Abstract
Staphylococcus aureus is one of the major causes of foodborne diseases and its presence in food products may poses a public health challenge. The aims of this study were to assess in vitro the capacity of S. aureus isolates from foods to form biofilm and to determine their antibiotic susceptibility. A total of 80 S. aureus isolates were characterized. The slime production ability was evaluated by congo-red agar (CRA) and the biofilm formation was carried out by microtiter-plate method (MPM). Resistance of isolates to eight antibiotics was determined using disc diffusion method. Sixty-four (80%) of the isolates were slime producers on congo-red agar. However, all isolates were biofilm producers on microtiter-plate method. The highest resistance profiles were ascribed to penicillin G (91.25%) and tetracycline (41.25%). Twelve isolates were methicillin-resistant (MRSA) harboring the mecA gene. All of these MRSA isolates were negative for the genes of the Panton Valentine leukocidine (lukF/S-PV). Typing of the MRSA isolates indicated that they belonged to three spa-types including t024, t450 and t688. The presence of biofilm producers and multidrug resistant isolates (MRSA) in food samples can represent a risk for public health. Therefore, an efficient control and effective measures were needed along the production chain to ensure the food safety.
Introduction
Staphylococcus aureus is a Gram-positive commensal opportunistic and life-threating pathogen bacterium (Parastan et al., 2020). It is the causative agent of variety of diseases ranging in severity from slight skin infections to more severe diseases, such as pneumonia, endocarditis, osteomyelitis, septicemia or toxic shock syndrome, among others (Lowy, 1998). Conversely, S. aureus is the causative agent of staphylococcal food poisoning (SFP), which is a common foodborne disease, resulting by the ingestion of staphylococcal enterotoxins (SEs) produced by enterotoxigenic strains of S. aureus (Strommenger et al., 2018).
In recent years, antimicrobial resistance has become a major issue at world scale, not only in human medicine but also in the animal sector (Madec and Haenni, 2018). For example, methicillin-resistant Staphylococcus aureus (MRSA) emerges when S. aureus acquires a specific genetic element called the staphylococcal chromosomal cassette (SCCmec) (Pantosti, 2012). This mobile genetic element contains the mecA gene encoding an alternative penicillin-binding protein, PBP2a (or PBP2’), which exhibits a low affinity for most β-lactam antibiotics (Hartman and Tomasz, 1984; Lakhundi and Zhang, 2018).
Currently, MRSA has spread globally and poses a significant threat for public health due to its capacity to develop new forms of antibiotic resistance, thereby reducing available treatments options (Doulgeraki et al., 2017). MRSA initially emerged in healthcare settings, known as healthcare-associated MRSA (HA-MRSA). However, in recent decades, MRSA has also been detected outside hospital environments, referred to as community-acquired MRSA (CA-MRSA) (da Silva et al., 2020). Additionally, MRSA infections in the community can also be caused by livestock-associated MRSA (LA-MRSA) (Köck et al., 2013; Spoor et al., 2013), which differs from HA-MRSA and CA-MRSA in its genomic traits (Cuny et al., 2015). Transmission of MRSA to humans can occur through direct contact with infected animals and their environment or by consumption of foods of animal origin (Oniciuc et al., 2017). However, there is little information about the potential transmission of MRSA to humans through the food chain (Petinaki and Spiliopoulou, 2012). Therefore, the European Food Safety Authority (EFSA) proposed to reinforce the monitoring of MRSA both in foods and in food-producing animals (Aerts et al., 2019).
The food production facilities provide ideal conditions for many foodborne pathogens, allow them to adhere and to form biofilms on most materials and under almost all the environmental conditions (Bridier et al., 2015). S. aureus has been found to create biofilm on a wide range of biotic and abiotic surfaces. These include human body tissues, medical implants such as catheters, artificial heart valves, bone and joint prostheses, as well as food items and equipment used in food processing plants (Vázquez-Sánchez and Rodríguez-López, 2018). As known, microorganisms that form biofilms gain significant survival and propagation benefits compared to their free-floating planktonic cells (McCarthy et al., 2015). This sessile lifestyle offers protection against various threats, including cleaning processes, host immune responses, and antimicrobial treatments. As a result, biofilms can cause long-lasting contamination or infection diseases, presenting a challenge in both industrial and medical settings (Thiran et al., 2018). In the present study, we aim to investigate in vitro the biofilm formation ability of S. aureus isolates of food origin and to study their antibiotic susceptibility.
