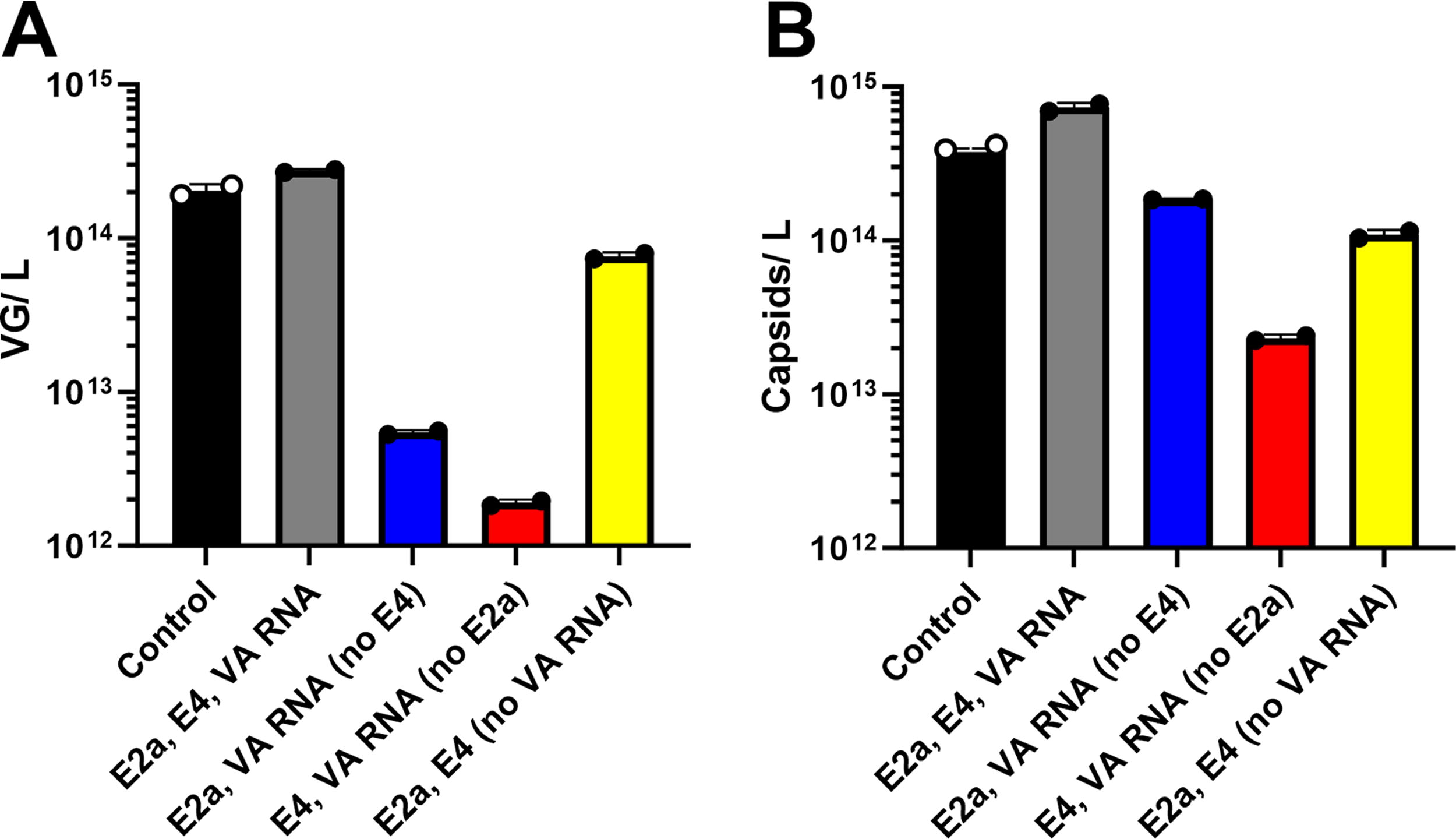
Productivity assessment of a full-sized control helper and individual helper gene plasmids. Cells were transfected with an OXB dual plasmid and helper plasmid(s) at equal molar ratios. Crude lysate samples were quantified for
Deletions within the E4 and E2a genes are feasible without decreasing AAV productivity
The individual helper plasmid system was used to introduce deletions into the E4 or E2a genes and screened for comparable productivity to the control helper plasmid. VA RNA is the smallest of the three genes at 731 bp; therefore, we focused our efforts on removing portions of the E4 and E2a genes (Table 1). For E4, three deletion constructs were generated: the first removed orf1-3 (E4Δorf1-3), the second removed orf1-4 (E4Δorf1-4), and the third removed orf1-4 as well as the remainder of the 3′ sequence following the orf6 stop codon (E4orf6 only) (Fig. 2A, Table 1). All three E4 deletion constructs produced equivalent or increased VG and capsid productivity compared with the control helper plasmid (Fig. 2B, Supplementary Fig. S1A). This suggests that E4orf1-4 are not critical for AAV production. The combination of largest deletion with the highest productivity led to the selection of E4Δorf1-4 for further development.
