Abstract
The emergence of SARS-CoV-2 necessitated the rapid deployment of tests to diagnose COVID-19. To monitor the accuracy of testing across the COVID-19 laboratory network in Thailand, the Department of Medical Sciences under the Ministry of Public Health launched a national external quality assessment (EQA) scheme using samples containing inactivated SARS-CoV-2 culture supernatant from a predominant strain in the early phase of the Thailand outbreak. All 197 laboratories in the network participated; 93% (n=183) of which reported correct results for all 6 EQA samples. Ten laboratories reported false-negative results, mostly for samples with low viral concentrations, and 5 laboratories reported false-positive results (1 laboratory reported false positives and false negatives). An intralaboratory investigation of 14 laboratories reporting incorrect results revealed 2 main causes of error: (1) RNA contamination of the rRT-PCR reaction and (2) poor-quality RNA extraction. Specific reagent combinations were significantly associated with false-negative reports. Thailand's approach to national EQA for SARS-CoV-2 can serve as a roadmap for other countries interested in implementing a national EQA program to ensure laboratories provide accurate testing results, which is crucial in diagnosis, prevention, and control strategies. A national EQA program can be less costly and thus more sustainable than commercial EQA programs. National EQA is recommended to detect and correct testing errors and provide postmarket surveillance for diagnostic test performance.
Introduction
Reliable and accurate diagnostic testing for SARS-CoV-2 is essential for COVID-19 prevention and control strategies.1,2 Real-time reverse-transcription polymerase chain reaction (rRT-PCR) is currently the “gold standard” for confirmation of SARS-CoV-2 nucleic acid detection in suspected cases. As SARS-CoV-2 is a novel virus, many in-house and commercial assays were developed rapidly. By March 2020, Thailand established the Emergency Decree on Public Administration in Emergency Situation B.E. 2548, 3 which limited the ability of people to move between provinces as a disease control measure. Therefore, to ensure access to timely rRT-PCR diagnostics for SARS-CoV-2 within each province, Thailand's Department of Medical Sciences (DMSc) under the Ministry of Public Health certified at least 1 laboratory in each province to provide SARS-CoV-2 testing. It announced the policy as “One Lab, One Province, 24-Hour Results.” This approach established at least 1 rRT-PCR testing laboratory in every province and required results to be reported within 24 hours. 4 The laboratory network for COVID-19 was thus established, and included nearly 200 government and private-sector laboratories.
Several external quality assessment (EQA) schemes for SARS-CoV-2 molecular testing have been published, which vary in design depending on national policies and guidelines and country-specific situations. Different EQA schemes were designed to assess various aspects of the testing process. For example, 1 EQA scheme used panels composed of 12 samples, including 6 positive for SARS-CoV-2, at different concentrations (to assess assay sensitivity) and 6 containing non-SARS-CoV-2 respiratory viruses (to assess assay specificity). 5 Another scheme assessed stages of the testing process and included 2 SARS-CoV-2-positive EQA samples containing purified SARS-CoV-2 RNA (to assess performance of PCR amplification) and 2 positive EQA samples containing SARS-CoV-2 pseudovirus (to assess both RNA extraction and PCR amplification). 6 A third scheme addressed test performance in different specimen types and included 4 upper and 4 lower respiratory track specimens spiked with decreasing concentrations of SARS-CoV-2 culture supernatant or non-SARS-CoV-2 human coronavirus culture supernatant. 7 Finally, another scheme assessed the performance of a SARS-CoV-2 rRT-PCR assay developed in-house that was widely used across India in a variety of different laboratory settings. 8
Given the rapid development of Thailand's COVID-19 laboratory network, a quality assessment was urgently needed to ensure the accuracy and reliability of test results. DMSc assured the accuracy of COVID-19 diagnostics through 2 primary strategies: (1) validation and authorization of diagnostic kits through the Government Gazette by the Thailand Food and Drug Administration before sale in Thailand and (2) establishment of an EQA scheme to assess the ability of laboratories within the COVID-19 network to accurately differentiate SARS-COV-2-positive specimens from SARS-CoV-2-negative specimens through qualitative rRT-PCR. Thailand's SARS-CoV-2 laboratory network certification requires that laboratories conducting SARS-CoV-2 molecular diagnostics enroll in EQA 9 ; therefore, the creation of a national SARS-CoV-2 EQA program was needed to eliminate any cost barrier to EQA participation. This article presents results and lessons learned from the first round of the SARS-CoV-2 EQA program in Thailand.
Materials and Methods
EQA Sample Preparation and Characterization
The SARS-CoV-2 strain BetaCoV/Nonthaburi/74/2020 (accession ID: EPI_ISL_403963) was cultivated at the DMSc National Institute of Health (Thai NIH). The virus was grown in Vero cells (ATCC CCL-81), from which a viral supernatant was harvested, diluted with viral transport medium, and inactivated by RA1 lysis buffer from the NucleoSpin RNA kit (Macherey-Nagel, Düren, Germany). The EQA panel included 6 samples with decreasing viral load and 1 negative sample. The negative sample (sample A) was made from human epithelial type 2 (HEp-2) cell supernatant (ATCC CCL-23) and served as an internal process control. Positive samples (B through F) contained serial 10-fold dilutions of inactivated SARS-CoV-2 culture supernatant, including a low viral load positive sample, near the limit of detection for many assays (Table 1). All EQA sample aliquots, of 500 μl each, were frozen at -70°C until shipment. A commercial courier service was used to transport EQA samples on dry ice for arrival within 24 hours to participating laboratories. User instructions accompanying the EQA samples recommended that samples be handled following the same workflow and biosafety procedures as any clinical sample received for SARS-CoV-2 rRT-PCR. To prevent consultation on results of EQA samples among participating laboratories, samples were given unique 4-digit codes for each laboratory.