Materials and Methods
Collection of bacterial isolates
This study was conducted on 80 S. aureus isolates recovered from various food products including raw milk, traditional milk products (rayeb, l’ben and butter), pastries and minced meat of beef collected between 2015 and 2019 from several randomly selected market points (butchers, creameries, cafeteria) located at Tizi Ouzou area (Algeria). The traditional milk products were produced using raw milk in factories located mostly in urban areas, and then distributed to market points. The collection of strains included 62 isolates from milk and traditional milk products, 15 isolates from pastries and 3 isolates from minced meat of beef. To isolates S. aureus, 10 mL (raw milk) or 10 g (traditional milk products, minced meat, pastries) of each sample was added to 90 mL of buffered peptone water (Conda Pronadisa, Madrid, Spain) and homogenized. Isolation was done by spreading 0.1 mL of each resulting suspension on mannitol salt agar (Conda Pronadisa, Madrid, Spain) or Baird-Parker base (Conda Pronadisa, Madrid, Spain) supplemented with 5% egg yolk and tellurite (Conda Pronadisa, Madrid, Spain), respectively. The obtained isolates were identified as S. aureus using Gram staining and conventional tests (catalase test, coagualase test and DNase test). Phenotypically identified S. aureus isolates were submitted to PCR simplex to detect the presence of 23S rRNA gene region specific for S. aureus according to Straub et al. (1999). After identification, all isolates were stored in brain heart infusion broth (BHIB) (Biokar, France) with glycerol (30%, V/V) at −20°C for further analysis.
Detection of biofilm formation in vitro
Congo red agar method (CRA)
The ability of S. aureus isolates to produce slime was assessed using congo red agar (CRA), as described by Freeman et al. (1989). Briefly, one colony from each S. aureus culture was streaked on CRA plates and incubated aerobically at 37°C for 24 to 48 h. After incubation, slime positive strains appeared as black colonies with dry crystalline consistency and slime negative strains remained red.
Microtiter plate assay (MPA)
The biofilm-forming ability of S. aureus isolates was assessed as described by Stepanović et al. (2007), with some modifications. Firstly, S. aureus isolates from the frozen stocks (−20°C) were grown overnight at 37°C in 5 mL of BHIB (Biokar, France) and then plated on brain heart agar (BHA) (Conda Pronadisa, Spain). After verifying purity of the cultures, two colonies from each isolate were inoculated into 5 mL of trypticase Soy Broth (TSB) (Conda Pronadisa, Spain) supplemented with 1% of Glucose (Sigma-Aldrich, Isère, France) and incubated overnight at 37°C without shaking. Overnight cultures were diluted 1:50 with TSB-1% glucose and 200 µL of cell suspensions were transferred to individual wells of a flat bottom 96-well polystyrene microtiter plate (ProLab Scientific Co Ltd, Zhejiang, China). The reference strain S. aureus ATCC 25923 was used as positive control and the medium TSB-1% glucose served as a negative control. After overnight incubation at 37°C, the non-adhered cells were removed from the wells by gently overturning the plate onto paper towels, and the wells were gently washed with phosphate-buffered saline (PBS) three times and air-dried. Adherent bacteria were fixed with methanol (Honeywell, Seelze, Germany) for 15 mn and stained with 150 µL of aqueous solution of crystal violet 0.5% (Biochem Chemopharma, Nièvre, France) for 15 min. After washing, the dye bound to the cells was eluted with 95% ethanol, and the biofilm formation was measured at 560 nm using microtiter-plate reader (Gentaur, Paris, France). The biofilm formation was expressed as optical density (OD) values. Each S. aureus isolate was studied in triplicate in a single experiment. The average OD value of all tested strains (ODs) and negative controls was calculated. Cut-off OD (ODc) is defined as three standard deviations above the mean OD of the negative control. The isolates were classified into four following categories based on the optical density: non-biofilm producer (OD test < ODc), weak biofilm producer (ODc < OD < 2X ODc), moderate biofilm producer (2X ODc < OD < 4X ODc), and strong biofilm producer (4X ODc < OD).