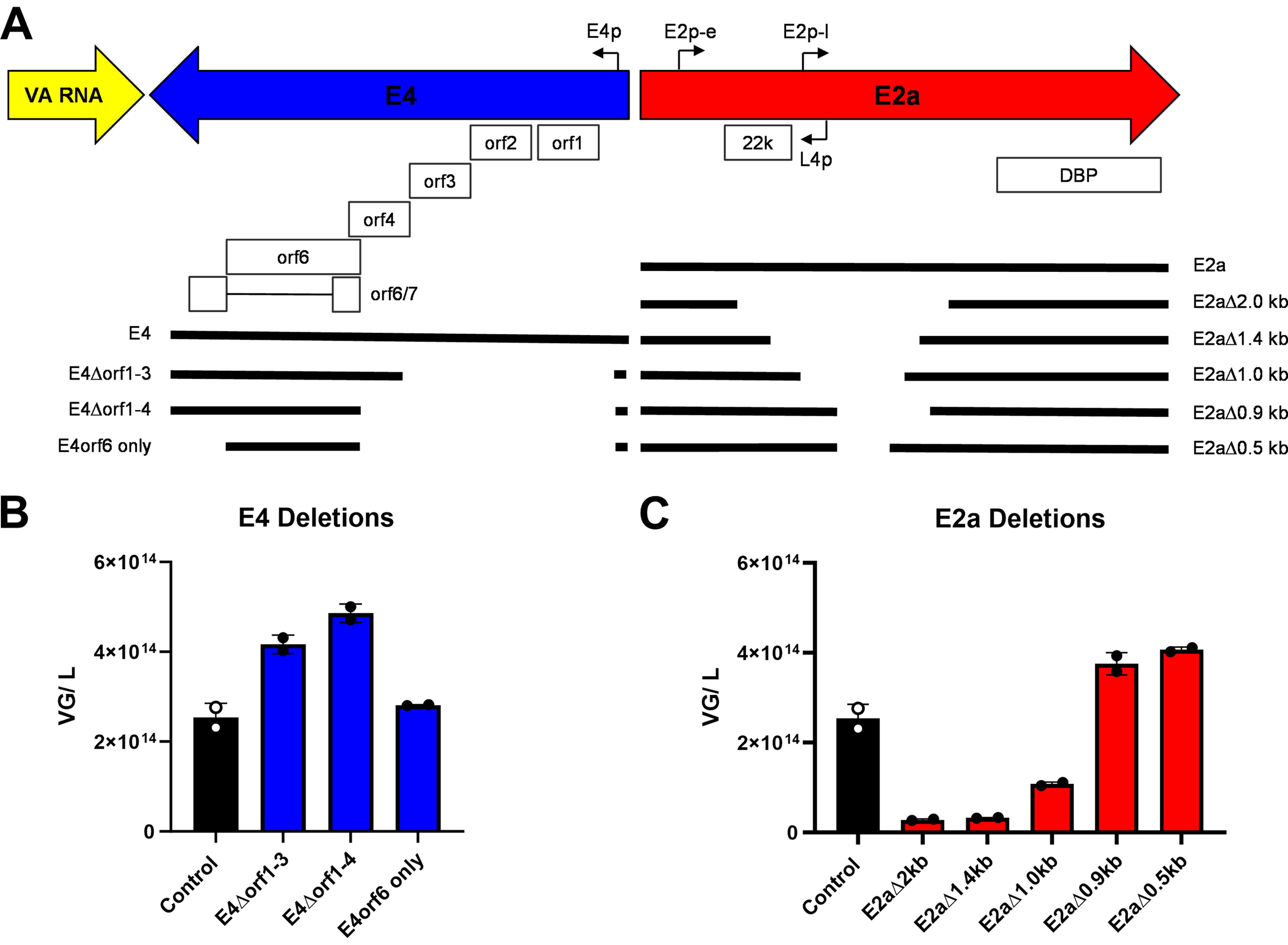
Deletions in E2a and E4 helper genes can result in equivalent VG productivity as the intact genes.
