Abstract
Global health security depends on effective surveillance for infectious diseases. In Uganda, resources are inadequate to support collection and reporting of data necessary for an effective and responsive surveillance system. We used a cross-cutting approach to improve surveillance and laboratory capacity in Uganda by leveraging an existing pediatric inpatient malaria sentinel surveillance system to collect data on expanded causes of illness, facilitate development of real-time surveillance, and provide data on antimicrobial resistance. Capacity for blood culture collection was established, along with options for serologic testing for select zoonotic conditions, including arboviral infection, brucellosis, and leptospirosis. Detailed demographic, clinical, and laboratory data for all admissions were captured through a web-based system accessible at participating hospitals, laboratories, and the Uganda Public Health Emergency Operations Center. Between July 2016 and December 2017, the expanded system was activated in pediatric wards of 6 regional government hospitals. During that time, patient data were collected from 30,500 pediatric admissions, half of whom were febrile but lacked evidence of malaria. More than 5,000 blood cultures were performed; 4% yielded bacterial pathogens, and another 4% yielded likely contaminants. Several WHO antimicrobial resistance priority pathogens were identified, some with multidrug-resistant phenotypes, including
F
Uganda, an inland East African country with a rapidly growing population estimated at 43 million people in 2017, has made progress in recent decades to improve life expectancy, reduce poverty and food insecurity, and expand access to immunizations and clean water. 7 Yet, as with many African countries, Uganda faces diverse health challenges in a weak health infrastructure that limits the rapid detection and confirmation of infections with epidemic potential. In the past 2 decades, Uganda has experienced outbreaks of emerging and reemerging infectious diseases including Ebola, Marburg, Crimean-Congo hemorrhagic fever, Rift Valley fever, yellow fever, hepatitis E, cholera, typhoid fever, plague, and anthrax.8-16
Effective laboratory capacity and disease surveillance are critical to global health security and the basis for the 2005 International Health Regulations (IHR) signed by all World Health Organization (WHO) member states. IHR compliance has proven challenging for many low-income countries, including Uganda.17,18 The Global Health Security Agenda (GHSA), a multisectoral and multilateral partnership intended to support countries toward IHR compliance, launched officially in 2014 (www.ghsagenda.org). The government of Uganda and the US Centers for Disease Control and Prevention (CDC) jointly implemented a demonstration project a year earlier, in 2013. The pilot project improved specimen referral networks and information systems associated with outbreak response and created an emergency operations center; these activities set the stage for multiple GHSA activities in the country. 19
Through GHSA-initiated partnerships, we introduced a cross-cutting surveillance approach to advance ability to detect unusual health events in Uganda. As conceived, this effort would provide comprehensive patient data and facilitate electronic systems infrastructure that could ultimately improve early detection of novel infections or outbreaks, define conditions causing human illness to inform appropriate and targeted laboratory capacity building efforts, and generate data for an antimicrobial resistance surveillance and intervention program in its infancy.
Methods
We modified an existing inpatient pediatric malaria surveillance system to provide patient data from all children 14 years old and younger who were admitted to 6 government hospitals, in concert with improved diagnostic ability to define causes of nonmalarial illness affecting the population. Five of the 6 designated hospitals have participated in ongoing pediatric malaria surveillance since 2010; 1 site in northwestern Uganda was added to improve geographic representation for this expanded system. Included hospitals reflected both district and regional hospitals, 2 tiers in the organizational structure of hospitals in Uganda. The Uganda Ministry of Health (designated sentinel hospital surveillance sites, Uganda Virus Research Institute, or UVRI, and the Uganda National Health Laboratory Services) partnered with a consortium composed of the Infectious Diseases Institute (IDI), Makerere University Department of Medical Microbiology, the Infectious Diseases Research Collaboration (IDRC), and the Health Information Systems Program (HISP-Uganda) for the capacity-building efforts. Standard operating procedures regarding data capture, phlebotomy, and specimen labeling, storage, and transport were developed or revised as necessary. Before implementation, plans were finalized by all partners, approved as public health surveillance/nonresearch by CDC (NCEZID #031416), and approved by the Director General of Health Services, Ministry of Health, Uganda.
Baseline Capacity Assessment, Training, and Site Activation
Surveillance for clinical and demographic data was ongoing at 5 of the 6 sites via the inpatient malaria surveillance system. Thus, routine and standard data collection for purposes of surveillance was a new activity for only 1 site. Site activation reflected collection of expanded information for surveillance, improved electronic surveillance infrastructure, and laboratory diagnostic capacity. Activation of sites occurred between July 2016 and October 2017. The 6 sentinel hospital sites and their associated system activation dates were: Jinja Regional Referral Hospital (Jinja RRH) (July 2016), Arua RRH (July 2016), Mubende RRH (August 2016), Kabale RRH (April 2017), Apac District Hospital (July 2017), and Tororo District Hospital (October 2017) (Figure 1).
