Abstract
Because of the medical importance of cumin as well as it being one of the food additives to many Saudi dishes, there was a need to study the fungal load of this type of spice. This study aimed to determine the mycological profile of the retail black and green cumin distributed in different markets at western region, Saudi Arabia, using the dilution plat method on dichloran 18% glycerol (DG18) agar and incubation at 25°C. Using morphological criteria and molecular markers (internal transcribed spacer sequence), 39 species belonging to 18 genera were collected from different black cumin (33 species belonging to 17 genera) and green cumin (25 species belonging to 9 genera). Alternaria alternata, Aspergillus flavus, A. niger, A. ochraceus, Cladosporium cladosporioides, and Stemphylium botryosum were the most prevalent. Black cumin harbors fungal counts reaching 545 colony-forming units (CFU)/g, while green cumin included 500 CFU/g. Also, the natural occurrence of aflatoxins and ochratoxin A was also measured. Seventy-two cumin samples (90% of tested samples) showed toxin contamination. Aflatoxins and ochratoxin A ranged from 9.35 to 3.9 PPB in black cumin samples and from 4.08 to 5.75 PPB in green cumin samples.
Introduction
The second most common spice in the world, after black pepper, is cumin (Cuminum cyminum L.), which is mostly grown in Arabia, China, India, and the nations that border the Mediterranean Sea (Hajlaoui et al., 2010). Cumin has several medical uses, such as a stimulant, carminative, astringent, and therapy for diarrhea, flatulence, and dyspepsia (Norman, 1990). Cumin essential oil (EO) has been shown to be effective against a variety of food pathogenic microbes in antimicrobial tests (Hajlaoui et al., 2010; Khosravi et al., 2011a,b; Mohammadpour et al., 2011; Singh et al., 2002).
The herbaceous plant known as black cumin (Nigella sativa L.) is a member of the Ranunculaceae family. Black cumin seeds have a variety of uses, as an antimicrobial and anticancer medicine product and as a flavoring component in cheese and pastry goods (Al Gaby, 1992; El-Kayati et al., 1995, El‐Dawy et al., 2019; Hanafy and Hatem, 1991; Mahmoud, 1993).
Spices that are susceptible to mycotoxin pollution are typically grown in tropical environments with high temperatures, high humidity, and variable rainfall. Furthermore, when the spices are being dried outside, unhygienic conditions encourage the development of mold and the production of mycotoxins (Martins et al., 2001). Under favorable storage circumstances that encourage fungal development, mycotoxigenic fungi can naturally produce harmful compounds in these commodities known as mycotoxins (Khodaei et al., 2021). The development of fungi and mycotoxin generation can also result from improper post-harvest handling and techniques, as well as poor field-to-market transportation. In the study by El Mahgubi et al. (2013), a few fungus taxa, including Aspergillus, Penicillium, and Fusarium, have the ability to contaminate dry goods and develop mycotoxins. Fusarium is a field fungus that is frequently discovered in crops before harvest. On the opposite hand, storage fungi like Aspergillus and Penicillium can proliferate in crops that are kept in storage and enter grains or seeds, while they are being stored (Afsah-Hejri et al., 2013). Aspergillus and Penicillium are examples of xerophilic fungi that generally thrive well in low-moisture environments (Fleurat-Lessard, 2017). While A. niger, A. carbonarius, and Penicillium verrucosum can produce OTA, A. flavus is the main producer of AFB1 and AFB2 (Bennett and Klich, 2003).
This study sought to ascertain the level of aflatoxins and ochratoxins present in samples of black and green cumin, which were sold in the Kingdom of Saudi Arabia’s western region. It is believed to be the first study to examine the fungal load of green and black cumin under consumption settings, based on the information currently available.
Materials and Methods
Sampling
Eighty cumin samples, including black cumin (n = 40) and green cumin (n = 40), were purchased randomly from retail markets in Jeddah and Taif cities (western region of Saudi Arabia). The sample size of each cumin was 250 g. The samples were saved in plastic bags and stored in freezer at 4°C until further analysis.
