Abstract
Aims:
Pathogenic variants in mitochondrial DNA are known to be associated with sensorineural hearing loss (SNHL) and aminoglycoside-induced HL. Among them, the m.1555A>G mutation is the most common. Thus, a rapid and easy companion diagnostic method for this mutation would be desirable to prevent HL caused by aminoglycoside therapy. In this study, we report an improved protocol for the single-stranded tag hybridization chromatographic printed-array strip (STH-PAS) method for identifying the m.1555A>G mutation.
Methods:
To evaluate the accuracy of a novel diagnostic for the m.1555A>G mutation we analyzed 378 DNA samples with or without the m.1555A>G mutation, as determined by Invader assay, and calculated the sensitivity, specificity, and false negative and false positive ratios of this new method.
Results:
The newly developed protocol was robust; we, obtained the same results using multiple DNA concentrations, differing annealing temperatures, and different polymerase chain reaction thermal cyclers. The diagnostic sensitivity based on the STH-PAS method was 0.99, and the specificity was 1.00. The false negative and false positive ratios were 0 and 0.01, respectively.
Conclusion:
We improved the genotyping method for m.1555A>G mutations. This assays will be useful as a rapid companion diagnostic before aminoglycoside use.
Introduction
Among the pathogenic variants in mitochondrial DNA, the m.1555A>G mutation is one of the most common genetic causes of maternally inherited hearing loss (HL) (Usami et al., 2000b; Jacobs et al., 2005; Yano et al., 2014; Usami and Nishio, 2018) and it is also known to increase the susceptibility to aminoglycoside antibiotics and the risk for aminoglycoside-induced HL (Prezant et al., 1993). In general, aminoglycoside-induced HL is bilateral and severe to profound, occurring within several days after the administration of aminoglycoside antibiotics (Usami and Nishio, 2018).
Approximately 3% of outpatients and 10% of cochlear implant patients have a mitochondrial m.1555A>G mutation (Usami et al., 2000b). In addition, 0.1-0.7% of the general population from various ethnic backgrounds, also carry this mutation (Usami and Nishio, 2018; Maeda et al., 2020). The mitochondrial m.1555A>G mutation results in conformational changes in the stem-loop structure of the aminoglycoside binding site of mitochondrial 12S rRNA similar to that of the bacterial ribosome. It also decreases protein synthesis in the mitochondria and causes sensorineural hearing loss (SNHL) (Hobbie et al., 2008; O'Sullivan et al., 2017). We previously found that the patients with the m.1555A>G mutation who were exposed to aminoglycoside showed more severe HL (Usami et al., 2000a; Lu et al., 2009). Currently, we distribute a warning card to family members who have not been exposed to aminoglycoside antibiotics (Usami et al., 1999). However, to prevent aminoglycoside-induced HL, a rapid companion genetic screening system is desired.
To date, various genotyping methods for mitochondrial m.1555A>G mutations, including the polymerase chain reaction with restriction enzyme fragment length polymorphism (PCR-RFLP) method (Usami et al., 1997, 1999), Invader assay (Abe et al., 2007), TaqMan genotyping (Huang et al., 2015), multicolor melting curve analysis (Wang et al., 2017), DNA microarray (Lévêque et al., 2007), and next-generation sequencing (Zhong et al., 2013), have been reported. However, most of these methods require special equipment or procedures, such as real-time PCR or next-generation sequencing. In addition, some methods are quite time consuming; for example, PCR-RFLP requires 24 h or more, and a more rapid diagnostic method is desired for companion diagnosis.
Recently, genetic diagnosis of this mutation based on a single-stranded tag hybridization chromatographic printed-array strip (STH-PAS) genotyping method has been reported. This method is easy and rapid, allowing genetic testing using only a PCR system with a step-up cycling function (Tian et al., 2014; Kumondai et al., 2018).
In this study, we report an improved protocol for this STH-PAS-based method for m.1555A>G mutations that enables it to be performed on a standard PCR system and show the accuracy of this method based on a large number of DNA samples collected in our laboratory.
Subjects and Methods
Subjects
A total of 378 DNA samples from SNHL patients were examined: 177 samples from patients with the mitochondrial m.1555A>G mutation and 201 without this mutation. All mutations were analyzed by Invader assay (Abe et al., 2007). This study was conducted in accordance with the Declaration of Helsinki and written informed consent was obtained from all subjects (their next of kin, caretakers, or guardians in the case of children) before participation. This study was approved by the Shinshu University Ethical Committee (approval number 576) and the ethical committees of all other participating institutions are listed in the Acknowledgments section.
