Abstract
T-cell receptor (TCR) immunotherapy uses T cells engineered with new TCRs to enable detection and killing of cancer cells. Efficacy of TCR immunotherapy depends on targeting antigenic peptides that are efficiently presented by the best-suited major histocompatibility complex (MHC) molecules of cancer cells. However, efficient strategies are lacking to easily identify TCRs recognizing immunodominant peptide-MHC (pMHC) combinations utilizing any of the six possible MHC class I alleles of a cancer cell. We generated an MHC cell library and developed a platform approach to detect, isolate, and re-express TCRs specific for immunodominant pMHCs. The platform approach was applied to identify a human papillomavirus (HPV16) oncogene E5-specific TCR, recognizing a novel, naturally processed pMHC (HLA-B*15:01) and a cytomegalovirus-specific TCR targeting an immunodominant pMHC (HLA-B*07:02). The platform provides a useful tool to isolate in an unbiased manner TCRs specific for novel and immunodominant pMHC targets for use in TCR immunotherapy.
Introduction
A
A crucial aspect for the outcome of TCR immunotherapy is the identification of TCRs that target peptide-MHC (pMHC) combinations efficiently expressed by cancer cells. Historically, TCR studies have focused primarily on the most frequent MHC allele HLA-A*02:01, which, depending on the patient population, is expressed by 18–47% of individuals. 6 TCRs restricted to HLA-A*02:01 are thereby applicable to only a limited number of patients. However, as antigens are expressed and degraded by the antigen-processing machinery of a cell, epitopes are generated that may fit into the binding groove of any of the six possible MHC class I alleles of the cell. Currently, available software algorithms fail to predict accurately which pMHC combinations are most efficiently presented at the cell surface to engage TCRs and induce strong T-cell responses. Therefore, strategies are required to enable unbiased identification of immunodominant pMHC combinations, taking into account all MHC alleles of a given donor.
For endogenous processing and presentation of pMHCs, we focused on two antigen-presenting cell (APC) types that have the capacity to stimulate T-cell responses. 7,8 Professional APCs (pAPC) induce the most potent T-cell responses. Expression of immune proteasomes is a key feature enabling pAPCs to increase their ability to generate epitopes and express cell surface pMHCs as targets for TCRs. Simultaneously, the accessory molecules CD80 and CD86 bind to CD28 on T cells, providing critical co-stimulatory signals. Mature dendritic cells (mDC) are potent pAPC to enrich antigen-specific T cells in vitro. 9,10 The human erythroleukemic cell line K562 can also be used as APC. It lacks natural surface expression of MHC I and MHC II alleles but does express β-2 microglobulin. 11 Thus, upon transgenic expression of an MHC allele, such as the HLA-A*02:01 alpha chain, these cells acquire functional antigen-processing and presentation capacity to form cell surface pMHCs. 12 –15 Such MHC-engineered K562 cells can form immunological synapses due to their co-expression of ICAM-1 and LFA-3 through which T cells can be activated. 15 Importantly, K562 cells do not contain viral sequences, 16 which could lead to unwanted responses, and express constitutive proteasomes that provide antigen processing typical of tumor cells. 17
This study used a multi-step platform approach for the identification, isolation, and characterization of TCRs using DC-based enrichment of antigen-specific T cells combined with the application of a K562-based MHC cell library. Using this platform, we identified a TCR specific for a novel human papillomavirus HPV16 E5 epitope presented by HLA-B*15:01 and a TCR specific for a human cytomegalovirus (CMV) pp65 epitope presented by HLA-B*07:02. Both TCRs may be candidates for clinical application of TCR immunotherapy.
Methods
In silico analysis of epitopes
To determine epitopes that have the highest binding affinity to a given MHC class I molecule, the online prediction server netMHCpan3.0 was used (
Cell lines
Suspension cells (lymphoblastoid cell lines [LCLs], K562) were cultured in RPMI1640 medium (Gibco) supplemented with 10% fetal bovine serum (FBS; Pan Biotech), 1 × minimum essential medium non-essential amino acids, 1 × sodium pyruvate, 1 × penicillin/streptomycin (all Gibco). LCLs (obtained from the International Histocompatibility Work Group) were seeded at 0.5 × 106/mL, and passaging was performed at a ratio of 1:2. K562 cells (ATCC CCL-243) were seeded at 0.1 × 106/mL and split twice a week at 1:5 to 1:20. Adherent HPV16-positive cervical carcinoma-derived cell lines CaSki (ATCC CRL-1550) and SiHa (ATCC HTB-35) and HPV16-positive head and neck cancer cell lines SCC090 and SCC152 (LGC Standards) were seeded at 1–2 × 106 cells/75 cm2 cell culture flask in 15 mL of Dulbecco's modified Eagle's medium (DMEM)-Ham's F12 (Gibco) supplemented with 10% FBS and 1x penicillin/streptomycin. Cells were split using 0.125% trypsin-EDTA (Gibco). The HG820-GALV packaging cell line (Eufets) 20 was cultured at 1–2 × 106 cells/75 cm2 flask in DMEM (Gibco) supplemented with 10% FBS and 1 × penicillin/streptomycin.
