Abstract
A prototype strain of Coxsackievirus A21 (CVA21) is being evaluated as an oncolytic virus immunotherapy. CVA21 preferentially lyses cells that upregulate the expression of intercellular adhesion molecule 1, which includes some types of tumor cells. CVA21 has an icosahedral capsid structure made up of 60 protein subunits encapsidating a viral RNA genome with a particle diameter size of 30 nm. Rapid and robust analytical methods to quantify CVA21 total, empty, and full virus particles are important to support the process development, meet regulatory requirements, and validate manufacturing processes. In this study, we demonstrate the detection of all four CVA21 capsid proteins, VP1, VP2, VP3, and VP4, as well as VP0, a surrogate for empty particles, using in-house-generated antibodies. An automated and quantitative capillary Western blot assay, Simple Western, was developed using these antibodies to quantify CVA21 total particles through VP1, empty particles through VP0, relative ratio of empty to full particles through VP0 and VP4, and the absolute ratio of empty to total particles through VP0 and VP1. Finally, this Simple Western method was used to support CVA21 cell culture and purification process optimization as a high-throughput analytical tool to make rapid process decisions.
INTRODUCTION
Oncolytic viruses are an emerging immunotherapy that specifically targets and kills tumor cells as well as enhances the adaptive immune response. 1,2 An oncolytic prototype strain of Coxsackievirus A21 (CVA21) binds to overexpressed cell surface receptor, intercellular adhesion molecule 1 (ICAM-1); this phenotype is displayed in various tumor cells. 3 As a result, CVA21 has been evaluated for the treatment of various tumors with upregulated ICAM-1, including bladder cancer, melanoma, breast cancer, non-small cell lung cancer, prostate cancer, head/neck cancer, and myeloma. 4,5 Clinical studies of CVA21 have demonstrated a good safety profile and viral targeted tumor cell death, indicating this therapeutic could be promising against cancers with upregulated ICAM-1. 6
CVA21 belongs to Picornaviridae family and is a small nonenveloped human enterovirus. A mature CVA21 virion is about 30 nm in diameter with an icosahedral capsid made up of 60 protein subunits that encase the single stranded positive-sense RNA viral genome of ∼7.4 kb. 7 In mature virions, each subunit consists of four viral proteins: VP1, VP2, VP3, and VP4, whereas an immature virion contains VP0, VP1, and VP3 viral proteins.
The assembled CVA21 capsid is made up of protomers and pentamers. Single monomers of VP1, VP3, and precursor protein, VP0, make up the composition of a “protomer,” and five protomers make up a “pentamer.” Twelve pentamers can either assemble into empty capsid (procapsids) that lack an RNA genome (i.e., immature virions), or they can encapsulate the viral RNA genome (provirion), initiate the cleavage of VP0 into VP4+VP2 and form mature virions. 8 –10 Herein, immature virions and mature virions may be referred to interchangeably as “empty particles” and “full particles,” respectively.
The production of CVA21 drug substance (DS) and drug product (DP) involves many unit operations and there is a need to characterize total particle concentration and empty to full particle ratio throughout the intermediate process steps. The total particle concentration is utilized during product development to quantify the mass yield of virus particles produced during both cell culture and downstream purification. But the noninfectious empty capsid particles that contain only VP1, VP3, and VP0 (37.2 kDa) 11 are considered product-related impurities and the majority of them must be removed. Therefore, the DS and DP must be purified to contain mostly infectious, mature, full particles consisting of VP1, VP2, VP3, and VP4 with theoretical molecular weights (MWs) of 33.2, 29.9, 26.5, and 7.3 kDa, respectively. 11
Several methods have been reported in the literature to characterize similar sized nonenveloped viral capsids such as adeno-associated viruses (AAV). Enzyme-linked immunosorbent assay (ELISA) and high pressure size exclusion chromatography (HPSEC) have been used to measure total AAV particles. 12,13 Likewise, cryo-electron microscopy (cryo-EM), ion exchange chromatography (IEX), analytical ultracentrifugation, capillary isoelectric focusing, and mass spectrometry have been used to characterize full versus empty AAV particles. 12,13 Similar methods have been utilized to characterize Coxsackievirus A16. 14
But there is limited quantitative characterization of CVA21 viral proteins and particles. Recently, we reported two analytical methods to quantitate total particles using HPSEC/multi-angle light scattering/refractive index (HPSEC/MALS/RI) 15 and full versus empty particles using reverse phase-high pressure liquid chromatography (RP-HPLC) 16 in CVA21. But these two assays cannot be performed in cell culture harvest samples or other impure process intermediates due to interference from cell culture media and host cell debris. Therefore, there remains a need for a more efficient analytical method for the characterization of CVA21 for this critical quality attribute.