Introducing a productive deletion into the E2a gene took two rounds. The first round included a 2.0 kb, 1.4 kb, and 1.0 kb deletion between the E2a early promoter and the DBP start codon (Fig. 2A). None of these constructs were as productive as the control helper plasmid, indicating that the deletions negatively impacted DBP expression or removed some other required element (Fig. 2C, Supplementary Fig. S1B). The second round of E2a deletions included a 0.9 kb and a 0.5 kb deletion that each keeps the E2a late promoter intact. Both constructs resulted in VG and capsid productivity that was greater than the control helper plasmid (Fig. 2C, Supplementary Fig. S1B), suggesting that inclusion of the E2a late promoter may be required for adequate DBP expression. E2aΔ0.9 kb is the largest deletion with the highest productivity and was selected for further development along with E4Δorf1-4.
Combination of individual deletions into full-sized helper plasmids
The E4Δorf1-4 deletion was introduced into the control helper plasmid, termed OXB-Helper_1, and the E2aΔ0.9 kb deletion was introduced separately into the control helper termed, OXB-Helper_2. A third full sized helper plasmid containing both E4Δorf1-4 and E2aΔ0.9 kb deletions, termed OXB-Helper_3, was also generated. In addition, all OXB helper plasmids contain a 1.0 kb deletion from the backbone of the plasmid, which did not alter the productivity (Supplementary Fig. S2). OXB-Helper_3 is the smallest of the helper constructs at ∼8.6 kb, which is over 3.4 kb smaller than the control helper plasmid (Supplementary Table S2).
Interestingly, OXB-Helper_1 and OXB-Helper_3 both contain the E4Δorf1-4 deletion, and both produced VG titers ∼2-fold higher than the control (Fig. 3A). We previously saw this trend when testing E4 deletions with the individual gene plasmids (Fig. 2A) but were unsure if that productivity increase would translate in the context of a consolidated helper or if plasmid input quantities were responsible. This increase in VG and capsid productivity indicates that the E4Δorf1-4 deletion either removes some inhibitory factor of orf1-4 or causes an alteration in orf6 expression that is beneficial for vector production (Fig. 3A, B). Conversely, OXB-Helper_2 was comparable with the control plasmid, which suggests that the 0.9 kb E2a deletion does not alter the functionality of DBP expression. As the E2a deletion is innocuous and the productivity of OXB-Helper_1 and OXB-Helper_3 is equivalent, it would appear that the E4 deletion is the driver of the productivity increase. The percentage of calculated full capsids in the crude lysate was relatively similar across all helper plasmids; however, there was a minor decrease in both E4Δorf1-4 bearing helpers in this experiment (Fig. 3C).
