Abstract
Filipino American women (FAW) have high incidence of coronary heart disease and high low-density lipoprotein cholesterol (LDL-C). The distribution of rs11206510 proprotein-convertase subtilisin-kexin type 9 (PCSK9) and rs1122608 low-density lipoprotein receptor (LDLR) single nucleotide polymorphisms (SNPs), known for genetic influences on LDL-C, is unknown in this population. The objective of this study was to examine the genetic determinants of LDL-C, their association with LDL-C, and effects of statins on LDL-C given the genetic determinants in this high-risk population. Data were obtained from the FAW Cardiovascular Study (N = 338) of women ages 40 to 65 years from four major U.S. cities between 2011 and 2013. Roche Modular methodology and Luminex-oligonucleotide ligation assay procedure were used for allele frequency, genotype, LDL-C, and lipid analysis. Analysis of variance was used to determine differences between genotype groups. Genotype and statin effect on LDL-C were tested using the generalized linear model procedure of SAS. The distribution of rs11206510 PCSK9 genotypes was 88% TT, 11% TC, and 1% CC, and the rs1122608 LDLR genotype distribution was 83% GG, 17% GT, and 0% TT. These SNPs showed no association with mean LDL-C in this cohort. FAW on statin medications had lower LDL levels regardless of their PCSK9 or LDLR genotypes. Most FAW had a gain-of-function allele of PCSK9 and LDLR. This predominance in FAW cohort may account for the high percentage of subjects with elevated LDL-C. In a population at high risk for hypercholesterolemia, optimal treatment with statins should be considered where appropriate.
Introduction
Filipino Americans, the second largest Asian American subgroup, are at a high risk for cardiovascular disease (CVD)1,2 and risk factors such as dyslipidemia/overweight,3–5 metabolic syndrome, 6 diabetes,7,8 hypertension, 6 and health risk behaviors. 9 Coronary heart disease is found to occur at a significantly higher rate and at much younger ages in Filipinos compared to non-Hispanic Whites (NHW).1,2,10 Metabolic syndrome, a known precursor for coronary heart disease, is similarly prevalent in both Filipino Americans (18.3%) and native Filipinos (18.6%),10–12 a concept that is also demonstrated among native Filipino women and Filipino American women (FAW) (52% vs. 55%, p = 0.08). 13
While Filipino men have a greater proportionate mortality ratio (PMR) from ischemic heart disease (PMR = 1.15) compared to NHW and Filipino women (PMR <1), 1 as well as an increased rate of metabolic syndrome (21.1% Filipino American men, 15.7% FAW),11,12 FAW continue to have 66% higher odds of coronary heart disease in comparison to NHW. 2
Many Filipino Americans have elevated low-density lipoprotein cholesterol (LDL-C) levels with 70% higher odds of high LDL-C compared to NHW. Hyperlipidemia is an established coronary heart disease risk factor, and therefore, prevention and treatment should include screening efforts followed by aggressive treatment if necessary for elevated LDL-C according to the latest American College of Cardiology/American Heart Association Guideline on the Treatment of Blood Cholesterol to Reduce Atherosclerotic Cardiovascular Risk in Adults. 14 In the Filipino population, no surrogate biomarker for LDL-C has been validated at this time. While both Filipino men and women were found to have high C-reactive protein (CRP) and high prevalence of low high-density lipoprotein cholesterol (HDL-C), 75% in men and 73% in women, respectively, 15 no study on LDL-C and HDL-C association with CRP has been done. In addition, despite the finding that FAW were shown to have more coronary artery calcium progression in comparison to their White counterparts, 16 no literature currently exists describing the association of high LDL-C or low HDL-C with coronary artery calcium in the Filipino population.