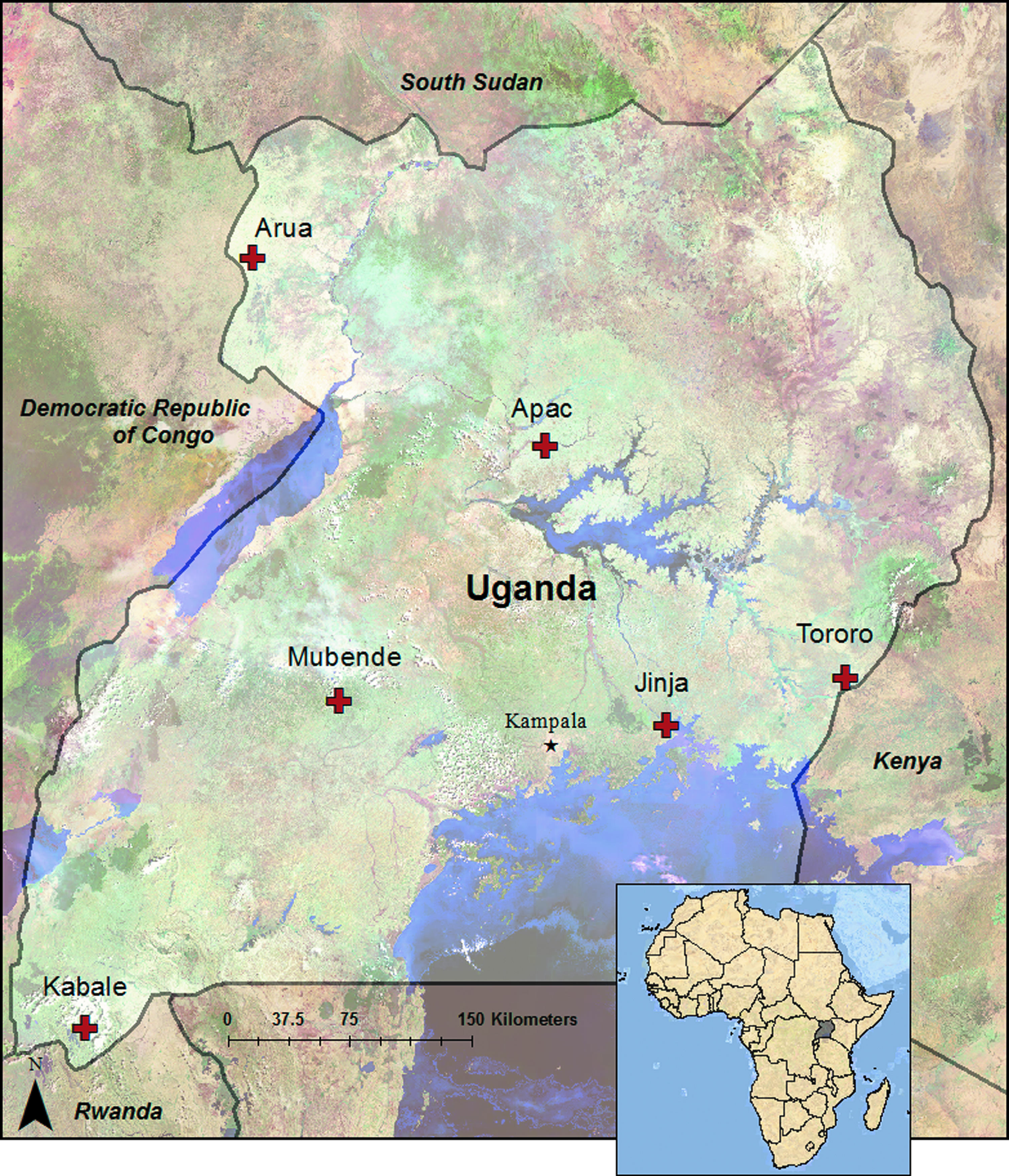
Geographic location of 6 sentinel site hospitals engaged in case-based surveillance and laboratory capacity-building efforts in Uganda
Before activation of this system at each selected government hospital, a baseline rapid assessment was conducted to determine: (1) existing on-site laboratory capacity for microbiology; (2) potential for expansion of on-site microbiologic capacity in terms of workforce, training needs, equipment needs, and available space; (3) biosafety and biosecurity practices and gaps; (4) infection prevention and control practices and gaps; and (5) existing organizational and management structures and their associated ability to support modified practices.
