Abstract
Background:
Lactate dehydrogenase (LDH) is a marker of injury and disease as it is expressed extensively in numerous cell types and tissues. Moreover it is released during tissue breakdown, and is elevated in cancerous tissues. However, the clinical significance and prognostic value of LDH as a tumor marker have been subject to considerable discussion.
Objective:
In this study, clinical serum LDH data from patients with cervical cancer (CC), CC microarray data, and RNA-seq data were integrated to assess the expression of LDH in CC.
Methods:
A total of 204 patients with newly diagnosed CC and 204 age-matched healthy controls were included to evaluate serum LDH levels in CC and non-cancer samples. External microarrays and RNA-seq datasets were collected for the differential expression analysis of LDH in CC and non-cancer tissue samples. Kaplan-Meier survival curves of the prognostic value of LDH for CC were plotted for RNA-seq data. Functional enrichment analysis was performed for the genes co-expressed with LDH.
Results:
The data from our in-house clinical cases as well as the data extracted from microarrays and RNA-seq databases demonstrated significant overexpression of LDH in CC samples. Elevated LDH expression levels were associated with poor overall survival in CC patients. The genes co-expressed with LDH were significantly correlated with the biological processes and pathways, associated with nuclear division, the condensed chromosome, protein serine/threonine kinase activity, and the cell cycle.
Conclusion:
In conclusion, LDH upregulation might serve as a therapeutic and prognostic biomarker for CC.
Introduction
Cervical cancer (CC), the fourth most common cancer and the main cause of cancer-related deaths among women globally, resulted in 569,847 new cases and 311,365 deaths in 2018 (Cohen et al., 2019). In the United States, CC is the second leading cause of cancer-related death in women aged 20 to 39 years and causes 10 premature deaths per week in this age group (Siegel et al., 2020). In China, CC is one of the most common cancers in women and has a high burden (Wei et al., 2020). CC is usually asymptomatic in the early stage, and nonspecific clinical manifestations, such as pelvic pain, weight loss, abnormal vaginal bleeding, and postcoital bleeding, occur at an advanced stage (Canavan and Doshi, 2000; Issah et al., 2011; Vu et al., 2018).
Over the past decades, the incidence and mortality of postcoital bleeding in women with CC have decreased owing to early detection through screening (Tarney and Han, 2014; Sawaya et al., 2015). Currently, there are only two types of diagnostic tests for CC screening: the Papanikolaou (Pap) smear and the human papillomavirus (HPV) test (Tsikouras et al., 2016). Pap test is performed to detect potentially precancerous and cancerous processes in the cervix. On account of the correlation between HPV infection and CC, the American Cancer Society (ACS) recommended early detection of cervical cancer in average-risk asymptomatic adults primarily with cervical cytology by Pap test.
And women aged 30-65 years should preferably be tested every 5 years with both the HPV test and the Pap test (Smith et al., 2017). Nevertheless, both Pap test and HPV test have their limitations. HPV test can be done to find out whether a woman has an HPV infection and is, therefore, at greater risk of cervical cancer but cannot detect abnormal cell changes. Pap test may suggest cervical cell abnormalities when there is none and may not detect cervical cell abnormalities even though they are present (Anorlu et al., 2003; Johnson et al., 2020).
Lactate dehydrogenase (LDH) is an enzyme composed of four subunits. LDH-M and LDH-H are the most common subunits, encoded by the LDHA and LDHB genes, respectively. LDH is a marker of injury and disease as it is expressed extensively in various cells and body tissues and released during tissue breakdown (Klein et al., 2020). In cancer, tumor cells rely on aerobic glycolysis, which results in lactate production even under conditions of sufficient oxygen, in a process known as the Warburg effect (Warburg, 1956). In this study, clinical serum LDH data of patients with CC, microarray data, and RNA-seq data from tissue samples of CC were integrated to assess the upregulation of LDH in CC.
Materials and Methods
Patient selection
From January 2017 to August 2020, serum LDH data of patients with newly diagnosed CC and age-matched healthy controls were included in this case-control study at the First Affiliated Hospital of Guangxi Medical University. Within 2 days of admission, peripheral venous blood samples were collected from patients with CC on an empty stomach in the early morning. The analysis for serum LDH was performed on an automatic analyzer (Hitachi Automatic Analyzer 7600, Tokyo, Japan) by standard IFCC (International Federation of Clinical Chemistry) methods.
The inclusion criteria were newly diagnosed CC without surgery or chemoradiotherapy, no other cancer, coronary artery disease, rhabdomyolysis, myoglobinuria, hemolytic anemia, or fracture.
This study has been approved by the First Affiliated Hospital of Guangxi Medical University Institutional Review Board (IRB) for the Protection of Human Research Participants. IRB Number: NO.2022-KY-E-(066).
LDHA expression in CC tissues from external RNA-seq and microarray data sets
To explore LDHA expression in CC and noncancer cervical tissues, external RNA-seq and microarray data sets obtained from The Cancer Genome Atlas (TCGA), Genotype-Tissue Expression (GTEx), Gene Expression Omnibus (GEO), Oncomine, ArrayExpress, and Sequence ReadArchive (SRA) databases were used for expression analysis. A fragments per kilobase per million (FPKM) expression matrix of LDHA in CC and noncancer cervical tissues and the transcripts per million (TPM) expression value of LDHA in 11 normal cervical tissues were downloaded from the TCGA and GTEx databases and merged into the TCGA_GTEx data set after log2(TPM +0.001) normalization.
Microarrays containing LDHA expression in CC and noncancer cervical tissues were searched with the following keywords: (“CESC” OR “cervical cancer” OR “cervical carcinoma”) in GEO, Oncomine, ArrayExpress, and SRA databases. Then, all microarrays were processed by log2 transformation and normalized between CC and noncancer cervical tissues. Expression and diagnostic data of LDHA in CC and noncancer cervical tissues were extracted from each data set to construct a Forest plot of the standard mean difference (SMD) using the meta package of R and summary receiver operating characteristic (SROC) curves using Stata v.14.0.
