Abstract
In 2022, the U.S. Food and Drug Administration (FDA), the Centers for Disease Control and Prevention (CDC), and state partners conducted a sample-initiated investigation of a multistate outbreak of Salmonella Senftenberg illnesses linked to peanut butter. Twenty-one illnesses and four hospitalizations were reported in 17 states, with a significant epidemiological signal for peanut butter from Firm A. Whole genome sequence (WGS) data from a Salmonella-positive environmental swab sample collected at Firm A in 2010 yielded the outbreak strain that was a match to the WGS data from the 2022 clinical isolates. Lot code information collected from patients indicated Firm A’s facility in Kentucky as a common manufacturing source, and FDA and state partners initiated an inspection. In 2021, Firm A installed two new roasters with at least one of the cooling air supply vents leaking, allowing unfiltered air and rainwater to enter the cooling section after the roasting process. Investigators noted the limitations of Firm A’s finished product testing program to identify contamination. Investigative partners from five states collected and analyzed 14 product samples, and FDA collected 205 environmental swabs, and all were negative. Although the exact source and route of the contamination were not determined, epidemiological and traceback evidence confirmed peanut butter consumed by patients was produced by Firm A. Firm A voluntarily recalled all implicated products and provided a plan for corrective actions and restart to FDA. This was the first major domestic investigation of a multistate-foodborne illness outbreak linked to peanut butter since 2012. This investigation demonstrates the importance of caution with reliance on finished product testing, taking appropriate corrective actions when detection occurs, and potential benefits for industry to incorporate WGS as a tool in their environmental monitoring program.
Introduction
The United States leads the world in peanut butter consumption, and it is found in 90% of households (Sithole et al., 2022). In 2020, approximately 87.9 million people in the United States reported consuming at least one jar per month (Statista, 2021). Salmonella spp. contamination of low-moisture foods, such as peanut butter, is a challenging food safety risk (Sithole et al., 2022). In the United States, there have been three notable outbreaks associated with peanut butter and peanut products caused by Salmonella Tennessee, Typhimurium, and Bredeney infections in 2006, 2008, and 2012, respectively (Cavallaro et al., 2011; Sheth et al., 2011; Viazis et al., 2015). These led to approximately 1471 illnesses, 269 hospitalizations, and 9 deaths, combined.
In 2022, federal, state, and local partners investigated a multistate outbreak of Salmonella Senftenberg illnesses linked to peanut butter. Here, we describe a sample-initiated retrospective outbreak investigation (SIROI), where the investigation began with a possible vehicle based on historic laboratory evidence (Wellman et al., 2023), and was combined with epidemiological and investigational findings to identify peanut butter as the source (U.S. Centers for Disease Control and Prevention, 2022; U.S. Food and Drug Administration, 2022b).
Materials and Methods
Outbreak detection
Surveillance by the Food and Drug Administration (FDA) of whole genome sequencing (WGS) of nonclinical bacterial isolates with single nucleotide polymorphism (SNP) analysis and the National Center for Biotechnology Information’s Pathogen Detection database (NCBI) was used to identify clinical isolates matching peanut butter and environmental isolates (ncbi.nlm.nih.gov/pathogens) that were previously submitted to the database (Sayers et al., 2021). The NCBI and PulseNet databases were used to determine the clinical isolates included in the outbreak definition (U.S. Centers for Disease Control and Prevention, 2016).
Within PulseNet, sequences from clinical and nonclinical isolates were compared in BioNumerics v 7.6.3 database using a predefined core genome multilocus sequence typing (cgMLST) allele scheme to identify genetic relatedness. All clinical isolates were serotyped and subtyped using WGS by state public health laboratories and uploaded to NCBI and PulseNet (National Center for Biotechnology Information, 2016).
Epidemiological investigation
FDA notified Centers for Disease Control and Prevention (CDC) partners of the WGS analysis (Wellman et al., 2023). State and local health officials interviewed patients with state enteric and outbreak-specific questionnaires to gather demographics, health outcomes, food exposure information, and details on peanut butter exposures.
Traceback investigation
Traceback activities were based on brand and lot codes collected from patients compared to the lot code scheme applied by Firm A.
Firm investigations
FDA and state partners investigated Firm A in 2022, which included interviews with key personnel, observation of the firm’s operations, records review, and collection of environmental swab samples for microbiological analysis.