The mainstay of hyperlipidemia treatment has been statin medication but the demonstration that genetic mutations in proprotein-convertase subtilisin-kexin type 9 (PCSK9), as well as low-density lipoprotein receptor (LDLR), which cause an increase or decrease in LDL-C levels has opened up other options for the treatment of elevated LDL-C. 17
PCSK9 is a 692 amino acid glycoprotein that is mainly synthesized in the liver and secreted into the blood. 18 It is a member of the subtilisin-like serine convertase superfamily involved with proteolytic processing of protein precursors. Physiologically, PCSK9 regulates cholesterol homeostasis by binding to the LDL-C at the cell surface, directing it to intracellular lysosome-like components for degeneration of the receptor. Loss-of-function mutations increase LDLR expression, thereby increasing LDL clearance. 19 In a study by Sirois et al., nonsense PCSK9 mutations in sub-Saharan Africans are hypothesized to protect against emerging CVD. 18 Conversely, gain-of-function mutations of PCSK9 genes reduce LDLR expression on the cell surface, decreasing LDL clearance, effectively increasing serum levels of LDL cholesterol.
Another gene locus identified influencing LDL-C, LDLR gene, involves a functional mutation on the LDL receptors. LDL receptors are located principally on liver cells. Impairment of the function of LDL receptors results in reduced clearance of LDL particles from the circulation, thereby increasing plasma LDL cholesterol levels.
In this study, we looked at the rs11206510 PCSK9 and rs1122608 LDLR gain-of-function mutations of the major “T” allele and “G” allele, respectively, two mutations that have been reproducibly associated with high LDL cholesterol levels and myocardial infarction.20,21 Conversely, carriers of the rs1122608 “T” minor allele were shown to have a favorable lipid profile and lower risk for coronary artery disease. 20
Monoclonal antibodies that inhibit PCSK9, PCSK9 inhibitors, are a new group of lipid-lowering medications approved in 2015 by the U.S. Food and Drug Administration. Treatment for hyperlipidemia includes use of statin therapy, as well as PCSK9 inhibitors, for statin-refractory patients and for those with familial hypercholesterolemia. FAW are at an increased risk for elevated LDL-C 3 and, therefore, may require additional lipid-lowering therapy despite diet and optimal statin therapy, at which point the use of these new agents may be considered.
Genetic loci for mutations associated with LDL-C hypothesized to influence cardiovascular risk, including PCSK9 and LDLR single nucleotide polymorphisms (SNPs), were identified by Akao et al. in 2012. 22 Several studies since then have investigated the effects of the PCSK9 and LDLR polymorphisms in LDL-C. A meta-analysis by Zhang et al. suggested that both PCSK9 and LDLR gene polymorphisms were associated with coronary heart disease in Caucasians but not in Asians, with Asian studies mainly investigating the Chinese population. 23 While these results could be attributed to genetic heterogeneity between the Asians and Caucasians and innate ethnic differences, the relatively small sample size of the study, as well as other literature in Han Chinese 24 showing an association of rs11206510 with early-onset coronary heart disease, suggests the need for further high-powered investigations to elicit possible genetic association in the Asian cohorts. In addition, it is important to note that CVD and survival among diverse Asian American cohorts are known to be heterogeneous, 1 thus limiting generalization across all Asian groups. While there are no formal studies available investigating the effect of Western diet on cardiovascular risk in Filipino Americans compared to native Americans, it is believed that both the native Filipino and the Filipino American population have an increased risk for CVD given the composition of both traditional Filipino and Western diet. Traditional Filipino cuisine is high in sodium and fat and is also coupled with the addition of rice in each meal; Western diet, in contrast, is high in fat and sugar. 10
With the knowledge that FAW are at increased risk for elevated LDL-C, it is important to determine if PCSK9 and LDLR mutations contribute to this elevated risk. Moreover, it is important to determine if statin treatment is effective in this population of FAW. To begin examining these questions, we identified the distribution of the gain-of-function rs11206510 PCSK9 “T” allele and LDLR rs1122608 gain-of-function mutation of the “G” allele mutation in this cohort of FAW. Furthermore, we examined the association between the PCSK9 and LDLR risk alleles and LDL-C levels and investigated the influence of both SNPs in relation to LDL levels and statin therapy.