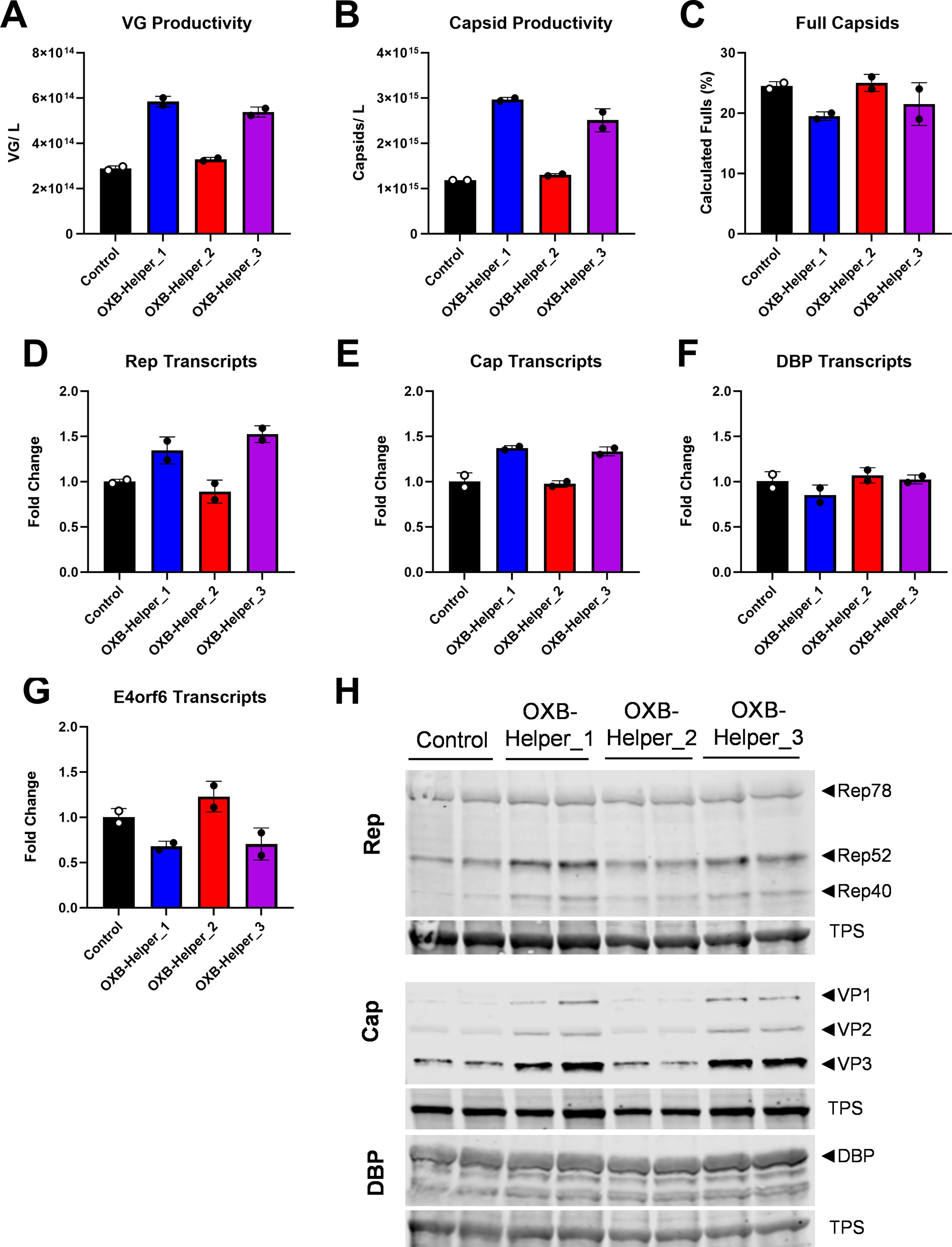
Evaluation of full-sized helper plasmids containing a deletion in E4 and/or E2a. Cells were transfected with the control helper, OXB-Helper_1, OXB-Helper_2, or OXB-Helper_3 and a dual plasmid at equal molar ratio and crude lysate was quantified for
Analysis of transcripts by reverse transcription quantitative PCR (RT-qPCR) demonstrates an increase in Rep and Cap transcripts for OXB-Helper_1 and OXB-Helper_3 that corresponds with the increase in VG and capsid productivity for these constructs (Fig. 3D, E). DBP transcripts were relatively unchanged across each of the four helper plasmids (Fig. 3F). Protein analysis by western blot supported the transcript trends with increased Rep, specifically Rep52, and Cap expression for OXB-Helper_1 and OXB-Helper_3 and consistent DBP expression for all constructs (Fig. 3H). OXB-Helper_2 transcripts were comparable to the control helper for all assayed genes. Protein expression for Rep, Cap, and DBP all aligned with the transcript levels. Unfortunately, our attempt to generate an E4orf6 polyclonal antibody was not successful so we were unable to assay protein levels, however E4orf6 transcripts decreased in OXB-Helper_1 and OXB-Helper_3, which both contain the E4Δorf1-4 deletion (Fig. 3G). As OXB-Helper_1 and OXB-Helper_3 performed similarly in terms of productivity and showed similar transcript and protein outputs, we selected the smaller of the two, OXB-Helper_3, to proceed with additional characterization.