Nonclinical microbiological investigation
State and FDA laboratories used the FDA Bacteriological Analytical Manual method to test for Salmonella spp. from product and environmental samples (Andrews et al., 2018). Opened and unopened product samples were collected by state partners from patients, and FDA collected environmental swabs.
Results
Outbreak detection
In April 2022, FDA bioinformaticians identified five clinical isolates from 2022 in NCBI as closely genetically related to numerous isolates that were collected between 2014 and 2017 and uploaded by a third-party laboratory. These isolates were labeled as “peanut butter” or “environmental” with a location of Kentucky. FDA engaged CDC investigators and identified a 2010 FDA environmental isolate of Salmonella Senftenberg recovered from the Kentucky manufacturing facility of Firm A; sequence analysis was conducted concurrent to the initiation of the epidemiologic investigation. Analysis revealed that the 2010 isolate was genetically related to the clinical isolates by 14–17 SNPs (Fig. 1) (Pightling et al., 2018). Based on 12 years between the sample collection and the clinical isolates, and with an assumption of genetic evolution at 1–2 SNPs per year, it was concluded that the 2010 sample was closely genetically related to the clinical cases (Pightling et al., 2022). The earlier timeframe of 2014–2017 predates routine use of WGS for clinical isolates; rather, illness cluster detection was accomplished by pulsed-field gel electrophoresis. Despite this difference in subtyping methods, there were no illness clusters likely associated with these nonclinical isolates at the time they were uploaded to NCBI based on serotype and epidemiological association.
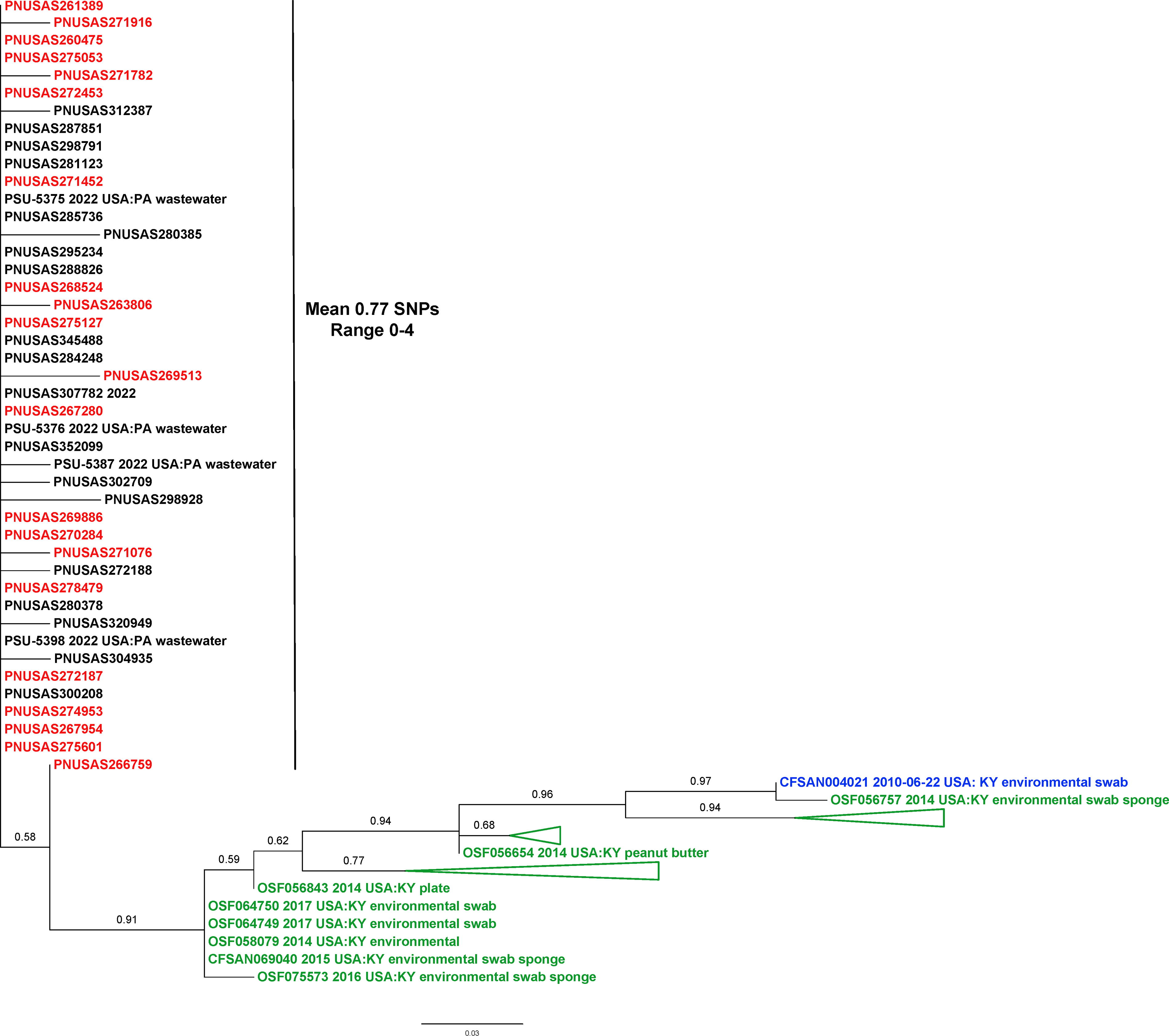
Phylogenetic analysis of Salmonella Senftenberg whole genome sequence data. Clinical isolates are labeled in red, an environmental isolate that was collected in 2010 is labeled in blue, and environmental and food isolates that were collected in 2014 through 2017 are labeled in green. Triangles represent clades of environmental and food isolates collected from 2014 through 2017 that were collapsed. Sequence data were analyzed with the Center for Food Safety and Applied Nutrition (CFSAN) SNP Pipeline. SNP, single nucleotide polymorphism.
