Abstract
The Food and Drug Administration (FDA), the Centers for Disease Control and Prevention (CDC), and state health and regulatory partners have investigated outbreaks of Salmonella enterica serovar Newport and Typhimurium infections in 2012, 2020, 2022, and 2023 linked to cantaloupes and/or watermelons from Southwest Indiana. In 2023, an outbreak of Salmonella Newport infections included 11 ill people and five hospitalizations reported in six states. The FDA and CDC collaborated with state partners to investigate the outbreak. The traceback investigation did not identify a single point of convergence for cantaloupe or watermelon. Products and environmental samples collected by the FDA and state partners from farms, public land, distributors, and points of service yielded isolates of multiple Salmonella serovars, including Salmonella Typhimurium and Salmonella Newport, that matched by whole genome sequencing to the 2020, 2022, and 2023 outbreak strains. Due to limitations with the traceback investigation and lack of convergence, a specific type of melon or source of contamination was not identified. However, the laboratory findings and historical outbreak information provided additional evidence to support a finding of ongoing contamination issues for cantaloupes and watermelons grown in Southwest Indiana. This is the fourth outbreak of salmonellosis confirmed to be linked to cantaloupes and/or watermelons from this area since 2012. These outbreaks of reoccurring and persisting strains of Salmonella illustrate the urgent need to determine the source, pathway, and extent of environmental contamination in the melon-growing region of Southwest Indiana and for outreach and education to help promote practices to reduce the contamination of melons.
Introduction
In 2012, the Food and Drug Administration (FDA), the Centers for Disease Control and Prevention (CDC), state, local health and regulatory partners investigated an outbreak of Salmonella enterica serovars Typhimurium and S. Newport infections linked to cantaloupe grown in Southwest Indiana, resulting in 261 total reported ill people and 94 hospitalizations in 24 states (U.S. Food and Drug Administration, 2013).
Several outbreaks linked to melons, which include cantaloupe and watermelon and are hereafter referred to as “melons,” grown in Southwest Indiana have occurred since 2012, posing investigative challenges for federal and state partners and food safety challenges to local growers. In 2020, there was another investigation of an outbreak of S. Newport infections linked to melons from Southwest Indiana, including 80 ill people and 18 hospitalizations reported in 15 states (Jenkins et al., 2023). In 2022, there was an investigation of an outbreak of S. Typhimurium infections linked to cantaloupe from Southwest Indiana, including 87 illnesses and 32 hospitalizations reported in 11 states (Schwensohn et al., 2024).
The response to these outbreaks included on-farm investigations, whose common limitation was that they were conducted after growing, harvesting, and postharvest activities had ceased, thus limiting the FDA’s and other regulators’ direct observations of these processes. Investigators have had difficulty in determining why S. Newport and Typhimurium outbreaks routinely occur in this region. During the recurring outbreaks and investigations, progress has been made in identifying underlying contributing factors, and steps have been taken by federal and state partners to minimize their impact (Jenkins et al., 2023; Seelman Federman et al., 2024; U.S. Food and Drug Administration, 2023a). Through whole genome sequence (WGS) analyses of isolates recovered from product and environmental samples, these investigations have identified a geographic clustering of related isolates, suggesting that these strains may be endemic to Southwest Indiana. CDC has designated S. Newport and S. Typhimurium as Reoccurring, Emerging, and Persisting (REP) strains related to this area (U.S. Centers for Disease Control and Prevention, 2023).
In 2023, a REP strain outbreak of S. Newport linked to melons grown in Southwest Indiana resulted in 11 reported ill persons and five hospitalizations reported in six states. Here, we report the details of this investigation conducted by the FDA, CDC, and state partners with a focus on the sample collection and analysis that linked multiple past and new outbreak strains to the same geographic area.
Materials and Methods
Outbreak detection and epidemiological investigation
WGS analysis of Salmonella spp. isolates from clinical samples was conducted by PulseNet, the national molecular subtyping network for foodborne disease surveillance, utilizing BioNumerics version 7.6. PulseNet uses a general cluster detection threshold of approximately 0–10 allele differences between clinical isolates, but other factors, including serovar, cluster size, and temporal or geographical relatedness, are considered when detecting clusters (Besser et al., 2019). An illness was defined as a laboratory-confirmed infection with the outbreak strain of S. Newport, with clinical isolates genetically related by core genome multilocus sequence typing (cgMLST), and illness onset beginning on or after June 26, 2023. Local and state public health partners interviewed ill people using routine enteric disease questionnaires and/or a melon-focused questionnaire to collect details for foods of most interest.
To collect food history and purchase information more rapidly, CDC actively queried the PulseNet database for this strain to identify illnesses and facilitate interviews of the ill people as soon as possible. Ill people are routinely interviewed using a questionnaire that focuses on melons and asks for detailed purchase information to inform traceback activities.
Traceback and farm investigations
A traceback investigation was initiated as per standard FDA traceback practices (Council to Improve Foodborne Outbreak Response, 2014; Irvin et al., 2021). Traceback investigations use information collected from ill people and trace commodities of interest through the supply chain, identify convergence in firms, and guide inspection locations.
The FDA conducted one packinghouse and two farm investigations, identified in traceback, per standard FDA practices (U.S. Food and Drug Administration, 2022). State partners accompanied FDA investigators during one of the two on-farm inspections.
Laboratory investigation
Clinical isolates
Clinical samples from ill people were cultured for Salmonella, serotyped, and subtyped by WGS at state public health laboratories using standard methods (Hassan et al., 2017; Hassan et al., 2019). WGS data was analyzed by PulseNet.