Mycobiota determination
Total fungal counts were performed on dichloran 18% glycerol (DG18) agar (glucose 10 g; peptone 5 g; KH2PO4 1 g; MgSO4 7H2O 0.5 g; glycerol 220 g; agar 15 g, dichloran 2 mg; chloramphenicol 100 mg; and H2O 1 L), a medium that has lower water activity (aw = 0.955) and favors xerophilic fungi development (Pitt and Hocking, 1997). Quantitative enumeration was performed by the surface-spread method. Ten grams of each sample were homogenized in 90 mL of peptone water solution. Serial dilutions were made, and 0.1 mL aliquot was inoculated in duplicate onto the culture media. All the plates were incubated in the dark at 25°C for 5–7 days. Only plates containing 15–150 colony-forming units (CFU) were used for counting and the results were expressed as CFU per gram of sample. All colonies were transferred for subculturing to plates of malt extract agar. Taxonomic identification of the different genera and species was made according to macroscopic and microscopic criteria with appropriate keys (Klich, 2002; Nelson et al., 1983; Pitt and Hocking, 1997; Samson et al., 2000).
Molecular identification of fungal isolates
All isolated fungi were subjected to confirmation by molecular analysis. The genomic DNA was isolated using the Gashgari et al. (2010) approach. The internal transcribed spacer (ITS) region of ribosomal DNA (rDNA) was amplified by PCR using primers ITS1-F (CTTGGTCATTTAGAGGAAGTAA) and ITS4 (TCCTCCGCTTATTGATATGC) (Gardes and Bruns, 1993; White et al., 1990). In a final volume of 50 μL, PCR amplifications were carried out using 2 μL of DNA with 0.5 μM of each primer, 150 μM of dNTP, 1 U of Taq DNA polymerase (Promega), and reaction buffer. Amplification was performed in a thermal cycler with an initial denaturation of 3 min at 94°C, followed by 35 cycles of 1 min at 94°C, 1 min at 50°C, 1 min at 72°C, and a final extension of 10 min at 72°C. Aliquots of PCR products were electrophoresed on a 1% agarose gel, stained with ethidium bromide, and observed using UV transillumination. ExoSAP-IT (USB Corporation, under license from GE Healthcare) purified the PCR products following one manufacturer’s directions. The purified products were sequenced using an automated DNA sequencer (ABI PRISM 3700) and the BigDye Deoxy Terminator cycle-sequencing kit (Applied Biosystems) according to the manufacturer’s instructions. Sequences were submitted to GenBank through the NCBI website (http://www.ncbi.nlm.nih.gov). The sequences acquired in this investigation were compared to the GenBank database using the BLAST software available on the NCBI website (http://www.ncbi.nlm.nih.gov/BLAST).
Detection of natural occurrence of total aflatoxins and ochratoxin A in cumin samples
The natural occurrence of total aflatoxins and ochratoxins in 80 samples was determined by using a slightly modified method based on the Association of Official Analytic Chemists (AOAC) method reported by Trucksess et al. (1991). Methanol:water (80:20), solvent (100 mL), and 5 g NaCl were added to 100 g of ground each sample, and the mixture was shaken in a blender at maximum speed for 3 min. Filtration process occurred through fluted filter paper (Whatman 2V; Whatman plc), and the filtrate was diluted (1:4) with water and refiltered through glass‐fiber filter paper. Two milliliters of the glass‐fiber filtrate were placed on an AFs or OTs Test RWB SR Column (VICAM) and allowed to elute at 1–2 drops/s. The columns were washed two times with 5 mL water, and aflatoxin or ochratoxins were eluted from the column with 1 mL high‐performance liquid chromatography (HPLC)‐grade methanol. A bromine developer (1 mL) to the methanol extract was added, and the total AFs or OT concentrations were read in a recalibrated VICAM Series‐4 fluorometer set at 360 nm excitation and 450 nm emission. The modified fluorometry method had ≥85% recovery and a 1 ppb limit of detection (Lewis et al., 2005).
Results and Discussion
Mycobiota of cumin samples
Fungal growth may occur before or during drying, storage, or shipping of spices, as reported by several researchers (Elshafie et al., 2002; Freire et al., 2000; Hashem and Alamri, 2010, Gherbawy et al., 2015; El‐Dawy et al., 2019).
The prevalence and population density of the mycobiota isolated in this study are shown in Figure 1. The moisture content mains were 6.47 and 8.15 from black and green cumin samples, respectively. In India, Moorthy et al. (2011) reported the moisture content of green cumin ranged from 7.41 to 9.31, while 4.45 to 8.47 for black cumin.