Generation of antigen-expressing DCs and stimulation of T cells
HPV16 E5- and CMV pp65-encoding genes were molecularly cloned into the expression plasmid pcDNA3.1(–) (Invitrogen) under the control of a T7 promoter. Plasmids were linearized by restriction enzyme digestion at the 3′ end of the transgene. ivtRNA was generated using in vitro synthesis of capped RNA followed by poly-A tailing (Ambion). Mature dendritic cells (mDC) were generated from plate adherent monocytes, as described. 21,22
Generation of MHC cell library
To generate cell lines expressing single Mhc alleles, cDNA gene sequences of different Mhc alleles were linked to reporter genes GFP or CFP via an internal ribosomal entry site (IRES). Resulting gene cassettes were molecularly cloned into the γ-retroviral vector MP71 for the generation of viral particles. 23 K562 cells in exponential growth phase were incubated with viral supernatant in the presence of 4 μg/mL of protamine sulfate (Sigma–Aldrich) followed by 2 h of spinoculation at 800 g and 32°C. Transduced K562 cells were sorted upon surface MHC expression using magnetic bead separation. To confirm expression of the target MHC transgene cassette, RNA/cDNA was generated after several passages followed by polymerase chain reaction (PCR) amplification of the transgene cassette and Sanger sequencing. Resulting electropherograms were analyzed to confirm single-target MHC expression. To assess the gene sequence expressing the exact epitope recognized by the E5 TCR, K562-B*15:01 cells were transduced with truncated gene segments (minigenes) of E5. To do so, HPV16 E5 reference sequences of different length were amplified by PCR, and resulting minigenes were cloned into the γ-retroviral vector MP71, which was used to generate viral supernatant for transduction.
Identification of antigen–MHC-specific T-cell responses
K562 cells of the MHC cell library were transfected with 10 μg of antigen ivtRNA of HPV16 E5 or CMV pp65 using electroporation. Four to six hours after pulsing, K562 cells were harvested and co-cultured with peripheral blood mononuclear cells (PBMCs). Two assays were chosen to determine antigen–MHC-specific reactivity of T cells within PBMC samples. First, supernatants of co-cultures were used to measure interferon (IFN)-γ release by enzyme-linked immunosorbent assay (ELISA; BD Biosciences). Second, T cells were labeled with antibodies to determine the expression of the T-cell activation marker CD137 and co-stained for CD8. The antigen–MHC specificity of T cells was regarded as positive when signals were detected in both assays and showed elevated levels compared to other antigen–MHC combinations.
Isolation of antigen-specific TCRs
Upon detection of specific T cells, PBMC samples frozen after one stimulation cycle were thawed and re-stimulated for 14 days to be used in a co-culture assay only with E5-specific T cells (K562-B*15:01) or pp65-specific T cells (K562-B*07:02). CD137- and CD8-positive T cells (between 500 and 10,000) were directly sorted in cell lysis buffer to purify whole-cell RNA. For efficient analysis, it was necessary to introduce an amplification step of TCR gene sequences. Unknown TRAV and TRBV gene segments precluded them from classical PCR amplification. Therefore, the SMARTer RACE cDNA amplification kit (Clontech) was used, according to the manufacturer's instructions. PCR products were separated on a 1.5% agarose gel, and bands representing amplified TRA and TRB chains were excised and purified using the Invisorb Fragment CleanUp Kit (Stratec). DNA was eluted in 30 μL of elution buffer. Purified PCR fragments were used for ligation into the pCR4Blunt-TOPO vector using the Zero Blunt TOPO PCR Cloning Kit for Sequencing (Invitrogen). After ligation of the RACE PCR products into the TOPO cloning vector, DNA preparations of the vectors were used for further processing. Suitable primers were designed (Eurofins Genomics), which contained sequences complementary to flow cell oligos, barcoding sequences, and binding sites for the TOPO cloning vectors. Primers were used to amplify TRAV and TRBV repertoires from the TOPO cloning vectors, and resulting PCR products were used for paired-end sequencing on a MiSeq machine (Illumina). After filtering reads, which fulfilled quality criteria (Eurofins), reads were clustered (>98% sequence similarity) and assigned to sequence representatives (Eurofins). TCR repertoire analysis was performed using web-based IMGT/HighV-quest (
Wild type TRAV and TRBV sequences of TCR chains, which were found to be dominantly expressed in antigen-MHC-specific T cells, were fused to codon-optimized murine TRAC and TRBC regions, and each TRA and TRB gene was ligated separately into the MP71 vector to enable combinatorial expression of different TRA and TRB chains. After identification of TRA and TRB chain pairs, which formed a functional TCR, modified TCR transgene cassettes for improved expression in primary human PBMCs were constructed as described. 26,27 To increase transgenic TCR expression further, codon optimization was performed (GeneArt). 28
Generation of γ-retroviral particles
Calcium phosphate transfection of HG820-GALV packaging cells with MP71 plasmids (18 μg) carrying the Mhc or TCR transgene cassettes, respectively, was used to generate virus particles. Viral supernatant was harvested 48 h after transfection and used for transduction of K562 cells. For transduction of PBMCs, viral supernatant was harvested after 48 h and used for the first transductions. Then, fresh medium was added to the packaging cells, and after another 24 h, the supernatant was harvested and used for a second transduction. 26,29
Transduction of PBMCs
PBMCs were isolated from blood samples of healthy donors, with written informed consent and approval of the Institutional Review Board of the Max Delbrück Center for Molecular Medicine Berlin. To isolate PBMCs, 30 mL of heparinized blood was mixed with phosphate-buffered saline, and gradient centrifugation was performed using Biocoll separating solution (Millipore). PBMC transduction was carried out as described. 26
IFN-γ release assays
Co-culture assays were performed to assess antigen-specific reactivity of T cells to different sets of target cells (K562-MHC, LCLs, and HPV16-positive tumor cell lines). LCLs were pulsed with HPV16 E5 ivtRNA using electroporation 4–6 h before adding TCR-transduced T cells. GFP electroporation was performed in parallel to assess transfection efficiency. HPV16-positive tumor cell lines were pretreated with 100 ng/mL of IFN-γ (Peprotech) for 24 h before they were trypsinized, washed extensively, and transfected with E5 ivtRNA. Tumor cell lines were seeded in plates for co-culture for up to 6 h prior to adding T cells. T cells stimulated with PMA and ionomycin were used as a positive control. To assess peptide specificity, K562-MHC cells were pulsed with different peptides (Biomatik) at a concentration of 10–6 M. To do so, peptides were added to the cell suspension of K562-MHC cells and TCR-transduced T cells. For peptide titration assays, K562-HLA cells were pulsed with different molar amounts of peptide, and TCR-transduced T cells were added for an overnight co-culture. The alanine scan was performed, as described before, with minor changes. 30 Nine peptides were used each having one amino acid substitution to alanine at a different position. Peptides were added at a concentration of 10–8 M to the co-culture of K562-B*15:01 cells and TCR-transduced T cells. IFN-γ release of T cells was determined as indication for antigen-specific TCR engagement by ELISA.