Epidemiological investigation
PulseNet identified 21 patients from 17 states (Fig. 2). Isolates were closely genetically related within 0–2 allele differences by cgMLST. An illness was defined as infection with Salmonella Senftenberg, closely related by WGS within 0–2 allele differences, and isolation dates between February 20 and May 24, 2022. Reported illness onset dates ranged from February 19, 2022, to May 23, 2022 (Fig. 3). The median age of patients was 59 (range of <1 to 85 years), and 75% were female. Of 13 patients with outcome information, 4 were hospitalized, and no deaths were reported. Of 13 patients with exposure information, all (100%) reported consuming Brand A peanut butter from Firm A prior to illness onset.
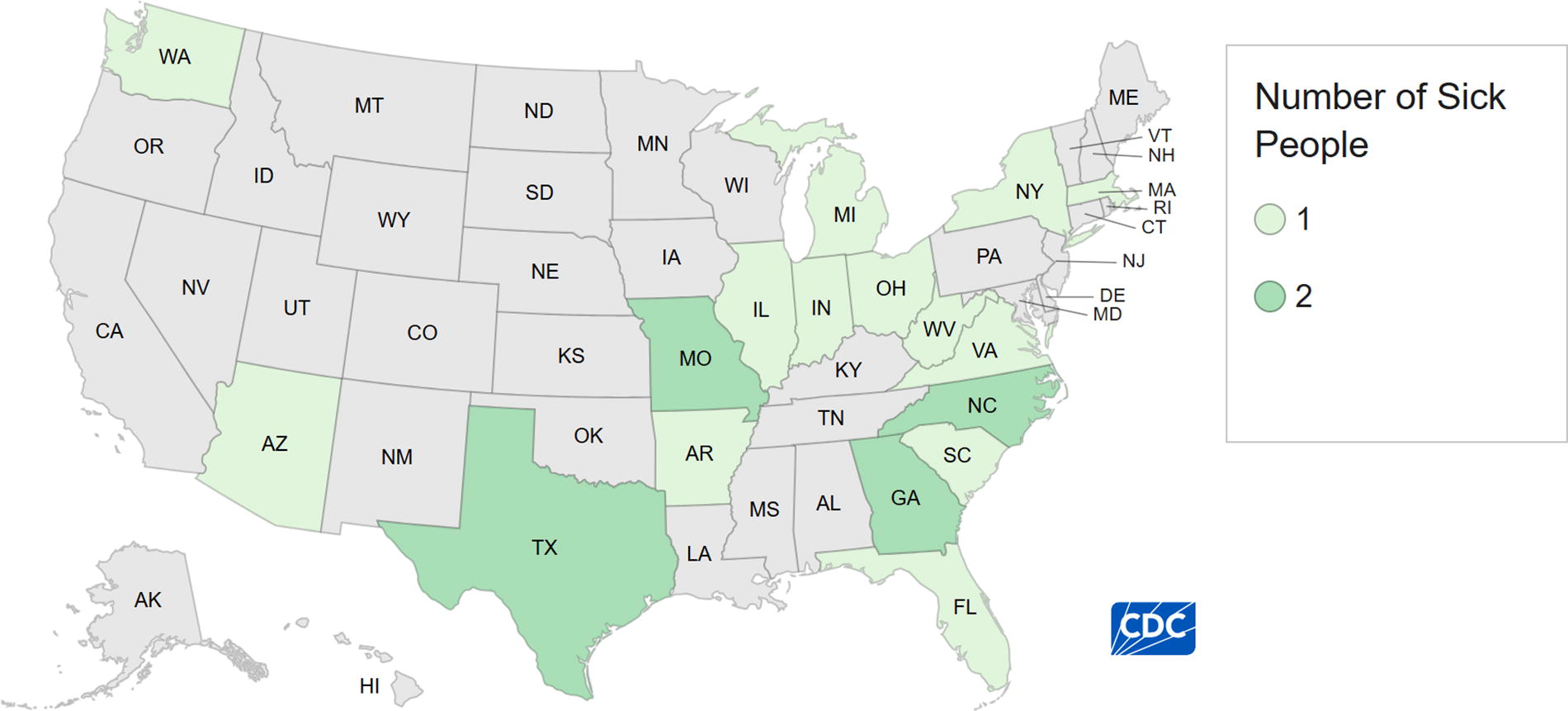
Patients infected with the outbreak strain of Salmonella Senftenberg (n = 21) by state of residence, United States, 2022.
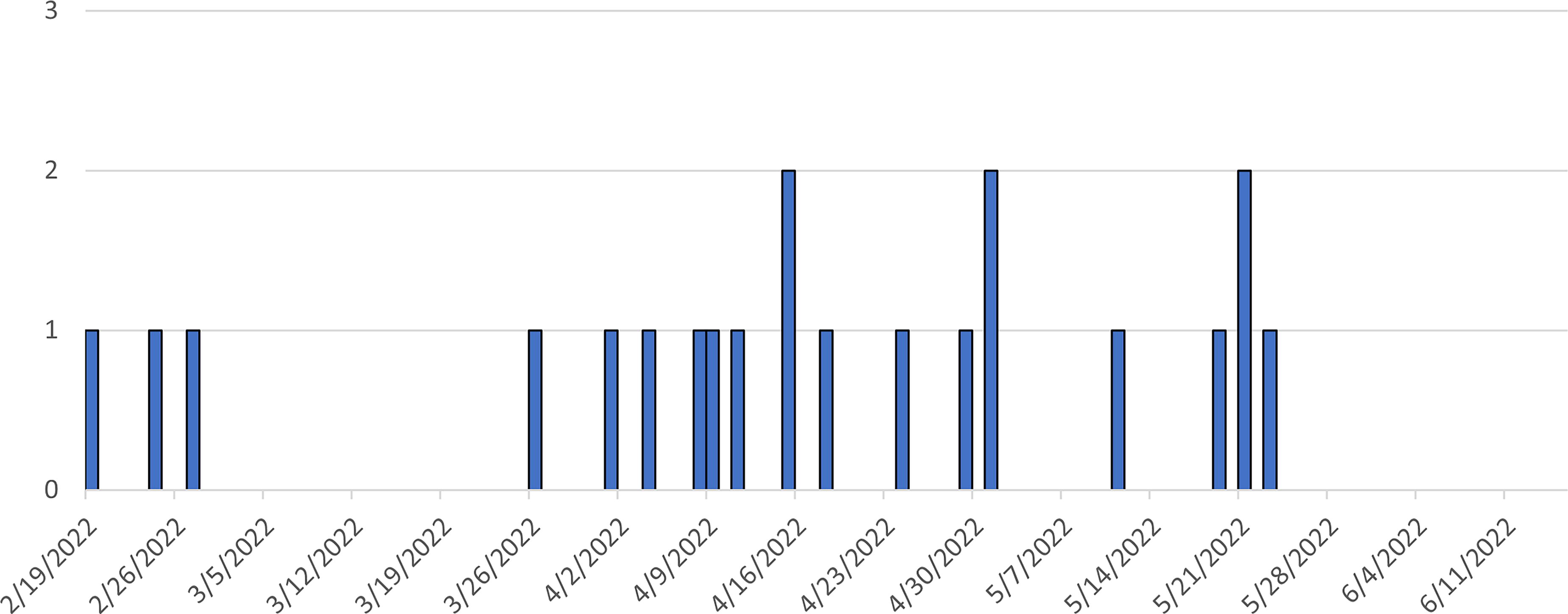
Patients infected with the outbreak strain of Salmonella Senftenberg (n = 21) by date of illness onset, United States, 2022. Some illness onset dates have been estimated from other reported information.