Gene Ontology and Kyoto Encyclopedia of Genes and Genomes enrichment analysis
All expression matrices of CC and noncancer cervical tissues in RNA-seq and microarray data sets were collected and subjected to differential expression and correlation analyses. The intersection of significantly upregulated genes (log2FC >1, adj.p < 0.05) and genes positively correlated with LDHA (r > 0, p < 0.05) and that of significantly downregulated genes (log2FC<−1, adj.p < 0.05) and genes negatively correlated with LDHA (r < 0, p < 0.05) were selected as genes coexpressed with LDHA in CC. Biological functions, pathways, and disease enrichment of the coexpressed genes were analyzed using the ClusterProfiler package in R v.3.5.2. p < 0.05 was considered statistically significant.
Statistical analysis
Statistical analyses were performed using SPSS v22 and GraphPad Prism v8.0. The Kolmogorov-Smirnov test was used to determine the distribution of continuous variables. Normally distributed continuous variables are reported as the mean value ± standard deviation. Non-normally distributed continuous variables are presented as the median value and interquartile range. The Mann-Whitney U test was used to evaluate the differences in quantitative data between patients and healthy controls. The Kruskal-Wallis test was used to evaluate the association of serum LDH level with tumor stage.
Differential expression analysis was conducted using the Limma package in R v.3.6.1 for the normalized FPKM or TPM expression matrix and the Voom algorithm for the count matrix. Correlation analysis was performed using the psych package of R v.3.6.1. p < 0.05 was considered statistically significant. The prognostic significance of LDHA in CC from the RNA-seq data set was calculated using Kaplan-Meier survival analysis with survminer and the survival package in R v.3.6.1. The cutoff value to separate patients with CC into different survival groups was the median log2(TPM +0.001) expression value of LDHA.
Results
Expression and clinical value of serum LDH levels in patients with CC
As a ubiquitous enzyme, LDH is present in high concentrations in liver, kidney, myocardium, skeletal muscle, and red blood cells and plays a key role in the Warburg effect, which seems to be a common characteristic of malignant cells for tumorigenesis (Jurisic et al., 2015). This hospital-based study included 204 patients with newly diagnosed CC (tumor-node-metastasis [TNM]: 58 stage I, 94 stage II, 25 stage III, and 27 stage IV) and 204 age-matched healthy controls. The median age at the diagnosis of CC was 53.0 years (range: 30-84 years). The median serum LDH level for the entire cohort was 184.0 U/L (range: 102.0-648.0 U/L).
The relationships between serum LDH levels and clinical characteristics in the patients with CC are given in Table 1. The serum LDH level was slightly higher in patients in stage III or IV than in those in stage I or II. The serum LDH levels in patients with CC were higher than those in the controls. The receiver operating characteristic (ROC) curve shows the capacity of serum LDH levels to discriminate between TNM stages I-II and stages III-VI. The area under the curve (AUC) of the ROC curve was 0.73 (95% CI: 0.46-0.62) with a pooled sensitivity of 0.90 and specificity of 0.54 (Fig. 1).
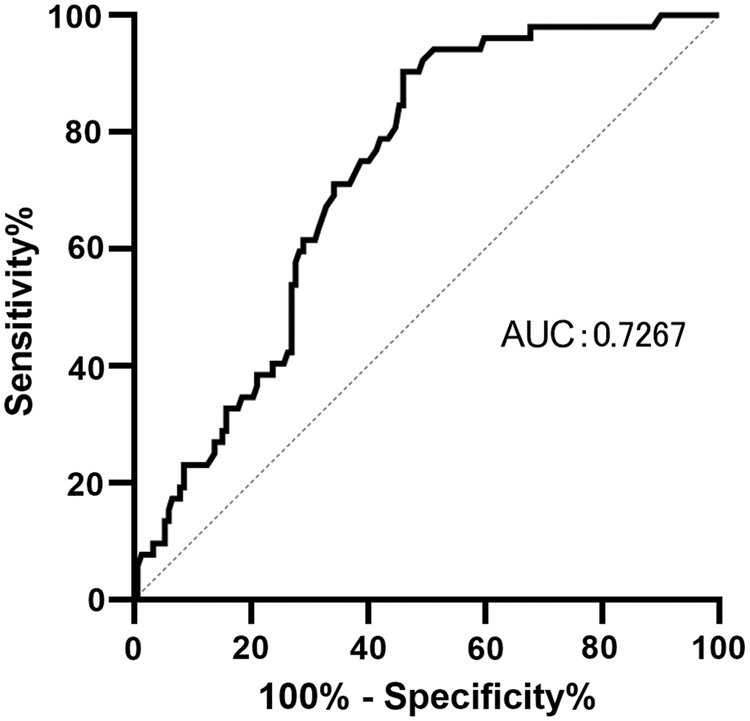
ROC curve for the ability of serum LDH in differentiating CC patients of TNM stages I-II from CC patients of TNM stages III-IV. AUC, area under curve; CC; LDH, lactate dehydrogenase; ROC, receiver operating characteristic; TNM, tumor-node-metastasis.
Relationship of Serum Lactate Dehydrogenase Levels with Clinical Characteristics in 204 Cervical Cancer Patients
Quantitative data are presented as median (interquartile range).
The p value was calculated by Mann-Whitney U tests.
The p value was calculated by Kruskal-Wallis tests.
LDH, lactate dehydrogenase; TNM, tumor-node-metastasis.
Expression value and prognostic significance of LDHA in CC based on RNA-seq and microarray data sets
In the GEO database, 15 microarrays were eligible for survival analysis. All microarrays and RNA-seq data sets included 704 CC and 181 noncancer cervical samples. Violin plots and ROC curves for LDHA expression in each data set are displayed in Figures 2 and 3. Violin plots denoted that the expression of LDHA was significantly upregulated in CC samples in comparison with that in noncancer samples for microarrays and data sets GSE138080, GSE27678-GPL570, GSE39001-GPL201, GSE52904-GPL6244, GSE63678, GSE7410, GSE89657, and TCGA_GTEx.