Antimicrobial susceptibility of S. aureus isolates
All S. aureus isolates were tested for their antimicrobial susceptibility using the disc diffusion method on Mueller-Hinton agar (Conda Pronadisa, Spain) according to the guidelines of the Clinical and Laboratory Standards [Clinical and Laboratory Standard Institute (CLSI), 2020]. Eight antimicrobial agents were tested (µg/disk): penicillin G (10 UI), cefoxitin (30), gentamicin (10), tetracycline (30), erythromycin (15), ofloxacin (15), chloramphenicol (30) and trimethoprim/sulfamethoxazole (1.25/23.75). The strain S. aureus ATCC25923 was used as control strain in susceptibility testing. After incubation at 37°C for 24 h, the isolates were classified as susceptible, intermediate or resistant according to breakpoints of the clinical and laboratory standards [Clinical and Laboratory Standard Institute (CLSI), 2020].
Detection of the mecA, mecC, spa and lukS/F-PV genes
The genes encoding methicillin resistance (mecA and mecC), Panton-Valentine leucocidin (lukS/F-PV), and the staphylococcal protein A (spa) were detected using a multiplex PCR as described by Stegger et al. (2012), with modifications. DNA ladder 1 kb (Promega) was used as molecular weight standard. Three reference strains were used as positive controls: S. aureus LGA251 (carrying mecC gene), S. aureus ATCC 25923 (carrying pvl gene) and S. aureus MU50 (carrying mecA gene).
Molecular characterization of MRSA isolates by spa-typing
The polymorphic X-region of the protein A gene (spa) was amplified by PCR using primers spa-1113f (5′-TAAAGACGATCCTTCGGTGAGC-3′) and spa-1514r (5′-CAGCAGTAGTGCCGTTTGCTT-3′) (Aires de Sousa et al., 2006). The PCR products were then separated by electrophoresis on a 2% agarose gel and visualized using a Bio-Rad Gel Doc EQ system. We obtained DNA sequences for both strands through Eurofins sequencing services (Esberg, Germany). Finally, spa-types were identified by analyzing these sequences with BioNumerics 7.1 software (Applied Maths, SintMartens-Latem, Belgium).
Results
Biofilm formation ability of tested isolates was phenotypically assessed by the CRA method, where the isolates appeared as typical black colonies (slime producing) on CRA plates. In total, 64 (80%) of the isolated S. aureus were found to be slime-producing (Table 1). All S. aureus isolated from minced meat were slime producing, whereas 50%, 75%, 66.7%, 77.8%, and 93.3% of isolates from butter, l’ben, rayeb, raw milk and pastries showed slime production, respectively (Table 1). Further, the isolates were subject to confirm their ability to form biofilm in MPA, and it was observed that all isolates showed the ability to produce biofilms, including 44 (55%) isolates that showed strong biofilm formation, 32 (40%) isolates that showed moderate biofilm formation and 4 (5%) isolates that showed weak biofilm formation (Table 2).