Materials and Methods
Study design and recruitment of participants
Approval of the university institutional research board (IRB) was obtained before the conduct of this study. This was a descriptive cross-sectional study conducted in a cohort of Filipino women (N = 338) who participated in an all women cardiovascular health screening event from four cities of the United States between 2010 and 2013. Subjects were between 40 and 65 years of age. Health screenings were performed in various U.S. cities: Jacksonville and Tampa, FL; Chicago, IL; and San Francisco, CA. Women who self-reported as Filipinos between ages 40 and 65 years and had fasted for at least 8–12 h were eligible for the study. Potential participants who had severe arthritis, any autoimmune disorder, recent cancer diagnosis, and/or presented with any infection or severe inflammation were excluded from the study. A written informed consent was obtained after the purpose of the study was thoroughly explained.
Screening protocol
A demographic and clinical information questionnaire, including participants' medical history and current medications, was completed for each subject by trained research staff. The clinical information questionnaire included family history, smoking history, participant's medical history, and current medications. Vital signs and morphometric measurements were measured using a developed standard study protocol. A licensed phlebotomist drew 5 mL of blood through venipuncture from each participant for a series of tests, including the cardiovascular lipid, inflammatory, and genetic biomarkers. The lipid panel assay included total cholesterol (TC), triglycerides (TG), HDL-C, and LDL-C, Roche Modular methodology, performed at Berkeley HeartLab, Inc. (Alameda, CA). The National Cholesterol Education Program (NCEP) Adult Treatment Panel III guidelines were used to define the cholesterol normal reference values, namely: TC should be ≤200 mg/dL, TG ≤150 mg/dL, HDL-C ≥50 mg/dL, and LDL-C ≤100 mg/dL. 25 All specimens were sent to a CLIA certified laboratory where clinical standard protocols of these assays were performed.
Genetic analysis
Celera research reagents were used to genotype the SNPs of each subject in a single-tube assay using the Luminex-oligonucleotide ligation assay (OLA) procedure, which involved amplification of genomic DNA (∼3 ng) by multiplex PCR followed by multiplex OLA. The ligation products were hybridized to Luminex® xMAP microspheres (Luminex Corporation, Austin, TX) and later labeled by the reporter molecule Streptavidin Phycoerythrin (SA-PE). The xMAP microspheres were analyzed on a Luminex 100 or Luminex 200 system. Genotypes were then determined using the Celera allele calling software as described by Iannone et al. 26
Data analysis
Data were analyzed using the Statistical Package for the Social Sciences (Version 9.4; SAS Inst., Inc., Cary, MC) software. Means and standard deviations were determined for all continuous variables, and number and percentage were determined for categorical variables. Analysis of variance with Tukey's post hoc tests was used to determine differences between groups for these variables. Statistical significance was set at p < 0.05. The effects of PCSK9 and LDLR genotypes and statin medication on LDL levels were tested using the generalized linear model procedure of SAS (Version 9.4; SAS Inst., Inc.). The model included the fixed effects of both PCSK9 and LDLR genotypes and statin medications on the outcome variable LDL-C and their interactions.
Results
Table 1 presents the demographic and relevant clinical data of the study population (N = 338). The mean age of the participants was 53 years, and 97% were born in the Philippines. Many participants did not have optimal lipid levels. Forty-seven percent had elevated TC (≥200 mg/dL), 22% had elevated TG (≥150 mg/dL), 65% had elevated LDL-C (≥100 mg/dL), and 27% had low HDL-C (≤50 mg/dL).
Demographic and Lipid Profile Data for the Filipino American Women Study (N = 338)
Elevated values: TC ≥200 mg/dL, TG ≥150 mg/dL, LDL-C ≥100 mg/dL, HDL-C ≤50 mg/dL.
TC, total cholesterol; TG, triglycerides; HDL-C, high-density lipoprotein cholesterol; LDL-C, low-density lipoprotein cholesterol.