Flow cytometry analysis
Surface expression of transgenic TCRs was detected using an antibody directed against the murine constant TCR β region (mTRBC; Biolegend) and flow cytometric analysis. Anti-CD8 and anti-CD137 antibodies (Biolegend) were further used for T-cell characterization. HLA-DR (MHC class II), CD80, CD83, and CD86 for phenotyping of mDCs as well as the HLA-A,-B,-C antibody for measuring surface MHC expression on K562-HLA cells were purchased from Biolegend. Before antibody staining, mDCs were incubated for 10 min with a human Fc blocking antibody. Sytox blue (Thermo Fisher Scientific) was used for live/dead staining of cells, and flow cytometry measurements were performed at a MACSQuant (Miltenyi). Fluorescence-activated cell sorting (FACS) was performed on an Aria II (BD).
Results
T-cell expansion on antigen-expressing DCs
T-cell responses were investigated to the HPV16 oncogene E5,
31
which is a potential target for immunotherapy of HPV-associated malignancies such as cervical as well as head and neck cancers. To date, isolation of E5-specific TCRs has not been reported.
32
E5 was found to be frequently expressed at the RNA and protein levels in HPV-induced tumors.
33,34
Epitope prediction for E5 revealed 61 epitopes as potential binders to at least one of the six MHC alleles of the PBMC donor used in this study (Supplementary Table S1; Supplementary Data are available online at
To identify T-cell clonotypes responding to E5 and pp65, mDCs expressing MHC II, CD80, CD86, and the differentiation marker CD83, were prepared from plate-adherent monocytes (Supplementary Fig. S1A). mDCs were transfected with ivtRNA of the antigens to ensure endogenous processing and MHC presentation 22,36 (Fig. 1A). Transfection efficiency was >90%, as measured by GFP ivtRNA expression (Supplementary Fig. S1B). Autologous PBMCs were stimulated twice for 14 days with antigen-expressing mDCs to induce proliferation of antigen-specific T-cell clones.
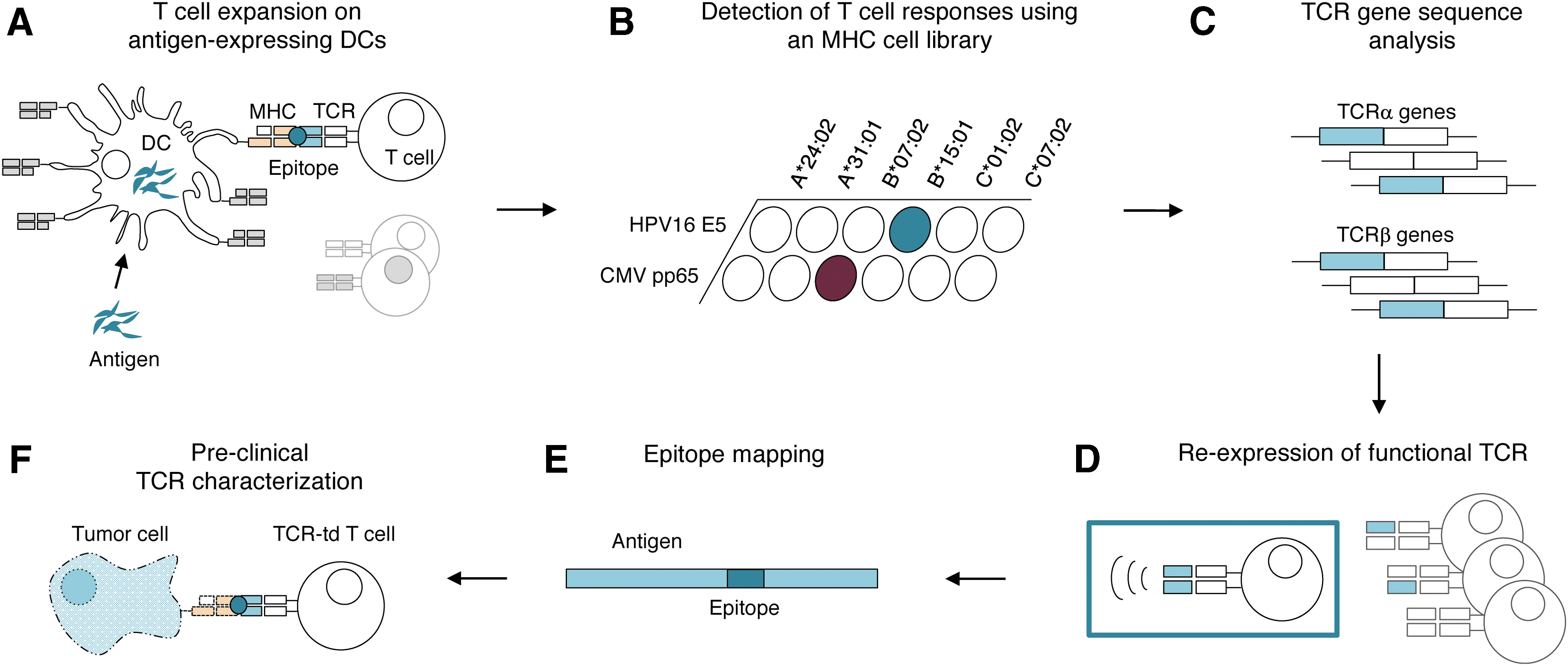
Multi-step approach for the identification, isolation, and re-expression of peptide major histocompatibility complex (pMHC)-specific T-cell receptors (TCRs).