Briefly, assessment was conducted using the Potter and Brough Capacity Pyramid approach, which assesses capacities at various structural levels, all within the local context, including tools, skills, staff and infrastructure, structures, systems, and roles. 20 Standards and tools used or adapted included those of the WHO SLIPTA program (Stepwise Laboratory Improvement Process Towards Accreditation), the WHO laboratory assessment tool, and CDC infection control assessment tools.21-23 These assessments revealed extensive gaps including inactive hospital-based infection prevention and control programs, lack of supplies and infrastructure to promote basic hand hygiene, lack of microbiology services, poor access control to laboratory rooms, inadequate sample transportation for microbiologic samples to a central reference laboratory, lack of defined systems for pharmaceutical tracking, weak information technology and data management, and inadequate interface between clinicians and hospital laboratories. Prior to activation, ongoing routine diagnostic testing was unavailable for infectious conditions other than malaria and HIV.
The process of surveillance site activation included inception meetings with varied hospital and department administrators and staff; provision of equipment and supplies for phlebotomy, infection control, clinical specimen processing and storage, and internet connectivity according to needs identified on assessment. At each surveillance site, multifaceted training was aimed at pediatricians, nurses, and laboratory technicians, but it included additional cadres of Ugandan government healthcare workers staffed in each hospital. This training was delivered through interactive lecture and hands-on practice. Content included aseptic technique, appropriate sample volumes for laboratory tests, and the importance of good record keeping for surveillance and improved clinical care. Training also included the importance of investigating nonmalarial causes of illness and incorporation of bacterial blood culture into routine clinical care. Initial emphasis focused on improving diagnosis of nonmalarial illnesses by reserving blood culture for febrile children without evidence of malaria by microscopy or rapid diagnostic test (RDT). However, clinicians were also encouraged to use blood culture as a diagnostic tool in any circumstance deemed appropriate.
Continued regular site support visits occurred following activation to provide ongoing training in phlebotomy, troubleshoot staff concerns, identify areas for improvement, and provide summary data and feedback. Concerns and issues that arose were addressed and incorporated into ongoing training efforts at all surveillance sites. When clustering of contaminated cultures in space and time occurred, investigation and intervention followed. Areas dedicated to phlebotomy were created on all wards to improve the likelihood of aseptic draws in otherwise difficult hygienic conditions.
Laboratory System Strengthening
Blood cultures were made available at the time of surveillance site activation. With a focus on improving surveillance and diagnostic capacity for severe illness in the pediatric population, most blood culture bottles purchased and provisioned to hospitals were aerobic pediatric bottles for blood volumes up to 3 mL according to patient age and weight. Before installation of BD BACTEC™ (Becton Dickinson, Franklin Lakes, NJ, USA) machines in sentinel hospital laboratories, all inoculated blood culture bottles were transported daily at ambient temperature to the Department of Medical Microbiology (DMM) at Makerere University in the capital city of Kampala for processing. Upon successful installation of BACTEC machines in sentinel site hospitals and associated training, those hospital laboratories incubated all blood culture bottles locally for up to 7 days. Negative results were provided directly back to hospital clinicians. Bottles read as positive by the machines were usually subjected to Gram stain at the hospital laboratory in order to provide timely and clinically relevant information before transportation to DMM for subculturing, identification, and antimicrobial susceptibility screening. Identification at DMM was performed using conventional, phenotypic identification algorithms down to species and subspecies or serotype when appropriate reagents were available. Classification of isolated organisms as pathogens or likely contaminants was made presumptively by DMM based on general knowledge regarding pathogenicity of various organisms and presented accordingly here. Most antimicrobial susceptibility testing was performed using the Kirby-Bauer disk diffusion method according to CLSI standards and interpreted according to 2014 breakpoints.
We instituted serologic tests for patients with a history of febrile illness of at least 4 days' duration, a restriction designed to assure improved sensitivity for detection of IgM antibodies for certain conditions. Tests were instituted at each site as they became available in Uganda and following adequate staff training, independent of site activation. A lateral flow assay to detect IgM antibody to a variety of
Surveillance and Laboratory Data Capture and Management
In each sentinel hospital pediatric ward, a standardized medical record form previously developed for malaria surveillance 25 was adapted to incorporate additional variables relevant to expanded diagnostic capacity with stakeholder and clinician input. This form serves as a medical chart for all admissions, allowing providers to record symptoms, signs, treatments, and outcome at discharge.