Slime Production in S. aureus Isolates (n = 80)
Biofilm Formation among S. aureus Isolates in Microtiter Plates (n = 80)
Antibiotic susceptibility testing showed that the majority of food related S. aureus isolates were resistant to penicillin G (n = 73; 91.25%), followed by tetracycline (n = 33; 41.25%). However, low resistances were observed against cefoxitin (n = 12; 15%), ofloxacin (n = 9; 11.25%), erythromycin (n = 4; 5%) and chloramphenicol (n = 2; 2.5%). All isolates were susceptible to sulfamethoxazole/trimethoprim and gentamicin (Fig. 1). Twelve (15%) isolates were multidrug resistant and three phenotypes of multidrug resistances were observed (Table 3). Twelve (15%) MRSA isolates we identified (resistant to cefoxitin), and harbored the mecA gene. Six isolates were recovered from raw milk and six from acidified milk (rayeb). None of them carried the Panton Valentine leukocidine (lukF/S-PV) genes (Table 4). Typing of the MRSA isolates indicated that they belonged to three spa-types including t024, t450 and t688. Nine isolates from raw milk and l’ben belonged to spa-type t024, two isolates from rayeb to t688 and only one isolate to t450 (Table 4).
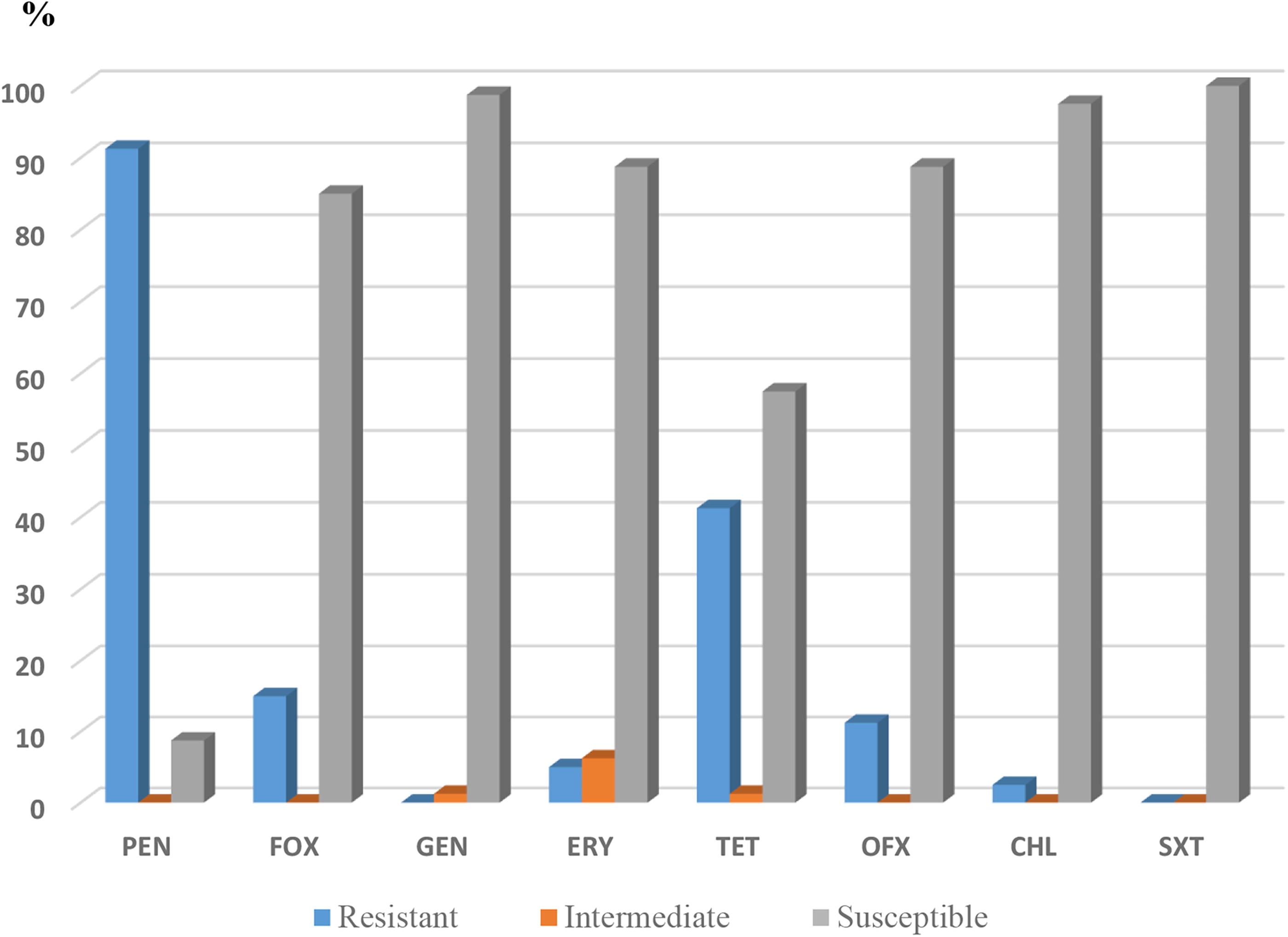
Antibiotic susceptibility of S. aureus isolates from food products (n = 80). PEN, penicillin G; FOX, cefoxitin; GEN, gentamicin; ERY, erythromycin; TET, tetracycline; OFX, ofloxacin; CHL, chloramphenicol; SXT, trimethoprim/sulfamethoxazole.