Detection of T-cell responses using the MHC cell library
To decipher antigen-specific T-cell responses, an MHC cell library was generated consisting of K562 cell lines expressing single MHC alleles. MHC gene sequences were stably transduced in K562 cells, which were then sorted for homogenous MHC surface expression. The MHC cell library, comprised of K562 cells expressing >40 different MHC alleles, provided the cellular reagents representing each of the six MHC I alleles of the study donor (Fig. 2A). E5-expanded donor T cells were co-cultured individually with each of the six different MHC-expressing K562 cell lines that were additionally transfected to each express one of the two study antigens. Expression of CD137 activation marker of CD8-positive T cells (Supplementary Fig. S2) and cytokine secretion of IFN-γ upon stimulation were correlated to form a concise picture of T-cell responses (Figs. 1B and 2B). T cells showed clear recognition of E5-expressing K562-B*15:01 cells, while no reactivity to any other E5-MHC combination was observed. PBMCs enriched for CMV contained T cells solely specific for pp65 in combination with B*07:02 (Figs. 1B and 2B). T-cell donors seropositive for CMV were described to have very high precursor frequencies of pp65-specific T cells 37 and although 200 possible pp65-specific pMHC combinations were calculated, T cells targeting pMHCs other than B*07:02 were not detected. This indicated that pp65-B*07:02 was also an immunodominant TCR target. 38
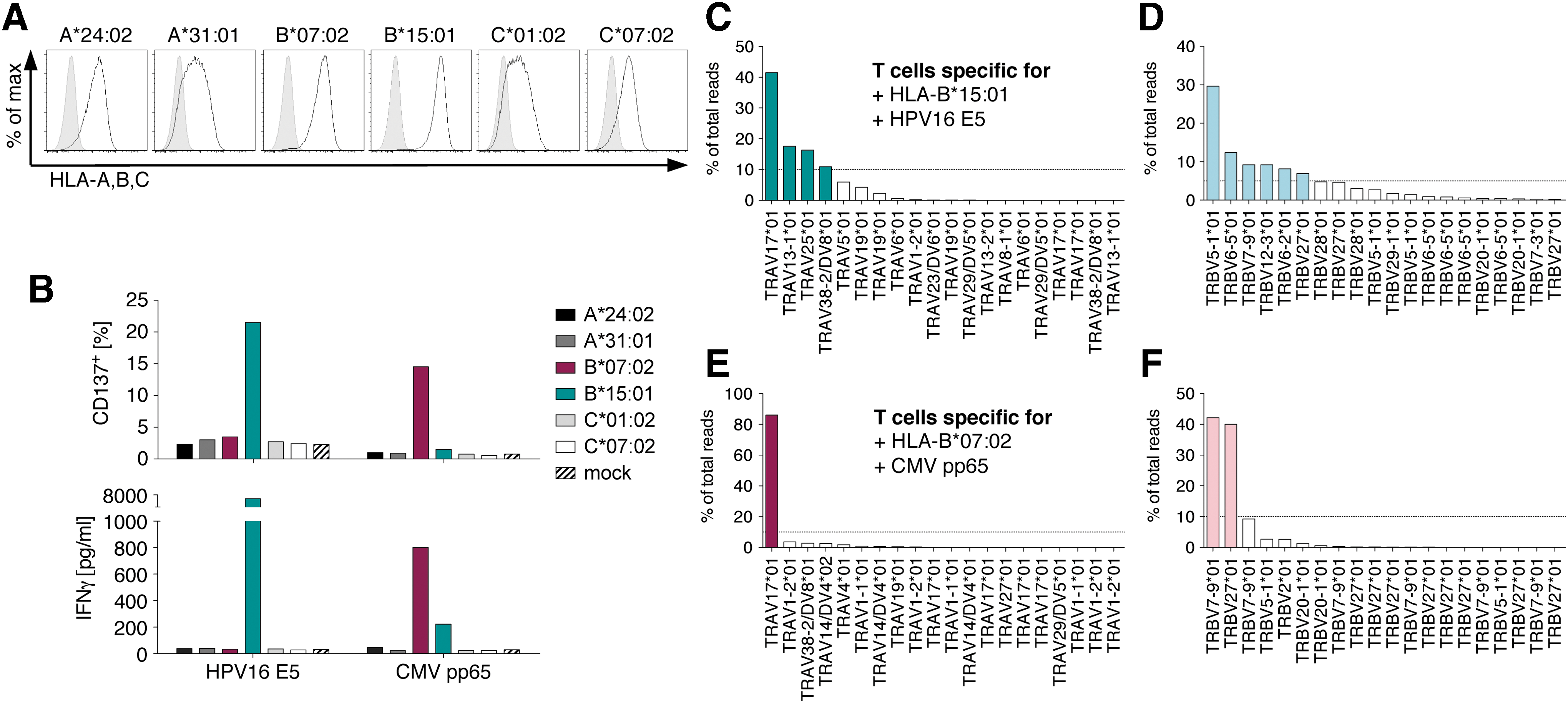
Detection of T-cell responses using an MHC cell library and TCR gene sequence analysis.