Primer design and genotyping of the m.1555A>G mutation
The primer sets for PCR amplification used in this study are listed in Table 1. Each reverse primer was labeled with a tag-spacer sequence at the 5′ end for detection with PAS, whereas the forward primer was labeled with biotin at the 5′ end (Fig. 1).
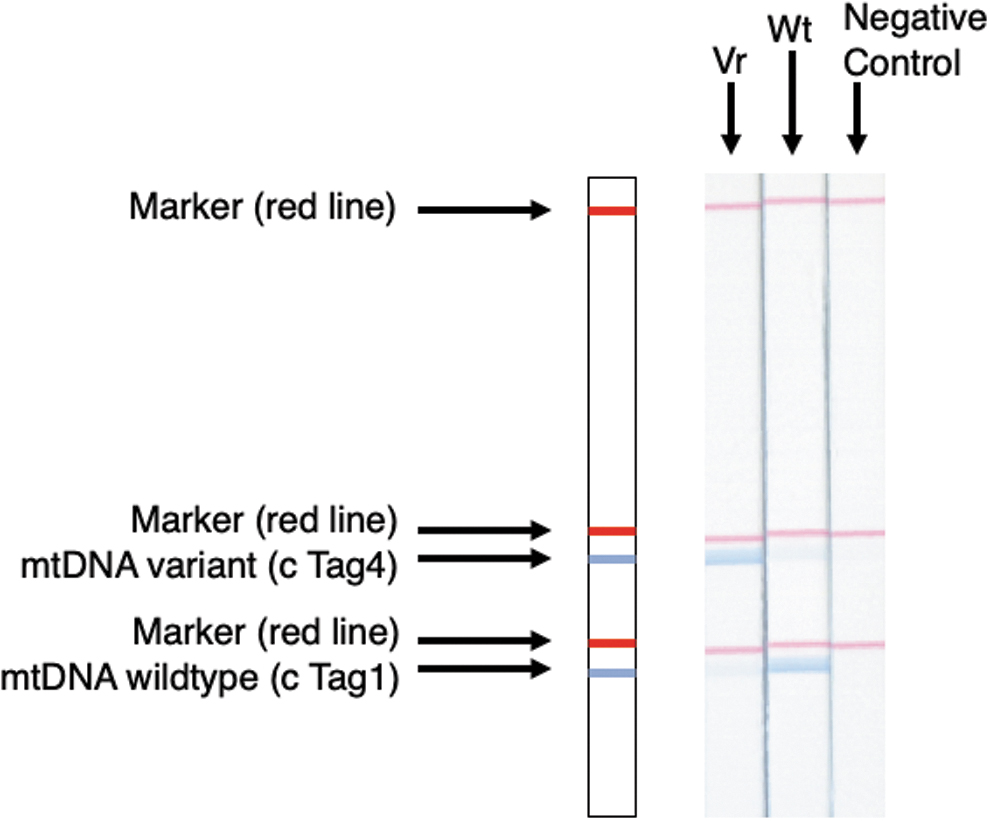
Schematic diagram of the STH-PAS-based genetic analysis and results of polymerase chain reaction amplicon product signals detected by STH-PAS. Wt: DNA sample without mutation (A residue), Vr: DNA sample with a mitochondrial m.1555A>G mutation (G residue). STH-PAS, single-stranded tag hybridization chromatographic printed-array strip.
Polymerase Chain Reaction Primers Used in This Study
PCR amplification was performed with 6.15 μL distilled water, 1.0 μL of 10× PCR buffer (with MgCl2), 0.8 μL of dNTP Mix (2.5 mM each), 1.5 μL of primer mixture (10 pM each of Bio_mt_1265F, F-1_mt1555A_Wt, and F-4_mt1555G_Var), 0.05 μL of Takara Taq (TaKaRa Bio, Shiga, Japan), and 0.5 μL of genomic DNA samples (concentrations of the DNA tested here were 250 ng/μL). The PCR temperature profiles involved an initial denaturation step at 94°C for 2 min; 30 cycles of denaturation at 94°C for 5 s, annealing at 60°C for 20 s, extension at 72°C for 10 s; and a final extension at 72°C for 2 min. The PCR was performed using a TaKaRa PCR Thermal Cycler Dice TP650 (TaKaRa Bio). STH-PAS diagnosis was performed using 20 μL of reaction solution containing 8.0 μL of distilled water, 10 μL of separation buffer (Tohoku Bio-Array, Sendai, Japan), 1.0 μL of streptavidin-coated blue latex suspension (Tohoku Bio-Array), and 1.0 μL of PCR product. The C-PAS4 membrane stick (Tohoku Bio-Array) was dipped into the mixture for 5 min at room temperature. The detailed mechanism for visualization with STH-PAS is reported in Kumondai et al. (2018). The diagnosis of each sample was based on the STH-PAS results (Fig. 1). The blue signals in different positions showed different genotypes in the mitochondrial 1555 residue. Some samples showed double signals on the PAS (in which blue bands were observed for both the wild type and variant), reflecting the heteroplasmy of this mitochondrial DNA position.