Since the first introduction of commercial capillary Western blotting technology 17 and its first application in vaccine was reported, 18 the technology has grown into a very practical and important tool to analyze vaccine products in development, including identity, concentration, enzyme kinetics, stability, and reagent screening. 19 –23 In this study, we report the development of an automated and quantitative capillary Western blotting method to measure the total virus particles, empty virus particles and empty to full particle ratio during CVA21 oncolytic Coxsackievirus production and development.
MATERIALS AND METHODS
Coxsackievirus production and commercial reagents
CVA21 was produced from infected MRC-5 cells cultured in a microcarrier bioreactor using a virus seed derived from a CVA21 Kuykendall prototype strain (ATCC VR-850). Lysates were harvested and clarified through depth filters to remove cell debris. The clarified harvest was purified through an affinity chromatography step and two polishing IEX steps to concentrate and clear residual impurities and empty capsids. Purified CVA21 product was exchanged into a stabilizing buffer and passed through a 0.2 μm filter to generate the final DS.
Secondary antibodies (goat anti-rabbit-horseradish peroxidase [HRP] conjugate and goat anti-mouse-HRP conjugate), dithiothreitol (DTT), MW fluorescent standards, luminol-S, hydrogen peroxide, 10 × sample buffer, antibody diluent II, wash buffer, 12–230 kDa plate prefilled with stacking matrix and separation buffer, and capillaries were purchased from Protein Simple (Santa Clara, CA).
4 × loading dye and 10 × reducing agent were purchased from Bio-Rad (Hercules, CA). Twelve percent acrylamide Bis-Tris NuPAGE gel and Pierce silver stain kit were purchased from Thermo Scientific (Waltham, MA).
Viral capsid peptides and antibodies production
Custom VP1, VP2, VP3, and VP4 peptides were synthesized at Lifetein, LLC (Hillsborough, NJ). The following capsid peptides were made: VP1 (1–76 amino acids (aa)), VP1 (221–298aa), VP1 (21-mer aa linked to KLH), VP2 (1–33aa linked to KLH), VP2 (34–76aa linked to KLH), VP2 (194–272aa), VP3 (1–80aa), VP3 (169–240aa), VP4 (2–69aa), VP4 (myristylated-2-69aa), and VP4 (34–52aa linked to KLH). They were all produced with >90% purity. Rabbit polyclonal anti-VP1, anti-VP2, anti-VP3, and anti-VP4 antibodies were generated by immunizing animals with the synthesized VP1, VP2, VP3, and VP4 peptides.
Two animals were used for each peptide immunization study. All animal work and sera collection were performed at Lifetein, LLC. The sera were affinity-purified against their corresponding peptides. The purified polyclonal antibodies (pAb) were lyophilized and rehydrated with MilliQ water to target final concentration of 1 mg/mL. All purified pAb were screened against CVA21 and their corresponding peptides to confirm reactivity and specificity.
Mouse monoclonal anti-VP1 was generated by immunizing animals with a synthesized VP1 peptide. Individual B cells were isolated and screened against CVA21 and their corresponding peptides. Reactive B cells were selected and hybridoma fusion was performed. Animals were immunized with selected hybridomas for ascites production. The ascites fluid was affinity-purified against the corresponding peptide. The purified anti-VP1 monoclonal antibody (mAb) was lyophilized and rehydrated with MilliQ water to target final concentration of 1 mg/mL.
Sodium dodecyl sulfate-polyacrylamide gel electrophoresis analysis of CVA21 samples
CVA21 samples were prepared neat and mixed with 4 × loading dye and 10 × reducing agent before heating at 70°C for 10 min. Samples were added to a 10-well 12% acrylamide Bis-Tris NuPAGE gel and run at 200V for 60 min. Gels were stained using Pierce silver stain kit and imaged using a gel imager (BioRad).
Capillary Western Detection of individual capsid proteins: VP1, VP2, VP3, and VP4
CVA21 purified DS was diluted in 1 × sample buffer to detect VP0, VP1, VP2, VP3, and VP4. Then, 2 × master mix containing DTT and sodium dodecyl sulfate (SDS) was added into diluted CVA21 samples at a 1:1 volume ratio. The samples were heated for 10 min at 70°C. The samples were equilibrated for 5 min at room temperature, spun down for 2–3 s, and transferred into a 12–230 kDa plate. Working concentrations of rabbit pAb (VP1, VP2, VP3, and VP4) and mouse mAb (VP1) were established to ensure consistent raw signal responses over the lifetime of the assay. The incubation time of all primary antibodies was 30 min and secondary HRP conjugated antibody incubation was 30 min. Data were analyzed using Compass software. Appropriate exposure times were set to ensure maximum signal while avoiding burnout.