The previous experiment compared all helper plasmids at equal molar plasmid ratios; however this may not be the optimal transfection condition. We tested 2:1, 1:1, 1:2, and 1:3 (dual:helper) molar plasmid ratios for the control helper and OXB-Helper_3. VG productivity increased for plasmid ratios favoring more of either helper but plateauing between 1:2 and 1:3 (Fig. 4A). The capsid productivity for OXB-Helper_3 also followed the VG trends, but the control helper produced less capsids as the helper ratio increased (Fig. 4B). Full capsids in the crude lysate were comparable between the control helper and OXB-Helper_3 with a slight trend toward lower values particularly at the ratios favoring more helper plasmid (Fig. 4C). Quantification of DBP and E4orf6 transcripts demonstrated a dose response linked to the amount of input helper plasmid for both the control and OXB-Helper_3 (Fig. 4D, 4E). There was a substantial reduction in E4orf6 transcripts from OXB-Helper_3 compared with the ratio-matched control that is apparent in all ratios and was consistent with the previous study. RNA sequencing of the 1:1 samples showed a slight decrease in the DBP transcripts and a substantial decrease in the E4orf6 transcripts for OXB-Helper_3 compared with the control helper (Supplementary Fig. S3). These findings align with the RT-qPCR results, confirming consistent transcript levels across both methodologies. Western blot analysis showed a slight increase in Rep expression for the OXB-Helper_3 conditions and a more obvious increase in Cap expression that supports that capsid ELISA data (Fig. 4F). The DBP western also shows a dose response for both the control and OXB-Helper_3 that supports the RT-qPCR data (Fig. 4D). Lastly, to confirm that differences in plasmid input due to the discrepancy in the size of the helper plasmids was not a contributing factor, we determined the proportion of cells expressing GFP from the dual plasmid genome as a measure of transfection efficiency. All conditions resulted in ∼60–70% GFP positive cells, and there was no difference between the two helpers despite the size variance (Supplementary Fig. S4).