Accuracy of the STH-PAS-based method for the mitochondrial 1555 A>G mutation
To evaluate the accuracy of this method, a total of 378 DNA samples from SNHL patients (177 samples with the mitochondrial m.1555A>G mutation and 201 without this mutation) were analyzed using the STH-PAS-based method and the results were compared with those obtained by Invader assay under randomized blind conditions. Invader assay is used as the gold standard to detect this mutation in Japan and is also used for social health insurance-based genetic testing. The sensitivity, specificity, and false negative and false positive ratios were calculated by comparing the results obtained by the STH-PAS-based method and Invader assay.
Results
Pilot studies showed that this STH-PAS-based method works well even under different concentrations of template DNA, different annealing temperatures, and using different PCR thermal cyclers (Supplementary Fig. S1). The results of the comparison between the STH-PAS-based method and Invader assay for the mitochondrial m.1555A>G mutation are summarized in Table 2. Most of the results obtained with this STH-PAS-based method were correct. There were two false negative but no false positive cases. One of the two false negative cases carried the m.1555A>G mutation as heteroplasmy in very low frequency. A double signal for both the wild-type and mutated sequence was observed in five samples. All of these five samples carried this mutation and the wild type with various heteroplasmy ratios, as confirmed by Invader assay. Among 177 samples with the mitochondrial m.1555A>G mutation, 6 samples were heteroplasmic. The sensitivity of this method was 0.99 and the specificity was 1.00. The false negative and false positive ratios were 0 and 0.01, respectively.
Results of a Randomized Blind Comparison of the STH-PAS-Based Testing and Invader Assay
STH-PAS, single-stranded tag hybridization chromatographic printed-array strip.
This STH-PAS-based method provided results within a 2-h turnaround time including the extraction of DNA from blood or oral epithelial cells, the preparation of the master mix, and the PCR amplification and visualization processes.
Discussion
In this study, we developed a rapid, robust, and easy-to-use diagnostic method for the mitochondrial m.1555A>G mutation. Based on the results of our randomized blind comparison, we concluded that this method is sufficiently reliable for clinical diagnosis and is robust enough for use in different clinical settings. Thus, it will be useful as a companion diagnostic tool for aminoglycoside antibiotics use.
Among the 177 positive control samples, 5 showed double signals (a mutation signal and a wild-type signal) on the STH-PAS analysis, and all 5 of these cases carried the mutation at various heteroplasmy ratios, as confirmed by Invader assay. In previously reported cases, the m.1555A>G mutation was identified as homoplasmic in most cases and only a limited number of patients carried this mutation as heteroplasmy (del Castillo et al., 2003; Lu et al., 2009).
In this study, we could not detect the mutation in two samples. One had a heteroplasmic mutation with a lower mutation frequency, whereas the other case had a homoplasmic mutation. A total of 6 of the 177 positive samples carried this mutation as heteroplasmy. The heteroplasmic ratio of the six samples analyzed here were 1.4%, 7.6%, 8.1%, 23.6%, 27.9%, and 43.5%, with one false negative sample carrying this mutation at a 1.4% heteroplasmic ratio, so we considered this false negative was caused by this very low heteroplasmic ratio under the detection threshold for the STH-PAS-based method. A previous report showed that there is variability in the heteroplasmic ratio of this mutation and this variability is proposed as one of the causes for its different clinical characteristics (Rossignol et al., 2003). In general, patients with this mutation at a lower heteroplasmic ratio would be regarded as at lower risk for aminoglycoside-induced HL. However, the detailed threshold is unknown. As the heteroplasmic ratio may differ among tissues, and the DNA samples could not be extracted from the cochlea, it would be important to find patients with a lower heteroplasmic ratio and instruct them to avoid aminoglycoside use. For the other false negative sample with a homoplasmic mutation, we performed direct sequencing analysis but could not identify any mutation in the PCR-amplified region or its peripheral region. Although unknown factors may be involved in this particular case, this method shows sufficient reliability for companion genetic testing.