TCR gene sequence analysis
To analyze the TCR repertoire of responding T cells (Fig. 1C), CD137-expressing T cells were sorted on a FACS (Supplementary Fig. S3) and directly used to extract RNA and to clone TCR-specific cDNA into the TOPO vector. PCR products of vector preparations were used for paired-end deep sequencing to analyze variable TCRα (TRAV) and TCRβ (TRBV) gene segments, respectively. The quantity of reads of a specific TRAV or TRBV clonotype was indicative for the frequency of a T-cell clonotype within the sorted T-cell population. In all samples, a strong enrichment for very few chains was observed (Fig. 2C–F), showing that pMHC-triggered CD137 activation was a selective process involving only a few T-cell clonotypes. T cells responding to HPV16 E5 in combination with B*15:01 revealed four TRAV chains that appeared at a frequency of >10% of all reads (Fig. 2C). In the TRBV repertoire, six chains yielded a frequency of >5% and were chosen for further analysis (Fig. 2D). TCR gene sequence analysis of pp65-B*07:02-reactive T cells exhibited one dominant TRAV chain contributing to >80% of total reads (Fig. 2E) and two predominant TRBV chains (Fig. 2F).
Re-expression of functional TCRs
Next, single-chain TRA and TRB retroviral vectors containing predominant chains for combinatorial re-expression in primary T cells were generated to identify TRA and TRB chains forming functional, antigen-specific TCRs (Fig. 1D). Constant TRA (TRAC) and TRB (TRBC) gene segments were exchanged by murine codon-optimized counterparts to enhance TCR expression (Fig. 3A). 26 –28 Thus, surface expression of all TRA/TRB combinations became detectable using a monoclonal antibody directed against the murine TRBC chain (Fig. 3B and C). Four TRA chains and six TRB chains derived from E5-B*15:01-specific T cells were re-expressed in 24 different combinations in primary T cells. TCR surface expression on CD8-positive T cells was achieved for every combination at rates between 6.7% and 27.1% (Fig. 3B). Therefore, all 24 TCRs were available for testing antigen specificity for E5-B*15:01. Also, both TRA/TRB combinations from the predominant TCR chains found in pp65-specific T cells were expressed at the cell surface and used for testing the antigen specificity to pp65-B*07:02 (Fig. 3C). Strikingly, for E5, there was only one TRAV/TRBV combination, where T cells showed clear recognition of K562-B*15:01 cells while exhibiting no reactivity to target cells without E5 antigen (Fig. 3D). None of the other 23 TRA/TRB combinations provided T cells with antigen specificity that led to elevated levels of IFN-γ release. For pp65, one TRAV/TRBV combination formed a functional, antigen-specific TCR, leading to specific IFN-γ release of T cells upon encounter of K562-B*07:02 cells expressing pp65 (Fig. 3E). Expression of the second TRA/TRB combination completely abrogated cytokine release, demonstrating that only correct pairs of TRA and TRB chains led to selective pMHC specificity.
