Abstract
Doxycycline (DOX) is widely used as a pharmacological agent and as an effector molecule in inducible gene expression systems. For most applications, it is important to determine whether the DOX concentration reaches the level required for optimal efficacy. We developed a sensitive bioassay for measuring the DOX concentration in biological samples. We used a modified HeLa cell line with the luciferase reporter gene under the control of the DOX-inducible Tet-On system for regulation of gene expression. These HeLaDOX cells constitutively express a novel variant of the rtTA transcriptional activator protein that is highly DOX-sensitive. Incubation of the cells with a DOX-containing biological sample triggers luciferase expression, which can be quantitated by standard methods. This bioassay is sensitive, with a DOX detection limit of 22 ng/ml in plasma. The assay was used to determine the DOX concentration in plasma derived from DOX-treated rhesus macaques and mice. Furthermore, we found that the DOX concentration in murine cerebrospinal fluid is 31-fold lower than the concurrent plasma DOX level. This bioassay for the quantification of DOX concentration in biological samples has several advantages over high-performance liquid chromatography-based and microbiological assays: (1) multiple samples can be assayed in a single experiment; (2) only small sample volumes are required; (3) the assay has a low detection limit; and (4) the assay can be performed in any cell culture laboratory.
Introduction
Oral administration of DOX has proven to be safe and effective in most applications. However, bioavailability may vary between different animals and under different conditions. For this reason, it may be necessary to confirm that the required DOX concentration is reached. Several methods have been used to measure DOX in biological samples, including high-performance liquid chromatography (HPLC) (Pijpers et al., 1991; Liu et al., 2001), microbiological assays (Abd El-Aty et al., 2004; Womble et al., 2007), thin-layer chromatography (Weng et al., 1990), fluorometric methods (van den Bogert and Kroon, 1981), and photometric techniques (Lopez Paz and Martinez, 1993).
HPLC has been widely used for DOX measurement because it is a sensitive and specific technique. However, it also has some major disadvantages: (1) it requires the removal of protein from the samples, which makes it labor-intensive and therefore less suitable for large-scale sampling; (2) it requires large sample volumes, which are often not available; and (3) it requires HPLC equipment, which is often not present in a standard molecular biology/cell culture laboratory. Microbiological assays have also been frequently used. They are based on the inhibition of a microorganism (e.g., Bacillus cereus or Bacillus subtilis) in an agar well diffusion system. These assays are not specific because other antibiotics present in the sample may also inhibit bacterial growth. The fluorometric and photometric techniques are sensitive for interference by other compounds.
In this study, we have developed the HeLaDOX reporter cell-based assay that uses luciferase expression as a read-out. This HeLa-derived cell line stably expresses an rtTA variant that is highly DOX sensitive (Zhou et al., 2006). Incubation of these cells with DOX triggers luciferase expression from an integrated gene under the control of the Tet-On system. We used the HeLaDOX assay to measure the DOX concentration in various body fluids from DOX-treated rhesus macaques and mice.
Materials and Methods
Cell culture
HeLa X1/6 cells (Baron et al., 1999) are derived from the HeLa cervix carcinoma cell line and harbor chromosomally integrated copies of the CMV-7tetO-promoter/firefly luciferase construct pUHC13-3 (Gossen and Bujard, 1992). HeLa X1/6 cells and human embryonic kidney (HEK) 293T adherent cells were grown with 5% CO2 at 37°C as a monolayer in Dulbecco's modified Eagle's medium containing 10% fetal calf serum (FCS), minimal essential medium nonessential amino acids, penicillin (100 U/ml), and streptomycin (100 μg/ml) (all from Invitrogen, Breda, The Netherlands).
Generation of HeLaDOX cells
The SpeI–PstI fragment of the lentiviral vector plasmid TRECMVR2 (pRRL cPPT TRE d2eGFP CMV rtTA2S-S2 PRE LTR SIN) (Markusic et al., 2005), which contains the rtTA2s-S2 gene, was inserted into the plasmid pBluescript SK+ to form a shuttle vector. The rtTA-V10 and rtTA-V16 sequences were isolated as XcmI–SmaI fragments from the corresponding pCMV-rtTA plasmids (Zhou et al., 2006) to replace the rtTA sequence of the shuttle vector. These rtTA sequences were subsequently cloned into TRECMVR2, using SpeI and PstI sites. All constructs were verified by sequence analysis.