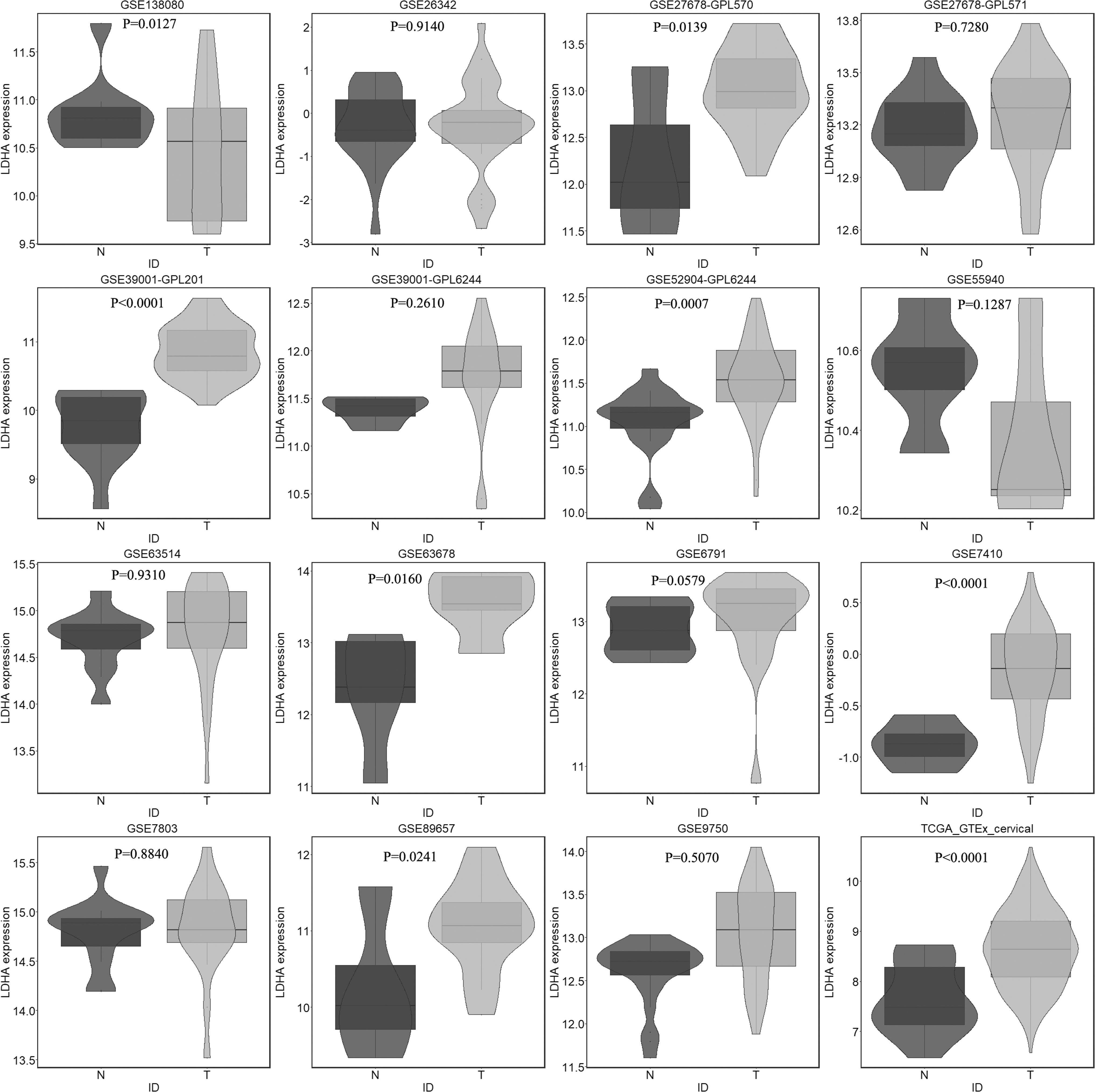
Violin plots of differentiating ability of LDH expression in CC and noncancer samples from all included microarrays and data sets (GSE138080, GSE26342, GSE27678-GPL570, GSE27678-GPL571, GSE39001-GPL201, GSE39001-GPL6244, GSE52904-GPL6244, GSE55940, GSE63514, GSE63678, GSE6791, GSE7410, GSE7803, GSE89657, GSE9750, and TCGA_GTEx). Blue represents noncancer samples and red represents CC samples. TCGA_GTEx, The Cancer Genome Atlas_Genotype-Tissue Expression.
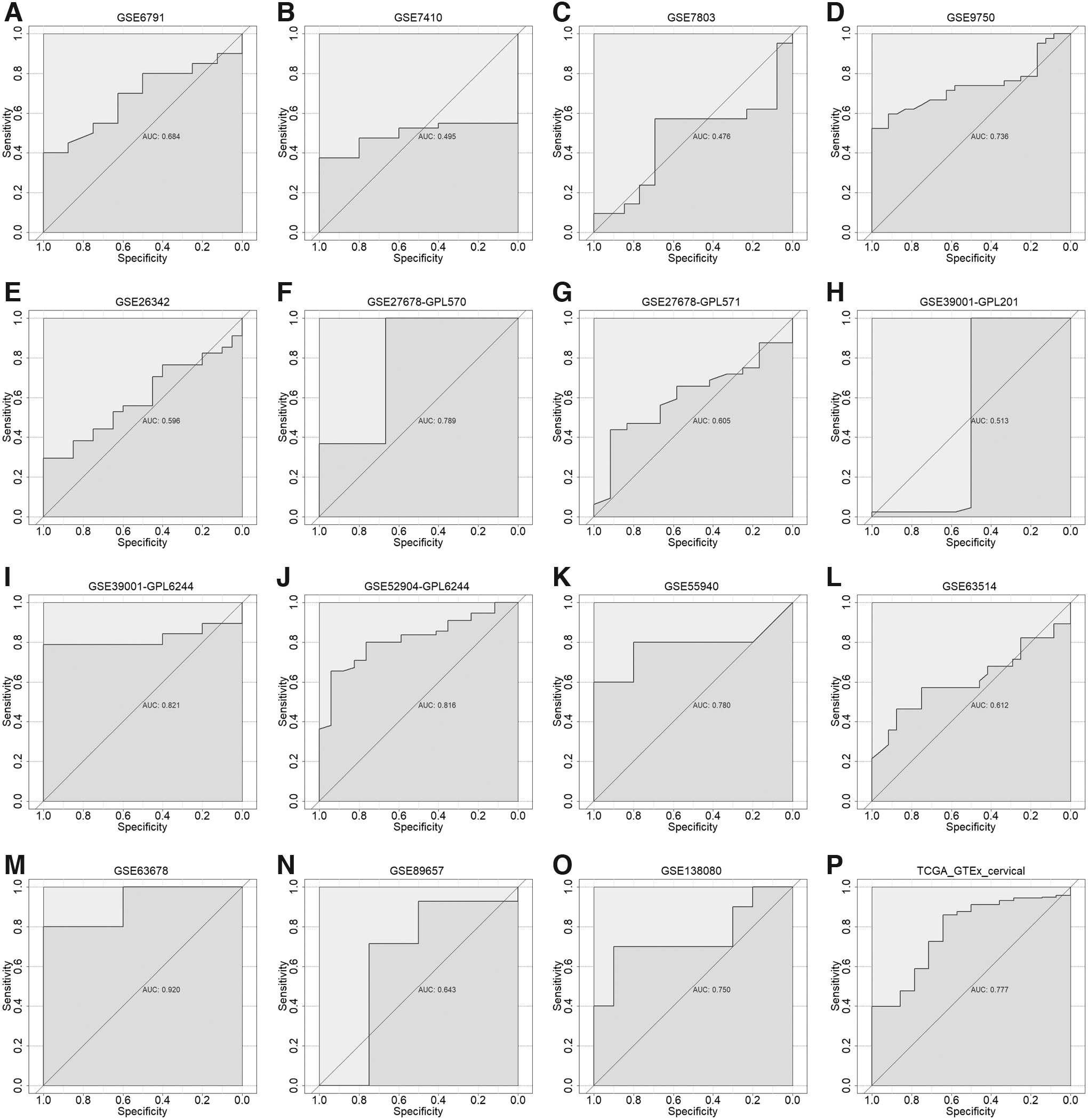
ROC curves of differentiating ability of LDH expression for CC.