Characterization of EQA Samples and Qualitative Results From 197 Laboratories Participating in Thailand's First SARS-CoV-2 EQA Scheme
Abbreviations: Ct, cycle threshold; ddPCR, droplet digital polymerase chain reaction; EQA, external quality assessment; Hep-2, human epithelial type 2 cells; SD, standard deviation.
The Thailand National Institute of Health determined the average viral copy number of each positive EQA sample (B through F) by testing 2 aliquots of each by droplet digital PCR (ddPCR) method using the DMSc SARS-CoV-2 assay targeting the RNA-dependent RNA polymerase gene 10 and the One-Step RT-ddPCR Advanced Kit for Probes (Bio-Rad, Hercules, CA).
The homogeneity and stability of each EQA sample were tested as follows: for homogeneity, 10 aliquots of each EQA sample were selected at random using a function of Microsoft Excel and tested in duplicate by extraction with the NucleoSpin RNA kit and amplification by rRT-PCR using the DMSc SARS-CoV-2 assay. 10 To test the impact of storage conditions on EQA sample stability, 2 EQA sample sets were stored at -70°C until all participant result reports were received and then tested for stability using the DMSc SARS-CoV-2 assay.
Participant Laboratories and Result Reporting
EQA panels were sent to all 197 laboratories (comprising 114 public laboratories and 83 private laboratories) in the COVID-19 laboratory network on July 1, 2020; results were returned to the EQA provider by July 24, 2020. Most participating laboratories in the Bangkok Metropolitan Region were from the private sector, while most provincial laboratories were from the government sector (Table 2). If the laboratory routinely used more than 1 type of rRT-PCR assay, it was asked to test the panel with each kit and report all results. In addition to the rRT-PCR result for each EQA sample, laboratories were asked to submit the raw data files from the rRT-PCR instrument, information on testing protocols, extraction kit(s), amplification kit(s), target genes, cycle threshold for positive results, and interpretation of results. Participating laboratories submitted all data online through the DMSc-developed EQA program website and a preliminary performance report was sent by the EQA provider to all participating laboratories within 2 days following the deadline for result return. The respondents from each participating laboratory were asked to complete a survey about their satisfaction with the overall national EQA process (eg, communication and coordination between the EQA provider and participating laboratory, condition of EQA panel upon arrival, turnaround time of preliminary individual laboratory performance report, and any other suggestions for program improvement). About two-thirds (n=130, 66.0%) of the respondents were medical technologists; the rest were lab managers/chief lab officers (n=58, 29.4%), lab supervisors/quality control specialists (n=6, 3.1%), or and 3 university teachers (n=3, 1.5%).
Laboratories Participating in the First Round of the SARS-CoV-2 EQA Scheme
The government sector included 14 regional medical sciences centers, 1 laboratory of the Department of Medical Sciences, and 10 laboratories of the Office of Disease Prevention and Control, the Institute for Urban Disease Control and Prevention, Bamrasnaradura Infectious Diseases Institute, the Bureau of Tuberculosis, the Central Institute of Forensic Science, the Queen Sirikit National Institute of Child Health, and the Thai Red Cross Emerging Infectious Diseases Health Science Centre (King Chulalongkorn Memorial Hospital).
International public health organizations included the Thai Ministry of Public Health and US Centers for Disease Control and Prevention Collaboration and 4 Armed Forces Research Institute of Medical Sciences laboratories (3 in Bangkok and 1 in Kamphaeng Phet Province).
Laboratories reporting incorrect results were asked to conduct an internal investigation and report findings back to the DMSc EQA provider using the corrective action report as required by the ISO 15189 quality standard. 11 The DMSc EQA provider then sent additional EQA sample aliquots (recoded relative to the first EQA panel to ensure the laboratories could test the second set of EQA samples blindly) to these laboratories and asked that they test again and return results to the EQA provider by the established deadline.
Respondents from the participating laboratories provided informed consent and data were anonymized for analysis. The protocol for this study was reviewed and waived by the Research Ethics Committee of the DMSc, Thailand (Study Code No. 6/2021). US Centers for Disease Control and Prevention staff were not involved in data/specimen collection and did not have access to any personally identifiable data.
Data Analysis
Data received from participating laboratories were stored in a Firebase Realtime Database as JSON objects and converted to CSV files for data analysis using Python. We used a chi-square (significance level at .05) to test the relationship between 2 variables, the extraction kit/rRT-PCR kit combination and the qualitative result reported. We used the Fisher exact test when the sample size for a given extraction kit/rRT-PCR kit combination was less than 5 and the odds ratios (OR) to measure the probability of reporting an “incorrect” result using a specific extraction kit/rRT-PCR kit combination. Independent variables were the extraction/rRT-PCR method combination, and in a separate analysis, the individual reagent associated with reporting an incorrect result.