FDA samples
FDA laboratories used the FDA Bacteriological Analytical Manual method to isolate Salmonella spp. from melon product and environmental samples collected by investigators at various points in the supply chain, including points of service (POS) and distribution centers (Andrews et al., 2018). Environmental swabs and drainage ditch samples, consisting of 336 subsamples, a subset of the whole sample used in laboratory analysis, were collected using the grab and dead-end ultrafiltration (DEUF) methods (McClure et al., 2023; Mull and Hill, 2009; Mull and Hill, 2012). Serotyping and phylogenetic analyses of WGS data (Davis et al., 2015) were conducted to characterize the isolates and compare them to clinical and historical product sample isolates (Andrews et al., 2018; Crowe et al., 2017). WGS analyses were conducted for Salmonella spp. isolates recovered from product and environmental samples using high-quality single nucleotide polymorphisms (hqSNPs), and determination of whether isolates were a match (i.e., likely deriving from a recent common source) was guided by factors including the number of single nucleotide polymorphism (SNP) differences and tree topology (Pightling et al., 2018).
Results
Epidemiological investigation
A total of 11 illnesses were reported in six states. Of eight ill people with outcome information, five (63%) were hospitalized, and no deaths were reported (Fig. 1). S. Newport isolates were highly genetically related by cgMLST within 0–7 allele differences and genetically related to an S. Newport REP strain historically linked to consumption of domestically grown melons. Illness onset dates ranged from June 26 to August 31, 2023 (Fig. 2). Ill people ranged in age from 1–86 years (median 69), and seven of the 11 ill people (64%) were female. Overall, eight ill people were interviewed with a standardized questionnaire to assess melon consumption in the 7 days prior to illness onset, and five reported consuming any melon. Among these five ill people, one person consumed only cantaloupe, two consumed only watermelon, and two consumed both watermelon and cantaloupe.
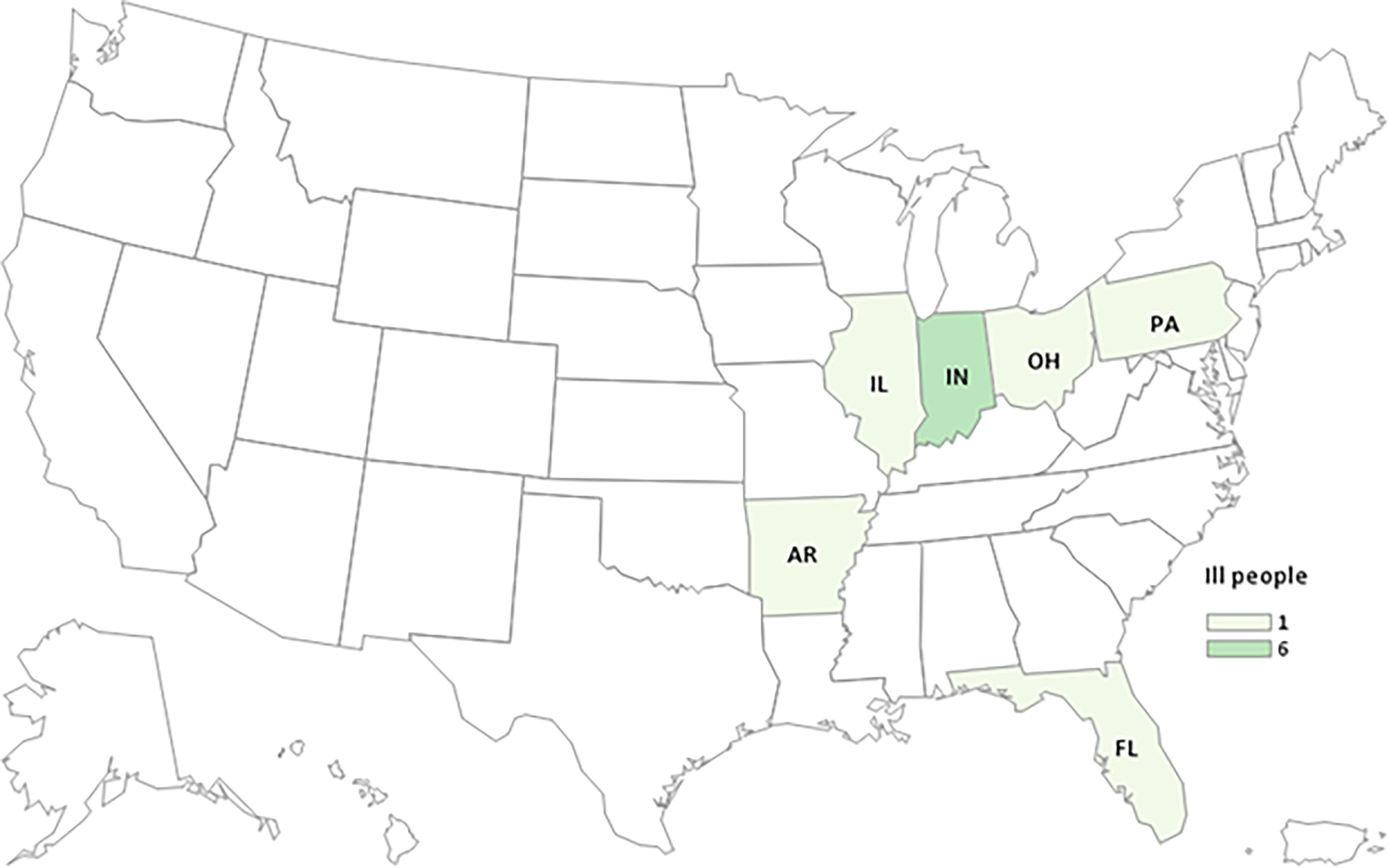
Distribution of reported illnesses in the Salmonella Newport melon-associated outbreak, by state, with illness onset dates from June 26, 2023 to August 31, 2023 (n = 11).