The SMD Forest plot shows the significant overexpression of LDHA in CC samples (SMD = 0.71, 0.32-1.10; Fig. 4) and the SROC curves show the moderate differentiation capacity of LDHA expression for CC (AUC = 0.68, 0.64-0.72; Table 2 and Fig. 5). Kaplan-Meier survival curves for overall survival show that patients with CC with lower LDHA expression had a longer overall survival time than those with higher LDHA expression (p = 0.007; Fig. 6).
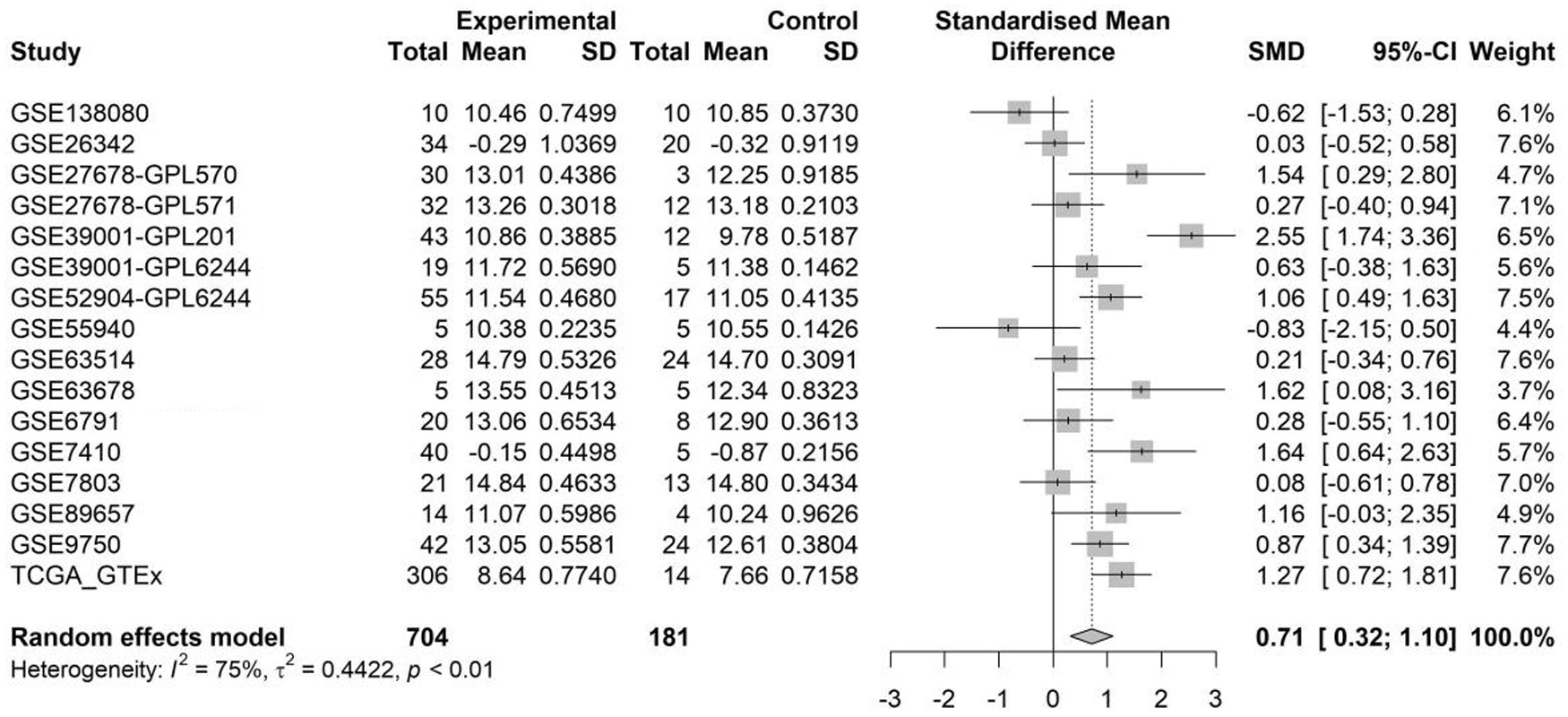
Forest plot of SMD for summarized LDH expression in CC for GSE138080, GSE26342, GSE27678-GPL570, GSE27678-GPL571, GSE39001-GPL201, GSE39001-GPL6244, GSE52904-GPL6244, GSE55940, GSE63514, GSE63678, GSE6791, GSE7410, GSE7803, GSE89657, GSE9750, and TCGA_GTEx data set. SD, standard deviation; SMD, standard mean difference.