Phenotypic Resistance Patterns among Multidrug-Resistance S. aureus Isolates
Phenotypic and Genotypic Characterization of MRSA Isolates
Discussion
In food processing facilities, microorganisms frequently colonize and form biofilms on both food-contact surfaces and non-food-contact equipment (Alvarez-Ordóñez et al., 2019). As known, bacteria’s capacity to form biofilms is a crucial factor in their pathogenicity. S. aureus, like many bacteria, has the ability to form biofilms as part of its normal life cycle. When in a biofilm state, S. aureus gains significant advantages over its planktonic mode of growth (Chen et al., 2020). In this study, we have used two methods to evaluate the capacity of isolated strains to produce biofilm in vitro. Among all tested isolates, 64 (80%) were biofilm producers by CRA-plate method. Our results agree with those of Ballah et al (2022) and Achek et al (2020), who reported a rate of 89% and 100%, respectively. Souza et al. (2024) have also reported a high incidence of slime-producers isolates of S. aureus from artisanal unripened cheeses, with a value of 70%. The Congo Red Agar (CRA) method was used for its reasonable sensitivity and specificity in detecting biofilm formation, despite not being regarded as the most sensitive method available (Ballah et al., 2022; Jain and Agarwal, 2009). As reported by Lee et al. (2016), multiple factors, such as glucose and chloride, affect biofilm formation of Staphylococcus spp. CRA directly interact with certain polysaccharides, forming colored pigments, and some metabolic reactions that form secondary products with the dye can influence the formation of dark colonies. Consequently, the CRA test could be used as the presumptive test for the biofilm formation (Avila-Novoa et al., 2018). Further, we have used a MPA method, and it was observed that all isolates have the ability to form biofilm in polystyrene plates. Our results were in accordance with those of previous studies (Achek et al., 2020; Ballah et al., 2022; Bissong and Ateba, 2020). The discrepancies observed between biofilm formation in polystyrene surface and the production of slime on CRA can be attributed to the fact that the biofilm formation phenotype is highly sensitive to in vitro conditions, and some staphylococcal isolates have the capacity to form biofilms but lack the ability to produce slime (Souza et al., 2024). In food industries, biofilm is an important source of microbial contamination that can lead to spoilage and transmission of foodborne pathogens (Filipello et al., 2019). In most cases, biofilms represent a challenge of great concern, since it could be origin of food cross-contamination, thereby diminishing the competency of food processing procedures and eventually compromising the quality of food and its safety standard (Olanbiwoninu and Popoola, 2023). Moreover, the high density of microorganisms within biofilms create an environment that facilitates the exchange of genetic material, including genes encoding for antibiotic resistance (Madsen et al., 2012).