Quantitation of VP1, VP0/VP4 ratio, and VP0 for virus particles
VP1 peptide (1–76aa, N-terminal, purity >95%) was used as a reference standard and was serially diluted in 1 × sample buffer to construct a calibration curve ranging from 6.3 × 109 to 1.0 × 1011 particles/mL (31–500 ng/mL). Mass concentration was converted to particle concentration by selecting a CVA21 sample with a known particle concentration measured through HPSEC [13]. Purified CVA21 DS was diluted to target concentration in 1 × sample buffer. The working concentrations of rabbit anti-VP1 pAb and mouse anti-VP1 mAb were 10 and 1 μg/mL, respectively.
For measuring the relative VP0 to VP4 ratio, CVA21 samples were probed with anti-VP4 pAb, as it detects VP0 and VP4, and the VP0/VP4 peak area ratio from a sample were normalized to the VP0/VP4 peak area ratio from a reference standard. The reference standard was a CVA21 sample that contained a mixture of empty and full particles. The working anti-VP4 pAb concentration was 20 μg/mL.
For VP0 quantitation, purified CVA21 DS was diluted to target concentration in 1 × sample buffer. VP0 protein purified from CVA21 was used as reference standard and was serially diluted in 1 × sample buffer to construct a calibration curve ranging from 0.78 to 25 ng/mL. Anti-VP4 pAb (20 μg/mL) was used as detection for VP0.
The sample preparation and capillary Western conditions described in Capillary Western Detection of individual capsid proteins: VP1, VP2, VP3 and VP4 section were used. Other parameter conditions were applied according to vendor default recommendations. Peak integrations were performed using Compass software. Peak area calculations and data analysis were performed using SigmaPlot 14.0 and Microsoft Excel software.
RESULTS AND DISCUSSION
VP0, VP1, VP2, VP3, and VP4 Identity in CVA21
Antibodies are important critical reagents for immuno-based assays, so specific antibodies against each of the four capsid proteins, VP1, VP2, VP3 and VP4, needed to be identified. A total of 33 commercial antibodies were screened from various vendors with only 6 having some reactivity against CVA21 VP1, VP2, and VP3 (see Supplementary Table S1 in Supplementary Data). No commercial antibodies could be identified that reacted strongly against VP2 or VP4 and all of the commercial antibodies that generated an adequate response against VP1 and VP3 were in very limited supply from the vendors so they could not be used. Therefore, custom antibody generation was initiated.
pAb were generated in rabbits against N-terminal and C-terminal peptides from each of the capsid proteins except for VP4. The VP4 sequence is only 68aa long so the entire peptide was synthesized in one strand. In addition, in wild-type CVA21, VP4 is myristylated so VP4 pAbs were generated against both a myristylated peptide and a non-myristylated peptide. All of the purified pAb were analyzed and screened against CVA21. The antibodies directed toward the N-terminal sequences of VP1, VP2, and VP3 and non-myristylated VP4 gave the best chemiluminescence signals and were selected for further assay optimization (Supplementary Table S2 and Supplementary Fig. S1 in Supplementary Data).
An overview of the antibody response and specificity for each capsid protein, VP1 (∼41 kDa), VP2 (∼37 kDa), VP3 (∼32 kDa), and VP4 (∼12 kDa) against CVA21 are illustrated in their respective electropherograms (Fig. 1A, B). In addition, VP0 (∼44 kDa) was observed in this sample, which was not surprising as it was expected to contain some immature virions. The observed MWs in the Simple Western were slightly higher than the reported values in the literature using manual western or sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) gel likely because of the different separation gel compositions. 22,23 Peak identities were further confirmed by running one sample of CVA21 that contained high amounts of mature virus and another sample with >95% empty particles (Fig. 1C). As expected, the mature virus contained VP1, VP2, VP3, and VP4, whereas the immature virus contained only VP0, VP1, and VP3.