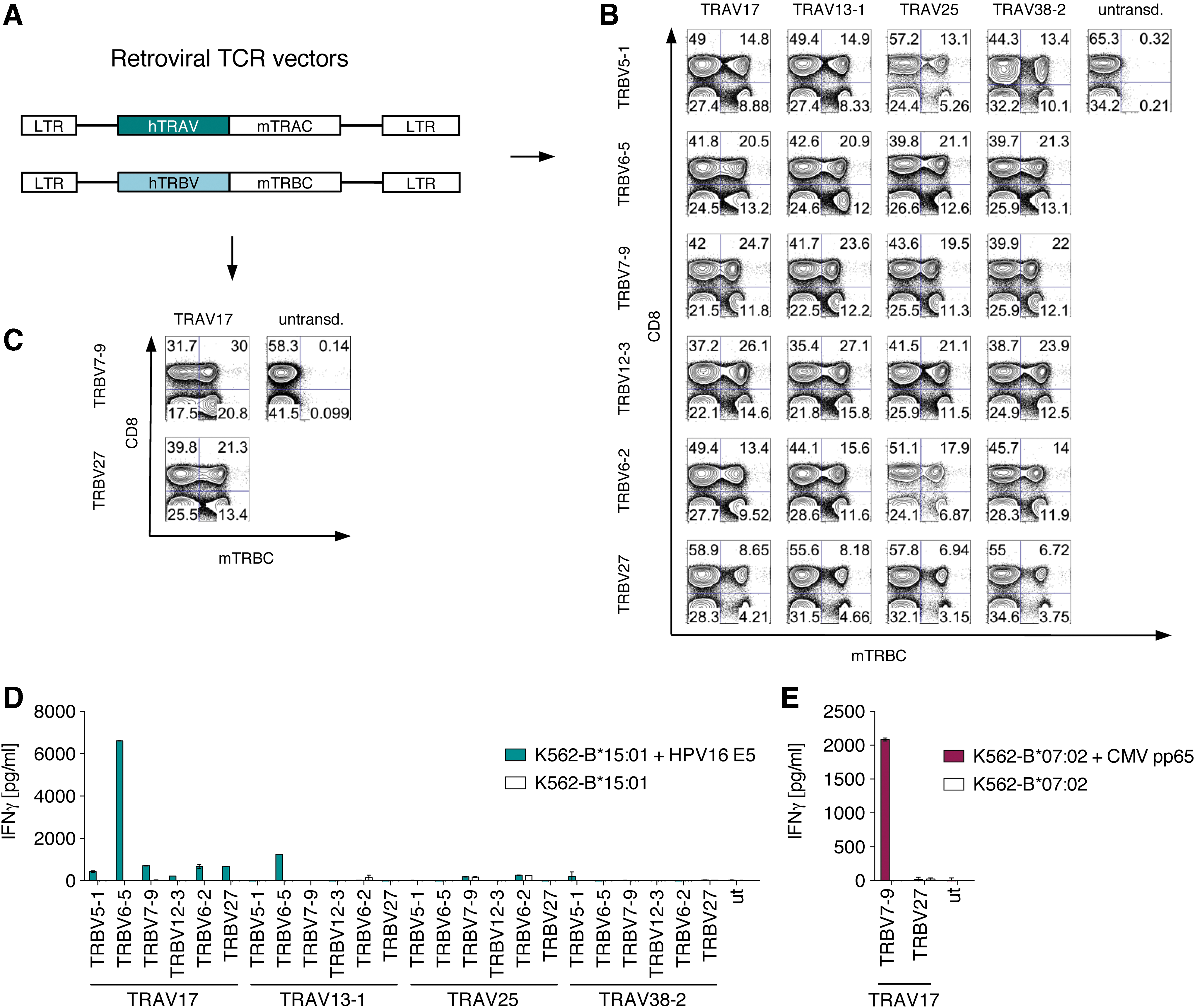
Re-expression of TCR chains in peripheral blood lymphocytes and analysis of TCR function.
To enable reproducibly high TCR transduction rates of primary T cells, TCR transgene cassettes were further optimized by linking TRB genes via a P2A element with TRA genes in the retroviral vector MP71 and applying codon optimization (Supplementary Fig. S4A). 26,27 Retroviral transduction of primary T cells with optimized TCR transgene cassettes yielded overall TCR transduction rates of around 40% (Supplementary Fig. S4B), and T cells were functional, as they specifically recognized E5-B*15:01- and pp65-B*07:02-expressing K562 cells (Supplementary Fig. S4C). T cells transduced with optimized TCR transgene cassettes were used for all subsequent experiments.
Epitope mapping
Functional TCRs were selected for recognition of immunodominant pMHC combinations without prior knowledge of epitope specificity. To map the exact epitope sequence recognized by the E5 TCR (Fig. 1E), 3′-truncated minigenes of the E5 oncogene were expressed in K562-B*15:01 cells using MP71 retroviral vectors. Additionally, a codon-optimized variant of E5 was used to test if the target epitope was expressed from an alternative reading frame (Fig. 4A). 39 T cells transduced with the optimized E5 TCR recognized target cells carrying the 189 nt short minigene of E5, but did not recognize target cells expressing minigenes shorter than 126 nt (Fig. 4A). Thus, the E5 TCR was specific for an epitope translated from the 126–189 nt coding region of E5, corresponding to the amino acid (aa) sequence 42–63. NetMHCpan3.0-based online binding prediction of all possible 9-mer and 10-mer peptides from the E5 oncoprotein returned six epitopes as potential binders to B*15:01 in the aa stretch 42–63 (Table 1). The top three predicted peptides of this aa stretch were used to pulse K562-B*15:01 target cells (Fig. 4B). E5 TCR-transduced T cells did not recognize target cells pulsed with the LLWITAASAF peptide and un-pulsed target cells, while releasing high amounts of cytokine upon encounter with target cells displaying the SAFRCFIVY (SAF) 9-mer and its corresponding 5′-elongated ASAFRCFIVY (ASAF)10-mer epitope (Fig. 4B). Thus, the E5 TCR specifically recognized the minimal-required SAF 9-mer epitope presented by B*15:01, revealing a previously unknown immunogenic HPV16 target for T cells. For pp65, it had been described that the TPRVTGGGAM (TPRV) epitope presented by B*07:02 is a frequent target for pp65-specific T cells. 40 To assess if this epitope was also the target for the pp65 TCR, K562-B*07:02 cells were pulsed with this TPRV 10-mer epitope (Fig. 4B) and co-cultured with pp65 TCR-transduced T cells. Indeed, high amounts of IFN-γ release confirmed that the TCR was specific for the TPRV epitope and that the approach yielded a functional TCR specific for an immunodominant pMHC combination (Fig. 4B).