Results
EQA Sample Characteristics
Viral RNA concentrations for each positive EQA sample ranged from 1,000 to 5,540,000 copies/mL (Table 1). Cycle threshold (Ct) values of 10 randomly selected replicates for each EQA sample were within the acceptable range for homogeneity. The EQA sample with the lowest viral RNA concentration (F, 1,000 copies/mL) had the most observed variability in Ct values across replicates (SD=0.7). For stability testing, qualitative results were the same, and Ct values were within ±2 SDs before and after storage (data not shown).
SARS-CoV-2 Testing Protocols Reported by EQA Participating Laboratories
The SARS-CoV-2 molecular testing protocol is divided into 2 main steps: RNA extraction and rRT-PCR amplification. Participating laboratories reported using 28 different nucleic acid extraction kits and 21 different rRT-PCR assays (Tables 3 and 4). Nineteen laboratories reported using more than 1 rRT-PCR assay for molecular detection. Almost 86% (187 of 218) of the participating laboratories used commercial rRT-PCR assays while the remaining laboratories used in-house rRT-PCR assays. Of the 197 laboratories, 22 (11.2%) reported using 1 or more fully automated detection systems for SARS-CoV-2, including QIAstat-Dx respiratory panel (QIAGEN, Germany), Xpert Xpress SARS-CoV-2 (Cepheid, California), and cobas SARS-CoV-2 6800 system (Roche, Switzerland).
Extraction Kits Used by the Participating Laboratories
Fully automated detection system, which incorporates RNA extraction and amplification in 1 closed system.
The 5 most commercial nucleic acid extraction kits were magLEAD (ANALIS, Belgium); QIAgen including, QIAamp Viral RNA Mini Kit, and RNeasy Mini Kit (QIAGEN, Hilden, Germany); abGenix (AITbiotech, Singapore); FavorPrep (Favorgen, Taiwan); and MagNA Pure (Roche, Germany).
Amplification Kits Used by the Participating Laboratories
Note: The 5 most frequently used SARS-CoV-2 rRT-PCR assays were Sansure (Sansure Biotech, China), Allplex (Seegene, South Korea), DAAN (DaAnGene, China), DMSc in-house assay, and VIASURE (CerTest Biotec, Spain).
Fully automated detection system, which incorporates RNA extraction and amplification in 1 closed system.
Abbreviations: DMSc, Department of Medical Sciences; E, envelope; N, nucleocapsid; NS, nonstructure polypeptide of ORF1a; ORF1ab; open reading frame: ORF1a and ORF1b; RdRp, RNA dependent RNA polymerase; S, spike.
Performance of Participating Laboratories
All 197 laboratories returned results including 218 unique EQA reports (20 laboratories submitted more than 1 result report, since more than 1 rRT-PCR assay was used in their laboratory). Of these, 177 (89.9%) laboratories used only 1 rRT-PCR kit/assay, while 19 (9.6%) laboratories used 2 assays, and 1 (0.5%) laboratory used 3 assays. The laboratories using more than 1 assay showed no discrepant qualitative results across assays.
Overall, 183 (92.9%) laboratories reported correct qualitative results for all 6 EQA samples. Five laboratories reported a false-positive result for the negative EQA sample, and 10 laboratories reported a false-negative result for 1 or more of the SARS-CoV-2 positive samples; 1 laboratory reported both false-positive and false-negative results (Table 1). One laboratory reported incorrect results for 3 samples, 4 laboratories reported incorrect results for 2 samples, and 4 laboratories reported incorrect results for 1 sample.
Variation in Ct values for the same EQA sample was observed when tested using different rRT-PCR assays or modified versions of the same assay (Figures 1, 2, and 3). However, Ct variation did not lead to incorrect qualitative interpretation of results. Of the 19 laboratories that reported using the DMSc assay, 14 reported using the original rRT-PCR protocol while 5 laboratories reported using a modified rRT-PCR protocol. The modified protocol included additional cycling conditions that resulted in lower Ct values compared with the original protocol when testing the same EQA sample (Figure 3). However, the qualitative result did not differ for any EQA sample between the original and modified protocol.