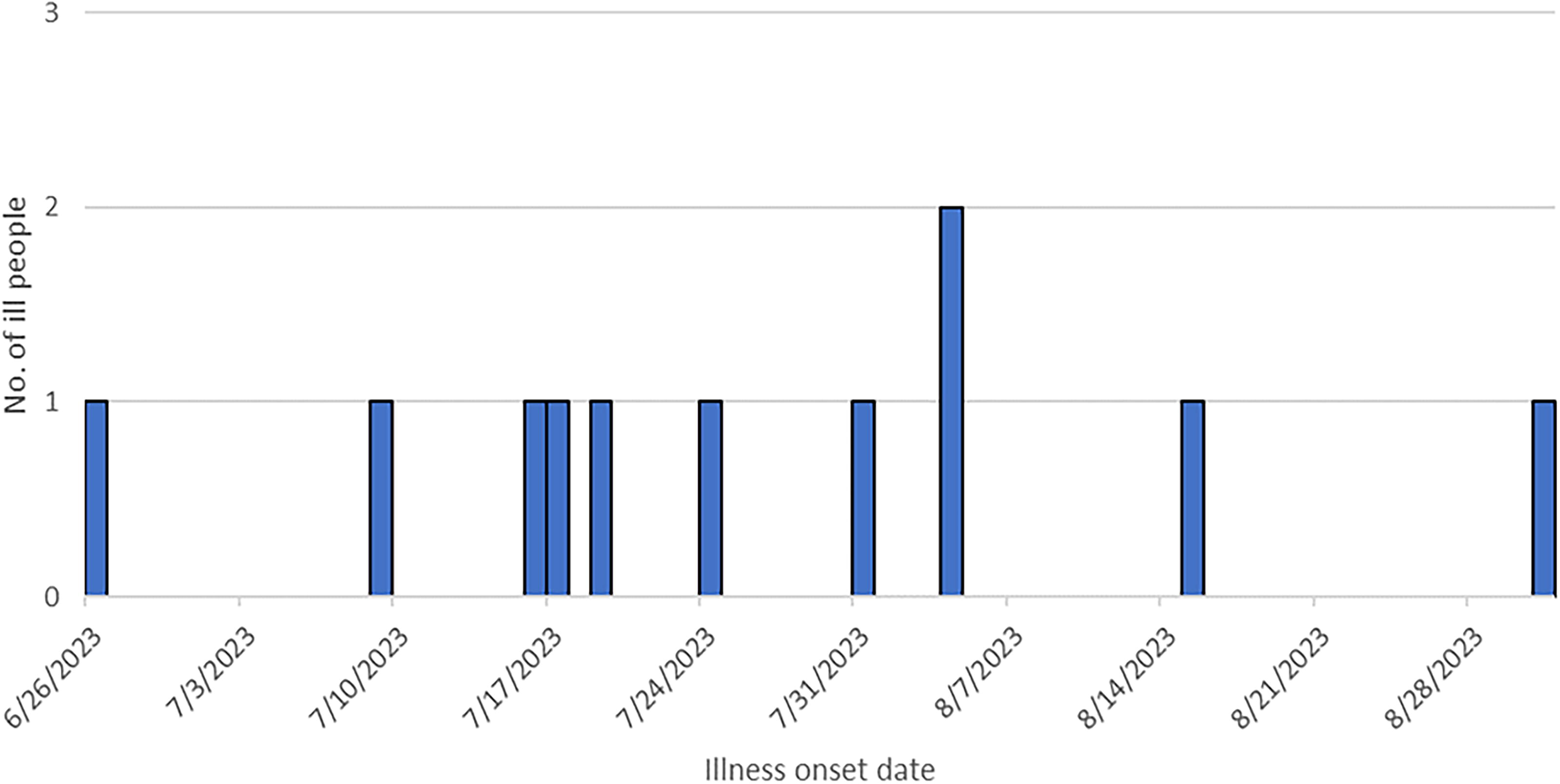
Number of reported illnesses in the Salmonella Newport melon-associated outbreak, by onset date from June 26, 2023 to August 31, 2023.
Traceback investigation and farm investigations
Based on available epidemiological information, FDA and state partners initiated traceback investigations for both cantaloupe and watermelon. Four ill people in Indiana reported purchasing melons from three POS. POS A and B were smaller, local POS, and POS C was a national retailer in the same state (Figs. 3 and 4). For the three POS locations, POS A and C were included in both the cantaloupe and watermelon traceback analyses, while POS B was only included in the cantaloupe traceback. Based on timeline information for when shipments were received at the POS during the traceback timeframe of interest, June 25 to July 17, 2023, cantaloupe supplied to POS A and B and watermelon supplied to POS A were most likely sourced from farms in Southwest Indiana. POS C sourced cantaloupe from Arizona and watermelon from Georgia and Florida.