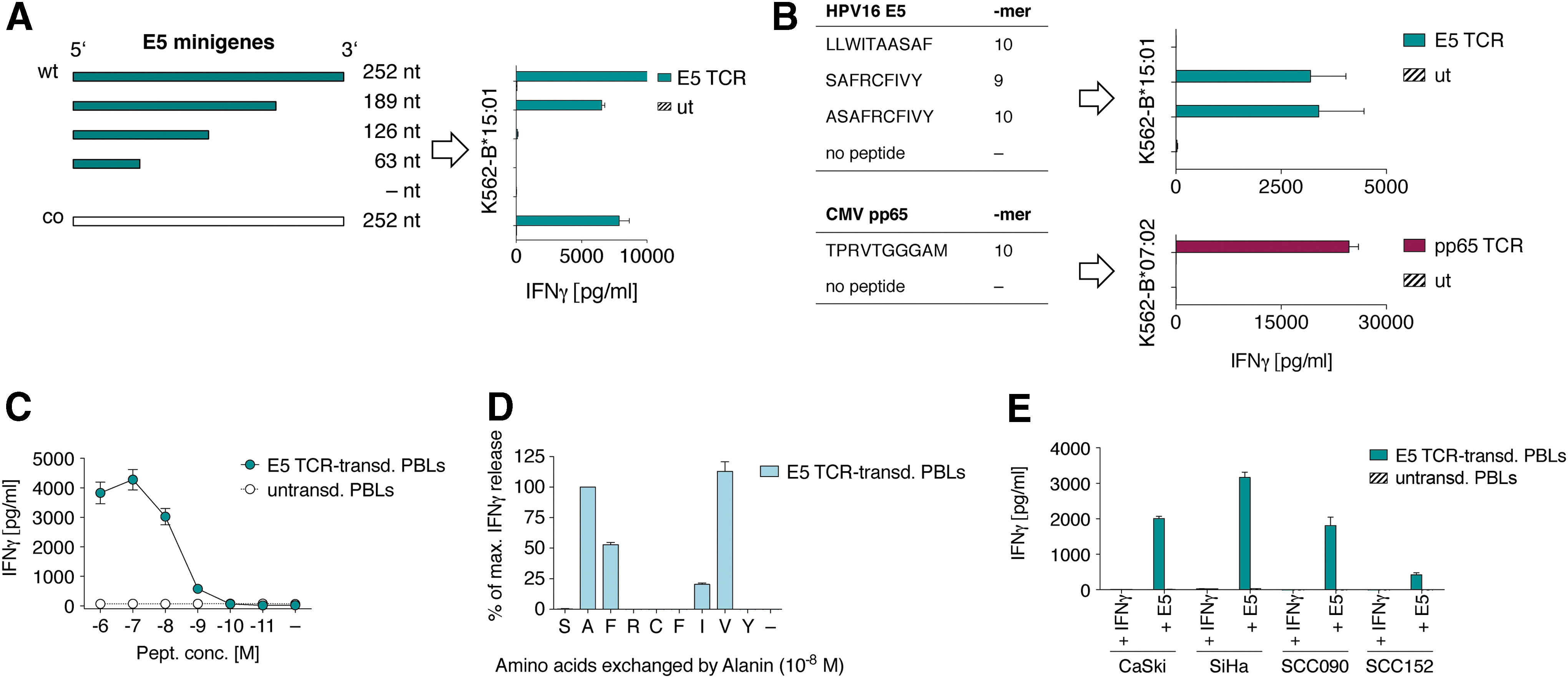
Epitope mapping and TCR characterization
HPV16 E5 binding prediction to HLA-B*15:01
nM, nanomolar affinity of peptide to MHC; rank, percentage rank compared to training database of 180,000 peptides; SB, strong binder; WB weak binder; bold, epitope recognized by E5-TCR.
Preclinical TCR characterization
The E5 TCR was assessed for its pMHC peptide sensitivity as a reflection of TCR affinity (Fig. 1F). Optimal affinity is a key characteristic of a useful TCR, which, upon expression in T cells, contributes to overall T-cell avidity that in turn impacts on clinical efficacy. Very high affinities/avidities contributed to off-target toxicities in clinical trials, 41,42 whereas lower affinities/avidities reduced efficacy of TCR immunotherapy. E5 TCR affinity to pMHC was measured in a peptide titration assay. TCR-transduced T cells recognized K562-B*15:01 target cells at peptide concentrations as low as 10–9 M. Half maximum IFN-γ release was determined between 10–8 and 10–9 M, indicating a naturally high TCR affinity to pMHC 43 (Fig. 4C). Although no results indicated potential cross- or background reactivity of the E5 TCR, we investigated whether sequences in the human proteome might be potential off-targets. To identify the aa of SAF essential for TCR recognition or MHC binding, an alanine scan was performed, whereby each aa of the SAF epitope was substituted by alanine (A). Residues at positions p2 (already A), p3 (F), p7 (I), and p8 (V) could be replaced by alanine without fully abrogating TCR-mediated antigen recognition (Fig. 4D). The Expitope online webtool 19 was used to find epitopes in the human proteome similar to SafRCFivY, where lowercase positions were allowed to be replaced by any aa. The Expitope algorithm found two epitopes with five of nine fixed aa shared with SAF. Both epitopes had very low overall expression scores in human tissues and very low MHC binding affinities to B*15:01, which most likely excludes them as potential off-targets for the E5 TCR (Supplementary Table S3).