The number of subjects with each of the PCSK9 rs11206510 genotypes and their use of statin medication is presented in Table 2. For the PCSK9 rs11206510 SNP, the genotype frequencies were 88% TT, 11% TC, and 1% CC; thus, 99% of this cohort had at least one copy of the T gain-of-function allele. The TT and TC groups without statin use had similar mean LDL-C levels (p = 0.9; Fig. 1). The CC genotype was not included in the analysis due to low subject numbers. The majority of both the TT (73%) and TC (78%) groups did not take statins although the mean concentration of LDL-C was ∼120 mg/dL for both groups (Fig. 1). However, the mean LDL-C concentration was significantly lower (p < 0.01) in TT statin compared to the TT non-statin group. Although the LDL-C mean was lower for the TC statin group, the difference was not statistically significant between the TC statin and TC non-statin groups (p = 0.13), likely due to the smaller sample size.
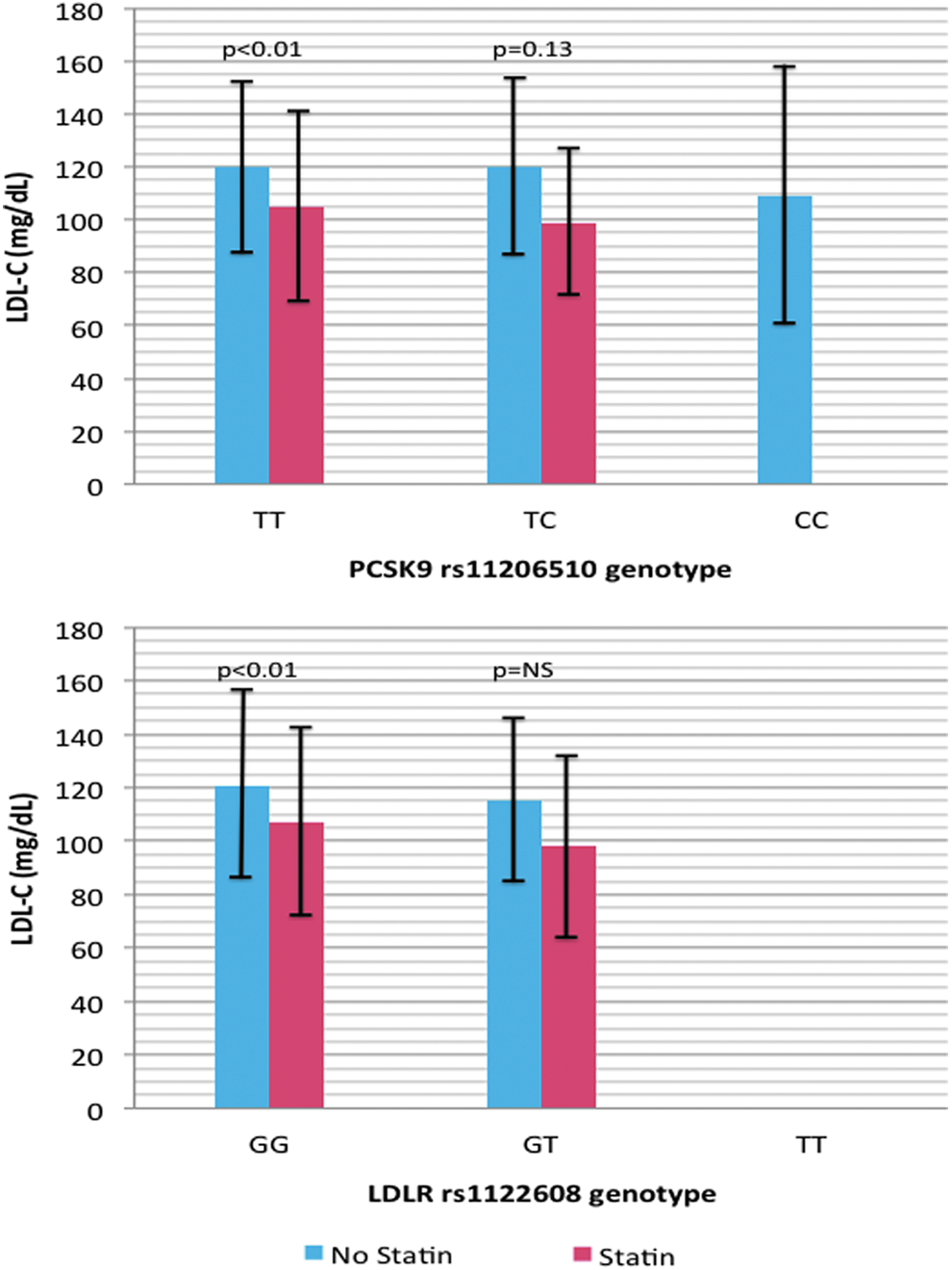
Mean LDL-C levels ± standard deviation as a function of genotype and statin use. LDL-C, low-density lipoprotein cholesterol.