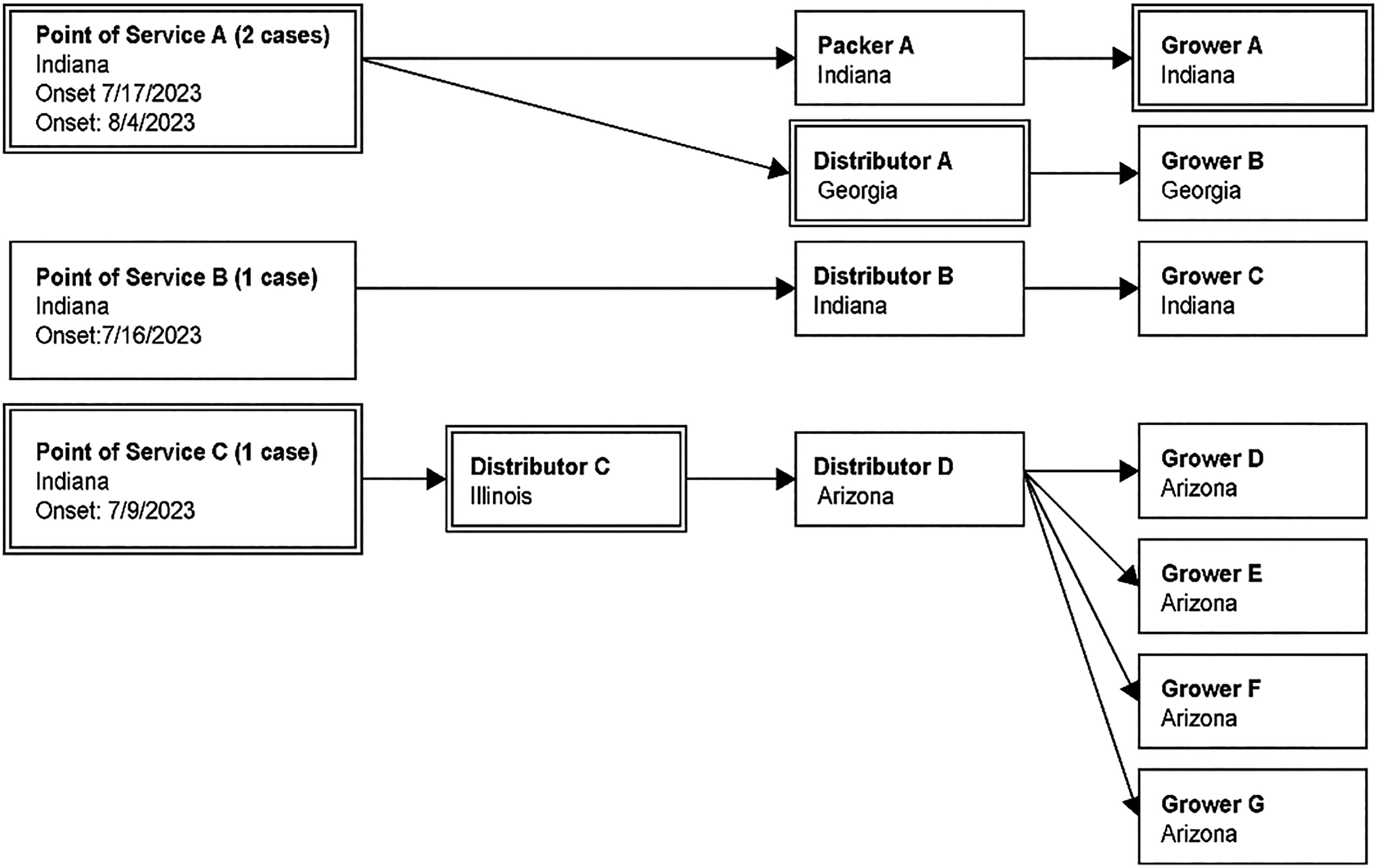
Traceback diagram for multistate outbreak of Salmonella Newport infections associated with consumption of cantaloupe in 2023. Purchases of products of interest are traced from the point of service, through the distribution chain, to distributors and importers. Product originates from the growers or fields, denoted on the right side of the diagram. Firms with double lined boxes indicate that those firms are present in both cantaloupe and watermelon traceback investigations.
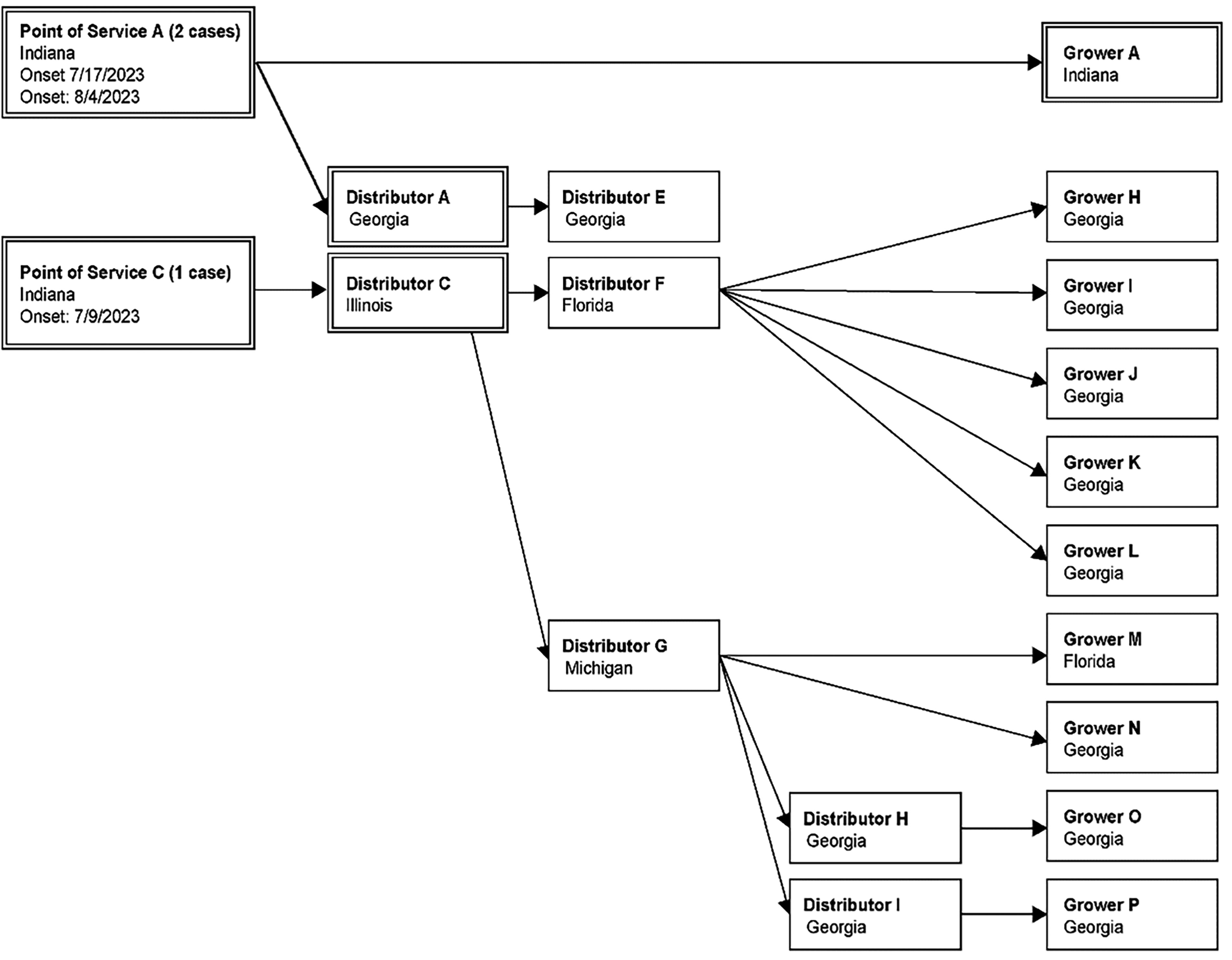
Traceback diagram for multistate outbreak of Salmonella Newport infections associated with consumption of watermelon in 2023. Purchases of products of interest are traced from the point of service, through the distribution chain, to distributors and importers. Product originates from the grower, denoted on the right side of the diagram. Firms with double lined boxes indicate that those firms are present in both cantaloupe and watermelon traceback investigations.
Based on traceback information and due to this outbreak strain being a CDC REP strain associated with previous outbreaks linked to melons, farm and packinghouse inspections were initiated by the FDA at Grower A, Grower C, and Packer A on September 11, 2023, September 7, 2023, and September 12, 2023, respectively. FDA initiated the farm inspection at Grower C with Indiana state partners. At the time of inspection, the farms had harvested their crops, and only watermelons too damaged to sell were still present at Grower C. Grower A was processing watermelons for shipment at the time of the inspection, and product samples were taken. No significant observations of objectionable conditions or practices were noted during the three inspections, both growers reported that they did not use biological soil amendments of animal origin (BSAAO), and both growers denied sharing equipment with other firms.
Laboratory investigation