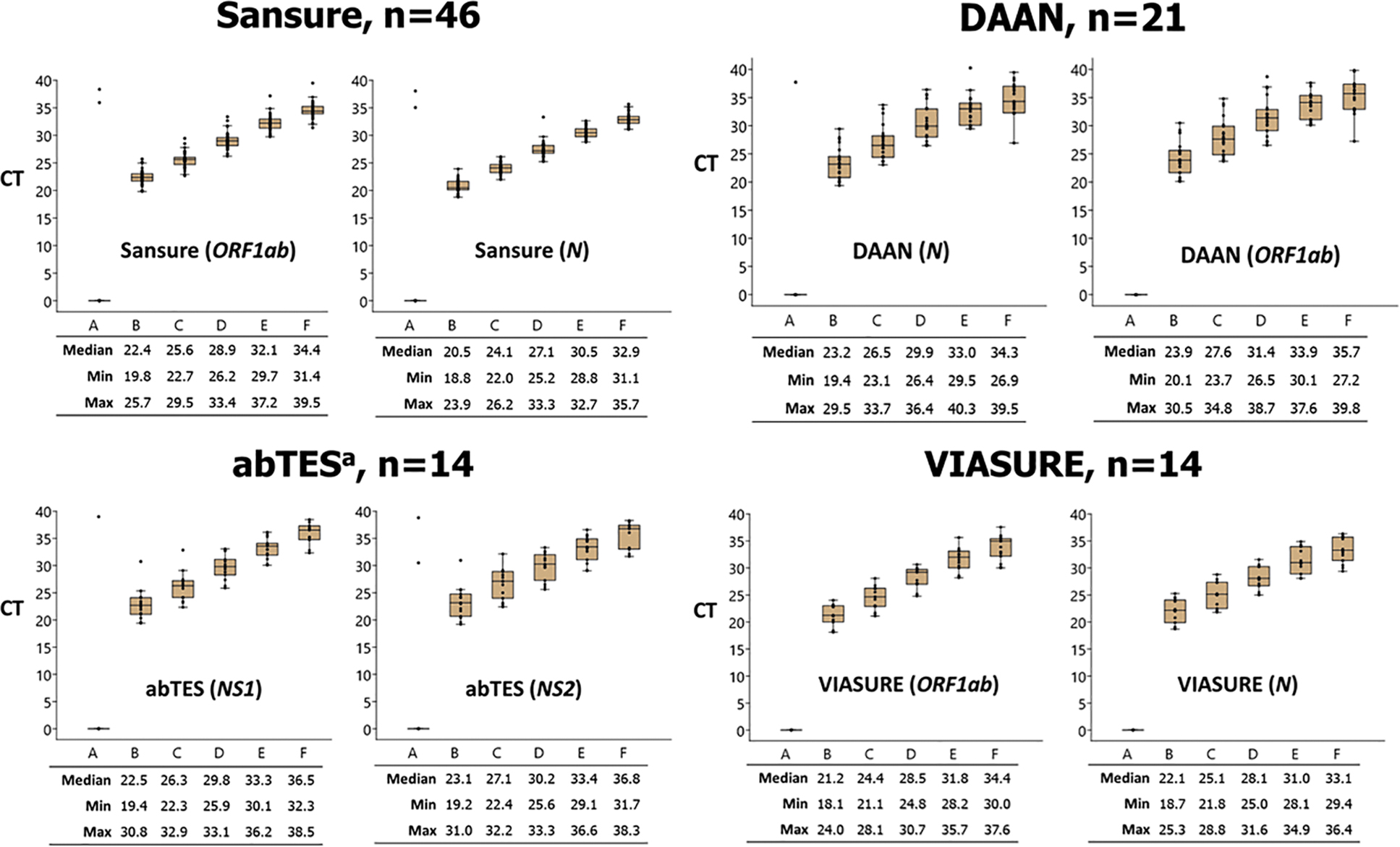
Ct values of the EQA samples tested by different rRT-PCR assay displayed in box plot. Note: n=number of laboratories using the given rRT-PCR assay (SARS-CoV-2 gene target); horizontal line within each box denotes the median Ct value; top and bottom of box indicates the third and first quartiles, respectively; error bars indicate minimum and maximum values; dots indicate outlier results.
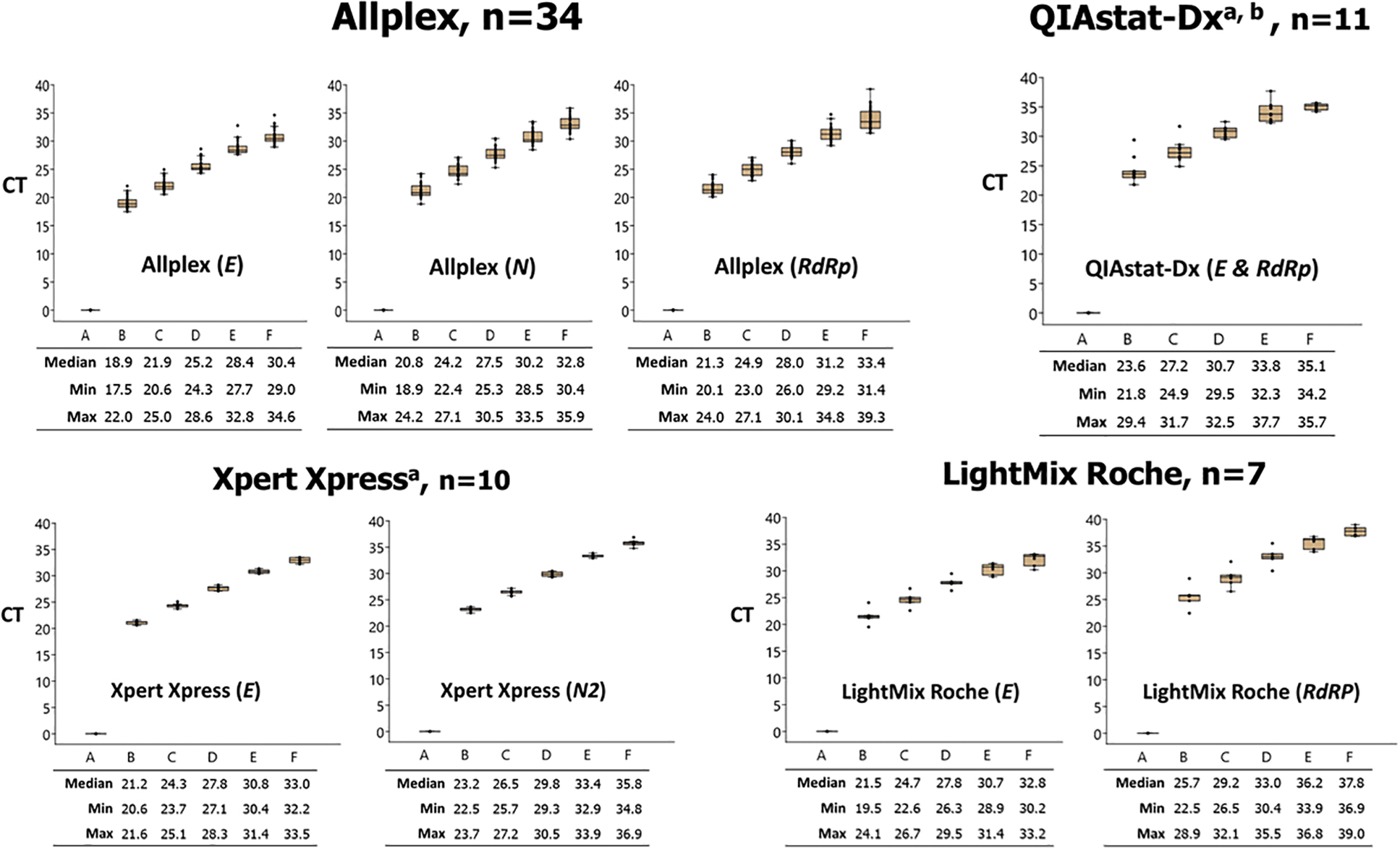
Ct values of the EQA samples tested by different rRT-PCR assay displayed in box plot. Note: n=number of laboratories using the given rRT-PCR assay (SARS-CoV-2 gene target); horizontal line within each box denotes the median Ct value; top and bottom of box indicates the third and first quartiles, respectively; error bars indicate minimum and maximum values; dots indicate outlier results.