Traceback investigation
Product lot codes were provided by 9 of 13 patients asked, with 5 patients reporting exposure to multiple lot codes. Multiple sizes and varieties were reported across all patients (Table 1). Purchases were made from at least seven different retail chains. The codes provided had manufacturing dates from June 2, 2021, to February 27, 2022. The patient who had product with the earliest manufacturing date in June 2021 was also exposed to a manufacturing date in December 2021. Manufacturing dates in January 2022 explained six of the nine ill people with lot code information. Two manufacturing dates were reported by two patients each: January 17, 2022, and January 26, 2022. Variety, size, and manufacturing information contained in the printed lot code were evaluated and aligned with the firm investigation as roasters are not exclusive to variety or size.
Summary of Product Information from the Patients
Each state represented includes a single patient whose lot codes were available, some with multiple peanut butter purchases.
First seven digits of lot codes. These digits represent the year, day of manufacture, and the manufacturing location code. Full lot codes have additional information, including a production timestamp.
All samples tested by the states were negative for Salmonella.
Not reported.
Firm investigation
From May 19 through June 9, 2022, FDA investigators and state partners inspected Firm A’s manufacturing facility as part of a full-scope preventive controls inspection and discussed with management potential routes of contamination. The firm’s hazard analysis for their ready-to-eat (RTE) peanut butter did not consider the hazard of recontamination with environmental pathogens, such as Salmonella spp., at post-roasting processing steps from blanching to sorting and milling. In-process RTE peanuts were exposed to the environment and did not receive further treatment or include a control measure that would significantly minimize the pathogen.
FDA collected and reviewed the firm’s finished product testing records from January 1, 2021, to February 23, 2022. The firm detected Salmonella in RTE peanut butter on two production dates in late-2021 and five production dates in early-2022 (Fig. 4). The firm indicated root cause investigations were conducted, and they implemented corrective actions consistent with their environmental monitoring procedures (U.S. Food and Drug Administration, 2023). In mid-February 2022, the firm identified and repaired a leak in the air intake vent of the cooling chamber of a roaster, identified as a source of unfiltered air and rainwater entering their equipment. After the repair was completed, Salmonella was detected in product lots using the firm’s routine testing program. Firm A’s practice following a lot testing positive in their routine testing program was to conduct intensified sampling of the lots produced immediately before and after the positive lot and continue this process until positive lots were bracketed by lots that tested negative by intensified testing. On multiple occasions, product that had initially tested negative in the routine testing program, tested positive when subjected to the intensified sampling plan. Firm A destroyed all production lots that tested positive as well as the adjacent lots that tested negative by the intensified sampling plan in their attempt to bracket and eliminate any potential contamination. The firm concluded they had all adulterated finished product under their control and therefore did not contact the FDA regarding the positive samples.