A full-time surveillance data officer was stationed at each hospital. The primary functions of this person were to engage with clinicians and laboratory staff to ensure that samples were collected, labeled, and transported properly and that clinical and laboratory records were complete; to enter demographic, clinical, and laboratory data into the electronic database; and to facilitate ongoing communication among the surveillance team and hospital staff and administrators. Initially, data were entered into a Microsoft Access® database, but following improvements in connectivity, data entry was subsequently transitioned to a web-based data management system on the District Health Information System (DHIS-2) platform (www.dhis2.org). DHIS-2 is a flexible, open-sourced information system used by Uganda's Ministry of Health for aggregate public health data management and by the Public Health Emergency Operations Center for recording and tracking case-based data related to outbreak response and specimen collection, transportation, and testing. The system was tailored to allow case-based routine surveillance data transmission, with multiple data entry points on the same individual record to enable linking of patient data with central laboratory results, including antimicrobial resistance patterns. The system permits site-based staff, central laboratory staff, and ministry of health personnel to view data on individual patients and associated laboratory results in real-time, thereby improving data quality and use for case management, specimen tracking and testing, and disease surveillance.
Results
Clinical and Demographic Data
Between July 2016 and December 2017, patient data were collected for 30,500 admissions in 6 hospitals (Table 1). Median patient age was 2 years (range: 1 day to 14 years); 56% were male. Of admitted patients, 74% (
Characteristics of pediatric patients admitted to 6 sentinel site hospitals according to frequency of clinical characteristics and select laboratory diagnostics—Uganda, 2016-2017
Includes presumptive positive alphavirus and flavivirus based on IgM as well as confirmed arboviral infection.
Among 30,500 total admissions electronically captured between July 2016 and December 2017 at all 6 hospitals, 21,199 admissions (70%) occurred after blood culture capacity had been established in each respective hospital and additional associated variables were being collected. Accordingly, this reflected all admissions from the first sites activated in July and August 2016 and only a fraction from sites activated later in July and October 2017 (Table 1). Among these 21,199 admissions, 20% (
Laboratory Data and Mortality
A total of 5,001 single bottle blood cultures were performed at sentinel site hospitals through December 2017. Among these, 412 (8%) yielded bacterial growth; 187 were deemed pathogenic bacteria by the processing laboratory. Of these 187 cultures, 160 had species identities available in the web-based surveillance system at the time of writing (Table 2). An additional 225 bottles grew likely contaminants, specifically
Pathogenic organisms isolated from blood culture at 6 sentinel site hospitals—Uganda, July 2016-December 2017
Considered a pathogen rather than contaminant here.
A broad range of pathogens was isolated. The most commonly isolated pathogens were
Jinja RRH, east of Kampala, is the largest sentinel site in terms of patient volume and yielded the greatest diversity of pathogens (Table 2). Several of the pathogens identified to date originated commonly or exclusively from Jinja RRH, including
Following initiation of hospital-based leptospirosis testing, 6% of patient samples screened showed IgM reactivity on rapid test, with some potential for geographic clustering (Table 1). Arboviral screening of 622 sera yielded 18 (3%) confirmed and presumptive-positive samples (Table 1). Fourteen samples were presumptive positive for recent flavivirus infection (eg, dengue, West Nile, yellow fever, or Zika). However, remaining sample volume was insufficient for PRNT confirmation of infecting virus. Positive samples originated from Jinja, Mubende, Arua, and Kabale. Of 4 samples with evidence of acute alphavirus infection, 2 were confirmed as o'nyong nyong virus infection, both originating from Arua. None of the sera screened for antibodies to
Based on data from the processing laboratory, resistance to commonly used antimicrobials was observed in several bacterial species; a subset of these findings is presented in Table 3. Among
Select antimicrobial susceptibility patterns, number of fully susceptible isolates/total screened (%) a
Data not shown where fewer than 3 isolates were screened or for antimicrobials infrequently screened.
Overall, all-cause mortality among pediatric admissions at sentinel hospitals following availability of blood culture was 4%, with a range across hospitals of 1% to 9% (Table 1). The case-fatality rate was 10% among those with positive blood cultures, 3% among those that had negative blood cultures, and 4% among those with confirmed malaria.