Lentiviral vector was produced as previously described (ter Brake et al., 2006). Briefly, HEK 293T cells were transfected with the lentiviral vector plasmid and packaging plasmids pMDLg/pRRE, pRSV-REV, and pVSV-G, using Lipofectamine 2000 (Invitrogen). Lentiviral vector-containing supernatant was collected, filtered, and stored at −80°C.
Lentiviral vector stocks were titrated on HeLa X1/6 cells at 1000 ng/ml DOX (Sigma, Zwijndrecht, The Netherlands). Subsequently, HeLa X1/6 cells were transduced at a multiplicity of infection (MOI) of 0.15 and cultured with DOX (1000 ng/ml). Three days posttransduction, live cells were isolated by fluorescence-activated cell sorting (FACS) with a FACSAria cell sorter interfaced to the Diva software (BD Biosciences, San Jose, CA). Single green fluorescent protein (GFP)-positive cells were selected and expanded to generate cell clones.
Sample collection
Rhesus macaques (Macaca mulatta; 3.9–4.9 kg) were housed and maintained in accordance with Home Office guidelines for the care and maintenance of primates. Fifty milligrams of DOX (Centaur, Somerset, United Kingdom) in fruit juice was administered orally once per day. Before and after 5 days of DOX administration, blood samples were collected in EDTA bottles by femoral venipuncture.
BALB/c H-2d Rag2−/−γc −/− immunodeficient mice (Weijer et al., 2002) were bred and maintained in isocages in the animal facility of the Academic Medical Center (Animal Research Institute of Amsterdam, Amsterdam, The Netherlands), and were fed autoclaved food and water. At the onset of DOX treatment, the food was replaced with DOX-supplemented food (6 g/kg food; Bio-Serv, Frenchtown, NJ) for 9 days. The DOX-supplemented food was refreshed once per week. At the end of the DOX treatment, blood samples were collected from the saphenous vein, using Microvette CB300 lithium heparin-coated tubes (Sarstedt, Etten-Leur, The Netherlands) and cerebrospinal fluid (CSF) was sampled from the cisterna magna under anesthesia with isoflurane. In most cases 3–5 μl of CSF could be obtained.
The plasma was harvested after centrifugation of blood samples at 1400 × g for 8 min and stored at −80°C.
HeLaDOX assay
HeLaDOX cells were seeded in a 96-well plate at a density of 10,000 cells per well in a total volume of 180 μl of culture medium. The next day, plasma and CSF samples were diluted in culture medium to fit in the linear part of the regression curve and 20 μl of sample was added to the cells. All measurements were performed in duplicate. To generate a standard curve, cells were supplemented with 20 μl of culture medium containing DOX at 0–1000 ng/ml for each 96-well plate. Cells were cultured for 2 days and washed with 200 μl of phosphate-buffered saline (PBS) and lysed in 50 μl of passive lysis buffer (Promega, Leiden, The Netherlands) for 20 min on a shaker. Firefly luciferase activity was determined with a luciferase assay kit (Promega). DOX concentrations were interpolated from the DOX standard curve by linear regression analysis. The effect of plasma on luciferase activity measurements was analyzed by two-way analysis of variance (ANOVA) followed by Bonferroni analysis. A p value < 0.05 was considered to be statistically significant.
Results and Discussion
We generated a HeLa-based reporter cell line that expresses luciferase in a DOX-dependent manner. For this purpose, we transduced HeLa X1/6 cells, which already contain a chromosomally integrated tetO-promoter/luciferase-reporter construct, with the TRECMVR2 lentiviral vector. This vector contains an rtTA expression cassette and rtTA production is driven by the constitutively active immediate-early enhancer/promoter of cytomegalovirus (CMV) (Fig. 1). We tested two new rtTA variants that exhibit increased activity and greatly improved DOX sensitivity (rtTA-V10 and rtTA-V16) (Zhou et al., 2006). The TRECMVR2 lentiviral vector also contains a tetO promoter/GFP reporter gene that allows the selection of GFP-positive transduced cells on pretreatment with DOX (Fig. 1). Transduced cells were cultured with DOX (1000 ng/ml) for 2 days. Subsequently, fluorescence-activated cell sorting (FACS) was used to determine intracellular GFP expression and we collected individual cells that exhibited high GFP expression. As shown in Fig. 2A and B, transduction efficiency was comparable for the rtTA-V10 and rtTA-V16 vectors (18.5 and 16.6% GFP+ cells, respectively). Individual cells were expanded for 2 weeks in medium without DOX to obtain clonal cell cultures. DOX-dependent luciferase expression was determined for each clone after culturing the cells for 2 days with DOX at 0–100 ng/ml (Fig. 2C and D). Luciferase expression was low for nearly all cell clones without DOX and gradually increased with increasing DOX levels. No major differences were apparent between the V10 and V16 clones, although the mean background activity without DOX was a little higher for the V16 cells.