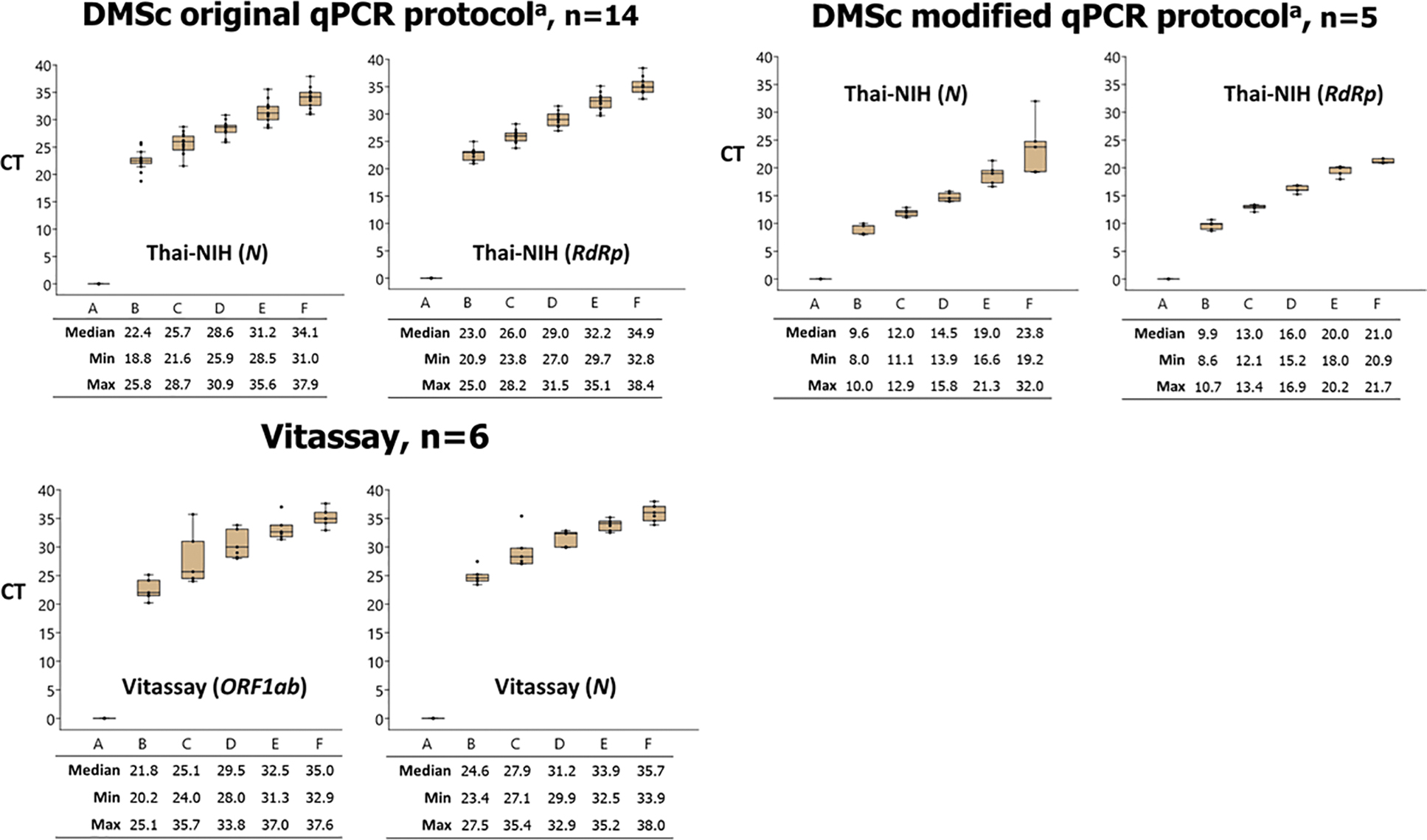
Ct values of the EQA samples tested by different rRT-PCR assay displayed in box plot. Note: n=number of laboratories using the given rRT-PCR assay (SARS-CoV-2 gene target); horizontal line within each box denotes the median Ct value; top and bottom of box indicates the third and first quartiles, respectively; error bars indicate minimum and maximum values; dots indicate outlier results.