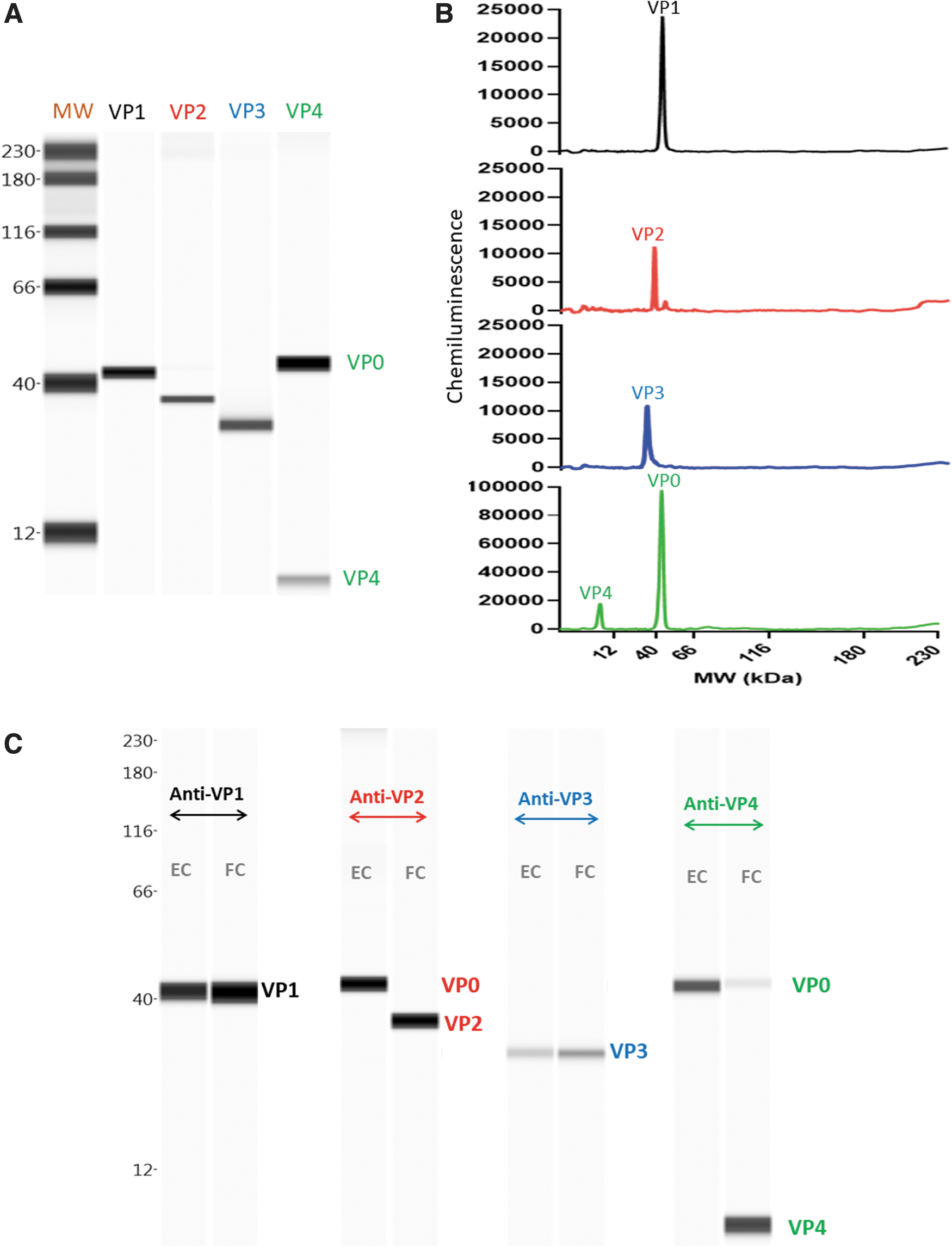
Simple Western Identity of VP0, VP1, VP2, VP3, and VP4 proteins in CVA21.
Anti-VP1 and anti-VP3 could be used for total particle quantitation and anti-VP2 and anti-VP4 could be used for empty versus full particle characterization. Anti-VP1 and anti-VP3 generated identical responses; however, anti-VP1 generated a more Gaussian shaped peak compared with anti-VP3; anti-VP4 generated a stronger response against VP0 and VP4 compared with anti-VP2 (Supplementary Fig. S2 in Supplementary Data). Therefore, anti-VP1 and anti-VP4 were selected for CVA21 total particle quantitation and full versus empty particle characterization studies.
Quantitative total particles concentration
Total virus particle concentration can be measured using HPSEC, but only purified samples can be analyzed due to matrix interference from cell debris and host cell proteins. Automated capillary Western avoids these matrix interference effects as it is denaturing and immuno-specific, so the method can analyze cell culture or nonpurified CVA21 samples. VP1 is known to be present in all particles; therefore, a quantitative VP1 Simple Western mass assay was developed to measure CVA21 total particles concentration during process development from cell culture to purification.
For absolute quantitation, a reference standard is required so, initially, a CVA21 DS batch with known infectivity measured by plaque assay and full/total particles measured by HPSEC was used in the VP1 Simple Western assay. However, there was limited availability of this CVA21 DS lot during early development, so there was a need to evaluate an alternative reference standard. The N-terminal VP1 peptide used in antibody generation was evaluated as a possible alternative reference standard. A standard curve was generated using the N-terminal VP1 peptide with exceptional linearity.
Furthermore, the VP1 peptide was very stable upon multiple freeze thaws and generated consistent chemiluminescence signals across the standard curve (%CV <20% for peak area, n = 6). Subsequently, the VP1 peptide reference standard mass concentration units, μg/mL, were converted to particle concentration units, particles/mL, by calibrating it to the known virus reference standard concentration, in particles/mL, measured from HPSEC. A typical standard curve using the calibrated VP1 peptide (∼12 kDa) reference standard demonstrates tremendous linearity with an R 2 > 0.99 (Fig. 2A).