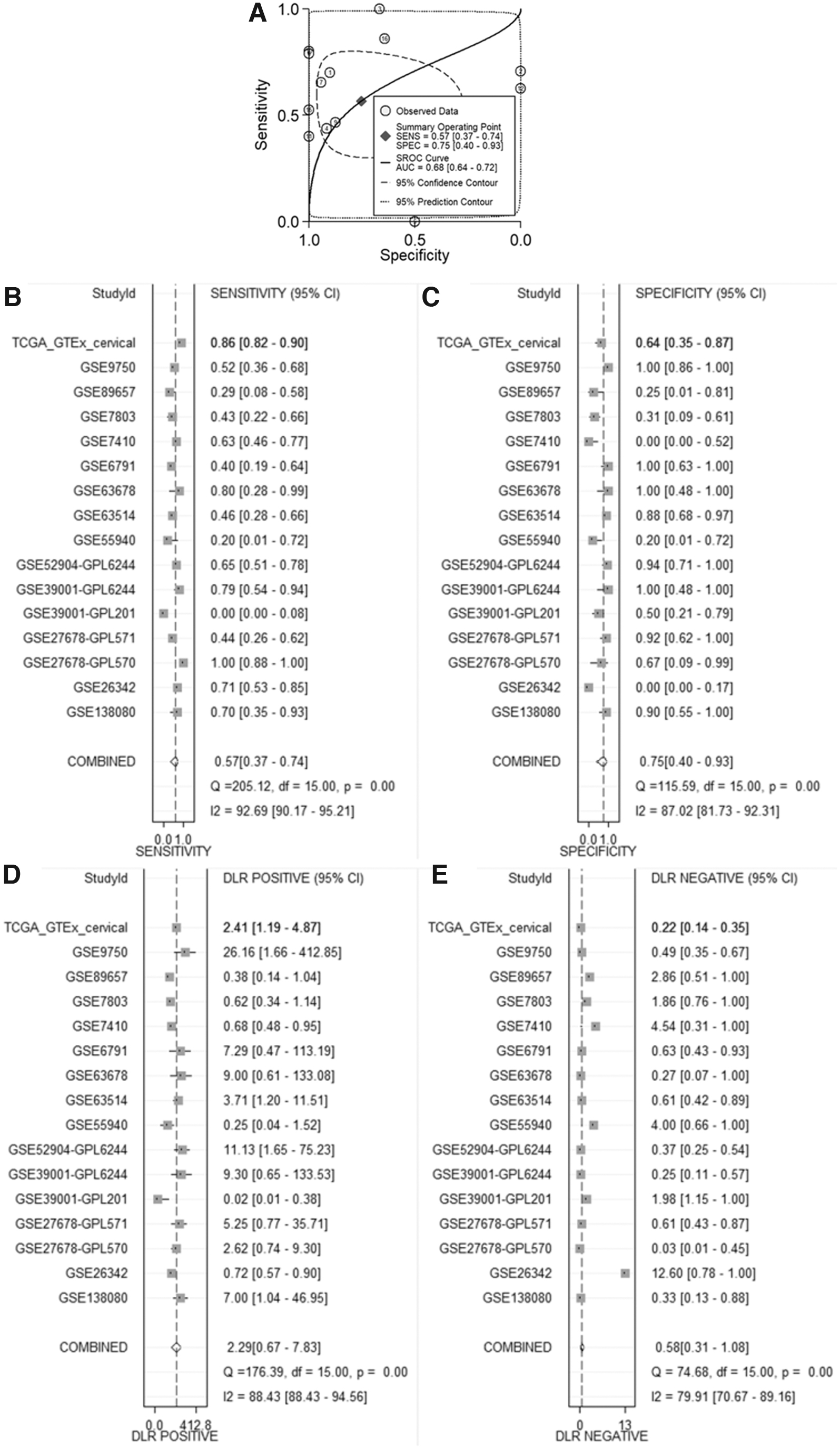

The summarized distinguishing ability of LDH expression for CC from all included microarrays and data set.