Number of Subjects and Percentage of Group Taking Statins as a Function of Genotype
LDLR, low-density lipoprotein receptor; PCSK9, proprotein-convertase subtilisin-kexin type 9.
The genotype frequencies of the LDLR rs1122608 SNP were: 83% GG, 17% GT, and 0% TT; thus, 100% of this cohort had the gain-of-function allele (Table 2). The GG and GT genotypes had similar mean LDL levels (p = 0.12; Fig. 1). The mean LDL-C concentration was significantly lower (p < 0.01) in the statin GG group (107 ± 35 mg/dL) versus the non-statin GG group (121 ± 35 mg/dL), but LDL-C concentration was not significantly lower in the GT statin group versus the GT non-statin group possibly due to the smaller sample size. The majority of both the GG (75%) and the GT (68%) did not take statin medications although the mean statin concentration was higher than the recommended LDL-C concentration for both genotypes.
Table 3 presents the results of the analysis of the combined effects of the PCSK9 and LDLR genotypes and statin medications on the outcome variable LDL-C. This analysis demonstrated no three- or two-way interactions as a function the LDLR or PCSK9 polymorphism on LDL-C concentration. Statin medication use was associated with reduced LDL-C (p < 0.05), independent of PCSK9 and LDLR genotype.
Individual and Combined Effects and Interactions of PCSK9 rs11206510 Mutation, Low-Density Lipoprotein Receptor rs1122608 Mutation, and Statins on Low-Density Lipoprotein Cholesterol
Pr >F, the significance probability value associated with the F value; SS, sum of squares.
Discussion
In this cohort of FAW, the majority carries the gain-of-function risk allele for PCSK9 (99%), as well as LDLR (100%). FAW participants who carry both the PCSK9 and LDLR gain-of-function risk allele (TT and GG) had significantly lower LDL levels while on statin therapy. By examining three- or two-way interactions involving the LDLR and PCSK9 genotypes and statin medication, further analyses showed that the two studied polymorphisms of LDLR and PCSK9 had no effect on LDL-C. Statin therapy lowered LDL-C, irrespective of the polymorphisms. Both the rs11206510 PCSK9 and the rs1122608 LDLR mutations did not affect lipid-lowering effect of statin medications.
Distribution of rs11206510 PCSK9 and rs1122608 LDLR
To our knowledge, this is the first time that the genotypic distribution of the rs11206510 PCSK9 and rs1122608 LDLR SNPs is being described for the Filipino population.