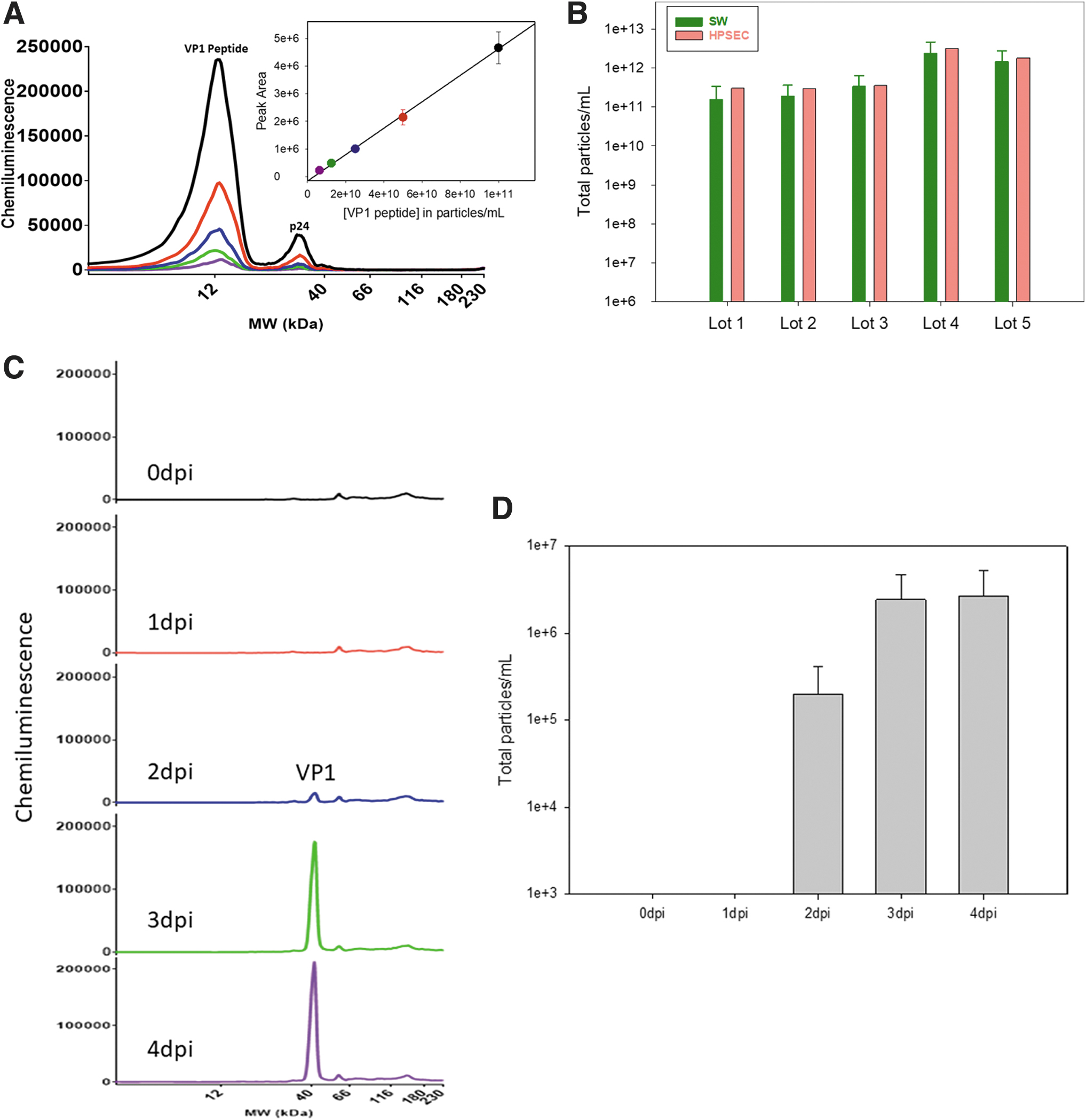
The measurement of total particles in CVA21 using the Simple Western with the new VP1 peptide reference standard was validated by running several CVA21 DS lots side by side with the established HPSEC method. As expected, the results suggested the total particle concentrations were comparable between the two methods providing confidence to the Simple Western method with the new VP1 peptide reference standard as it was orthogonal to the current HPSEC method (Fig. 2B).
The quantitative total particle concentration capillary Western assay using N-terminal VP1 peptide as the reference standard was applied to analyze samples in cell culture experiments to monitor the virus yield. A VP1 peak at 41 kDa was observed after 2 days postinfection (dpi) and plateaued at 3 dpi (Fig. 2C, D). The total particles measurement would be very difficult using HPSEC or RP-HPLC due to cell culture media interference effects and cellular debris. The ability of the Simple Western method to measure impure samples with a fast turnaround time (∼3–4 h) allowed it to be used as an analytical tool to quickly inform process decisions.