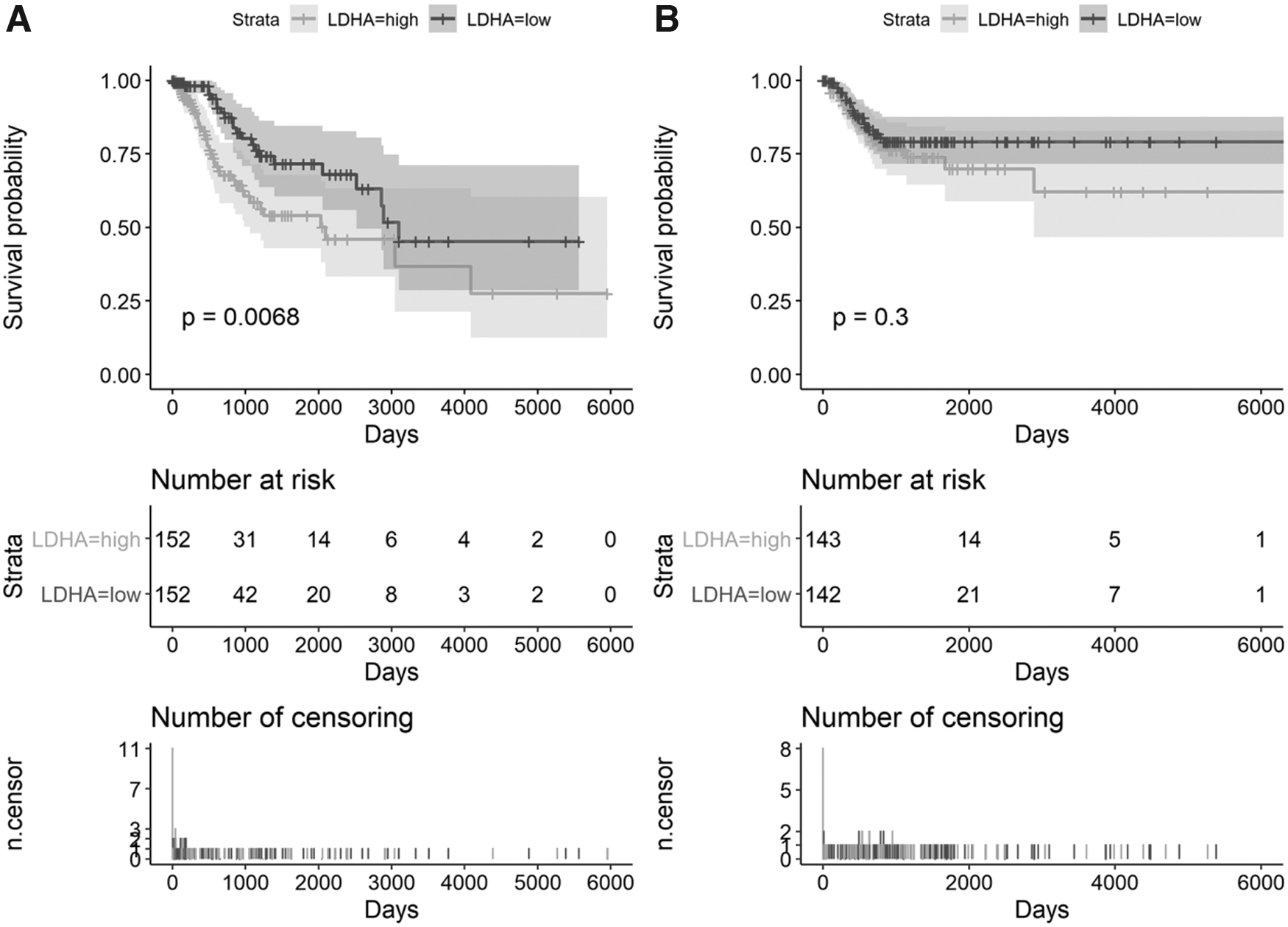
The prognostic value of LDH for CC based on RNA-seq data set.
Data Used to Plot the Summary Receiver Operating Characteristic Curves from All Included Microarrays and RNA-seq Data Set
FN, false negative; FP, false positive; TCGA_GTEx, The Cancer Genome Atlas_Genotype-Tissue Expression; TN, true negative; TP, true positive.
Functional enrichment analysis of genes coexpressed with LDHA
A total of 70 and 33 genes were positively and negatively coexpressed with LDHA, respectively. Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analyses were conducted using R packages. GO analysis consisted of three categories: biological process, cellular component, and molecular function. In the biological process group, the positively coexpressed genes were mainly enriched in nuclear division, organelle fission, mitotic nuclear division, and chromosome segregation. For cellular component group, the positively coexpressed genes were particularly enriched in the spindle, condensed chromosome, and chromosomal region. The top three significantly enriched terms were protein serine/threonine kinase activity, lyase activity, and carbon-oxygen lyase activity in the molecular function group.
Moreover, the cell cycle, oocyte meiosis, and cellular senescence were significantly enriched KEGG pathways of positively correlated genes (Fig. 7). The negatively coexpressed genes were mainly clustered in biological processes, such as second messenger-mediated signaling, platelet degranulation, and bone mineralization, and KEGG pathways, including aldosterone-regulated sodium reabsorption, malaria, and arachidonic acid metabolism (Fig. 8).
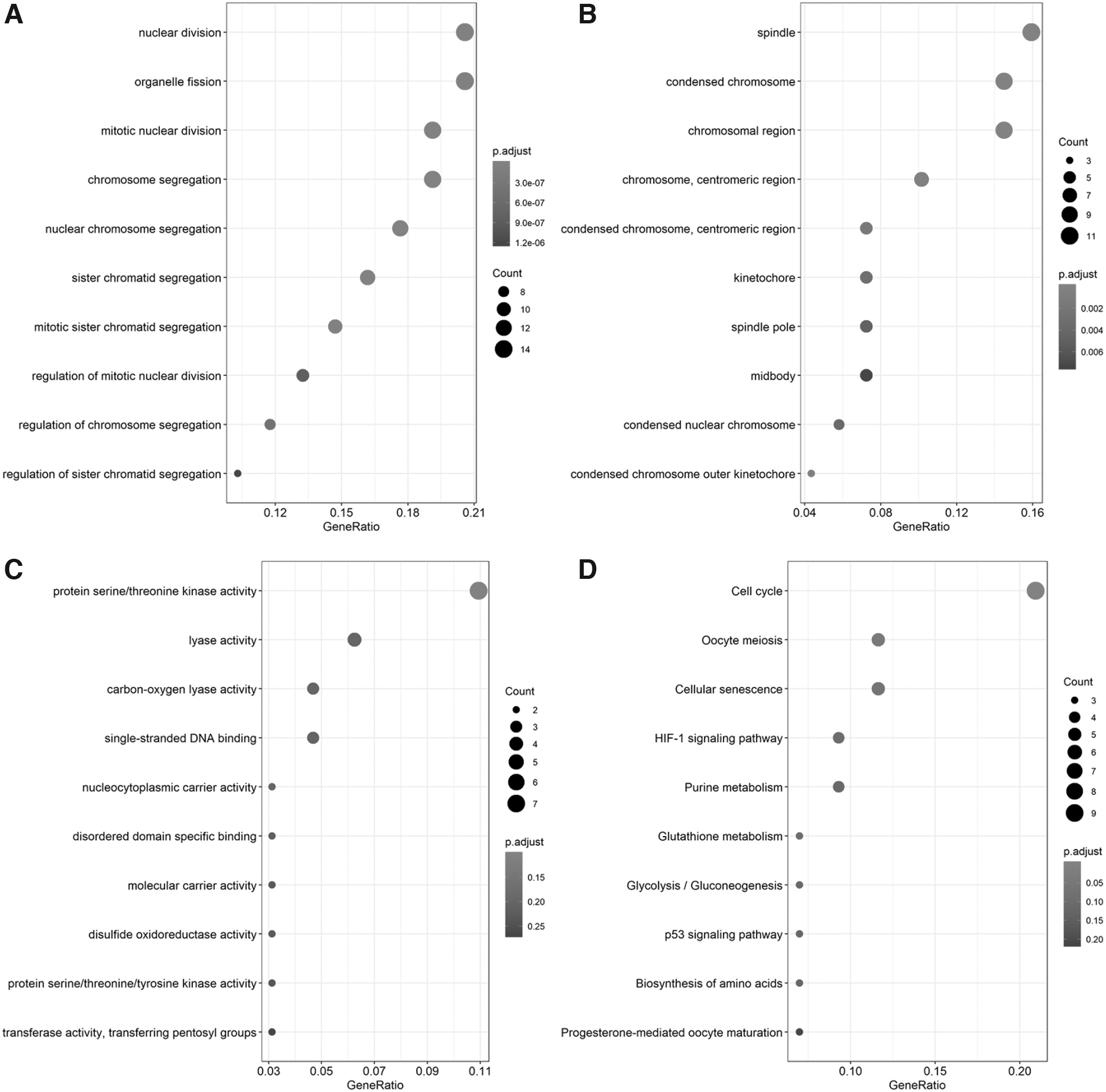
Functional enrichment analysis for positively coexpressed genes of LDH.