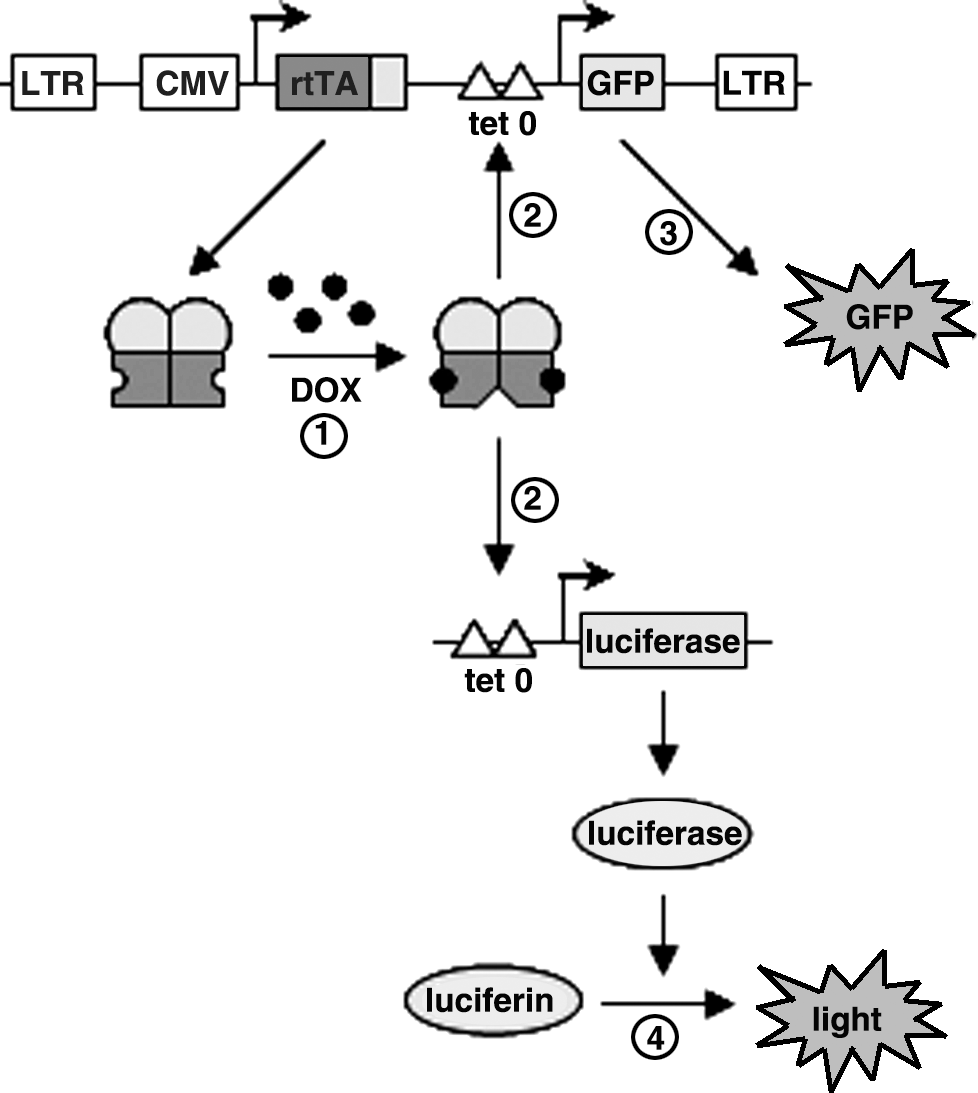
Detection of DOX using HeLaDOX cells. HeLa X1/6 cells, which contain a chromosomally integrated tetO promoter/luciferase construct, were transduced with the TRECMVR2 lentiviral vector expressing rtTA under the control of the CMV immediate-early enhancer/promoter and GFP under the control of a tetO promoter. These cells constitutively express rtTA. Addition of DOX (1) leads to binding of rtTA to the tetO promoter, which initiates expression of luciferase and GFP (2). GFP expression can be measured by FACS or fluorometry (3) and luciferase expression can be measured in a luciferase assay by luminometry (4). Comparison of the luciferase activity observed on sample addition with the activity obtained with a standard DOX range allows the quantification of DOX in the sample.