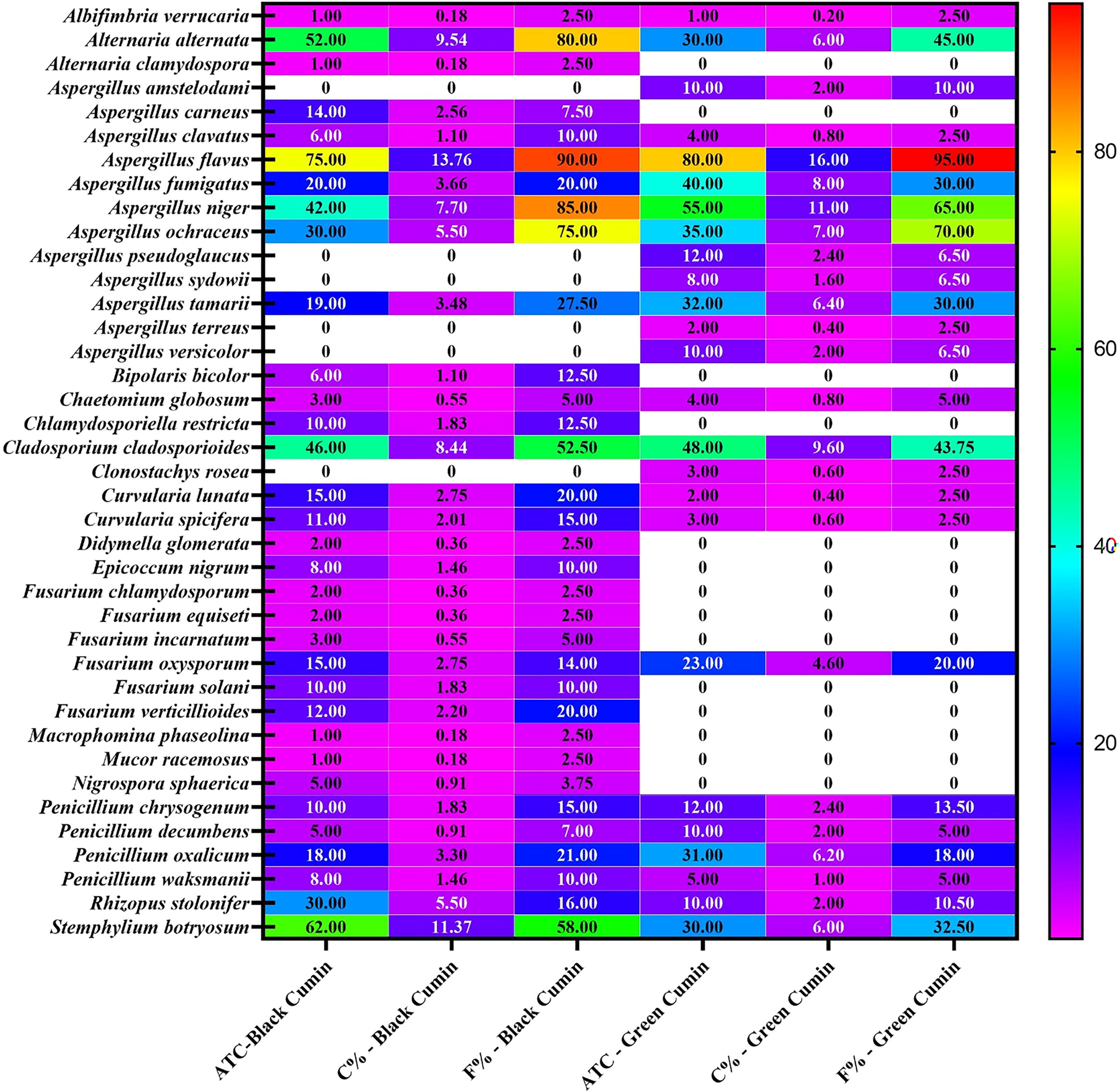
Average total counts (C), percentage of C%, frequency (F%) of fungal species isolated from 80 samples of black and green cumin on dichloran 18% glycerol (DG18) agar and incubation at 25°C.
The average total counts of collected fungi were 545 and 500 CFU/g from black and green cumin samples. There is high correlation between the average total counts of isolated fungi from both black and green cumin and the correlation between them was 0.7 (Fig. 2). Bokhari (2007) reported that the average of total counts of collected fungi isolated from black cumin was 680.3 CFU/g (out of 5 samples), while it was 1634.6 CFU/g from green cumin in Jeddah, Saudi Arabia. Also, the average total counts of fungi isolated from cinnamon, green cumin, pepper, green cardamom, caraway, and cloves were from 100 to 1000 CFU/g in Aseer region, Saudi Arabia (Hashem and Alamri, 2010). The gross total counts for 41 and 34 fungal species isolated from cumin samples in Egypt using glucose—and cellulose—Czapeks agar, were 51,950 and 47,550 CFU/g, respectively (Saleem et al., 2013). The total fungal counts that contaminated green and black cumin in Upper Egypt were 1,933.32 and 4,349.99 CFU/g (El‐Dawy et al., 2017).