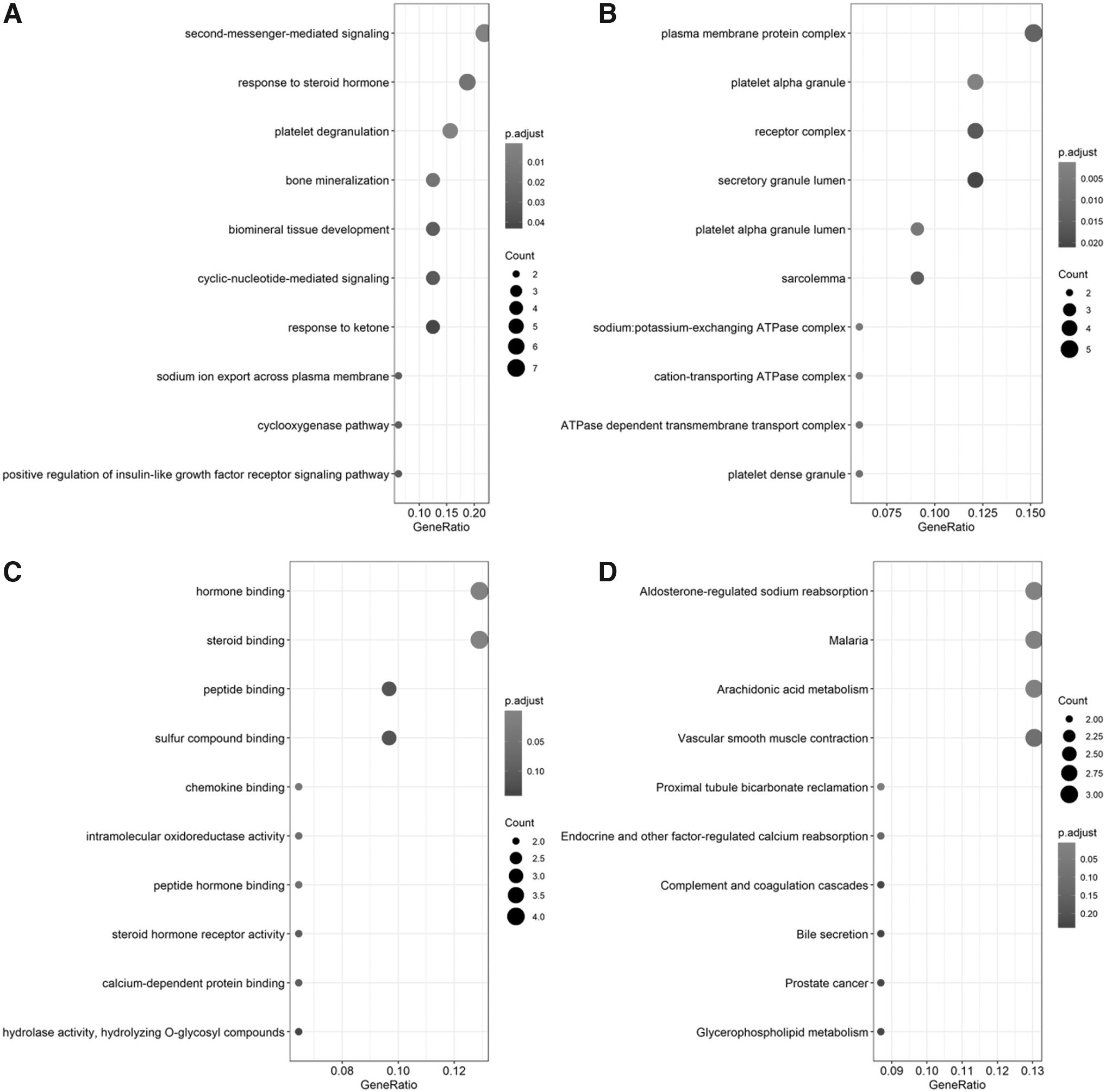
Functional enrichment analysis for negatively coexpressed genes of LDH.
Discussion
The aim of this study was to determine the expression of LDH in the serum of patients with CC and CC tissues in RNA-seq and microarray data sets. The large pool of 908 CC and 385 noncancer cervical samples and use of multiple detection methods ensured the reliability of our results. In this study, the results of Kaplan-Meier survival curves showed that patients with CC with higher LDHA expression had poor outcomes.
It is interesting that CC patients have better survival rate if LDHA expression was low, but it does not affect the recurrence of the CC or its progression among them according to our study. It may count on the late identification of CC patients. The most important result of this study was that both mRNA and protein LDH levels were upregulated in CC compared with those in healthy controls, and higher LDH expression indicated a worse overall survival in patients with CC.