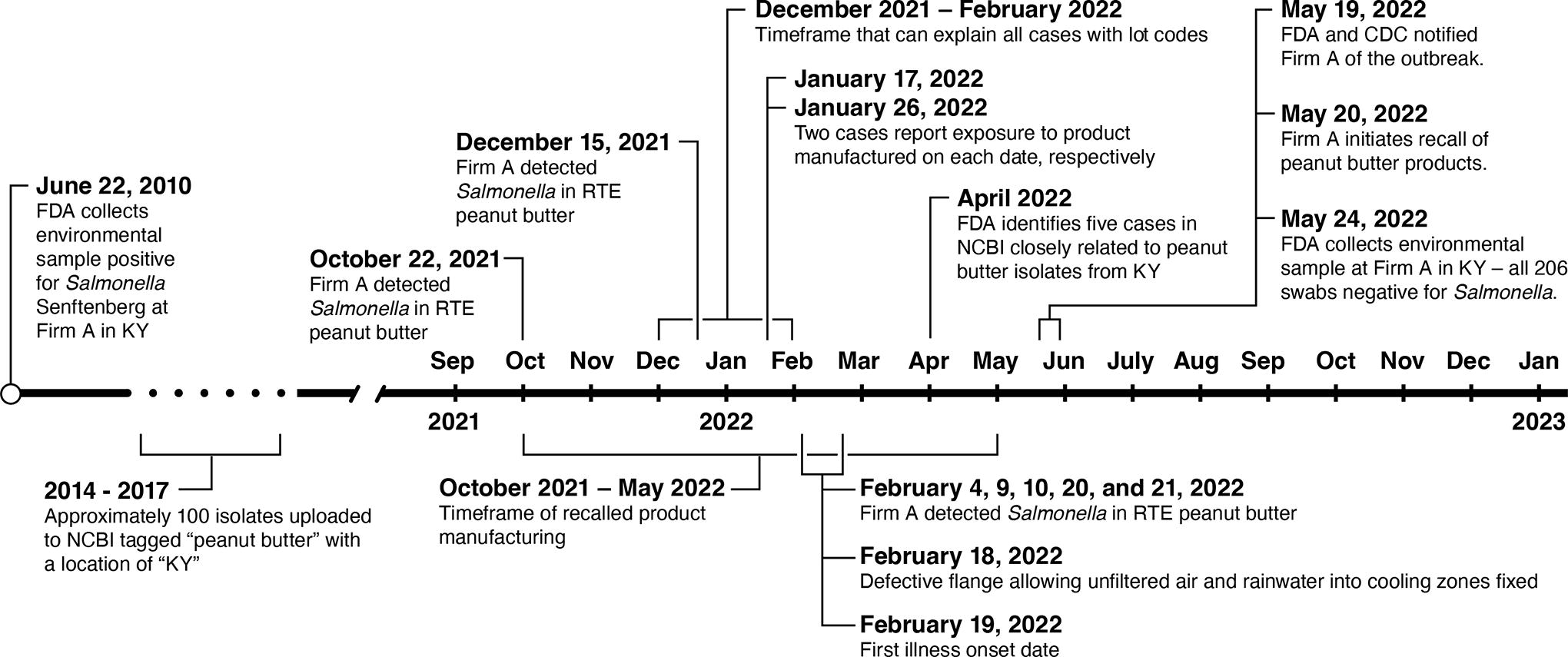
Timeline of events that took place, including the outbreak identification, firm activities, regulatory actions, inspections, recalls, and results of product testing. CDC, Centers for Disease Control and Prevention; FDA, Food and Drug Administration; KY, Kentucky; NCBI, National Center for Biotechnology Information; RTE, ready-to-eat.
Microbiological investigation
FDA collected 205 environmental swabs throughout Firm A’s manufacturing facility, and five state partners analyzed at least 14 product samples from patients. No samples yielded any Salmonella spp. isolates.
Investigation outcomes
On May 20, 2022, Firm A agreed to voluntarily recall peanut butter produced at their facility manufactured between October 1, 2021, and May 20, 2022, which precipitated several downstream recalls. Firm A’s products were also distributed to multiple countries and investigators worked with the International Food Safety Authorities Network on international distribution and awareness. FDA issued a Warning Letter to the firm (U.S. Food and Drug Administration, 2023).
Public communication
Firm A’s recall notice was published on FDA’s website (U.S. Food and Drug Administration, 2022a) on May 20, 2022, and an outbreak advisory was posted on May 20, 2022, by FDA (U.S. Food and Drug Administration, 2022b), and on May 21, 2022, a food safety alert was posted by CDC (U.S. Centers for Disease Control and Prevention, 2022). FDA and CDC’s communications were updated through July 27, 2022, to include advice to consumers detailing updated recall information. FDA and CDC included recommendations for pet owners not to feed recalled peanut butter to their pets in their advisories, social media, and a CDC-issued newsletter.
Discussion
Significance of outbreak
This multistate Salmonella Senftenberg SIROI identified peanut butter as the source of illness, underscoring the importance of the SIROI process to assist in identifying outbreaks earlier compared to traditional outbreak identification by providing a focused hypothesis when collecting epidemiological information (Wellman et al., 2023). Although other recent outbreaks (Hassan et al., 2019; Hassan et al., 2018; U.S. Centers for Disease Control and Prevention, 2014) have been associated with the broader category of “nut butters,” this was the first U.S. outbreak exclusively linked to peanut butter since 2012, with previous outbreaks in 2008 and 2009 (Cavallaro et al., 2011; Sheth et al., 2011; Viazis et al., 2015). All three past peanut butter outbreaks were characterized by two common findings: (1) the detection of Salmonella from products and/or the processing environment over a prolonged period, ranging from 6 months for two firms to 3 years for one firm, which demonstrated a continued failure to control the pathogen in the processing environment, and (2) all three firms failed to take the appropriate corrective actions when identifying Salmonella during product testing. Similarly, Firm A’s corrective action procedures did not ensure appropriate action taken, when necessary, to reduce the likelihood that environmental contamination would recur when they detected Salmonella in their facility’s environment.