Discussion
This sentinel surveillance and laboratory improvement effort in 6 Ugandan government hospitals demonstrates a pragmatic approach to identifying and tracking nonmalarial causes of febrile illness among hospitalized children aged 0 to 14 years. Early results from this effort led to the generation of more than 5,000 blood cultures, 4% of which yielded bacterial pathogens, including several WHO antimicrobial resistance priority pathogens, some with multidrug-resistant phenotypes, including
Although malaria is a common cause of fever in sub-Saharan Africa, including Uganda, there is increasing recognition that nonmalarial etiologies comprise a substantial portion of febrile illness.25,26 During the first several months of this effort in 6 hospitals, malaria accounted for approximately half of all pediatric febrile admissions. Infectious etiologies among hospitalized children at these 6 government hospitals in Uganda are similar to that from other reports from the region, including varied bacterial blood stream pathogens, arboviruses, and leptospirosis.3,5,27,28 Variability in the suite of pathogens across sites was evident, and further analyses should help identify baseline trends according to geography and season. Despite enhancement of laboratory capacity at these hospitals and centrally, the cause of illness for a large percentage of febrile admissions remain unknown. With over half of screened samples demonstrating reactivity to spotted fever group
Challenges in interpretation of laboratory data in this setting remain evident. Antibiotic self-treatment was reported for 15% of pediatric admissions throughout Uganda during this 18-month period, a frequency similar to that from other reports from Africa.27,30 Antibiotic pretreatment may have decreased sensitivity of blood cultures. Additionally, challenges obtaining adequate blood volumes, particularly from malnourished children, decreases the ability to subject clinical specimens to a wide array of diagnostic tests, potentially limiting diagnostic success. An investigation in early 2017 in Jinja RRH revealed that the majority of contaminants were from cultures drawn from children on the malnutrition ward. Some isolates deemed contaminants here could be causing illness, particularly in the malnourished or immunocompromised population. Moreover, some organisms isolated could represent hospital-acquired infections. However, the clinical and laboratory limitations inherent in this healthcare setting make it difficult to discern the clinical significance of these organisms. Despite these challenges, the overall proportion of pediatric blood cultures with evidence of bacterial pathogens was similar to that of other reports from Africa.3,5,28,30,31 Antimicrobial susceptibility results presented here must also be interpreted with care. Reproducible AST results depend on the reagents used. The panel of antimicrobials tested varied with availability of reagents (or pharmaceuticals) and in some cases was expanded when resistance to first line drugs was observed. While these results may facilitate individual patient management and indicate overall trends, minimum inhibitory concentration testing against a panel of clinically and epidemiologically relevant drugs would be required for a comprehensive analysis.
Antibiotic misuse and lack of antimicrobial stewardship programs threaten the ability to treat historically treatable bacterial infections.
32
Low- to middle-income countries may disproportionately feel the devastating consequences of drug-resistant bacteria, as therapeutic options beyond traditional first line therapy may often be limited. Absent bacterial culture and susceptibility testing, empirical antimicrobial therapy may result in underuse or misuse of antibiotics in bacteremia or bacterial/malarial co-infections, and overuse in other undiagnosed viral febrile illnesses, thereby fomenting antimicrobial resistance. Gentamicin was the most commonly prescribed drug in these hospitals, and resistance to gentamicin was evident, particularly among
Conclusion
Data on routine and rare etiologies of febrile illness and associated trends improve not only health outcomes, but also the ability to detect and respond to unusual events of potential public health consequence. 6 In Uganda, many disease-specific surveillance systems exist, such as for malaria, influenza, viral hemorrhagic fevers, and plague.15,33,34 A more comprehensive and integrated approach to disease surveillance and laboratory evaluation can generate information to minimize inefficiencies of disease-specific surveillance silos and assure that limited public health resources are leveraged appropriately to improve health security.
The system described here leverages prior surveillance investments to better identify common and rare causes of illness, generate antimicrobial susceptibility data, inform infection prevention and control interventions, and facilitate electronic surveillance infrastructure improvements. This approach to surveillance and laboratory systems improvement generates data and systems that can serve as the basis for responsive and sustainable public health surveillance in Uganda. Understanding of routine and rare causes of illness and development of appropriate laboratory capacity and associated systems is important to ensure systems in place are flexible and responsive to identify and detect epidemic-prone illness, whether existing or novel. In July 2017, a single case of
Footnotes
Acknowledgments
This work was supported through cooperative agreement number 5NU2GGH001744-02-00, funded by the US Centers for Disease Control and Prevention. Its contents are solely the responsibility of the authors and do not necessarily represent the official views of the Centers for Disease Control and Prevention or the Department of Health and Human Services.