CVA21 purification development support
The VP1 quantitative Simple Western assay was used to inform CVA21 process purification optimization development. As previously discussed, the VP1 assay total particle concentration measurements aligned with the HPSEC method, so it was used as a surrogate assay for titer step yield for each process step. The process step yield from harvest to sterile filtration was compared from the initial process called process 1 with an alternative process called process 2 (Fig. 3A). Both processes had similar overall VP1 yields, recovering about 25% of the total CVA21 particles present in the cell culture harvest.
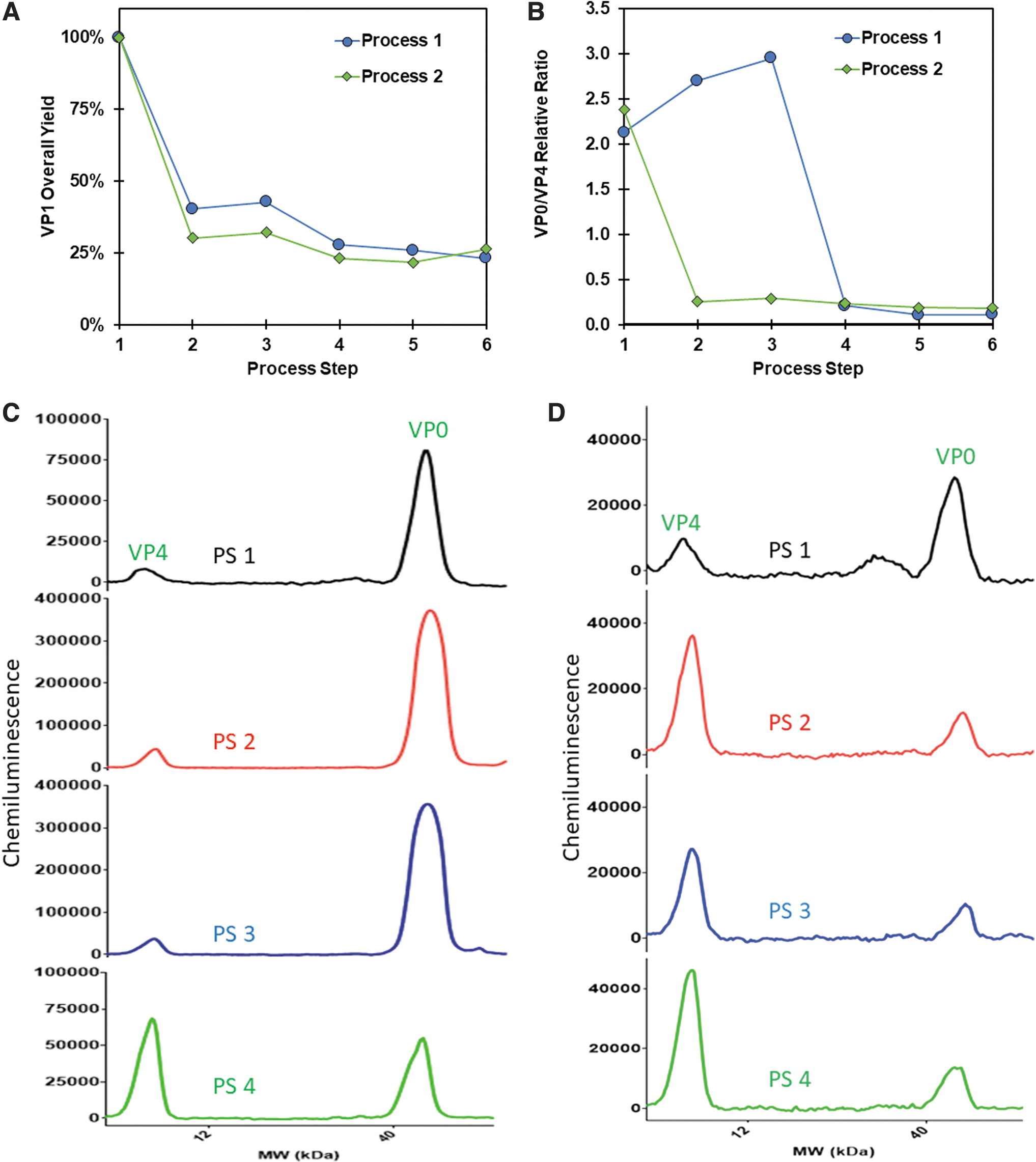
In addition, in both processes there was an ∼70% loss in total particles after an initial purification step (PS), but subsequent step yields remained consistent between the two arms. The VP0/VP4 relative ratio, a surrogate for empty (VP0) to full (VP4) particles, was measured and compared for the two processes (Fig. 3B). Process 1 had a VP0:VP4 relative ratio that was reduced about an order of magnitude in PS 4, suggesting a purification of full particles from empty particles.