Internal investigations of errors in laboratories reporting incorrect results revealed RNA contamination of the rRT-PCR assay and poor-quality RNA extraction as possible causes of error. Upon retesting by these laboratories using a new lot of RNA extraction reagents provided by the reagent distributor and new aliquots of the EQA sample provided by the EQA provider, 13 of 14 laboratories returned correct results. Additional follow-up interviews with the laboratory that had repeated incorrect results identified poor laboratory technique and switching of samples due to poor sample tracking throughout the testing process as likely causes of error.
Most (89%) of the participating laboratory respondents expressed overall satisfaction with the SARS-CoV-2 EQA process as indicated by their responses to the satisfaction survey that all participating laboratories were invited to respond to. Specifically, they expressed their appreciation for the fast turnaround time to receive the preliminary EQA performance report (all participating laboratories received their preliminary EQA performance report within 2 days of the deadline for laboratories to return their EQA results to the EQA provider). Some respondents recommended increasing the EQA sample volume to enable testing of the EQA samples using all SARS-CoV-2 rRT-PCR assays available in their laboratories.
Impact of RNA Extraction and rRT-PCR Assay Combinations on EQA Results
A total of 63 different extraction kit/rRT-PCR assay combinations were used by the participating laboratories. Figure 4 shows the relationship between RNA extraction kit/rRT-PCR assay combination and the accuracy of reported qualitative results. Used by 55 laboratories, magLEAD (Precision System Science, Japan) was the most common extraction kit; no laboratories using this kit reported an incorrect result. Sansure (Sansure Biotech Inc., China) was the most common rRT-PCR assay used (n=46 laboratories), with 2 laboratories reporting false-positive results. Allplex (Seegen, Inc., South Korea) was the second most used rRT-PCR assay (n=34 laboratories), with no laboratories reporting an incorrect result. The most common extraction kit/rRT-PCR assay combination was magLEAD/Allplex (n=31 laboratories) and none of the laboratories using this combination reported an incorrect result.
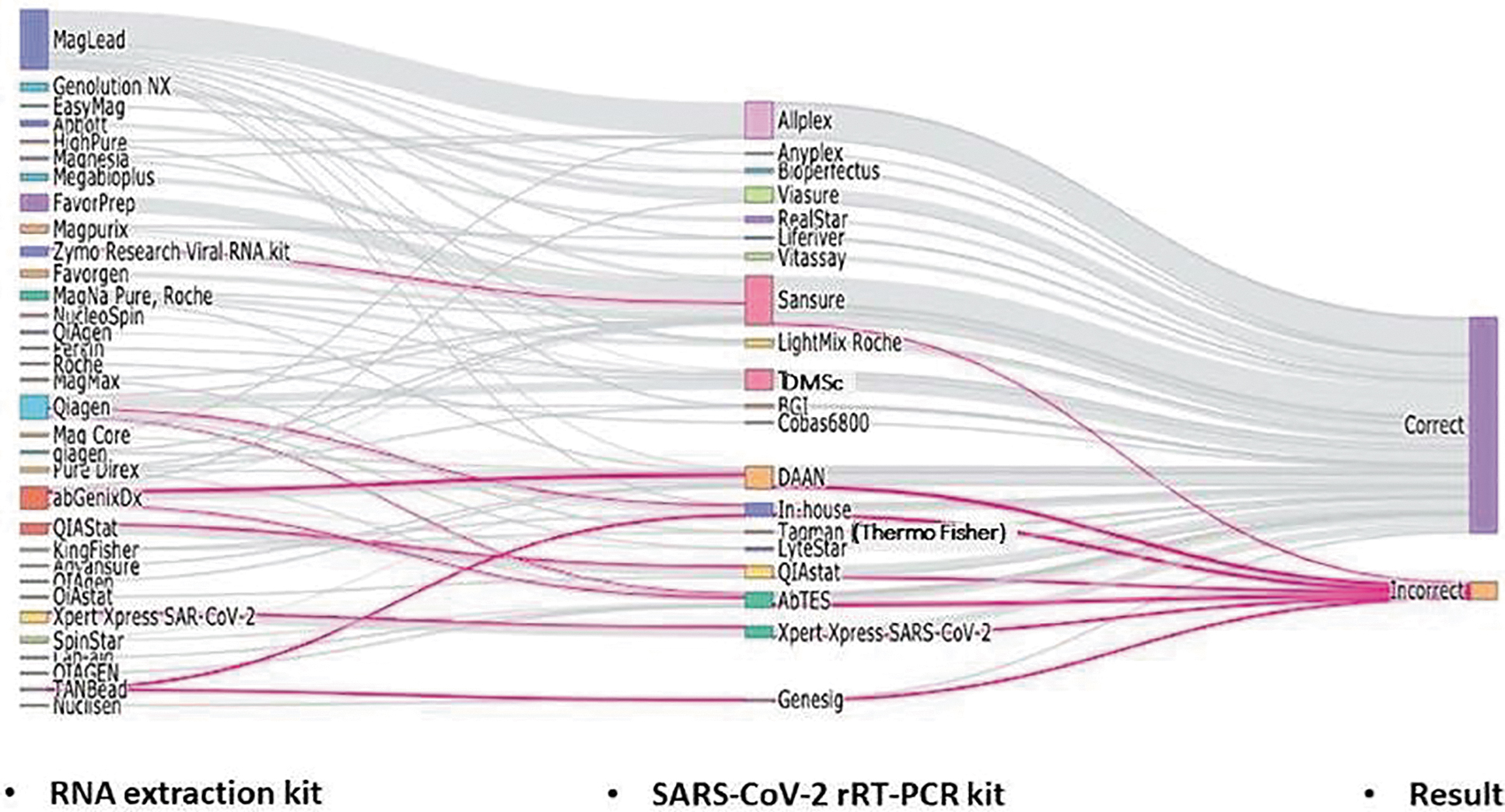
Relationship between RNA extraction kit/rRT-PCR assay combination and the accuracy of reported qualitative results. The weight of the lines reflects the number of laboratories using a particular extraction kit and rRT-PCR test. The pink lines highlight the extraction kit and rRT-PCR test combinations that were associated with incorrect results of some samples in some laboratories. Abbreviation: rRT-PCR, real-time reverse-transcription polymerase chain reaction.