The screening for antigen-specific T cells relied on the antigen-processing capacity of K562 cells. To exclude that the SAF epitope could only be artificially generated by K562 cells, a panel of B*15:01-positive LCLs was transfected with the E5 antigen and analyzed for recognition by E5 TCR-transduced T cells. Supplementary Figure S5 shows that all LCLs were recognized, while B*15:01-negative LCLs were ignored. Since the LCLs were derived from unrelated donors, the results also indicate that the SAF epitope was readily processed by cells of different genotypic origin.
HPV16 E5 is involved in early tumorigenesis, 44 which may explain why established tumor cell lines naturally expressing HPV16 E5 are not available. As an alternative, to analyze if HPV16-positive patient-derived cervical and head and neck cancer cell lines have capacity to process and present SAF naturally, they were transfected with E5 ivtRNA and B*15:01 ivtRNA, as all cells were negative for this allele. Figure 4E shows that cell lines CaSki, SiHa, SCC090, and SCC152 could each efficiently process and present the SAF epitope on B*15:01, thereby inducing IFN-γ release by E5 TCR-transduced T cells.
Discussion
Compared to the vast number of polypeptide sequences in a cell's proteome, only a small proportion of epitopes (1,000–10,000) is presented on the estimated 100,000 MHCs at the cell surface, 45,46 and presumably most epitopes are derived from self-proteins. Degradation of an antigen, TAP transport, peptide trimming, and binding affinity to the six available MHCs alleles influence the formation of immunogenic pMHC complexes eligible for T-cell attack. Reliable prediction programs for these different complex processing steps are lacking, and MHC binding prediction programs are less well trained on non-HLA-A*02:01 alleles. Considering all possible pMHC combinations that can be formed from one antigen sequence and six different MHCs, it remains a challenge to determine which pMHC(s) are finally adequately expressed and able to trigger TCR-dependent T-cell activation.
In general, several approaches are available to generate antigen-specific T cells and to isolate the respective TCRs for immunotherapy. First, allogeneic priming can be employed to obtain allo-restricted antigen-specific T cells. 9 A challenge in this approach is the separation of tumor antigen-specific T cells from those recognizing the allo-MHC, irrespective of the presented peptide. Second, T cells can be isolated from mice that recognize human target proteins. However, human T cells, which are genetically engineered with murine TCRs, may not persist in humans and may become a target for cytotoxic T cells or antibodies, as recently shown. 47 Third, a transgenic mouse carrying the full human TCR repertoire can be used to generate high-affinity TCRs. Here, the antigenic peptide that is used for immunization of the mouse has to differ from the murine counterpart in its amino acid sequence to induce an immune response. 48 Unfortunately, at present, this elegant approach is not available for other HLA alleles than HLA-A*02:01.
Here, a platform approach is presented that enables detection, isolation, and characterization of antigen-specific TCRs in the absence of predetermined target epitope and MHC binding information, instead allowing APC function to reveal the best-suited antigenic pMHC(s) for surface expression. The isolation of the E5 TCR specific for the SAF/B*15:01 pMHC showed the potential of this platform approach to identify a TCR specific for a previously unknown viral target pMHC. Seventeen epitopes other than SAF were also ranked in the best 0.6% of peptides predicted to bind to the six study MHCs compared to the training database of netMHCpan3.0 (Supplementary Table S1), but only SAF was found to stimulate T cells as an immunodominant epitope. Interestingly, the strategy described here would also enable detection of T-cell responses to proteasome-spliced epitopes—a class of epitopes that cannot be predicted by currently available algorithms. 49,50
This platform can be used to identify novel pMHC combinations as therapeutic targets for TCR immunotherapy. The authors are currently also developing an MHC II cell library to enable the identification of novel MHC II epitopes as targets for CD4-based TCR immunotherapy.
In the future, optimal personalized TCR immunotherapies will require the means to generate sets of TCRs targeting different pMHCs that are efficiently expressed on the surface of cancer cells. Patients for TCR immunotherapy can then be grouped according to MHC expression, allowing enlargement of patient groups eligible for treatment.
Footnotes
Acknowledgments
The authors wish to thank Christin Hesse, Kordelia Hummel, Mareen Kamarys (all MDC Berlin), and Barbara Mosetter (Helmholtz-Zentrum München) for their excellent technical assistance, and Hans-Peter Rahn (MDC) and his team for assistance with flow cytometry and sorting. Thanks to Matthias Leisegang (Charité Universitätsmedizin Berlin) for critical discussion. This work was supported by the following grants: Deutsche Forschungsgemeinschaft SFB-TR36 (D.J.S. and W.U.), Wilhelm Sander-Stiftung (D.J.S. and W.U.), and SPARK Berlin (Stiftung Charité; F.K.M.L.). F.K.M.L. was supported with an ATTRACT Early Stage Research Fellowship from the EU 7th framework program.
Author Disclosure
F.K.M.L., W.U., C.E., and D.J.S. are inventors on a patent covering the technology described in this study. At present, C.E., S.W. and D.J.S. are employees of Medigene Immunotherapies GmbH. W.U. is consultant of Medigene Immunotherapies GmbH. No competing financial interests exist for E.K. and M.L.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