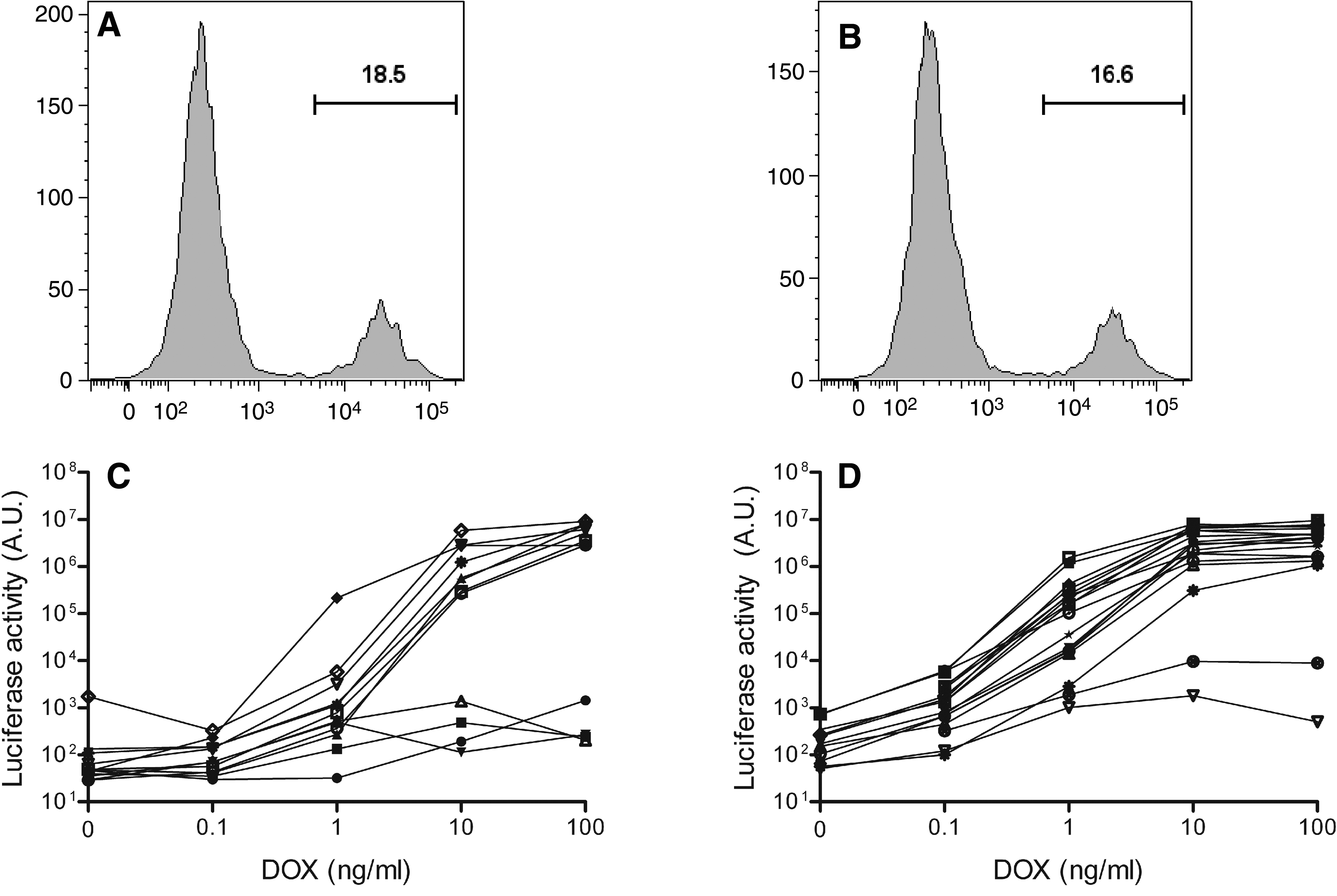
Selection of HeLaDOX cells. (
We selected a single V10 and V16 cell clone with low background activity and high DOX-induced luciferase activity. These clones demonstrate a similarly high luciferase level at 100 ng/ml DOX (Fig. 3A). The V10 cell line requires 3 ng/ml DOX for induction of luciferase expression, which peaks at 30 ng/ml DOX. Luciferase expression in the V16 cell line is activated already at 0.3 ng/ml DOX and reaches a maximum at 10 ng/ml DOX. The V16 clone is thus more DOX sensitive than the V10 clone, which is in agreement with the higher DOX sensitivity of rtTA-V16 (Zhou et al., 2006). For this reason, the V16 cell line (which we named HeLaDOX) was selected for further testing and development of the DOX bioassay. To demonstrate the specificity of this reporter assay, we cultured HeLaDOX cells with the DOX-related compounds tetracycline and minocycline, which do not or only marginally activate rtTA-V16 (our unpublished results). These two compounds did not or hardly induced luciferase expression as compared with DOX (Fig. 3B).