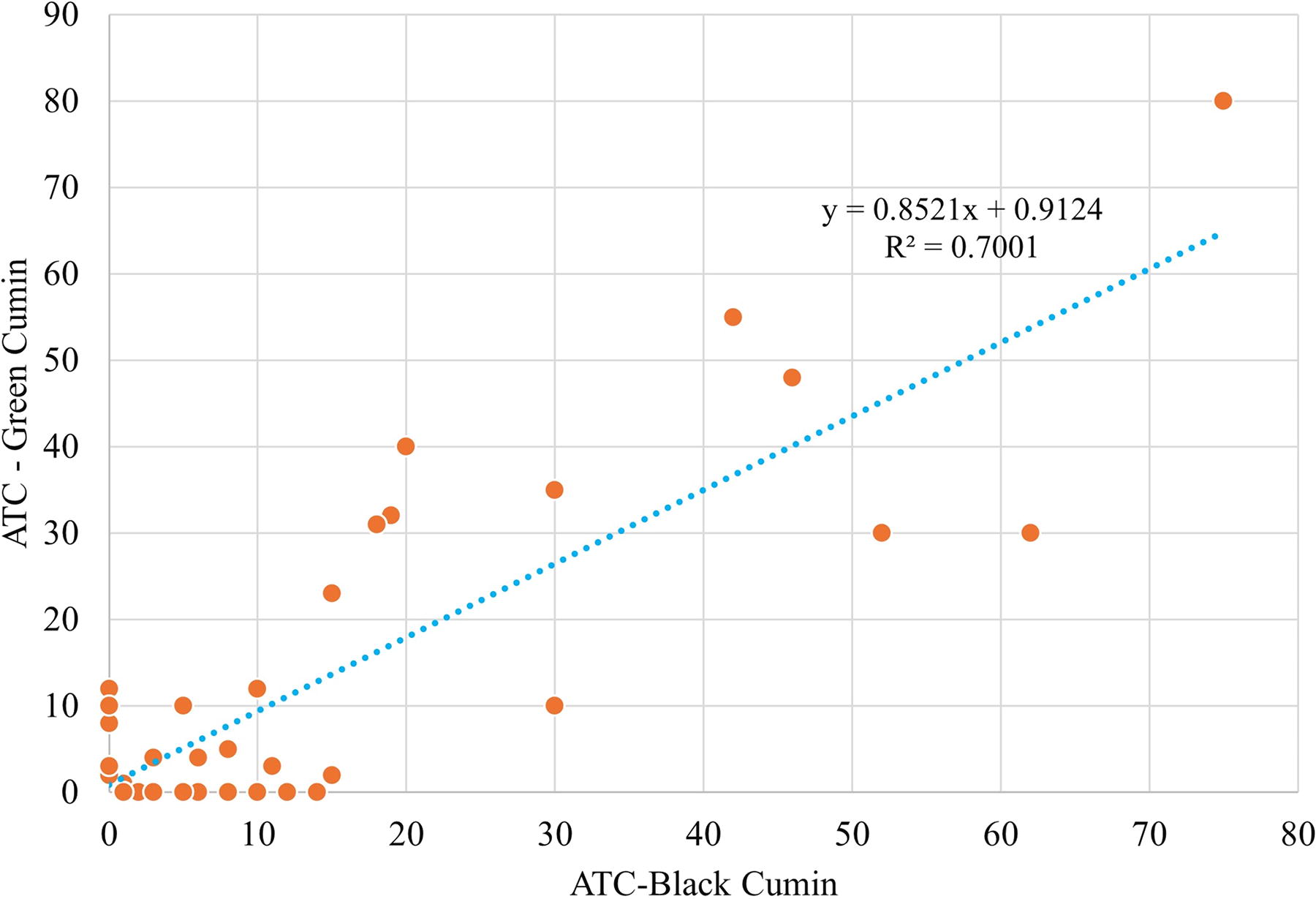
The correlation between average total counts of fungi isolated from black and green cumin samples.
Thirty-nine species belonging to 18 genera were collected from the different black cumin (33 species belonged to 17 genera) and green cumin (25 species belonged to 9 genera) samples and identified from their morphological criteria and ITS sequence (Fig. 1). Sequences were submitted to GenBank on the NCBI website (http://www.ncbi.nlm.nih.gov). The sequence results were deposited in GenBank under accession numbers from PP873058 to PP873096 (Table 1). Elwakil and Ghoneem (1999) isolated 37 fungal species that belonged to 20 genera from black cumin seeds in Egypt.