However, in Process 2, empty particles were cleared earlier in the process at PS 2. At the final PS 6, the VP0:VP4 relative ratio for both processes were similar, representing either process could be used to manufacture a consistent DS (Fig. 3B). These data were corroborated with additional orthogonal methods as demonstrated by Swartz et al that confirmed the identity of CVA21 particles present in cell culture harvest. 24 Notably, Swartz et al utilized ultracentrifugation, SDS-PAGE, RP-HPLC, infectivity, and cryo-EM assays in combination with the VP0/VP4 and VP1 Simple Westerns to characterize the clearance of empty particles in the manufacturing processes.
The associated electropherograms of VP0/VP4 relative ratios for process 1 and process 2 are shown (Fig. 3C, D). The electropherograms of VP0/VP4 in process 1 demonstrate a large change in relative ratio of VP0/VP4 at PS 4. The electropherograms of VP0/VP4 in process 2 illustrate a large change in relative ratio of VP0/VP4 at PS 2 and remain constant throughout the rest of the PS steps. This example demonstrates the ability of the Simple Western to characterize the relative VP0/VP4 ratio in process intermediates as a surrogate for the all-important quantification of empty versus full particles. The actual absolute VP0 concentration in final DS was quantified as described in the next section.
Quantitation of VP0 as a surrogate for empty particles
A known reference standard of VP0 was needed to quantitate the absolute amount of VP0 in the final CVA21 DS. VP0 protein was purified using RP-HPLC from one of the CVA21 process intermediate samples known to contain substantial amounts of VP0 (Supplementary Fig. S3 in Supplementary Data). The concentration of purified VP0 was measured using an SDS-PAGE gel (>95% purity) with bovine serum albumin as a reference standard (Supplementary Fig. S4 and Table S3 in Supplementary Data). The purified VP0 from CVA21 was used as a reference standard to generate a linear calibration curve using anti-VP4 antibody to measure the concentration of VP0 in final CVA21 DS samples (Fig. 4A).
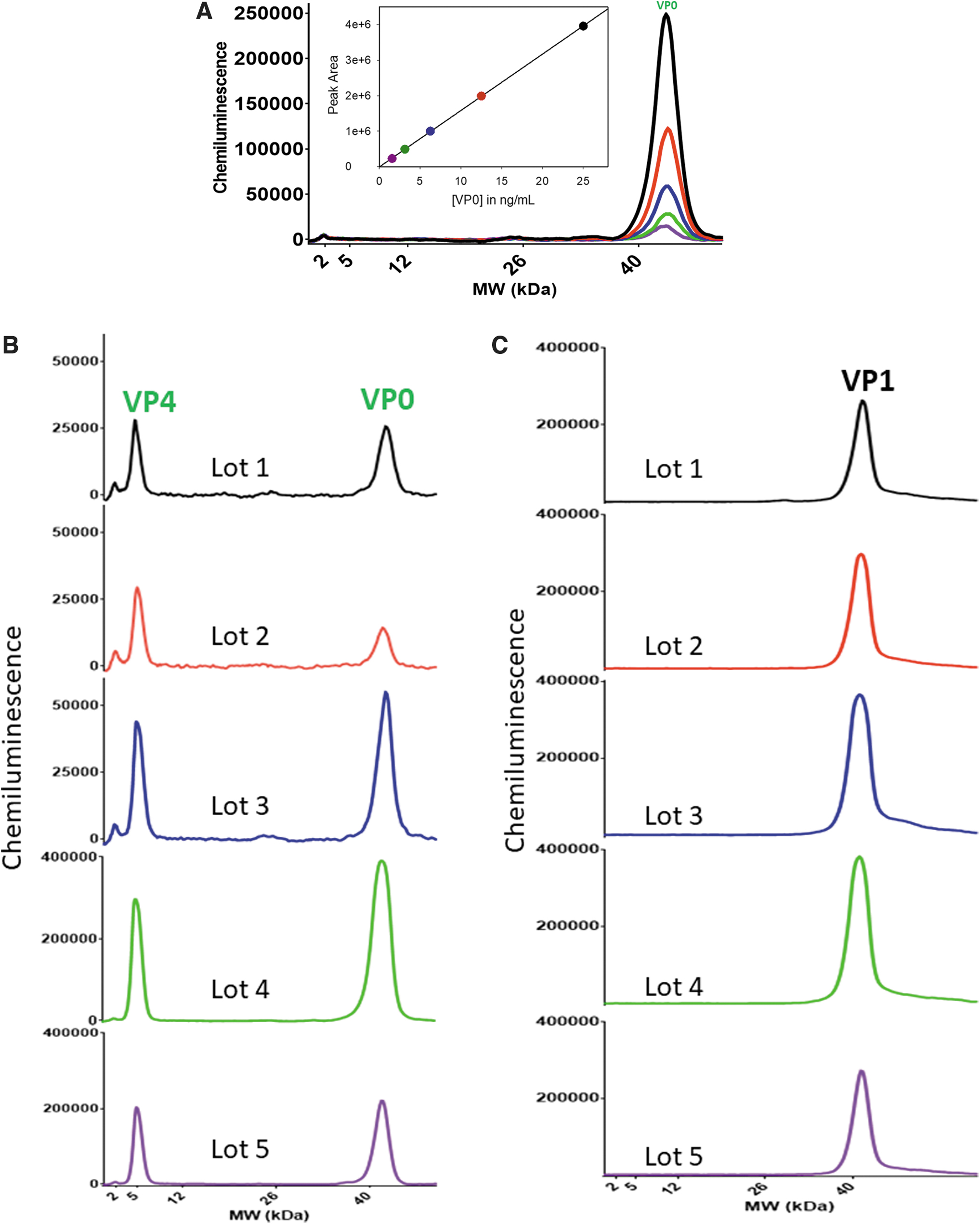
% Empty virions in five drug substance lots
Five different CVA21 lots of DS were measured for VP0 concentration (Fig. 4B). Expectedly, both VP0 and VP4 peaks were observed and the amount of VP0 was quantitated using the VP0 reference standard and converted to total empty virus particles assuming 60 copies of VP0 per virion. The same samples were also measured using the quantitative VP1 protein assay, a surrogate for total particles (Fig. 4C). In these experiments, lower MW range separation gel matrix (2–40 kDa) from Protein Simple was utilized for measurements.
The final results of measured VP0 (empty particles) and VP1 (total particles) are tabulated in Table 1 as particles/mL and the calculated percentage of empty particles was very low across all lots (<0.5% for all lots). This demonstrates the Simple Western method was able to measure the absolute amount of VP0 as a surrogate for empty particles and subsequent results provided evidence that the process is able to selectively remove empty particles efficiently.