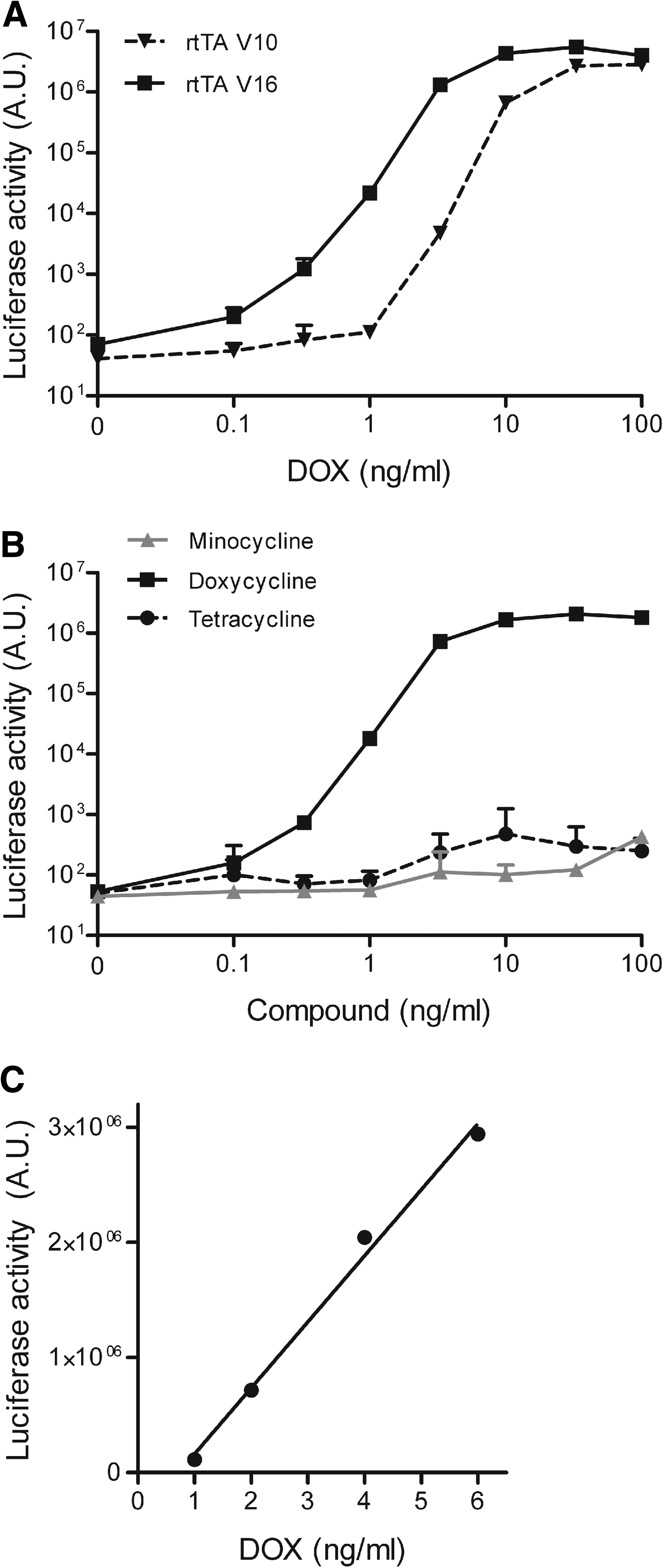
Sensitivity and specificity of the HeLaDOX assay. (
To measure DOX in biological samples such as cell culture medium or body fluids, the HeLaDOX cell cultures were supplemented with the test sample and analyzed for luciferase expression after 2 days. Luciferase expression was compared with the values obtained with a standard DOX range, which allows quantification of the DOX concentration in the sample. Linear regression analysis of the standard curve showed significant linearity from 1 to 6 ng/ml DOX (r 2 = 0.94, p < 0.0001) (Fig. 3C). It is important that luciferase reporter expression is not affected by other components present in the test sample (e.g., cytokines). This was tested with plasma samples derived from rhesus macaques. HeLaDOX cells were cultured in the presence of various amounts of plasma (final concentration in culture medium, 0–10%) and various DOX concentrations (final concentration, 0–10 ng/ml). Culturing with up to 4.5% plasma did not significantly affect background luciferase activity (at 0 ng/ml DOX) and DOX-induced luciferase expression. However, incubation of HeLaDOX cells with 10% macaque plasma significantly reduced luciferase production at 4 and 10 ng/ml DOX (Fig. 4A). For this reason, such high amounts of plasma (>4.5%) should be avoided in the DOX bioassay.