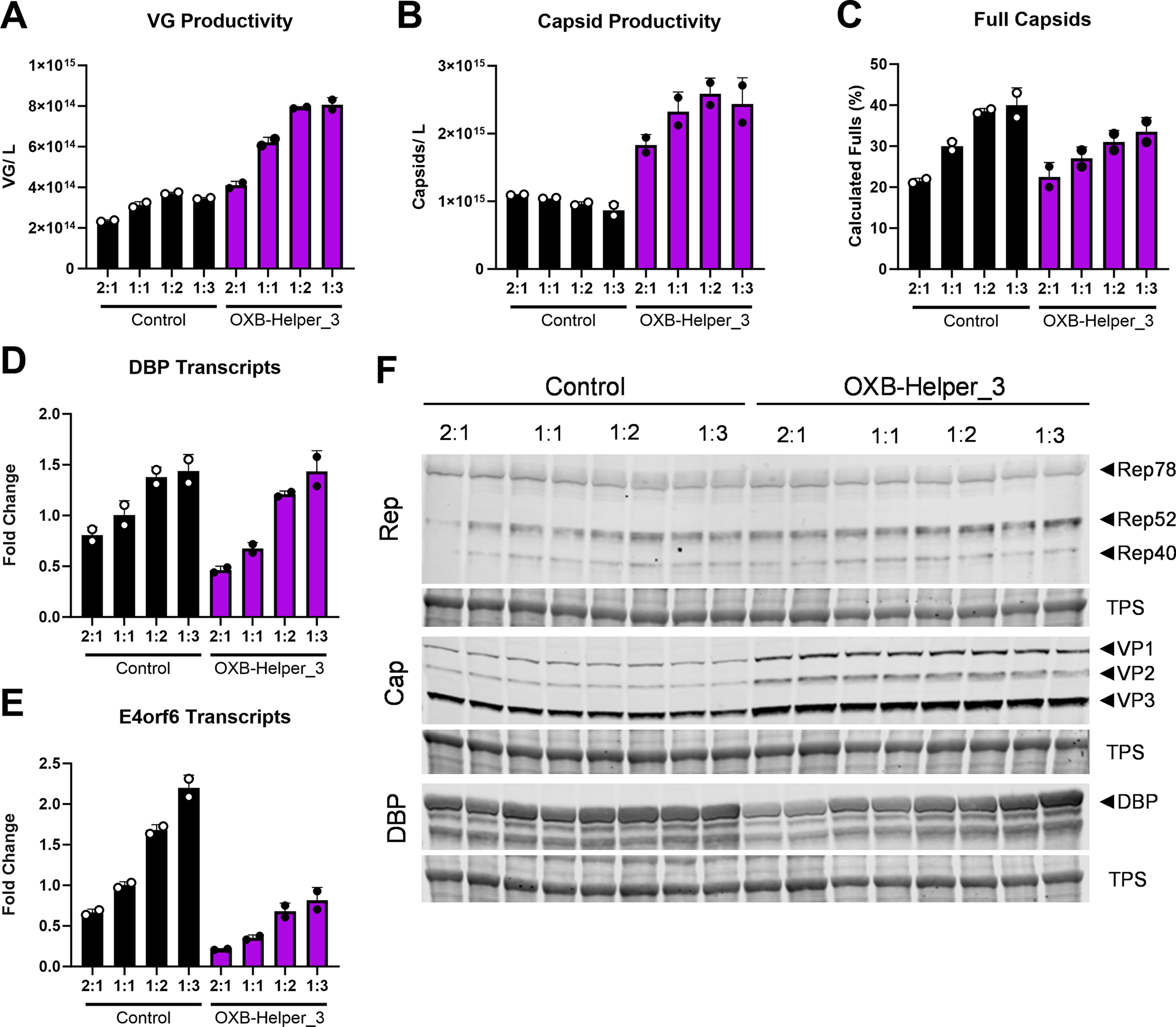
Comparison of plasmid ratios for the control helper and OXB-Helper_3. Cells were transfected with 2:1, 1:1, 1:2, 1:3 molar plasmid ratios (dual:helper) and crude lysate was quantified for
We have previously demonstrated the pOXB dual design does not increase residual RepCap or the presence of replication competent AAV despite the genome and RepCap sequences being located in cis on a single plasmid. However, to address concerns that OXB-Helper_3 could provide an additional risk compared with the control helper plasmid, purified vectors were assayed for residual RepCap by ddPCR (Supplementary Table S3). Vectors produced with either helper plasmid fell within the range of our historical data from over 40 individual lots.
OXB-Helper_3 improves productivity for various capsid serotypes and genome designs
To ensure the productivity increase demonstrated with OXB-Helper_3 was not specific to AAV9 or the GFP-IRES-Luciferase genome used in previous studies, we tested four additional capsid serotypes and genome designs. AAV2, AAV5, AAV6, and AAV8 capsids were evaluated with the GFP-IRES-Luciferase genome, and VG productivity increased by 1.5–3.5× using OXB-Helper_3 depending on serotype (Fig. 5A). This corroborates the ∼2× increase demonstrated previously with AAV9 and suggests that the improvement in productivity using OXB-Helper_3 is not serotype specific. Additional genomes ranging in size, promoter, and transgene were tested in a triple transfection platform to confirm the performance of OXB-Helper_3 is broadly applicable (Supplementary Table S1). The use of triple transfection for this experiment allowed evaluating whether the productivity improvement of OXB-Helper_3 was specific to the OXB dual design. OXB-Helper_3 increased the VG productivity by 1.4–2.0× across these four genome designs, suggesting all genomes will see a benefit from the use of OXB-Helper_3 but perhaps to different extents depending on the specific design (Fig. 5B). Capsid productivity trended with VG productivity as expected (Fig. 5C, D).