As noted above, the genotype frequencies for the PCSK9 SNP for our Filipino American cohort were 88% TT, 11% TC, and 1% CC. Most of the studied population had either one or two copies of the T gain-of-function allele, with proportions similar to the other studied populations of Czech Republic, Italy, Netherlands, Finland, and China, where more than 80% of the population carried the risk allele.20,24,27–29
As seen in Appendices A1 and A2, our results showed that the prevalence of PCSK9 and LDLR is similar to those studies conducted in a population in Italy and Netherlands that found the PCSK9 minor allele C in 20% and 19% of their subjects, respectively.20,27 A study by Vrablík et al. in 2012 of 1567 patients in Prague showed a genotype distribution as follows: 65% TT, 30% TC, and 5% CC, while Kunnas and Nikkari in 2015 described the genotype distribution in a Finnish population of 769 patients as follows: 73% TT, 26% TC, and 2% CC. Both studies demonstrated that the carrier risk T allele is present in the majority of the two populations, similar to the FAW in our study.28,29
In a study of a prospective cohort of 5482 patients in Netherlands, the minor allele frequency of the LDLR SNP was found to be 24%, similar to our findings of 17% in FAW. 20 A study by Martinelli et al. in 2010, including 983 patients in Italy, showed a genotype distribution as follows: 55% GG, 38% GT, and 6% TT, demonstrating that majority of this population carries the risk allele G, as in our study. 21 The rs1122608 LDLR genotype distribution, 49% GG, 42% GT, and 9% TT, was found in a study by Jamaldini et al. in 2013 that was conducted on a population of 274 patients in Iran, also similar to the results of this study. 30
In the analysis of genotype frequencies in the Filipino American population, it is important to note that this population has significant genetic admixture owing to its colonial (Spanish, Dutch, Portuguese, British, American, and Japanese) history. Thus significant genetic heterogeneity may be expected. In the future, genetic testing for identified risk alleles may play a role in further stratification of this heterogeneous population.
Effect of rs11206510 PCSK9 and rs1122608 LDLR SNPs on LDL cholesterol
Our results showed no effect of the PCSK9 and LDLR SNPs on LDL cholesterol. This is in contrast to previous studies such as that of Guella et al. that examined 1880 Italian patients with 1880 age-matched controls and showed a strong correlation between the rs11206510 PCSK9 SNP and LDL and TC. 27 A study by van de Woestijne et al. also showed association of both rs11206510 PCSK9 and rs1122608 LDLR mutations and LDL-C levels irrespective of lipid-lowering medication. 20 The Tampere Adult Population Cardiovascular Risk Study (TAMRISK) of 769 Finnish patients in 2015 showed that the minor C allele in rs11206510 PCSK9 SNP was associated with lower cholesterol levels in the blood; conversely, the TT genotype was associated with increased lipid levels and early coronary artery disease. 29 In the Chinese Han population, a study by Xu et al. in 2010 also demonstrated the significant association of the rs11206510 PCSK9 SNP and LDL; however, it was the minor C allele which appeared to be the risk allele in contrast to the T risk allele in white populations, attributed to likely ethnic genetic differences or a different true risk allele in linkage disequilibrium with the SNP. 24 In addition, the minor allele T of the rs1122608 LDLR SNP was shown to be associated with a favorable lipid profile and decreased risk for coronary artery disease according to a study by Martinelli et al. in 2010, confirming that G is the risk allele in the rs1122608 SNP. 21
While rs11206510 PCSK9 SNP and rs1122608 LDLR SNP have been associated with increased LDL levels and risk for myocardial infarction in other studies, this association was not evident in the FAW cohort in this study. It could be that the limited sample size of this study was insufficient to show a true significant association between the SNPs and LDL-C. Linkage disequilibrium is the nonrandom association of alleles, produced when two loci are closely linked together. In this scenario, the possibility that a true pathogenic SNP responsible for significant changes in LDL-C and CVD is only in linkage disequilibrium with either rs11206510 or rs1122608 SNPs could also be considered. Moreover, these findings could also be due to ethnic differences in this population, a presence of a different gene conferring a varied or modulatory effect, or difference in gene penetrance in the Filipino American population. It is possible as well that the multiple LDL-C related SNPs exert their effects collectively and that a significant association with LDL-C was not apparent in this study that only looked at two SNPs: rs11206510 PCSK9 and rs1122608 LDLR.