Antibiotic resistance has been widely recognized as one of the main global health concerns, threatening food security, and human and animal health, causing considerable economic losses (Caniça et al., 2019). In this study, the resistance rates of S. aureus isolates were higher for penicillin G and tetracycline, which is similar to previous findings (Chaalal et al., 2018; Ma et al., 2018; Mekhloufi et al., 2021). The global overuse and misuse of antibiotics in both human health care and animal farms has contribute to the rise of antibiotic-resistant bacteria, posing a significant threat to human and animal health worldwide (Vinayamohan et al., 2022). All isolates were susceptible to gentamicin and sulfamethoxazole/trimethoprim, which is not in agreement with the data reported in other studies (Ma et al., 2018; Wang et al., 2017). A slight resistance was observed to chloramphenicol, which suggests its use in veterinary medicine. The same results were obtained in previous study conducted by Patel et al. (2021). A high susceptibility was observed for erythromycin and ofloxacin, which agree with the results obtained by Mekhloufi et al. (2021).
In the present study, MRSA strains have been identified among isolated strains. All MRSA strains of our study were from raw milk and traditional dairy products (rayeb and l’ben). As reported by Titouche et al. (2022), milk and milk products have been identified as a source of MRSA demonstrating the potential food safety risk of contaminated milk and milk products entering the human food chain. However, MRSA was also isolated from various food items including, meat of different animal species such as beef, pork, veal, lamb/mutton, chicken, turkey, fowl and game (De Boer et al, 2009; Martínez-Vázquez et al., 2021), ready to eat products (Gelbíčová et al., 2022; Mekhloufi et al., 2021) and fishs from aquaculture and sea (Arfatahery et al., 2016). The prevalence of MRSA is on the rise, which is attributed to its capacity to form reservoirs across diverse hosts and environment, including humans, animals, and various ecological niches (Silva et al., 2022). A part from direct contact with animals or humans in contact with them, MRSA can also can be transmitted through food production systems. However, the risk of acquiring MRSA through food appears to be low (Crespo-Piazuelo and Lawlor, 2021).
The antibiotic susceptibility of MRSA strains showed that all these strains were resistant to antimicrobial agents other than β-lactams, such as tetracycline, aminoglycosides (neomycin), macrolides (erythromycin) and fluoroquinolone (ofloxacin), indicating a multidrug-resistant phenotype, as in other studies (Alghizzi and Shami, 2021; Chaalal et al., 2018; Parisi et al., 2016). As reported by Zhang et al (2005), multidrug resistance is common among MRSA isolates due to their ability to transfer staphylococcal cassette chromosome mecA elements and additional resistance determinants on plasmids. None of our MRSA strains carried the genes encoding the Panton Valentine leucocidin (lukF/S-PV), although these genes have been detected in MRSA strains isolated from various food products, including raw milk (Chaalal et al., 2018; Sadat et al., 2022), meat (Chaalal et al., 2018; Hanson et al., 2011) and ready to eat products (Chaalal et al., 2018). In this study, genetic typing of MRSA isolates from raw milk and traditional milk products (l’ben and rayeb) revealed that these isolates belonged to three spa-types: t024, t450 and t688. Our results agree with those of Herrera et al (2016), who isolated MRSA strains (t024) from raw milk fresh cheese in Colombia, and those of Parisi et al (2016), Basanisi et al (2017) and Zayda et al (2020), who isolated MRSA strains (t688) from milk and milk products.
Conclusions
This study revealed high percentages of antimicrobial resistance, particularly against penicillin and tetracycline. The ability of the isolates to produce biofilms highlights the risk of these potentially virulent strains, especially MRSA isolates, persisting in food processing. Thus, a continuous monitoring and improvement of hygienic quality of food products along the production chain are needed to control the presence of MRSA.
Footnotes
Authors’ Contributions
Y.T. and M.A.: Conceptualization, methodology, investigation, writing—original draft, writing—review and editing. Y.D., A.C., D.M., and L.B.: Formal analysis, writing—review and editing. A.F., Y.N., and J.-A.H.: Supervision, funding acquisition, writing—review and editing.
Author Disclosure Statement
Authors declare no conflict of interest
Funding Information
This work was supported by Laboratory for Food Safety, ANSES, Maisons-Alfort, France.