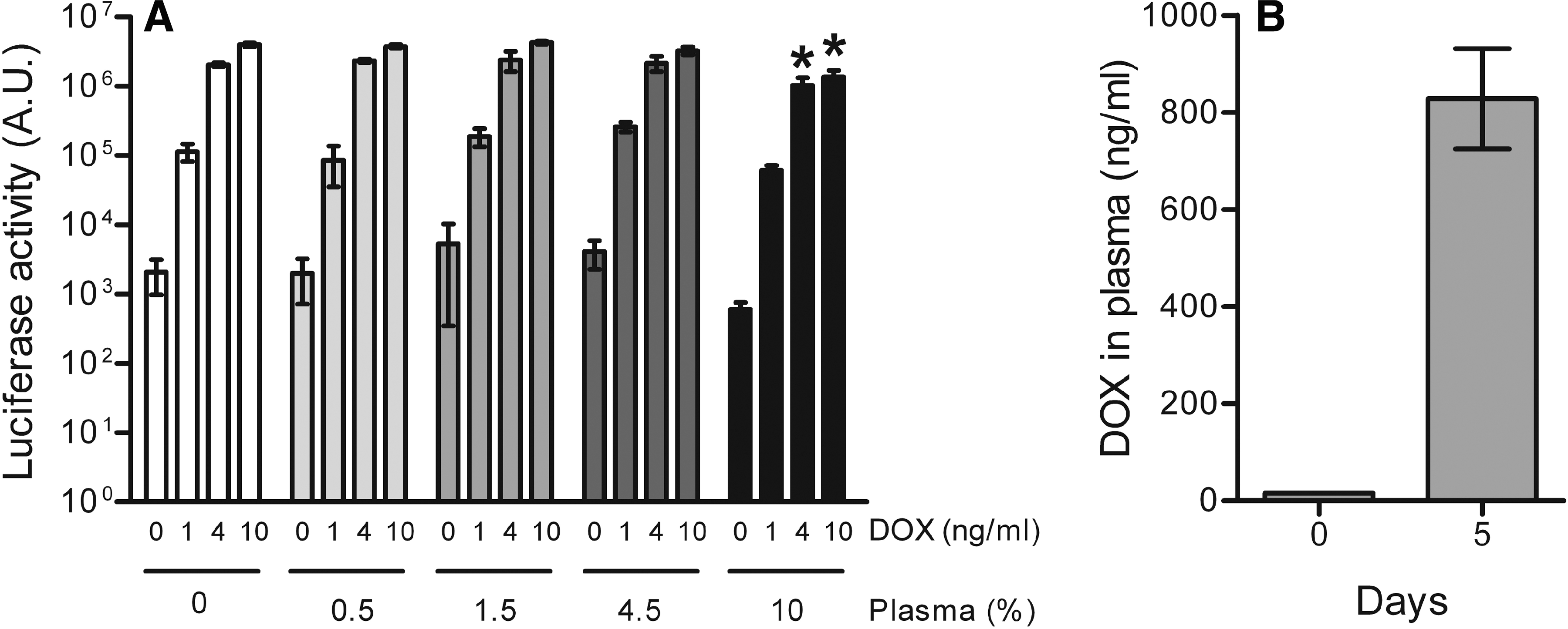
Determination of the DOX concentration in plasma of rhesus macaques. (
Because the HeLaDOX assay can detect 1 ng/ml DOX (final concentration in the cell culture; Fig. 3A and C) and the plasma should be diluted at least 22-fold (≤4.5% of the culture medium), our assay is able to detect down to 22 ng/ml DOX in plasma. This detection limit of the HeLaDOX assay outperforms other methods. For example, detection limits of 100–200 ng/ml DOX for HPLC-based methods (Karlsson et al., 1996; Axisa et al., 2000; Liu et al., 2001) and 200–250 ng/ml DOX for microbiological assays (Dotevall and Hagberg, 1989; Abd El-Aty et al., 2004) have been described.