Finished product testing
Neither FDA nor the states isolated Salmonella from samples associated with Firm A during this outbreak. Firm A identified Salmonella on multiple occasions, including lots for which Salmonella was not initially detected during routine sampling and testing. Firm A’s finished product testing procedures included holding product in the event a positive result was detected in their RTE peanut butter to allow the intensified testing of the lots produced immediately before and after the positive lot and continue this process until positive lots were bracketed by lots that tested negative by intensified testing. According to Firm A, this approach could ensure any contamination would be included in the held product, identified, and prevented from entering commerce; however, this approach did not identify all the contaminated lots leading to the outbreak. Therefore, this outbreak demonstrates the well-established shortcomings of the attempted use of finished product testing to ensure the absence of Salmonella when the contamination is present at very low levels and is not uniformly distributed.
Although the firm’s testing program kept some contaminated products out of the market, it is unknown whether this averted illnesses, particularly without further confirmation and characterization of the Salmonella identified. The manufacturing dates reported by cases clustered in January 2022, yet Firm A did not report any presumptive positives in their product during this month. This discrepancy may suggest the testing was inadequate during January 2022 to detect positive products. Two case-reported production dates, February 27, 2022, and February 26, 2022, were after the repairs had been made; however, it should be noted that neither of these products were tested to confirm contamination. Despite the low numbers of illnesses compared to previous outbreaks, peanut butter from Firm A was implicated in this outbreak, demonstrating a limitation of reliance on testing programs to identify contamination and prevent contaminated products from reaching consumers.
Potential factors leading to the outbreak
During the 2022 inspection, Firm A discussed several potential factors with FDA and state partners. These included the entry of unfiltered air/rainwater to the cooling section of the roaster, the potential reintroduction of the outbreak strain to the facility, and the potential of a resident pathogen harbored within the facility’s production environment.
This outbreak may have resulted, in part, due to physical gaps in the production process through a cooling air supply vent that was installed incorrectly on at least one roaster, thus allowing unfiltered air and rainwater to enter the cooling section where the peanuts were RTE. Preventing water entry in dry processing environments for low-moisture foods is essential to control environmental pathogens such as Salmonella (Finn et al., 2013). Although this scenario can set up a low-moisture food for a variety of microbial concerns, it does not directly explain the presence of the matching strain of Salmonella Senftenberg identified at Firm A in 2010, particularly since this portion of the facility and equipment was not present in 2010.
Firm A provided the continual receipt of contaminated raw peanuts from a common source carrying the same strain into the processing environment on multiple occasions as a potential explanation for why the same strain of Salmonella was previously identified at Firm A; however, no data was provided demonstrating the presence of the outbreak strain in raw peanuts received at the facility. For reintroduction to be a contributing factor in this outbreak, the contamination would have to repeatedly enter the facility on the raw side of the process and then be transferred to the RTE side of the facility (meaning a breakdown of hygienic controls to prevent such transfer would have had to occur) or the contamination would have to repeatedly survive the roasting process. Another possible explanation is that the outbreak strain had been a resident strain in the Firm A environment from a contamination event preceding its 2010 recovery in the facility.
Benefits of WGS use by industry
WGS is a powerful tool used to advance food safety. The FDA recommended that Firm A consider incorporating WGS as a tool to investigate pathogen isolates obtained in their environmental monitoring program and/or finished product testing program. FDA further stated the use of WGS to investigate any pathogen isolated can provide more detailed information to support verification programs, investigations, and corrective actions.
Using the NCBI database, FDA initially identified several 2022 clinical isolates as matches to over 100 nonclinical historic isolates. Most of these nonclinical isolates were labeled as “peanut butter” or “environmental” with a location of Kentucky, but the metadata associated did not directly indicate Firm A as the origin. Although a hypothesis was formed, additional information was necessary before FDA approached Firm A about the potential issue. Had the full metadata been included in NCBI initially, FDA could have alerted Firm A sooner about the potential issue, even prior to the epidemiological association later made. Additionally, Firm A could have self-identified a potential issue by recognition of clinical isolates that may be associated with their firm. Specifically, the first clinical cases associated with the outbreak occurred in February 2022, the same timeframe when the firm was identifying the leaking cooling vent on the roaster. If Firm A was using WGS to characterize positive samples, for example, those identified in December 2021 and February 2022, and comparing their sequences with those in the NCBI database, they may have had indications months earlier than the eventual recall that their corrective actions to address all sources and routes of contamination and reliance on sampling efforts to keep contaminated product off the market were not working. Had Firm A acted in February 2022, additional cases may have been averted.