Statistical analysis found that 7 of the 17 most frequently used extraction kit/rRT-PCR assay combinations were associated with incorrect qualitative results (Table 5). However, 2 combinations were significantly associated with incorrect results: abGenix (AITbiotech, Singapore)/DAAN (DaAnGene, China) (P <.001; OR 8.0; 95% CI, 3.2 to 20.0) and abGenix/abTES (AITbiotech, Singapore) (P =.018; OR 6.3; 95% CI, 1.7 to 22.4). To assess if either component of the combination (extraction kit or rRT-PCR assay) was significantly associated with reporting an incorrect result, we performed a chi-square analysis comparing the individual reagent with the incorrect result reported. The abGenix extraction kit, DAAN rRT-PCR assay, and abTES rRT-PCR assay were each significantly associated with reporting an incorrect result (P <.001; OR 6.7; 95% CI, 2.9 to 15.2; and P =.007; OR 3.8; 95% CI, 1.5 to 9.2; and P =.019, OR 3.8, 95% CI, 1.3 to 10.4, respectively;). The use of any of the 3 fully automated systems was not significantly associated with incorrect results, and likewise the use of a conventional workflow, Sansure rRT-PCR, and QIAGEN extraction kit were not significantly associated with incorrect results.
Performance of Participating Laboratories by SARS-CoV-2 Reagent Combination
Notes: Reagent combinations used by less than 4 laboratories are not shown. NS=not significant for P values >.05 and OR values; 95% CI are provided only for those samples with P <.05.
Values of EQA samples obtained using a droplet digital polymerase chain reaction assay: sample A containing no virus; samples B through F containing SARS-CoV-2 5,540,000; 682,000; 71,000; 7,050; and 1,000 copies/mL, respectively. bComparing reagent combinations with the inability to report correct results for each EQA sample. cFully automated systems. dFor combinations associated with an incorrect result, each reagent of the combination was compared with the inability to report correct result for each EQA sample.
Abbreviations: CI, confidence interval; DMSc, Thailand Department of Medical Sciences; EQA, external quality assessment; NS, not significant; OR, odds ratio; rRT-PCR, real-time reverse transcription polymerase chain reaction.
Discussion
This article describes the first round of EQA for SARS-CoV-2 rRT-PCR in Thailand. Lessons learned on how to improve Thailand's national EQA program can serve as a roadmap for other countries interested in implementing a national EQA program. Most (n=183, 92.9%) of the 197 laboratories within Thailand's COVID-19 network could produce accurate and reliable results as observed in this first round of EQA. Thailand's national EQA program used samples containing inactivated SARS-CoV-2 culture supernatant prepared from a predominant strain in the early phase of the outbreak (January 2020) in Thailand. This approach to EQA sample preparation more closely mimicked the true scenario for laboratories testing clinical specimens from cases in Thailand compared with EQA programs that use samples containing RNA transcripts or pseudoviruses based on reference SARS-CoV-2 sequences.
The national EQA program was provided free of charge to all SARS-CoV-2 testing laboratories in Thailand to reduce any financial barrier to participation in EQA and to encourage all COVID-19 testing laboratories to participate. The availability of free EQA for laboratories performing tests to confirm diseases of pandemic potential is an important factor to consider in pandemic preparedness. In this first round of Thailand's national EQA, the cost was US$150 per laboratory, including shipping, and was fully covered by the DMSc. Commercial EQA programs for SARS-CoV-2 rRT-PCR, such as that provided by Quality Control of Molecular Diagnostics, can cost as much as US$800 per laboratory, including shipping of the panel to Thailand. 12 However, national EQA programs require upfront investments as well as maintenance costs, especially for programs accredited by ISO 17043.13,14 To offset these costs, national EQA programs can consider a cost-recovery model, whereby private laboratories pay a fee to participate while public laboratories can participate in EQA free of charge.
The internal investigations conducted in each of the 14 laboratories reporting incorrect results revealed problems with RNA extraction, contamination of the negative RNA sample with SARS-CoV-2 RNA from the positive samples, and switching of EQA samples at some point during the testing process. These types of errors represent common laboratory errors and are not unexpected given the rapid expansion of COVID-19 rRT-PCR testing in laboratories with little to no experience with PCR testing. Many new laboratory staff were hired and quickly trained to keep pace with the demand for COVID-19 testing. The tremendous workload in these laboratories, with staff often working 24 hours a day/7 days a week, limited their ability to provide close supervisory oversight, which would be the norm when new testing capacities are introduced and optimized in the laboratory. Given these challenges and the risks they pose to accurate and reliable diagnostic testing, EQA became an even more critical strategy to assess the accuracy and reliability of the COVID-19 laboratory testing network in Thailand during the pandemic.