We applied the HeLaDOX assay to determine the DOX concentration in plasma of naive and DOX-treated rhesus macaques (n = 4). Plasma samples, which were isolated before and after 5 days of daily DOX administration (50 mg orally), were diluted 2.2- to 20-fold in cell culture medium and 20 μl was added to 180 μl of HeLaDOX cell culture (final plasma concentration, 4.5–0.5%). Luciferase expression was measured after 2 days and compared with values obtained with the standard DOX range. Plasma of the DOX-treated rhesus macaques contained 838 ± 103 ng/ml DOX and no DOX was measured in naive animals (Fig. 4B). These DOX values are consistent with previous studies that quantitated DOX by alternative methods. Using a microbiological assay, Kelly and colleagues (1992) measured a DOX level between 810 and 1960 ng/ml in serum derived from macaques receiving 25 mg of DOX every 12 hr via orogastric administration (Kelly et al., 1992). Furthermore, daily administration of 5 mg of DOX to Sprague-Dawley rats (approximately 10 mg/kg body weight) resulted in serum DOX concentrations of 900 ng/ml as measured by HPLC (Liu et al., 2001).
The mouse is frequently used as an animal model in a variety of biological and gene therapy studies that employ the tetracycline-controlled gene expression system, for example, for regulation of transgene expression (Corbel and Rossi, 2002; Toniatti et al., 2004). The measurement of DOX levels in mice is limited because of the small volume of body fluids that can be obtained. We therefore used the HeLaDOX assay to determine the DOX concentration in the plasma and cerebrospinal fluid (CSF) of naive and DOX-treated mice. Plasma (∼20 μl) and CSF (3–5 μl) were isolated from mice (n = 7) that had received DOX-supplemented food (6 g/kg food) or control food for 9 days. The plasma DOX concentration as measured in the HeLaDOX assay was high in the DOX-treated mice (12,187 ± 2227 ng/ml) and below the detection level in mice that never received DOX (Fig. 5A). The DOX level in CSF (394 ± 104 ng/ml) of DOX-treated animals was 31-fold lower than in plasma (Fig. 5B), which is in accordance with previous reports that indicated that the blood–brain barrier restricts DOX diffusion into the brain (Barza et al., 1975; Andersson and Alestig, 1976). These results demonstrate that the sensitive HeLaDOX assay allows the accurate quantitation of DOX levels when only small test samples are available.