Isolated and Identified Fungal Species with its Relationship to the Genus or Species and the Identity Percentage Found in the GenBank
Aspergillus was the most common genus in the different species tested. The average total counts of this genus in black and green cumin samples were 206 and 288 CFU/g, representing 37.79 and 57.6 of total fungi, respectively. Twelve species collected from Aspergillus. A. flavus, A. fumigatus, A. niger, A. ochraceus, and A. tamarii were the most prevalent. They emerged from 19–75 CFU/g in black cumin samples, contributing 9.2–36.41% of total aspergilli and 3.48–13.76% of total fungi. Their percentage of occurrence ranged from 27.5% to 90% for total frequencies of all isolated fungi (Fig. 1). Elwakil and Ghoneem (1999) studied the seed-borne fungi of black cumin in Egypt. They isolated A. carneus (20.5 CFU/g and 18% of total number of samples), A. clavatus (4.5 and 4%), A. flavus (80.7 and 71%), A. fumigatus (20.5 and 18%), A. niger (36.6 and 32%), A. ochraceus (29.5 and 26%), and A. tamarii (13.6% and 12%). Fourteen isolates were collected from samples of one cumin from China markets (Kong et al., 2014). In green and black cumin samples collected from Upper Egypt, Aspergillus spp. occurrence percentages were 58.19% and 85.05%. (El‐Dawy et al., 2017). On the other hand, the previously mentioned species emerged from 32 to 80 CFU/g of green cumin, contributing 11.11–27.77% of total aspergilli and 6.4–16% of total fungi, with 30–95% of total isolated fungal frequencies (Fig. 1). A. flavus and A. tamarii are well known as aflatoxin producers, as reported by several researchers, so more attention is given to their total counts and frequencies. The average total counts of A. flavus and A. tamarii were 75–19 and 80–32 CFU/g from black and green cumin, while their frequency percentages were 90–95% and 93.75–30%, respectively (Fig. 3). Gracia et al. (2001) did not detect aflatoxigenic molds (A. flavus or A. parasiticus) from cumin samples sold in Mexican markets. In Oman, the average total count of A. flavus in black cumin was 225.5 CFU/g (Elshafie et al., 2002). Using PDA medium, Hashem and Alamri (2010) noticed that the percentage of cumin samples in Aser region with A. flavus was 40%.
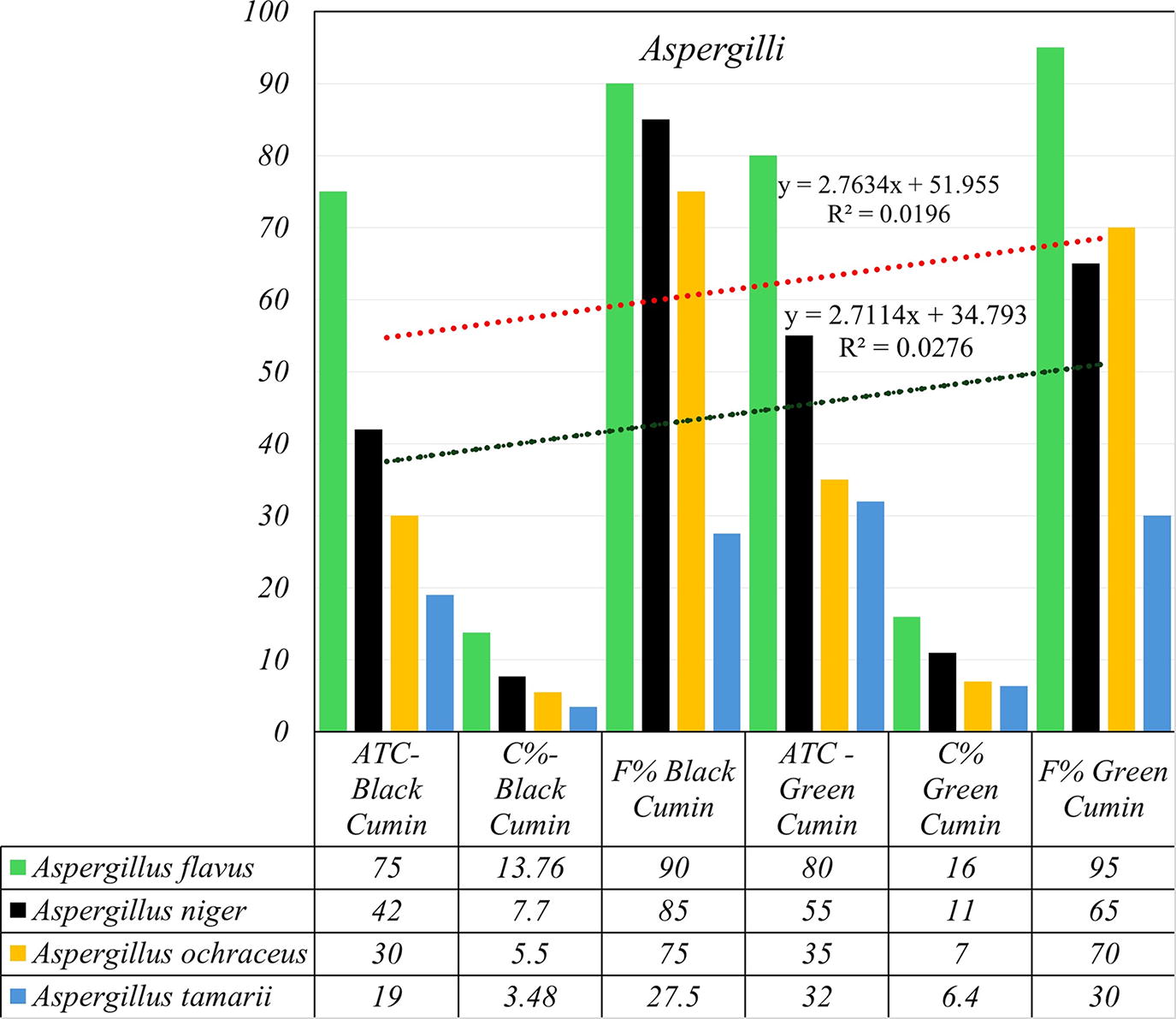
The correlation between aflatoxigenic (Aspergillus flavus and A. tamarii) and ochratoxigenic (A. niger and A. ochraceus) species-contaminated cumin samples.