WGS analysis by the firm of the positive Salmonella samples could have helped to confirm or refute hypotheses of root causes, including the potential of a resident strain, and provide data to determine whether these strains were the same or closely related to previously identified isolates associated with the facility or additional clinical isolates in the NCBI database. WGS information has been critical in identifying multiserotype outbreak investigations (Crowe et al., 2017; Whitney et al., 2021). In the short term, FDA recognizes that transparent WGS information could be a concern for some individual firms; however, FDA also recognizes the long-term benefits of information-sharing to public health and the potential for firms to prevent outbreaks through the use of WGS in their environmental monitoring program. FDA will continue to work with partners, including industry, toward transparency and stakeholder access to WGS information (U.S. Food and Drug Administration, 2021).
Wastewater has potential for community pathogen surveillance. For example, nearly a year after an outbreak of Salmonella Java occurred, the same strain was detected in wastewater in the same geographic region (Diemert and Yan, 2019). Similarly, in 2022, four wastewater isolates were submitted to NCBI that matched the Salmonella Senftenberg outbreak cluster (Goldblum et al., 2020). Although the exact relationship of these Salmonella Senftenberg isolates to the outbreak is unknown, their identification highlights the potential of WGS to continually provide context clues, support hypothesis generation, and ultimately paint a fuller picture of adverse public health events.
Communication challenges
The advisory and recall announcement for this outbreak were published on FDA’s website the day after notifying the firm of the issue. They included information about where the recalled product was manufactured to emphasize that not all product was recalled, only specific lots. The firm originally provided a larger range of lot codes, leading to consumer confusion. Additionally, some consumers incorrectly believed that the location that was listed as “distributed by” on the product label was also the manufacturing location. Each of these posed unique communication challenges requiring additional consumer messaging and clarification from federal partners.
Conclusion
Outbreaks associated with nut butter continue to occur, albeit smaller in scale compared to previous known peanut butter outbreaks. This outbreak highlighted the importance of water management in dry operations, identifying all processing steps where RTE product is exposed to environmental contamination, and the importance of design and maintenance of the manufacturing equipment and facility to prevent opportunities for pathogen entry and harborage. This outbreak also highlights potential benefits for industry to incorporate WGS as a tool in their environmental monitoring program and the importance of appropriate corrective actions to reduce the likelihood that environmental contamination will recur. The outbreak resulted in 21 infections of Salmonella Senftenberg and four hospitalizations in 17 states. Overall, epidemiological, traceback, and historic laboratory evidence identified the commodity and processing facility and informed actions that prevented additional illnesses.
Footnotes
Acknowledgments
The hard work of all federal and state partners involved in this outbreak investigation is acknowledged, including the Kentucky Department for Public Health. The investigation team’s prolonged efforts at the firm are recognized and highly appreciated. Thank you to the CORE Communications Team for all their efforts and resilience and to the CFSAN Graphics Team for assistance with figure design.
Disclaimer
The views expressed in this article are those of the authors and do not necessarily reflect the official policy of the Department of Health and Human Services, the Centers for Disease Control and Prevention, the U.S. Food and Drug Administration, or the U.S. Government.
Authors’ Contributions
B.W.: Investigation, conceptualization, and writing—original draft preparation. D.K.: Writing, reviewing, and editing. A.E.G.: Investigation, writing, reviewing, and editing. A.C.: Writing, reviewing, and editing. S.S.: Writing, reviewing, and editing. L.W.: Investigation, reviewing, and editing. J.M.: Investigation, reviewing, and editing. A.P.: Reviewing and editing. B.W.: Writing, reviewing, and editing. K.H.-G.: Investigation, reviewing, and editing. V.W.: Investigation, reviewing, and editing. A.P.: Investigation, reviewing, and editing. T.J.: Investigation, reviewing, and editing. L.G.: Investigation, reviewing, and editing. M.L.: Investigation, reviewing, and editing. L.G.: Investigation, reviewing, and editing. S.V.: Conceptualization, writing—original draft preparation, reviewing, and editing.
Disclosure Statement
All authors declare that they have no conflicts of interest.
Funding Information
The authors received no financial support for the research, authorship, and publication of this article.