The samples analyzed as part of this outbreak investigation are outlined in Table 1. With assistance from state partners, FDA collected a total of 21 product and environmental samples (one state and 20 FDA). The FDA collected five environmental samples, consisting of 90 subsamples from Grower C, one environmental sample, consisting of 83 subsamples from Packer A; one watermelon product, consisting of 10 subsamples from Grower A; seven environmental samples, consisting of 117 subsamples from Grower A’ and two environmental samples (soil and DEUF), consisting of six subsamples from public land near Grower A and Grower C. Two cantaloupe samples containing 10 subsamples each were collected from Distributor C, and another distributor of the same firm, supplied by Distributor D. State partners collected one cantaloupe product sample from POS A.
Food and Drug Administration and Indiana Department of Health Samples Collected during the Investigation of Salmonella Newport and Typhimurium Infections in 2023 and 2022 Linked to Melons, Including Sample Numbers and Number of Positive Subsamples, Collection Agency, Sampling Locations, Sample Descriptions, and Salmonella Serovar Identified
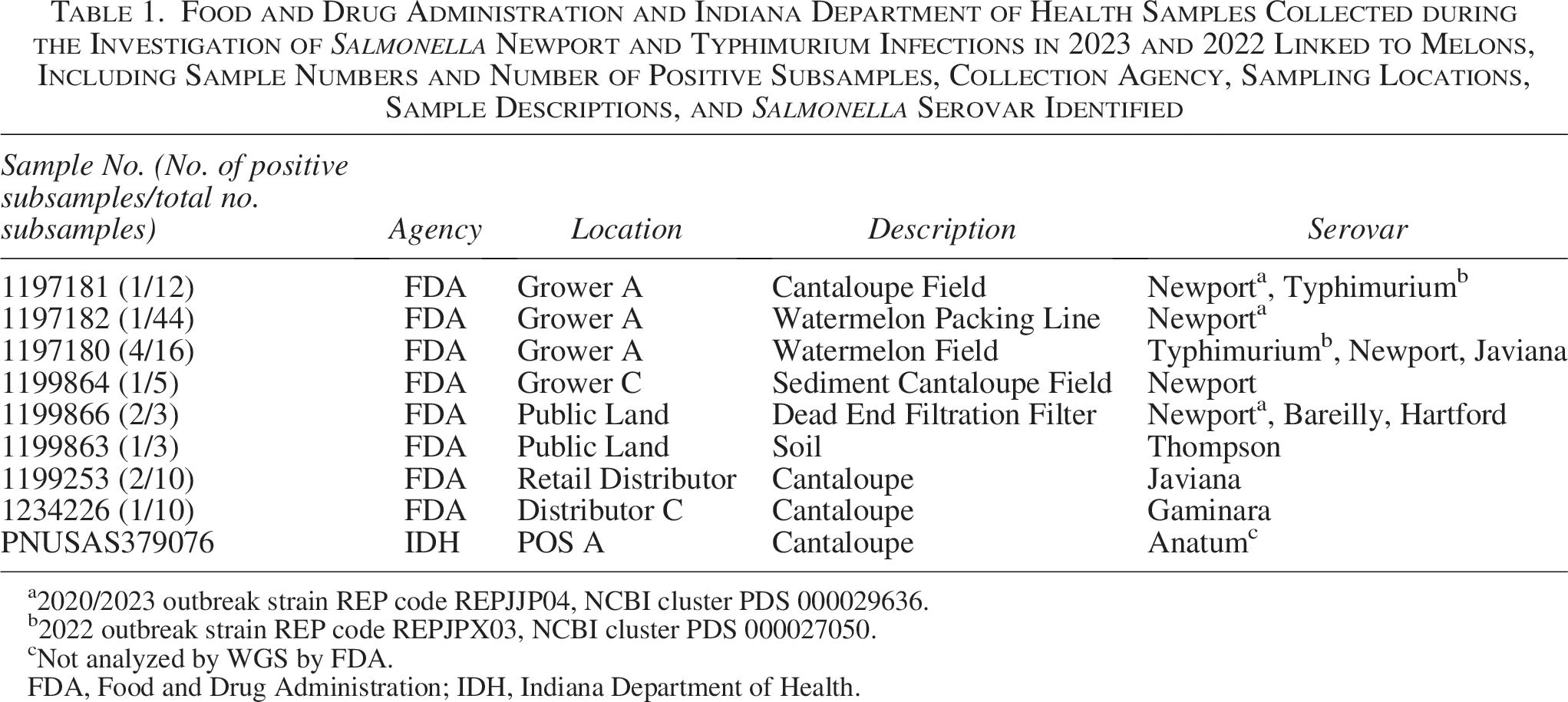
2020/2023 outbreak strain REP code REPJJP04, NCBI cluster PDS 000029636.
2022 outbreak strain REP code REPJPX03, NCBI cluster PDS 000027050.
Not analyzed by WGS by FDA.
FDA, Food and Drug Administration; IDH, Indiana Department of Health.
Nine of the 21 samples were positive for Salmonella spp., yielding over 100 isolates (Table 1). WGS analysis indicated these isolates clustered into six strains, including two clusters that match by WGS to isolates from large, multistate outbreaks (Fig. 5). Samples collected during the investigation yielded isolates that matched the 2012/2020/2023 S. Newport outbreak strain (National Center for Biotechnology Information [NCBI] cluster PDS000029636; Fig. 5A), including a DEUF sample from public land (#1199866), a drag swab from a cantaloupe field at Grower A (#1197181), and an environmental swab collected from the floor beneath the watermelon packing line from Grower A (#1197182). In addition to over 150 clinical isolates, 11 of which were from the 2023 S. Newport outbreak, this NCBI cluster includes environmental sample isolates collected by the FDA during a 2022 investigation of an S. Typhimurium outbreak associated with cantaloupe in this region, as well as isolates from a Missouri poultry facility (Seelman Federman et al., 2024).