A. niger and A. ochraceus were reported as ochratoxin producers. In this study, A. niger was isolated from 85% and 65% of black and green cumin samples with total average counts of 42 and 55 CFU/g. Also, A. ochraceus was isolated from 75% and 70% of both cumin samples, comprising 30 and 35 CFU/g average total counts (Fig. 3). The relative density of A. niger in cumin samples in China was 15.2 (kong et al., 2014). A. niger was detected in 80% of green cumin samples in Aser region, but A. ochraceus did not show up in any sample (Hashem and Alamri 2010). A. niger contaminated 13 samples (28%) of cumin from Mexican markets (García et al., 2001). In India, A. niger was one of the predominant species isolated in all the spices studied, with average contamination ranging from 2.6% to 19.6% (Jeswal and Kumar, 2015; Yogendrarajah et al. (2014); also, A. niger was found to be one of the predominant species in Piper nigrum from Sri Lanka. According to the authors, A. niger was present in 62% of the samples, with contamination levels higher than 1.3 × 103 CFU/g. In Oman, from black cumin samples, Elshafie et al. (2002) isolated A. niger and A. ochraceus with average total counts 111.7 and 111.1 CFU/g. A. carneus appeared in black cumin samples only, while A. amstelodami, A. pseudoglaucus, A. sydowii, A. terreus, and A. versicolor were isolated from green cumin samples (Fig. 1).
A. flavus, A. niger, and A. ochraceus sections were isolated with consideration for potential mycotoxin-producing fungi; of these, A. niger was significantly predominant. These findings are consistent with those of other studies (Abdel-Hafez and El-Said, 1997; Abdulkadir et al., 2003; El Mahgubi et al., 2013; El-Kady et al., 1992; Garrido et al., 1988; Hashem and Alamri, 2010; Hitokoto et al., 1979; Lee and Lim, 1985; Moharram et al., 1989; Pal and Kundu, 1972; Sharma and Sharma, 1984; Udagawa, 1982).
Dematiaceous hyphomycetes fungi ranked second according to their average total counts and frequency percentages. Alternaria alternata, Cladosporium cladosporioides, and Stemphylium botryosum were the most common species in this group. They were isolated from 80%, 52.5%, and 58% of back cumin samples (Fig. 1). The percentages of contamination of cumin samples from Aser region with A. alternata, C. cladosporioides, and S. botryosum were 40%, 10%, and 60% from tested cumin samples (Hashem and Alamri 2010).