CONCLUSIONS
We have developed a quantitative capillary Western method to analyze CVA21 during process development. Owing to the lack of commercial CVA21 antibodies against the capsid proteins, high affinity in-house rabbit pAb against VP0, VP1, VP2, VP3, and VP4, and a mouse monoclonal against VP1 were generated using truncated peptides as immunogens. The Simple Western method was capable of measuring the total particle concentration through anti-VP1 and VP1 peptide reference standard, empty to full particle ratio through anti-VP4 and empty particle concentration through anti-VP4 and VP0 protein reference standard.
During process development optimization, the Simple Western method provided valuable process characterization information about the two main downstream process candidates, including yield, through total particle concentration, and efficient removal of empty particles, through empty to full particle ratio and empty and total particle concentrations. The Simple Western method provided evidence that both processes generated similar yields and empty particle removal efficiency, but the majority of empty particles were removed during different steps in the two processes.
In short, the Simple Western method was an invaluable analytical tool providing timely information to help guide downstream process decisions and optimization. The developed quantitative Simple Western method presented in this article, including the framework for antibody generation, can have broader applications toward other types of oncolytic viruses with similar maturation life cycles as well as gene therapy area using AAVs.
Footnotes
ACKNOWLEDGMENTS
We also would like to extend our gratitude to our colleagues at Vaccine Upstream and Downstream Process Development teams for providing CVA21 materials. We would like to thank Dr. Josef Vlasak for early support. Authors would like to thank Drs. Anne Payne and Caroline McGregor for critical review of the article.
AUTHORs' CONTRIBUTIONS
Conceptualization, methodology, writing—original draft, and visualization by R.R.R. Investigation, writing—review and editing, and visualization by P.F.G. Investigation by L.S., J.R., A.P., N.C., M.D., J.Z.D., and S.W. Resources by J.O. and Y.S. Resources, writing—review and editing by A.R.S. Conceptualization, supervision, writing—review and editing by J.W.L.
AUTHOR DISCLOSURE
All authors are/were employees of Merck Sharp & Dohme LLC, a subsidiary of Merck & Co., Inc., Rahway, NJ and may potentially own stock and/or hold stock options in Merck & Co., Inc., Rahway, NJ.
FUNDING INFORMATION
Funding of this study was provided entirely by Merck Sharp & Dohme LLC, a subsidiary of Merck & Co., Inc., Rahway, NJ.
Supplementary Material
Supplementary Data
Supplementary Figure S1
Supplementary Figure S2
Supplementary Figure S3
Supplementary Figure S4
Supplementary Table S1
Supplementary Table S2
Supplementary Table S3
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