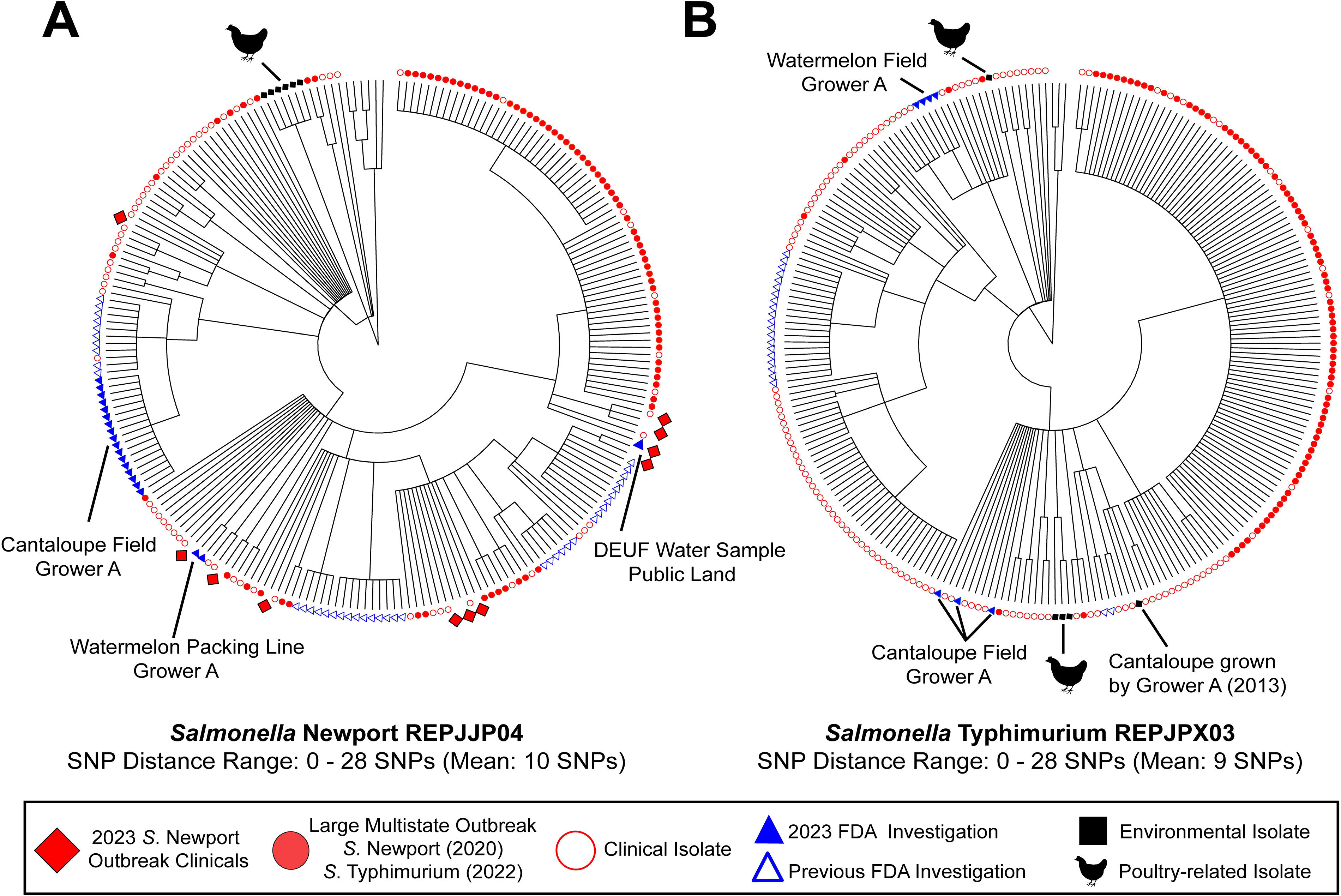
Maximum likelihood trees for
Samples collected during the investigation also matched an S. Typhimurium REP strain (NCBI cluster PDS000027050; Fig. 5B), including isolates from the Grower A cantaloupe field drag swabs (#1197181) and isolates from the Grower A watermelon field drag swabs (#1197180). These isolates matched over 200 clinical isolates, isolates from Grower A collected during the 2022 S. Typhimurium outbreak investigation, isolates collected during the 2020 S. Newport outbreak investigation, cantaloupe collected from Grower A in 2013, and four poultry samples collected by the FDA and USDA from Indiana in 2016 and 2022 (Jenkins et al., 2023; Seelman Federman et al., 2024).
Isolates from two samples matched another cluster of S. Newport isolates (PDS000004406), including one DEUF sample (#1199866) collected on public land near the growing area of interest and a soil sample (#1199864) collected from the cantaloupe field at Grower C. These isolates matched 34 clinical isolates; six were from one 2012 outbreak, and four were from another 2012 outbreak. These isolates also matched two cantaloupe isolates associated with one of the Indiana farms from the 2012 S. Newport and Typhimurium outbreak investigations, nine environmental isolates collected by the FDA in Indiana, and an additional MO cantaloupe isolate collected by the FDA.
Ten isolates from drag swabs collected from the watermelon field at Grower A (#1197180) recovered S. Javiana. These samples matched two isolates collected during the 2020 S. Newport outbreak investigation (Jenkins et al., 2023) and one clinical isolate (PDS000028584).
Isolates from a drag swab of the watermelon field at Grower A (#1197180) and a DEUF sample from public land (#1199866) matched a third strain of S. Newport (PDS000011406). These isolates matched nine clinical isolates, two cantaloupe isolates associated with Indiana, and three isolates from a sample taken as part of the 2020 S. Newport investigation into another melon grower in Southwest Indiana (Jenkins et al., 2023). Three isolates from a soil sample collected from nearby public land yielded S. Thompson, and these isolates matched two 2020 clinical isolates (PDS000032705).