The same findings are consistent with the research conducted by Saleem et al. (2013), which found that the common genera that contaminate anise and cumin seeds are Alternaria, Aspergillus, Emericella, Mucor, Penicillium, Stachybotrys, and sterile mycelia. Furthermore, El-Gali (2014) has demonstrated that Aspergillus spp., Penicillium spp., Alternaria spp., and Fusarium spp. were the most common fungal genera isolated from 14 species of spices, including turmeric, cumin, and ginger. A. flavus and A. niger were the most often isolated species from nine different types of spices, which also included ginger, turmeric, and cumin, according to Jeswal and Kumar’s (2015) research.
Natural occurrence of aflatoxin and ochratoxin in cumin samples
It has been reported that, under favorable conditions, some fungal species can synthesize toxic metabolites (mycotoxins), and most of the identified fungi, such as A. flavus section, A. niger section, and A. ochraceus section produce mycotoxins, namely aflatoxins (AFB1, AFB2, AFG1, and AFG2) and ochratoxins (OTA) (Romagnoli et al., 2007). Other previous studies have reported some mycotoxin contamination, especially AFB1, AFB2, AFG1, AFG2, and OTA in spices, aromatic and medicinal herbs (Bokhari, 2007; El-Dawy et al., 2019; Liu et al., 2012, Gherbawy et al., 2015; Zinedine et al., 2006).
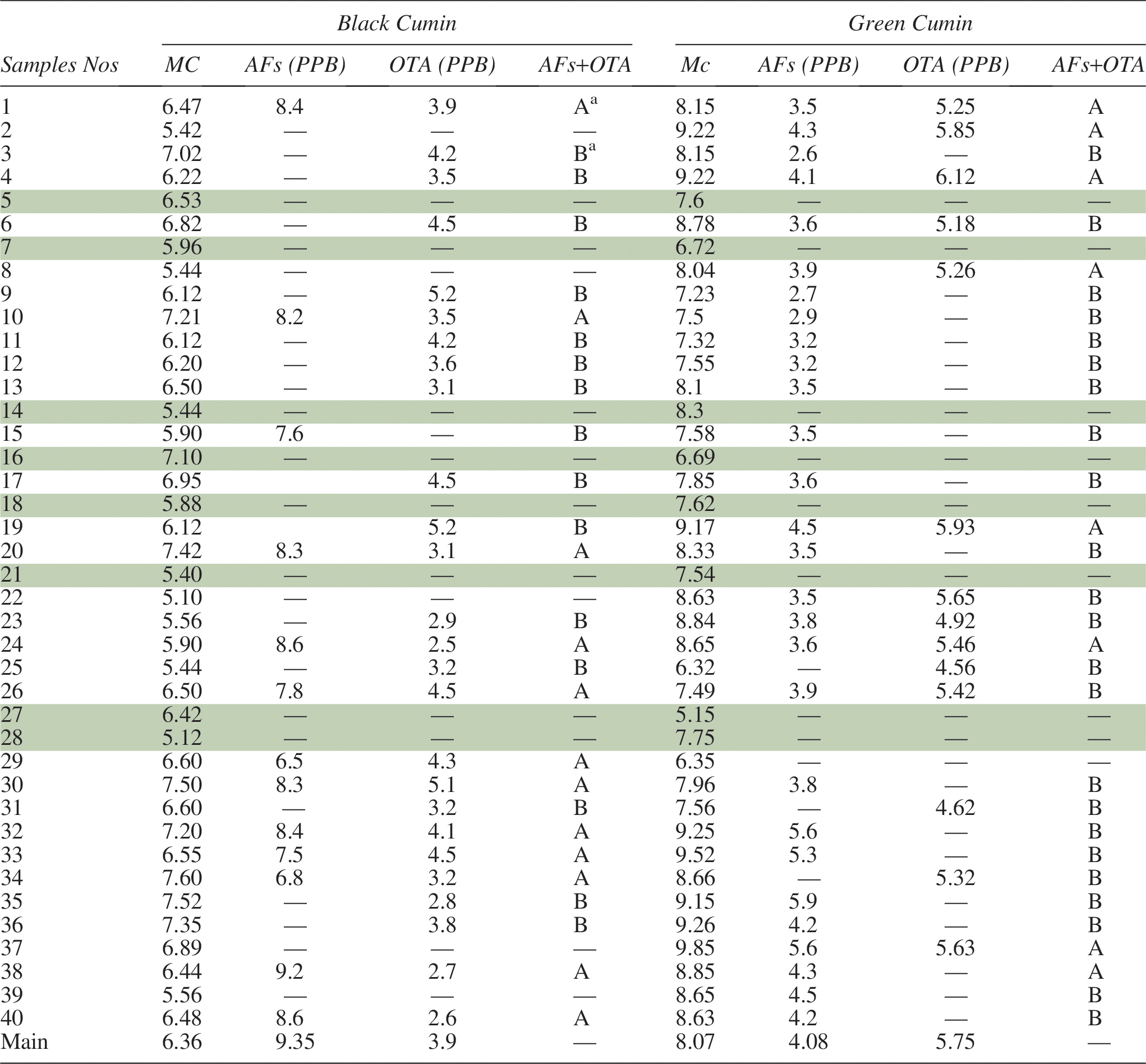
The main moisture content for black cumin samples was 6.36. Twelve samples (30% of the tested samples) showed total aflatoxin occurrence with percentage ranging from 6.5 to 9.2 PPB and 26 samples (65%) showed ochratoxin contamination with percentage ranging from 2.5 to 5.2 PPB. Twelve samples showed co-occurrence of both toxins, matching 30% of examined samples, while 15 samples (37.5% of total samples) showed single type of toxin detection (Table 2). In Morocco, Zinedine et al. (2006) noticed that 57% of cumin samples were contaminated with AFB1 and AFs, and the average contamination in the positive samples was 0.03 and 0.05 μg/kg, but the maximum levels were 0.08 and 0.18 μg/kg. In Upper Egypt, El-Dawy et al. (2019) investigated the natural occurrence of AFs and OTA in black and green cumin in Upper Egypt. Their results indicated that the main values from AFs and OTA were 8.2–1.9 PPB and 4.5–4.9 PPB, respectively.
Moisture Contents (MC), Total Aflatoxins (AFs), Ochratoxin A (OTA), and Co-Occurrence of AFs + OTA in Different Cumin Samples
A: Both AFs and OTA were detected in the sample.
B: One of the AFs or OTA was detected in the sample.
 Negative samples.
Negative samples.