Effect of rs11206510 PCSK9 and rs1122608 LDLR SNPs on statin efficacy
Significant interindividual variation to statin response in patients with documented compliance brings to light the role of genetics and genomics. 31 Statins have a higher magnitude of LDL-C lowering effect in Asians compared to Europeans. A meta-analysis by Li et al. showed that LDL-C lowering by >40% is achieved by atorvastatin 80 mg or rosuvastatin 40 mg in Europeans in contrast to a similar effect size in Asians that is obtained with atorvastatin 18.9 mg or rosuvastatin 14.1 mg. In addition, for reduction of coronary atherosclerotic plaque, a longer duration of 24 months of rosuvastatin therapy is required in Europeans compared to 7.8 months of atorvastatin 18.9 mg or 10.7 months of rosuvastatin 14.1 mg in Asians. 32 The cytochrome P450 enzymes, of which CYP3A4, CYP2D6, and CYP3A5 account for majority of the metabolic reactions, comprise the most important enzyme system in phase I metabolism. Statins are highly metabolized through this route and are therefore prone to change in concentration based on enzyme activity. Poor metabolizers are predisposed to higher concentrations of statins and, subsequently, carry higher risk for adverse events such as myopathy. 33
In this study, statin therapy was shown to be associated with lower LDL-C irrespective of the rs11206510 PCSK9 and rs1122608 LDLR SNPs. Both rs11206510 PCSK9 and rs1122608 LDLR mutations were not associated with differential lipid lowering by statin medications. These results are similar to the findings of Vrablík et al. in 2012 that did not show a significant association between the PCSK9 polymorphism and statin-induced plasma lipid changes. 28
Most FAW have been shown to have persistently elevated LDL-C levels despite statin therapy in this study. While noncompliance or suboptimal therapy could not be ruled out due to the inherent limitations of this study, a true predisposition for elevated LDL-C due to genetic factors remains a possibility. PCSK9 inhibitors, alirocumab and evolocumab, have been shown to be effective in LDL-C reduction both as monotherapy and as additional medication to statin therapy.34–37 When added to statin therapy, the addition of PCSK9 inhibitors caused similar LDL-C reductions regardless of baseline statin type, dose, or intensity. 36 In this study, we found the predominance of the gain-of-function allele of rs11206510 PCSK9 SNP. Further investigation needs to be done to assess if PCSK9 inhibitors have an additional role in lipid lowering in this at-risk population apart from current standard indications.
Study limitations
The study's cross-sectional design limits causal inference, especially with regard to statin therapy. Given that this was a cross-sectional study, the pre-statin level of LDL-C was not available, limiting interpretation of LDL-C lowering due to statin therapy. Further research is needed to conduct a longitudinal study to examine the effects of both SNPs of interest and cardiovascular outcomes to include CVD events such as myocardial infarction, stroke, and CVD death. In addition, our study was conducted in a sub-population of FAW excluding males due to funding and therefore gender differences could not be determined. To our best knowledge, these are the first documented data on PCSK9 and LDLR genotype data in this population. Sample size was limited, and therefore, our results could not be generalized to the Filipino American population. Moreover, given unknown compliance and nonuniform statin type and dosage in this study, owing to the self-reporting nature of data gathering, it would be difficult to ascertain if the elevated LDL-C levels seen were in the context of an optimal or suboptimal statin therapy or compliance. Thus, the utility of PCSK9 inhibitors in this setting could not be addressed. Finally, the possibility of a true causative variant in linkage equilibrium with one or both of these two SNPs could not be ruled out since only two of the many LDL-C related SNPs were examined.
Conclusions
The gain-of-function T allele is the predominant allele of rs11206510 PCSK9 SNP and the gain-of-function G allele is the predominant allele of the rs1122608 LDLR SNP in our cohort of FAW. Statin-associated LDL-C reduction was observed in groups with rs11206510 PCSK9 and the rs1122608 LDLR mutations. In a population at high risk for hypercholesterolemia, hypertension, diabetes, and cardiovascular heart disease, optimal treatment with statins is effective in lowering LDL-C and should be considered where appropriate.
Footnotes
Acknowledgments
The authors are most grateful to all our FAW participants and research study volunteers without whom this study would not have been possible. The authors are sincerely grateful to Celera Corporation for providing their genetic testing services. This study was partially funded by the University of North Florida Dean Professorship Grant (No. 10-1025).
Ethical Approval
All applicable international, national, and/or institutional guidelines for the care and use of animals were followed. All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards.
Informed Consent
Informed consent was obtained from all individual participants included in the study.
Author Disclosure Statement
No competing financial interests exist.