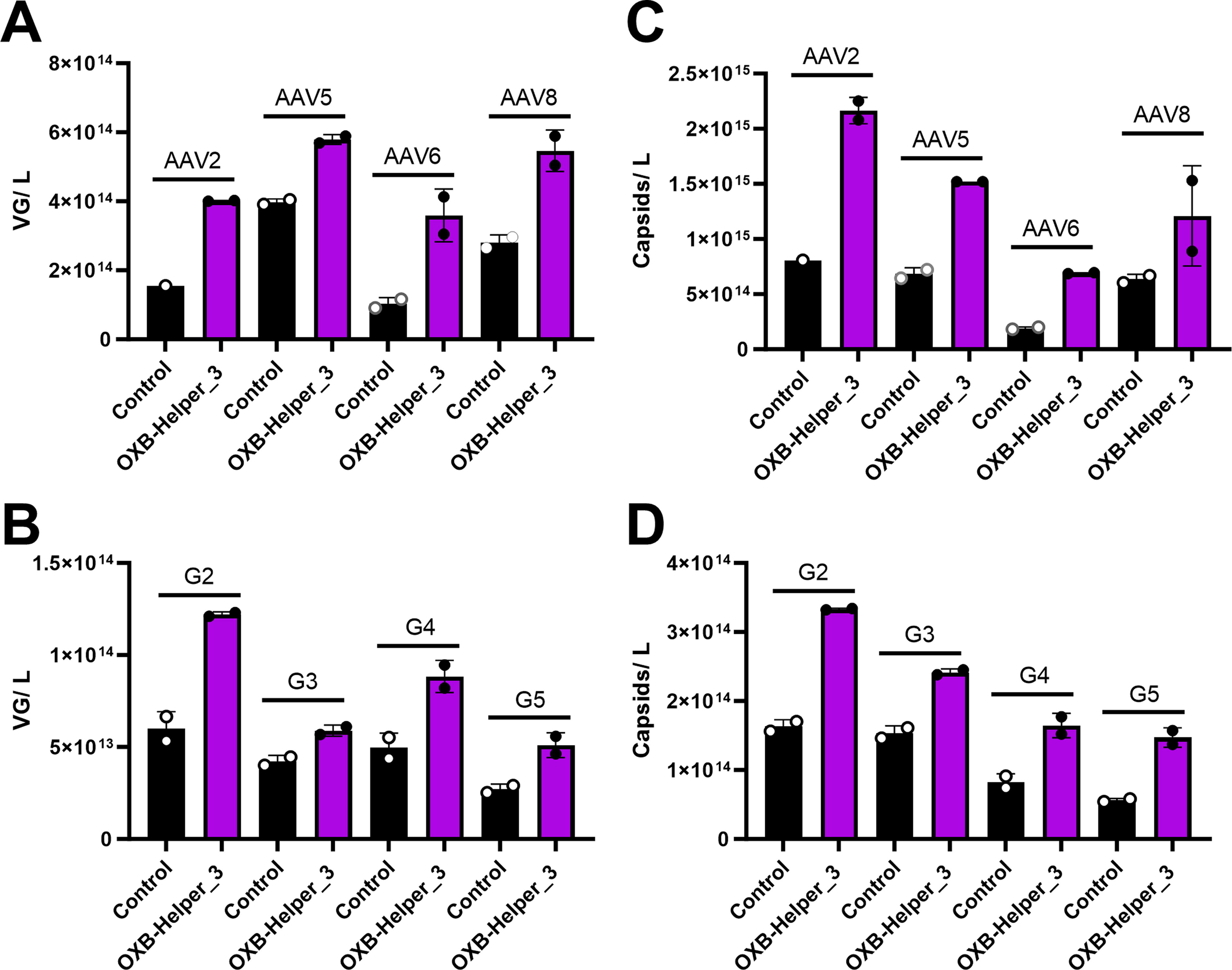
OXB-Helper_3 VG and capsid productivity evaluation with additional capsid serotypes and genome designs.