On the other hand, the main moisture content of green cumin samples was 8.07 and the positive samples in terms of mycotoxin occurrence were 31 samples comprising 77.5% of total examined samples. The total aflatoxins were detected in 28 samples (70%) ranging from 2.6 to 5.9 PPB. Also, ochratoxin was detected in 14 samples (35%) ranging from 4.56 to 6.12 PPB. The co-occurrence of both toxins was detected in eight samples of green cumin, matching 20% of total examined samples (Table 2). In Corum and Ankara cities, Turkey, Ozbey and Kabak (2012) studied the natural co-occurrence of aflatoxins and ochratoxin A in spices. Their results showed that the positive samples from cumin for total aflatoxin and ochratoxin were 4 (21.1%) and 1 (5.3%), the range of AFs and OTA were 0.32–0.97 and 0.63 mg/kg−1. Rani and Singh (1990) found that 89% of samples of fennel, coriander, and cumin were contaminated with aflatoxin B1 at the levels 3000, 1640, and 1580 mg/kg, respectively.
In eight of 80 cumin samples used in this investigation, none of the two mycotoxins was detected. The European Union has set maximum allowable limits for total aflatoxins in spices at 10 mg/kg and ochratoxin A at 15.0 μg/kg. Since none of the positive samples in this investigation exceeded the limitations set by the European Union, no treatment for the leftover censored data was carried out.
Conclusion
The results of the mycological investigations showed that Aspergillus spp. was the most common fungus and that the black cumin samples were severely contaminated. In comparison to black cumin, green cumin samples had a higher percentage of AF contamination. Compared to the number of positive samples from green cumin, there were more positive samples from black cumin regarding OTA contaminations. The chemical composition of the spices, their antimicrobial qualities, the ways in which they are handled by different people, and environmental influences are some of the possible causes of this variance in the microbiological and mycotoxin profiles of the two spices. With proper packaging and hygienic conditions during harvesting, processing, and storage, spices can be produced in an acceptable quality.
Footnotes
Acknowledgments
The authors would like to acknowledge the Deanship of Graduate Studies and Scientific Research, Taif University, for funding this work.
Author’s Contributions
Conceptualization, methodology, validation, formal analysis, investigation, data curation, writing—original draft preparation, writing—review and editing, visualization, supervision, project administration, and funding were performed by the author. The author has read and agreed to the published version of the article.
Data Availability Statement
All data generated or analyzed during this study were included in this article.
Disclosure Statement
No potential conflict of interest was reported by the author(s).
Funding Information
The author reported that the Deanship of Graduate Studies and Scientific Research, Taif University, was funding this work.