Although several studies have evaluated LDH expression in CC, these studies were confined by the limited size of samples and superficial investigation of the molecular mechanism of LDH in CC. For example, Subramanian et al. (2008) suggested that patients with moderately differentiated cervical squamous cell carcinoma (grade 2) have higher levels of total LDH in both serum and cervical tissue than healthy controls; however, patients with poorly differentiated cervical squamous cell carcinoma (grade 3) exhibit a reduction in the level of total LDH in serum and cervical tissue, which differed from the results presented here. The study by Subramanian et al. used only 50 CC samples; thus, owing to the small sample size, the results may have been skewed in some way, which would cause instability in the results.
Owing to the Warburg effect, cancer cells generate more lactate than normal cells under both aerobic and hypoxic conditions. As this reaction is catalyzed by the A subunit of LDH, the presence of cancer cells may result in an increase in LDH levels in both serum and tissues (Augoff et al., 2015). Monitoring serum LDH levels in patients with breast cancer was done as an effective prognostic indicator in a report by Agrawal et al. (2016), who showed that higher serum LDH levels indicate a poor outcome, such as recurrence or metastasis. The main source of serum LDH is the LDH5 isoenzyme (Koukourakis et al., 2003a).
Several studies by Koukourakis et al. have shown that approximately half of cases of nonsmall-cell lung cancer and operable colorectal cancer with high tissue LDH5 levels have normal serum LDH levels, even before surgery. In the study of nonsmall-cell lung cancer, they observed that serum LDH levels drop sharply after surgery, which indicates that the rise in serum LDH levels may be caused by tumor tissue. Furthermore, LDH5 overexpression in tissue is significantly linked to highly proliferating prostate carcinomas and biochemical failure, which ultimately lead to an unfavorable prognosis (Koukourakis et al., 2003b, 2006, 2009, 2014).
There is a correlation between the expression of LDH and many human cancers. Studies have suggested that higher LDHA levels owing to anaerobic glycolysis cause the accumulation of lactic acid, which results in an acidic tumor microenvironment and promotes the invasion and metastasis of malignancies (Goldman et al., 1964; Fantin et al., 2006). Another study demonstrated that LDH5 expression is an independent prognostic marker of endometrial cancer that is linked with the impaired host immune response and activation of VEGFR2/KDR receptors in both cancer cells and tumor-associated vasculature (Giatromanolaki et al., 2006).
Oxamate, a weak LDHA inhibitor, inhibits LDHA activity by targeting its pyruvate binding pocket. An in vitro study of T98 and U87 glioblastoma cell lines showed that LDHA gene silencing or exposure to oxamate (30 mM) results in enhanced radiation sensitivity, especially in the T98 radioresistant cell line, which indicates that blocking the LDHA glycolytic pathway may be of therapeutic importance by enhancing the efficacy of radiotherapy and temozolomide against glioblastoma (Koukourakis et al., 2017).
To gain a deeper understanding of the molecular basis of LDH in CC, we carried out a functional enrichment analysis of the genes coexpressed with LDH. Different enrichment analyses based on GO and KEGG pathways were generated for positively and negatively coexpressed genes and indicated that these genes play various roles in the LDH-related oncogenesis of CC. We noted that the positively coexpressed genes were significantly correlated with nuclear division, the condensed chromosome, protein serine/threonine kinase activity, and the cell cycle, all of which are closely associated with tumorigenesis in human cancers. Specifically, protein serine/threonine kinase activity is involved in many cancers (Brault et al., 2010).
Moreover, serine/threonine kinase inhibitors have been shown to confer certain benefits in the treatment of ovarian cancer (Ciccone et al., 2016). Cell cycle regulators are strongly implicated in the progression of CC (Matsuda et al., 2013; Kim et al., 2014; Xu et al., 2014). Therefore, LDH might influence the occurrence and development of CC through these biological processes and pathways. Furthermore, Fantin et al. (2006) showed that a reduction in LDHA expression inhibits the proliferation of tumor cells under hypoxic conditions through the stimulation of mitochondrial respiration and reduction of mitochondrial membrane potential. For negatively coexpressed genes, biological processes such as second messenger-mediated signaling, platelet degranulation, and bone mineralization were clustered.
In the KEGG pathway, aldosterone-regulated sodium reabsorption, malaria, and arachidonic acid metabolism were included. Interestingly, functional enrichment analysis showed no significance in the correlation between genes related to glycolysis and LDHA. Glycolysis-related genes included GLUT1, HK1, and HK2. These three genes had feature in common with LDHA that they are all involved in glycoprotein binding, which is a molecular function that is commonly found on tumor cells (Massari et al., 2016; Ding and Zhang, 2018).
Nevertheless, this study contains several limitations. First, heterogeneity among all included microarrays and data set might have a detrimental effect on subsequent analysis due to the different platforms and patients of different races. Secord, functional enrichment analysis in our study is only related at the gene expression level instead of transcriptional level in CC. Therefore, prospective large-scale clinical studies are needed to further validate our findings. Third, during the preparation of serum, however, LDH release also may occur from platelets during clotting and from erythrocytes as a consequence of hemolysis.
Some studies pointed out that the buffer composition of the IFCC-recommended lactate-to-pyruvate conversion method (pH 9.40 ± 0.05) influenced the integrity of platelets and erythrocytes, and the presence of cellular aggregates in the top layer of the plasma was the main determinant for the occurrence of the high frequency of duplicate errors in the IFCC-recommended LDH method (Bakker et al., 2003, 2006; Herzum et al., 2003).
Conclusions
In conclusion, LDH upregulation might serve as a therapeutic and prognostic biomarker for CC. Further studies, including in vitro and in vivo experiments, to validate the interaction between LDH and coexpressed genes and the biological effect of LDH on CC should be conducted in the future to overcome the limitations of this study.
Footnotes
Authors' Contributions
Y.Y., M.C., and X.C. collected clinical data and data from public data sets. Y.Y., M.C., and X.C. performed the statistical analysis. F.L., L.L., and Y.Y. participated in the design of the study. Y.Y. and M.C. drafted the article and analyzed GO terms and KEGG pathways. X.C. and J.X. assisted in the drafting of the article. All authors have read and approved the final article.
Author Disclosure Statement
No competing financial interests exist.
Funding Information
No funding was received for this article.