The mitochondrial m.1555A>G mutation increases the risk for aminoglycoside-induced HL, but aminoglycoside antibiotics are still used in many countries, especially for patients with tuberculosis, methicillin-resistant Staphylococcus aureus infections, or neonatal intensive care unit babies (Van Boeckel et al., 2014; Krzyżaniak et al., 2016). Meanwhile, newly developed aminoglycoside antibiotics have been proposed (Becker and Cooper, 2013). However, careful attention is needed even with the newly developed aminoglycoside antibiotics, and it has been reported that these drugs are also responsible for HL (Usami et al., 1998). Thus, a rapid, simple, and reliable companion diagnostic method before the use of aminoglycoside antibiotics is important.
This STH-PAS-based method for identifying the mitochondrial m.1555A>G mutation has several advantages in comparison with other methods. This method only requires a commonly used standard PCR thermal cycler (without step-up function). In addition, this method requires little technical experience and provides results within 80 min (including the time for preparation of the master mix and the visualization processes). In this study, although there was no amplicon product contamination, it would be possible to improve our STH-PAS-based diagnostics by employing dUTP and uracil DNA glycosylase for the PCR step to prevent such contamination.
In addition, as this method only requires a PCR thermal cycler, it has been reported that the cost of this method is ∼$7 per sample (Kumondai et al., 2018). We believe this method based on STH-PAS will be beneficial as a tool for companion diagnosis in many countries as this mutation was identified in many ethnic populations (e.g., the Japanese, Mongolian, Chinese, Turkish, Tunisian, Greek, Polish, Spanish, Italian, and American populations) (Usami and Nishio, 2018).
Footnotes
Authors' Contributions
All authors participated in the design and interpretation of the study and analysis of the data; Y.I., and S-y.N. performed the experiments; S-i.U. administered and supervised the project, and acquired funding; Y.I. wrote the original article; and S-y.N., E.H, M.H., and S-i.U. reviewed and edited the article.
Acknowledgments
We thank the participants of the Deafness Gene Study Consortium (see Nishio et al., ![]() ) for providing samples and clinical information. The Deafness Gene Study Consortium comprises the following institutions: Hokkaido University, Hirosaki University, Iwate Medical University, Tohoku University, Yamagata University, Fukushima Medical University, Jichi Medical University, Gunma University, Nihon University School, Nippon Medical School, Nippon Medical School Tama Nagayama Hospital, Jikei University, Toranomon Hospital, Kitasato University, Hamamatsu Medical University, Mie University, Shiga Medical Center for Children, Osaka Medical College, Hyogo College of Medicine, Kobe City Medical Center General Hospital, Wakayama Medical University, Okayama University, Yamaguchi University, Ehime University, Kyushu University, Fukuoka University, Nagasaki University, Kanda ENT Clinic, Miyazaki Medical College, Kagoshima University, and Ryukyus University.
) for providing samples and clinical information. The Deafness Gene Study Consortium comprises the following institutions: Hokkaido University, Hirosaki University, Iwate Medical University, Tohoku University, Yamagata University, Fukushima Medical University, Jichi Medical University, Gunma University, Nihon University School, Nippon Medical School, Nippon Medical School Tama Nagayama Hospital, Jikei University, Toranomon Hospital, Kitasato University, Hamamatsu Medical University, Mie University, Shiga Medical Center for Children, Osaka Medical College, Hyogo College of Medicine, Kobe City Medical Center General Hospital, Wakayama Medical University, Okayama University, Yamaguchi University, Ehime University, Kyushu University, Fukuoka University, Nagasaki University, Kanda ENT Clinic, Miyazaki Medical College, Kagoshima University, and Ryukyus University.
Author Disclosure Statement
No competing financial interests exist.
Funding Information
This study was funded by a Health and Labor Sciences Research Grant for Research on Rare and Intractable Diseases and Comprehensive Research on Disability Health and Welfare from the Ministry of Health, Labour and Welfare of Japan [S.U. H29-Nanchitou (Nan)-Ippan-031], and a Grant-in-Aid from the Japan Agency for Medical Research and Development (AMED) (S.U. 16kk0205010h0001, 18ek0109363h0001).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