Finally, one cantaloupe sample collected by state partners from POS A (PNUSAS379076) recovered S. Anatum (PDS000060321.21). Two cantaloupe samples collected from Distributor C (#1234226) and another retail distributor of the same retail chain (#1199253) recovered S. Gaminara (PDS000010687.8) matching one clinical isolate and S. Javiana, respectively. A distributor in Arizona, Distributor D, was identified as the supplier of these cantaloupes.
Investigational outcomes
On August 30, 2023, the outbreak was initially listed on the Coordinated Outbreak Response and Evaluation Network (CORE) Investigation Table (CIT) (U.S. Food and Drug Administration, 2020). On September 27, 2023, Distributor D issued a voluntary recall due to the recovery of Salmonella spp. from cantaloupe grown in Arizona, collected by the FDA at Distributor C and another retail distributor of the same firm (Table 1), although these did not match the outbreak strain. The outbreak strain was recovered from environmental samples collected during inspections at two Indiana growers on Grower A’s cantaloupe field, a non-food contact surface in their watermelon packing line, and on public land near both Grower A and Grower C; however, public communications were not issued because the product was no longer on the market.
Discussion
Impact and significance
This is the fourth outbreak of salmonellosis investigated by the FDA related to melons and linked to Southwest Indiana since 2012 (Jenkins et al., 2023; U.S. Food and Drug Administration, 2013). Despite the small number of illnesses in this outbreak, epidemiological information indicated melons were the vehicle of interest. Although the melon traceback investigation did not show a convergence on a specific grower, it did indicate geographic convergence with two of the three traceback legs to Southwest Indiana. Since the outbreak strain of S. Newport has historically been linked to the same growing region, the geographic convergence prompted inspections at the growers identified in the melon traceback investigation. The recovery of the outbreak strain from a grower identified in the traceback underscores the value of on-farm inspections and sampling of the growing environment for Salmonella.