DISCUSSION
The aim of this work was to investigate the E4 and E2a helper genes to identify whether full-length sequences are required for their effective helper function. In the event that portions of these genes were not required, the result could be a minimized helper plasmid where all transfected DNA played a functional role with respect to AAV production. We were able to remove 0.9 kb from E2a and 2.0 kb from E4 (E4orf6 only) without lowering VG titers (Fig. 2). Unexpectedly, a slightly smaller E4 deletion (E4Δorf1-4) resulted in increased VG and capsid production and was associated with a reduction in E4orf6 transcripts (Fig. 3G, 4E). Liu et al. previously found the removal of E4orf1-4 from adenovirus type 5 also increased VG productivity, which supports our findings in adenovirus type 2. 35 This suggests that the endogenous helper expression from the native adenovirus genes may not be optimal for AAV and merits further development to determine the ideal conditions for maximum AAV replication and packaging.
It became clear during the E2a deletion experiments that part of the intronic region was required for successful AAV production. At the time, we surmised this was related to the removal of the E2a late promoter altering DBP expression. However, since then we have demonstrated that unexpectedly DBP protein expression is unchanged in all the individual E2a deletions plasmids, despite the VG and capsid productivity being severely impaired with some constructs (Fig. 2C, Supplementary Fig. S5). Recently, there have been two independent publications demonstrating L4 22k is a required factor for AAV production. 36,37 Armed with this information, our data can be interpreted to support the same conclusion, in that it was not deletion of the E2a late promoter that was problematic but rather the removal of the L4 22k region that happens to be located in the E2a intron in close proximity to that promoter. Both of the successful E2a deletions (Δ0.9 kb and Δ0.5 kb) left L4 22k intact.
OXB-Helper_3 produces fewer E4orf6 transcripts; however, this does not alone explain the titer improvement. In the control helper and OXB-Helper_3 ratio experiment, the control at a 2:1 ratio is the overall lowest producer, but the E4orf6 transcript levels are similar to OXB-Helper_3 at 1:3, which is the top producing condition (Fig. 4A, E). DBP transcripts and protein output in this experiment were either similar or slightly reduced in the OXB-Helper_3 conditions compared with the control at the same plasmid ratios (Fig. 4D, F). Perhaps even minimal reductions in DBP expression are beneficial or more likely there is another unassayed factor at play, such as L4 22k.
In terms of product quality, calculated full capsids in the crude lysate suggest similar packaging performance between the control helper and OXB-Helper_3. This calculation is useful for general comparison; however, as it compounds potential variability from both the ddPCR and ELISA assays, it can be difficult to pinpoint minor changes. Future analysis of purified vector would be useful to understand if changes in E4orf6 expression have an impact on vector quality.
Helper plasmids have been relatively untouched since their inception, but recently, there has been active development to improve these plasmids as we push the frontiers of AAV manufacturing forward. Our data suggest that there is additional work to be done to modify the endogenous expression to be optimal for AAV production rather than relying directly on what has evolved for the adenovirus genome. Furthermore, our data support recent publications indicating that there is, in fact, a fourth adenovirus gene required for AAV production: E4, E2a, VA RNA, and L4 22k. This discovery comes over 25 years after a helper plasmid was first introduced, clearly demonstrating further characterization of helper functions for AAV manufacturing is warranted.
Footnotes
ACKNOWLEDGMENT
The authors thank all those who contributed to the analytical data presented in this article.
AUTHORS’ CONTRIBUTIONS
L.V.L.: Conceptualization, methodology, validation, writing—original draft, visualization, and supervision; S.O.: Investigation, formal analysis, and visualization; A.A.: Investigation and formal analysis; E.W.: Investigation and validation; K.C.-G.: Investigation and formal analysis; D.L.: Investigation, formal analysis, and writing—original draft; S.D.: Project administration and writing—reviewing and editing; M.S.: Project administration; I.I.: Project administration; D.G.: Conceptualization, writing—reviewing and editing, and supervision; J.Y.: Resources, writing—reviewing and editing, and project administration
AUTHOR DISCLOSURE STATEMENT
All authors are employed by Oxford Biomedica. L.V.L. and D.G. are inventors of a patent application related to this work.
FUNDING INFORMATION
This work was funded by Oxford Biomedica (US) LLC.
SUPPLEMENTARY MATERIAL
Supplementary Data S1
Supplementary Figure S1
Supplementary Figure S2
Supplementary Figure S3
Supplementary Figure S4
Supplementary Figure S5
Supplementary Table S1
Supplementary Table S2
Supplementary Table S3
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