In addition to human error, the observed poor RNA extraction in some laboratories that reported incorrect results could have been due to problems with specific lots of reagents. The rapid surge in demand for reagents to meet testing needs required companies to increase production, which may have resulted in undetected problems with some reagent lots. Analysis of data submitted by participating laboratories about the types of extraction and rRT-PCR reagents used found statistically significant associations between use of specific reagents and incorrect results. Among the most used reagents, the abGenix/abTES and abGenix/DAAN combinations were significantly associated with incorrect results; each alone was also significantly associated with an incorrect result. As part of postmarket surveillance, the DMSc reported these data without any laboratory-specific identifying information to the manufacturer of abGenix. The company provided a new lot of reagents and DMSc provided additional EQA sample aliquots to relevant laboratories for retesting. All laboratories, except 1, were able to report correct results for each EQA sample after implementing laboratory-specific corrective actions such as cleaning workspaces to reduce the risk of RNA contamination and using a new lot of reagents provided by the distributor. However, since laboratory-specific corrective actions and the use of a new lot of reagents were not tested independently, it is not possible to determine which was the underlying cause of the incorrect results. Additional information from the manufacturer on reasons for reagent failure were not available. This highlights the importance of collecting information from EQA participating laboratories on testing workflows and reagents used and sharing anonymized data with manufacturers for product improvement. Investigations in laboratories that reported incorrect results also found that these laboratories did not document key testing information required by their quality management system, such as lot numbers and expiration dates for the reagents used in their laboratories, and thus it was not possible to track the performance of a specific lot of reagents and its association with incorrect results. This observation suggests failure of the laboratory's quality management system, which must be rectified as part of overall laboratory quality improvement.
In concordance with a previous study, 8 more laboratories reported incorrect (false-negative) results for EQA samples with low viral concentration, close to the given assay's limit of detection, compared to EQA samples with high viral concentration. The limit of detection of approved assays can vary by as much as 10,000-fold, with the best assays demonstrating an ability to detect approximately 100 copies of viral RNA per mL. 15 In our study, the EQA sample with the lowest RNA concentration had approximately 1,000 copies of viral RNA per mL and 9 laboratories reported a false-negative result for this sample, suggesting that patients in the early course of disease would not be detected and could unknowingly transmit the disease to others. A previous study suggested that with each 10-fold increase in the limit of detection, assay sensitivity is lowered by 13%. 15
The first round of SARS-CoV-2 EQA in Thailand had some noted limitations, including the absence of EQA samples that were positive for non-SARS-CoV-2 respiratory viruses to assess assay specificity, only 1 negative EQA sample included in the panel, insufficient EQA sample volume to enable testing by all test systems for those laboratories that use multiple SARS-CoV-2 rRT-PCR kits, and failure to collect information from participating laboratories on reagent lot numbers and expiry dates for reagents used to test the EQA samples. These represent opportunities for improvement of Thailand's national EQA program.
Conclusion
Thailand's first EQA for SARS-CoV-2 rRT-PCR revealed that the COVID-19 diagnostic network could produce accurate and reliable results, which was critical to detect patients as early in the course of their disease as possible, reducing further transmission. Thailand's experience identified several important lessons that can improve future rounds of Thailand's EQA program and help other countries interested in establishing their own national EQA program. Key lessons learned were: (1) inclusion of sufficient numbers of positive EQA samples, including those with a low viral copy number near the limit of detection of commonly used assays, and negative EQA samples to assess a laboratory's ability to confirm (positive samples) and rule out (negative samples) the presence of the pathogen in a clinical specimen; (2) inclusion of EQA samples positive for non-SARS-CoV-2 respiratory pathogens to assess assay specificity; (3) sufficient volume of EQA samples to allow laboratories to report results for all platforms and reagent kits currently active in the laboratory; and (4) inclusion of addition questions on the participant questionnaire that accompanies the EQA panel to capture information about the laboratory's overall management of the testing process. While these lessons were learned through Thailand's first round of EQA for SARS-CoV-2, they are applicable to EQA for any emerging pathogen. Strengthening national EQA programs is an essential component of pandemic preparedness and should be prioritized to ensure accurate and reliable diagnostics, especially for diseases with pandemic potential.
Footnotes
Acknowledgments
We thank the SARS-CoV-2 RT-PCR laboratory network for participation; Ms. Thanutsapa Thanadachakul for determining viral load by droplet digital PCR; and Dr. Joshua A. Mott, Thai Ministry of Public Health–US Centers for Disease Control and Prevention Collaboration (TUC), for technical inputs. Thailand's Department of Medical Sciences supported the cost of this first round of the EQA scheme. This work was an extension of the influenza proficiency testing platform, which was supported by the TUC Influenza Program through Cooperative Agreement 1U01GH002084 with the Thailand Ministry of Public Health. The findings and conclusions in this report are those of the author(s) and do not necessarily represent the official position of the US Centers for Disease Control and Prevention. Use of trade names and commercial sources is for identification only and does not imply endorsement by the US Department of Health and Human Services.