The laboratory investigation of samples collected in Indiana yielded seven strains of Salmonella spp. from environmental and product samples in addition to the outbreak strain: three strains of Newport and one strain each of Typhimurium, Javiana, Anatum, and Thompson. All strains matched clinical isolates, indicating they had the potential to cause illness in humans, and two REP strains that have been identified as causing reoccurring outbreaks. An in-depth discussion of these previous outbreaks and WGS findings appears in Seelman et al. (Seelman Federman et al., 2024). It is notable that all strains, except the S. Thompson, have been recovered from the growing environment during previous FDA investigations at melon growers in Southwest Indiana prompted by the outbreaks in 2012, 2020, and 2022. The recovery of outbreak REP strains over several years suggests there exists a persistent Salmonella reservoir in the environment. This underscores the need for further research into transmission dynamics in this growing region to inform the development of effective risk reduction strategies. The ultimate source of these strains of Salmonella spp. remains unclear, though the presence of isolates from poultry facilities in the region in both REP strains indicates the need for a One Health approach to determine their association (U.S. Food and Drug Administration, 2024).
Investigational challenges and future opportunities
The challenges that investigators encountered during the traceback investigation were similar to those from past produce-associated (Irvin et al., 2021; Whitney et al., 2021) and melon-associated outbreaks (Jenkins et al., 2023). Records obtained from Indiana growers or farm stand retailers of interest yielded limited information for traceback analysis, and invoices were often handwritten carbon copies that included shorthand references to describe products supplied (e.g., “lopes” for cantaloupes). Rarely were any quantities or unique identifiers included in the obtained documentation. These traceability challenges could be alleviated by the Food Traceability Final Rule, which establishes traceability recordkeeping requirements for certain foods and provides a framework for documenting Critical Tracking Events (such as harvesting and initial packing) and Key Data Elements (such as harvest date and location and primary recipient[s]) that would preserve important traceability information across the supply chain and allow for expedient and targeted traceback and rapid removal of impacted product. It should be noted that while fresh melons are included on the Food Traceability List, which means certain producers will need to comply with the applicable requirements of the Food Traceability Rule by the compliance deadline, full or partial exemptions may also apply (U.S. Food and Drug Administration, 2021; U.S. Food and Drug Administration, 2023b). Therefore, traceability challenges as described during this investigation may persist for supply chains even if a food is included on the Food Traceability List.
Another challenge posed to investigators was the limited number of traceback legs used by the FDA to identify common distributors or growers of melons likely consumed by ill people. Investigators conducted melon traceback at three POS, where four ill people reported purchasing melons. Overall, eight of the 11 ill people agreed to be interviewed with the standardized questionnaire to assess melon consumption, and only four ill people provided enough information to initiate melon traceback. As a result, the four best traceback candidates were all from Indiana, which made a more geographically homogenous group. Supply chain convergence was not observed; the small number of cases and uncertainties with exposure information, such as the lack of confirmed purchase or consumption dates and cases reporting exposure to both cantaloupe and watermelon, further limited conclusions that could be drawn from the traceback.
A common limitation for all on-farm investigations since 2020 was that they were conducted after growing, harvesting, and postharvest activities had ceased, thus limiting direct observations of these processes. All four outbreaks in 2012, 2020, 2022, and 2023 were linked to melons from Southwest Indiana, but they were traced to melons grown on different farms, further supporting the regionality of the issue and that contamination is not specific to a single grower.
Outbreaks caused by melons are most likely the result of contamination during growing, packing, harvesting, or processing (Walsh et al., 2014). Potential prevention measures may include contamination detection through sampling melons early in the growing season, coordinating with local growers and agricultural cooperative extension programs on best practices to prevent environmental contamination (i.e., contaminated soil or BSAAO) on fields within their control and adjacent fields, and research aimed at developing methods to prevent contamination during melon harvesting and packing.
Conclusions
Epidemiological, laboratory, and traceback evidence confirm that whole cantaloupe and watermelon from Indiana were the vehicles of S. Newport infections among the 11 ill people that were part of this outbreak. The traceback investigation did not identify a single point of convergence for either cantaloupe or watermelon. Product and environmental samples collected by the FDA and state partners yielded isolates of multiple Salmonella serovars, including S. Typhimurium and S. Newport, that matched present and previous outbreak strains. Due to limitations with the traceback investigation and lack of convergence, specific melon products or sources of contamination were not identified. However, the laboratory findings and historical outbreak information provided additional evidence to suggest ongoing contamination issues for melons grown in Southwest Indiana. This outbreak investigation supports the need for additional research to determine the source and extent of environmental contamination in the melon growing region of Southwest Indiana.
Authors’ Contributions
N.C.: Investigation, conceptualization, writing—original draft preparation; C.S.: Investigation, writing, reviewing and editing; M.K.: Conceptualization, writing, reviewing and editing; E.J.: Reviewing and editing; T.J.: Investigation, reviewing and editing; M.D.: Writing, reviewing and editing; C.H.: Writing, reviewing and editing; D.B.: Writing, reviewing and editing; C.W.: Data analysis, reviewing and editing; K.N.: Investigation, reviewing and editing; A.B.: Investigation, reviewing and editing; T.J.: Investigation, reviewing and editing; B.L.: Investigation, reviewing and editing; R.B.: Investigation, reviewing and editing; T.H.: Writing, reviewing and editing; V.M.: Investigation, reviewing and editing; L.G.: Investigation, reviewing and editing; M.S.: Investigation, reviewing and editing; S.V.: Conceptualization, writing—original draft preparation, reviewing and editing.
Footnotes
Acknowledgments
The assistance of state partners, especially the Indiana state partners, was crucial in the collection and analysis of product samples, traceback documents, and epidemiological information. Special thanks to FDA’s emergency response coordinators as well as other FDA staff instrumental in the outbreak coordination for their tireless efforts and assistance.
Funding Information
No funding was received for this article.
Disclaimer
The findings and conclusions of this report are those of the authors and do not necessarily represent the official position of the Food and Drug Administration (FDA) and the Centers for Disease Control and Prevention (CDC).
Disclosure Statement
No competing financial interests exist.