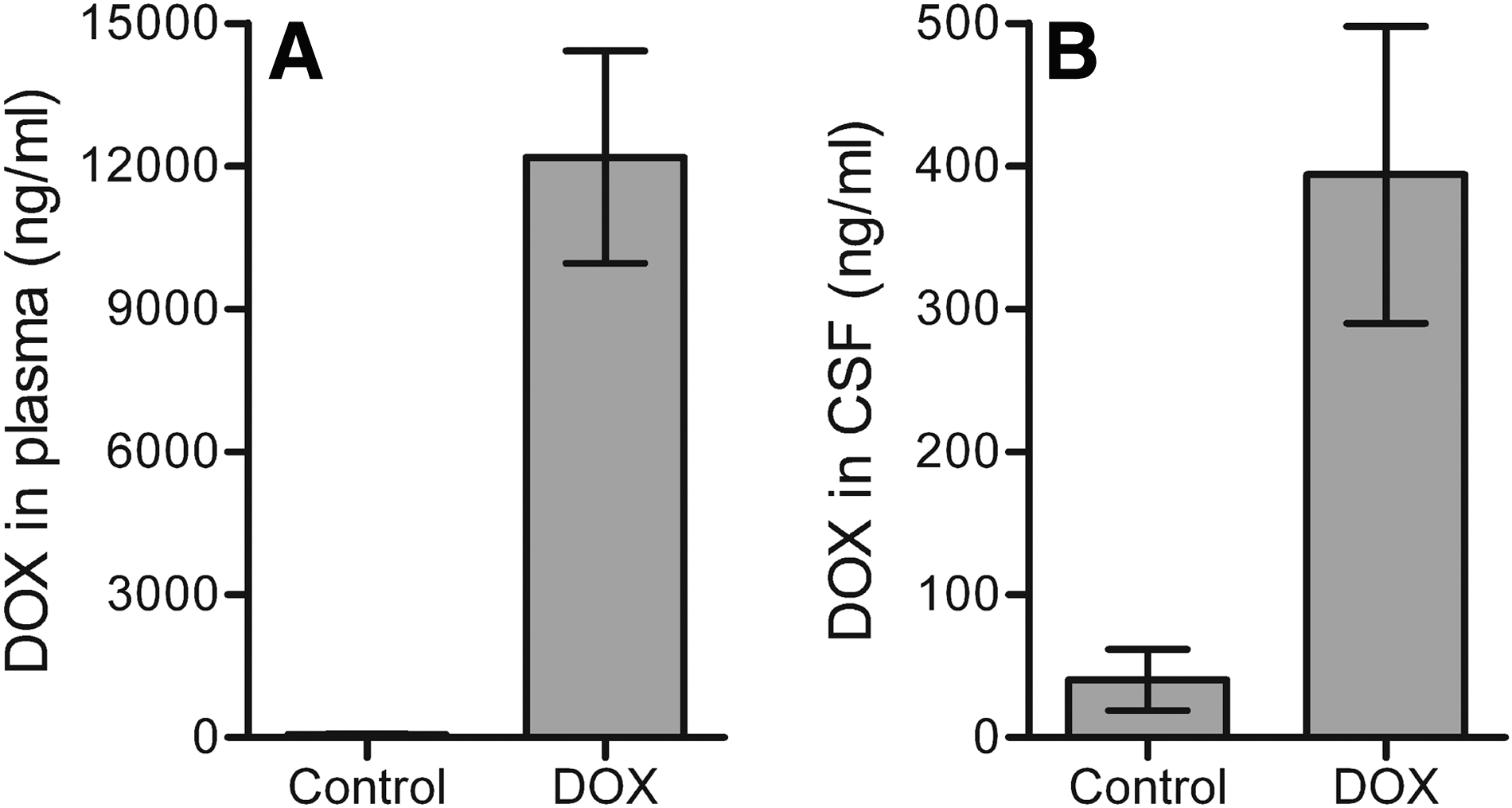
Determination of the DOX concentration in plasma and CSF of mice. The DOX concentration was determined in (
In conclusion, we have established the HeLaDOX cell line, which expresses luciferase on DOX induction. We here demonstrate that the luciferase activity measured in this DOX bioassay can effectively be used to determine the DOX concentration in small biological samples. Because these HeLaDOX cells also express GFP in a DOX-dependent way, measurement of GFP by fluorometry or FACS will also allow the quantification of DOX. These methods can be used in laboratories in which it is not possible to measure luciferase activity or if the biological sample contains a compound that specifically interferes with luciferase expression. Our HeLaDOX assay possesses major advantages as compared with conventional methods: (1) multiple samples can be assayed in a single experiment; (2) the assay requires only a small sample volume; (3) the assay has a low detection limit (22 ng/ml); and (4) the assay can be performed in any cell culture laboratory. This work forms a prelude to testing of our DOX-dependent variant of simian immunodeficiency virus (SIV-rtTA) in the rhesus macaque model (Das et al., 2007, 2008). The plasma DOX levels that we measured should be sufficient to drive the expression and replication of this unique virus, which will allow us to test the conditionally live virus vaccine approach and to study the immune correlates of protection.
Footnotes
Acknowledgments
The authors thank Christian Berens and Wolfgang Hillen (University of Erlangen, Germany) for their generous gift of HeLa X1/6 cells. This research was sponsored by the International AIDS Vaccine Initiative (IAVI) and the Technology Foundation STW (the applied science division of NWO and the technology program of the Ministry of Economic Affairs, Utrecht, The Netherlands). Mireille Centlivre is supported by a Marie Curie Intra-European fellowship (MEIF-CT-2007-039689). Nicolas Legrand is supported by the Bill and Melinda Gates Foundation, through the Grand Challenges in Global Health program (Human Vaccine Consortium).
Author Disclosure Statement
No competing financial interests exist.
